Abstract
Extensive application of technologies like phage display in screening peptide and protein combinatorial libraries has not only facilitated creation of new recombinant antibodies but has also significantly enriched repertoire of the protein binders that have polypeptide scaffolds without homology to immunoglobulins. These innovative synthetic binding protein (SBP) platforms have grown in number and now encompass monobodies/adnectins, DARPins, lipocalins/anticalins, and a variety of miniproteins such as affibodies and knottins, among others. They serve as versatile modules for developing complex affinity tools that hold promise in both diagnostic and therapeutic settings. An optimal scaffold typically has low molecular weight, minimal immunogenicity, and demonstrates resistance against various challenging conditions, including proteolysis – making it potentially suitable for peroral administration. Retaining functionality under reducing intracellular milieu is also advantageous. However, paramount to its functionality is the scaffold’s ability to tolerate mutations across numerous positions, allowing for the formation of a sufficiently large target binding region. This is achieved through the library construction, screening, and subsequent expression in an appropriate system. Scaffolds that exhibit high thermodynamic stability are especially coveted by the developers of new SBPs. These are steadily making their way into clinical settings, notably as antagonists of oncoproteins in signaling pathways. This review surveys the diverse landscape of SBPs, placing particular emphasis on the inhibitors targeting the oncoprotein KRAS, and highlights groundbreaking opportunities for SBPs in oncology.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
Transformation of a normal cell into a cancerous one involves several molecular events, including activation of oncogenes – either directly or indirectly – and/or loss of the tumor suppressor functions. These events occur in parallel with modifications in the regulation of gene expression leading to enhanced cell proliferation, replicative immortality, and reduced sensitivity to anti-tumor defense. In the pursuit of potent therapeutic solutions, a substantial assortment of targeted drugs has been developed. Predominantly, these drugs are of two types: monoclonal antibodies, which primarily serve as receptor blockers on tumor cells, and small molecules that predominantly inhibit protein kinases vital for the growth and invasiveness of cancer cells. Research in the field of antibodies conjugated with toxic inhibitors, as well as precise guiding of the cellular and viral antitumor agents is currently one of the most promising areas of modern molecular oncology. Challenges in the cancer treatment arise mainly from the tumor adaptive resistance, caused by molecular processes that bolster their aggressiveness. Genetic diversity of tumor cells often renders many drugs ineffective as they cannot target the entirety of the tumor cell population. The more selective an inhibitor is, the swifter resistance to it emerges due to the high mutation rate and epigenetic plasticity of cancer cells. Consequently, the potent drugs of the future might not be highly specific. Instead, they should address a broader category of interrelated target molecules, indicating a need for increasingly intricate therapeutic strategies.
In this review, we focus on oncological applications of the synthetic binding proteins (SBPs, also called mimetics or inaccurately referred to as antibody mimics). These SBPs include non-immunoglobulin scaffolds, with the term “scaffold” denoting a single-domain polypeptide with a typical molecular mass between 4-20 kDa. We also discuss the challenges of their development and production, a significant hurdle to their broader adoption in both scientific and medical areas, especially considering that a decade has already passed since the publication of the seminal review [1].
ADVANTAGES AND LIMITATIONS OF ANTIBODIES
Immunoglobulins stand out as the most multifaceted and diverse binding proteins in humans; their antigen-binding sites consist of six loops connecting β-strands in the variable domains of the light and heavy chains. At present, traditional antibodies are the mainstay among the protein drugs used both as tools in basic research and applied therapeutic targeting. It is crucial to balance pros and cons of immunoglobulins. Their preeminence stems from the several traits: their natural origins as major plasma proteins, their high affinity and specificity to chosen targets, established methods for large-scale production, availability of straightforward conjugation procedures, etc. Notable advancements have been made with camel and llama antibodies, as they maintain many properties even without light chains, which permits their intracellular applications under the name of nanobodies. Nonetheless, these still fall short when compared to numerous other SBPs based on alternative scaffolds, which are selected to precisely target intracellular oncoproteins, including some previously deemed “undruggable”, like certain GTPases. It is worth noting that while monoclonal antibodies have shown profound efficacy against the tumor cells and specificity to their targets in comparison with the low molecular weight chemical inhibitors, their substantial size (around 150 kDa) limits their potential in targeting intracellular proteins. Other disadvantages of antibodies restricting their application in several contexts include high production costs, presence of disulfide bridges complicating their use in intracellular reducing environments and their suitability in such procedures as tumor imaging or radioimmunotherapy due to their large size, slow blood clearance, and prolonged retention in non-tumor tissues. Reducing size could provide a number of advantages, such as increased accessibility to sterically hindered epitopes. Other enhancements, such as reducing hydrophobicity, reducing aggregation, and increasing stability of the scaffold irrespective of its redox potential, are also crucial. Therefore, there is an immediate need to engineer both protein and non-protein antibody analogs that retain high affinity but come with the advantages of reduced size, easier synthesis, and enhanced stability both inside and outside the cell.
GENERAL INFORMATION ON THE DIVERSITY OF NON-IMMUNOGLOBULIN SYNTHETIC BINDING PROTEINS
One must recognize the inevitable trade-offs when optimizing various properties. For instance, boosting affinity might require sacrificing other crucial parameters like specificity and often stability [2]. Such trade-offs have been especially well documented for antibodies [3]. Indeed, all monoclonal antibodies are inherently polyspecific, capable of binding to epitopes with significantly different geometries [4, 5]. When discussing alternative SBP scaffolds, understanding these trade-offs becomes crucial.
Occasionally, it is feasible to both anticipate and propose ways to enhance SBP properties. For instance, many SBPs have a high number of lysine and arginine residues. With antibodies, abundance of these residues worsens their properties [6]. Analogously, for the intracellular SBP applications, there is a likelihood of non-specific binding to nucleic acids. One might even consider whether it may be beneficial to reduce complexity of interactions by minimizing both the size of molecular scaffold and the size of interaction surface. Ideally, SBPs should be good building blocks for combinatorial approaches in the creation of multifunctional molecules. A number of excellent reviews on alternative SBP scaffolds can be recommended. It is worth mentioning only the most recent ones from the last four years covering: promising therapeutic SBPs [7], SBPs in developmental biology [8], SBPs for toxin neutralization [9], SBPs for pull-out followed by mass spectrometric analysis of the therapeutically important proteins and biomarkers [10]. The main aim of our review is to explore the development of diverse SBPs for oncological applications.
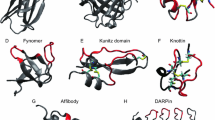
Search for the new variants of SBPs typically starts with the randomized library construction, moves to selection and clone analysis, and culminates in rational design. A robust SBP scaffold should tolerate substantial variation of the residues forming binding site of its target. Existing platforms (Fig. 1) can logically be grouped based on their binding surface formation into two groups: helix-chain type (DARPins, miniproteins including affibodies, affilins and others) and loop type (monobodies, anticalins, avimers, knottins, atrimers, fynomers and others, including also immunoglobulins). This distinction is very important because in the first type, residues with side chains exposed outwards are randomized to create libraries of new variants, whereas in the second type the scaffold itself is most often unaffected, but the loops may experience significant variations, even in their lengths. Apparently, except for the cyclic miniproteins, the N- and C-termini are free in all variants, allowing fusion with other sequences. To date, the elements like intrinsically disordered proteins [11] and, less commonly, PDZ domain [12] remain underexplored, as the primary requirement for good scaffolds is thermodynamic stability. Given the popularity of monobodies, DARPins, anticalins, and affibodies [13, 14], they warrant individual consideration. Other scaffolds are briefly touched upon here, noting that attempts to innovate using a novel scaffold might occasionally reach a deadlock, as evidenced by tendamistat [15].
Three-dimensional folds of SBP polypeptide chains in complex with their targets. a) Monobody (green) against fluoride channel (grey), 6BQO; b) DARPin (grey) against MTFP1 protein (green), 6FP7; c) lipocalin (grey) against carboxymethylcobactin (green), 1X8U; d) ZHER2 affibody (green) against HER2 protein (grey), 3MZW; e) crystallin affilin, 2JDG; f) monobody (green) against MLKL (grey), 7JXU; g) DARPin (green) against caspase 7 (grey), 4LSZ; h) monobody (green) against fluoride channel (grey), 6B2B.
Miniproteins are conventionally defined as proteins with masses significantly less than 10 kDa (closely associated and extremely popular affibodies are discussed separately). The characteristic small size of miniproteins also facilitates their chemical synthesis for preparative production; their diversity has been covered in a special review [16]. Here we should mention a prototypical example of miniproteins, the knottins [17]. Their size starts from 14 amino acid residues (a.a.), usually spanning around 30 a.a., and they are commonly stabilized by numerous disulfide bonds. Knottins possess antiparallel β-strands connected by loops of varying lengths, 6 cysteine residues are linked by three disulfide bonds. This composition offers remarkable stability even under harsh acidic or alkaline conditions. Avimers provide proof that the stable miniproteins, unlike nanophitins, affibodies, etc., can also originate from the human protein domains (LRP – VLDLR, etc.) of 35-a.a. length, in which 12 a.a. are conserved and also contain six disulfides [18]. Adhyron affimers originate from the consensus sequence of cystatin and stefin: such disulfide bond-free SBPs consisting of one α-helix and four β-strands with two variable loops display impressive thermal stability (Tm around 101°C) [19, 20]. Among other applications, they are considered as affinity tools for diagnosing carcinoembryonic antigen levels [21]. The Kunitz domain contains approximately 60 a.a. with three disulfide bonds and three loops suitable for randomization and variation, and its usefulness is supported by successful clinical applications of the kallikrein inhibitor, Kalbitor. Recent developments using this domain include the soybean trypsin inhibitor gastrobody tolerating pH 2 [22] and its sunflower homologue [23]. Diversity of the protease inhibitors with Kunitz domain among human proteins is quite high. For example, such domain from the amyloid precursor protein inhibitor (APPI) could be a good platform, three mutations are sufficient for transformation into an excellent inhibitor of mesotrypsin (Ki about 89 pM) [23], or into a proteolysis-resistant inhibitor of kallikrein-related peptidase 6 (KLK6) with Ki of about 160 pM and lifetime in the organism about 10 days [24, 25]. Potential of some developments like SPINK2 [26] or proteins from snake venoms [27] is currently difficult to foresee.
Antibodies of a non-immunoglobulin nature from jawless fish lay the groundwork for a modular scaffold of the so-called repebodies [28], demonstrating high level of expression in bacteria, thermodynamic stability, including under pH changes, and ability to inhibit the VEGFR type target in vivo [29].
The SH2/SH3 domains, which are widespread among the non-receptor tyrosine kinases of the Src family, are vital for cell cycle regulation. They have around 100 a.a. in their composition organized into a globule with a central β-sheet encircled by two α-helices with two loops that can be randomized. Fynomers, the most know representatives of this class, are derived from the tyrosine kinase Fyn [30], are cysteine free, stand out for their soluble expression in bacteria, commendable stability without significant aggregation (Tm ~ 70°C), while high degree of conservation suggests low immunogenicity [31].
In terms of stability, proteins and peptides from hyperthermophilic and hyperacidophilic microorganisms are of undoubted interest. Representatives of the so-called OB-domain originating from the DNA-binding polypeptides Sac7d and Sso7d of archaea of the genera Sulfolobus, Acidanus, Pyrobaculum, etc. under the names nanophytins and affitins have been significantly developed. Structurally, they comprise a short cylinder of five β-strands capped by an α-helix; they are remarkable for their good soluble expression in E. coli and high stability (up to 74°C in the pH range 0-12) [32-35].
The intracellular protein glutathione transferase, notable for its solubility, can be transformed into a universal scaffold glubody [36], but its development has been limited.
Affilins, originating either from the dimerized ubiquitin or γ-B-crystallin, exemplify the challenges in repurposing excellent natural proteins as SBP scaffolds (with a mass of about 20 kDa). While affilins boast impressive solubility and stability, their potential randomization sites are limited, making modification challenging: indeed, affilins are highly soluble and very stable (in 8 M urea, up to 75°C, pH range 1 to 9) due to their compact stacking (for example, the 76-a.a. long ubiquitin has 3.5 α-helixes and five β-strands). However, only 6 a.a. with chains exposed outward from the β-structure side that form a binding region can be used for randomization, introduction of variable loops significantly impairs the above advantages, and additional efforts are required to reduce rigidity of the secondary structure [37].
Continued exploration of novel scaffolds from human proteins, using refined screening methods with carefully formulated criteria for systematic screening from the database of three-dimensional structures (molecular mass in the range 10-25 kDa, monomer, presence of a structure with sufficiently high resolution (<3.0 Å), production in E. coli) followed by manual selection according to the additional criteria (different from the previously published protein scaffolds; low number of cysteine residues and disulfide bonds; no evidence of high toxicity and/or immunogenicity, no ligands or cofactors; high solubility, easy purification) yielded a new scaffold named ProBi (Protein Binder). It contains two surface-exposed sites suitable for randomization demonstrated by successful selection of variants binding to human interleukin-10 [38]. Given the large number of predicted structures, such methodologies could become even more fruitful.
Further development of the de novo design methods will inevitably be increasingly productive. For example, it has been shown for the loop-helix-loop and loop-helix-loop-coil-loop elements (LUCS) that even such simple module can generate entire families of proteins with customizable geometries including both natural and previously unknown sequences [39]. A good example is the scaffold Alphabody (with molecular mass of about 10 kDa, designed on the principle of triple helix connected by linkers rich in glycine residues, homologues of which do not exist in nature) that recently has been introduced as a potential therapeutic candidate [40].
MONOBODIES-ADNECTINS AND OTHER SYNTHETIC BINDING PROTEINS BASED ON THE FIBRONECTIN DOMAIN
The results of pioneering research on monobodies/adnectins were published in 2008 [41], 2011 [42], and 2013 [43]. By 2012, the potential of monobodies in oncology began to be rigorously assessed, as detailed in the review [1]. Gradual improvements in understanding of the problems of their practical use [44, 45] have propelled at least two monobodies into clinical trials [7].
Monobodies that originate from the tenth domain of fibronectin 3 (FBN3) were denoted, therefore, as 10Fn3. Initially, several different domains were compared and any of them could be used (pronectins originate from the domain 14 of fibronectin 3). The tenth domain was favored due to its unmatched stability among the human FN3 repeat domains boasting a mean denaturation point (Tm) of 84°C [41]. It also exhibited tolerance to mutations across three exposed surface loops. Thus, a monobody is a simple structure like a small β-sandwich of seven β-strands tolerating randomization and elongation of two or three surface-exposed loops. Remarkably, melting point of the scaffold is around 80°C, even in the absence of disulfide bonds [46]. Since cysteine residues are absent, stability of the monobody is weakly dependent on the redox potential, eliminating the possibility of disulfide-mediated aggregation in the intracellular environment. The traditional target of fibronectin is integrins, and interaction of FN3 with its target is facilitated by the loop flexibility and β-sheet dynamics to ensure optimal binding conformation to the target protein, in contrast to the rigidity of antibody β-sheets stemming from the internal disulfide bonds between heavy and light chains [30, 31]. Advantages of monobodies explain their use in diverse scientific applications: Kükenshöner et al. reported the use of monobodies as protein blockers, modulation of allosteric sites, and crystal chaperones [47]. Intracellular stability of monobodies has also been used for intracellular studies of the structure and function of proteins, where monobodies fused to the fluorescent protein (here called intrabodies against the endogenous neuronal proteins) have allowed visualization of the excitatory and inhibitory synapses in living neurons [48].
High stability of monobodies was further improved in the ultra-stable scaffold FN3con [49] also created by the consensus design (using 2123 sequence alignments); it is even suitable for grafting loops from other monobodies [50, 51]. It should be noted, however, that high stability may not always be desirable for therapeutic applications.
LIPOCALINS-ANTICALINS
Aesthetically, these are, probably, the most attractive scaffolds as they form an elegant bowl similar to flower calyx [52, 53] with central eight-strand antiparallel β-cylinder and lateral α-helix. The ligand binding pocket is formed by four loops that connect the β-strands protruding from the open end of the β-barrel, and these four loops can be used for variation. Typical anticalins comprise around 188 amino acid residues and at least one N-glycan with molecular mass of about 21-24 kDa and isoelectric points of approximately 7.1. Thus, the lipocalin/anticalin libraries have four structurally hypervariable loops atop of a highly conserved β-barrel. Although the lipocalin/anticalin is not a small peptide, it proved to be suitable for phage display. Moreover, they show good bacterial expression in terms of solubility in the form of monomers [54]. Using targeted random mutagenesis combined with molecular selection techniques, it is possible to change the shape of this loop region, creating pockets for tight binding of various ligands, from small molecules, peptides, to proteins. Origin of the scaffold from the naturally secreted and highly soluble human protein makes it particularly appealing. Lipocalin-2 (also known as LCN2, NGAL, oncogene 24p3, siderocalin secreted by neutrophils) is able to actively bind iron simultaneously with catecholate thus arresting bacterial growth. Its homologue is prealbumin from tear fluid, which is also an antibacterial agent due to its ability of iron sequestration. Unfortunately, this same property is exploited by some tumors, for example, in the process of metastasis to cerebrospinal fluid [55], and enhances aggressiveness of breast cancer [56] (reviewed in more detail in [57, 58]). However, on the other hand, expression of lipocalin-2 as a transgene can significantly enhance the action of oncolytic adenoviruses in some tumor models, such as pancreatic adenocarcinoma, and colorectal cancer cell lines [59, 60]; moreover, its expression is reduced in gastric cancer [61]. Given these diverse roles, therapeutic use of lipocalins demands meticulous examination, especially since they may influence cancer progression and correlate with cognitive impairment in some patients [62].
DARPINS
DARPins resemble an elegant comb; they exemplify an extremely successful development of an artificial scaffold based on ankyrin repeats. This design showcases efficacy of the consensus method in protein design, where consensus is derived from multiple alignments of the related sequences (unfortunately, although promising, this approach often does not yield successful results). In the human proteins, this repeat is common among the adaptor intracellular proteins and contains about 33 a.a. with two α-helices and one β-strand, hydrophilic side-exposed residues, and hydrophobic interior. A typical DARPin contains two or four repeats flanked by two capping repeats, thus giving a mass of about 14-21 kDa (denaturation temperature also therefore varies widely from 60 to 95°C). It should be pointed out that, in comparison with most of the SBPs mentioned here, DARPins present a more expansive binding surface area. This facilitates randomization of a considerable number of exposed residues. Presently, at least two DARPins are in the pipeline for clinical trials [7].
AFFIBODIES
For applications where the minimized molecular weight of an SBP is paramount, affibodies derived from the B domain of the IgG-binding site of S. aureus protein A have frequently emerged as the preferred choice among the various scaffolds. In their design, a small number of mutations have been shown to be sufficient to increase stability and loss of antibody binding ability. The three α-helices lacking cysteine residues are linked together forming a scaffold weighing approximately 6 kDa, where the exposed residues of α-helices 1 and 2 can be randomized.
It has now been possible to develop affibodies against many tumor-associated targets, and it is their compact size that is especially beneficial for diagnostic purposes using radioisotope-labelled affibodies. Nonetheless, it is important to note that the size advantage of an affibody diminishes, when it is fused with a partner protein (e.g., a toxin), and thus compactness becomes relatively less significant compared to other scaffolds. The same is true for bispecific affibodies [63].
PRINCIPLES OF CREATING AND SCREENING OF LARGE LIBRARIES OF SYNTHETIC BINDING PROTEINS
Fundamentally, the main methods for screening SBP libraries can be divided into cellular display (predominantly yeast), viral display (most commonly phage display, on the surface of the phages using the pVIII protein of the filamentous bacteriophage fd/M13), and entirely in vitro methods like RNA display (an improved version of ribosomal display). While phage display has proven to be effective for monobodies [41, 64], and is also suitable for many other SBPs including repebodies [29], current limitations of the real phage libraries, which are described in the excellent review [65] are significant. The fundamental approach to utilizing such SBP libraries is illustrated in Fig. 2.
Selection of synthetic binding proteins by cellular, phage, or mRNA display. Systems for expressing SBPs on the surface of a virus (a, most commonly bacteriophage), cell (b, most commonly yeast), or as nascent polypeptide chains synthesized but not dissociated from ribosomes (c, due to puromycin linker) allow screening of SBP libraries quite rapidly and efficiently, e.g., using targets immobilized on a solid phase. Typically, about three rounds of library selection and amplification are required before individual clones can be analyzed. Draft prepared with biorender.com.
Problems with production of synthetic binding proteins. Quite often rather interesting projects with recombinant proteins are set aside due to their poor stability and, consequently, low expression. This is an issue that antibodies seldom face.
Monobodies are often very successfully produced in E. coli. Couple of examples could be mentioned: monobodies for which the authors use the term intrabodies [66, 67], and also monobodies described in the recent work [68]. Interestingly, it was also observed during analysis of a number of random clones that although monobodies are usually produced in satisfactory to commendable levels (4-20 mg/liter culture), their stability and solubility do not correlate with overall expression levels [69], i.e., a substantial fraction of clones yields a significant amount of aggregated protein.
Since the efficiency of monobodies and other non-immunoglobulin scaffolds often lags behind the antibodies, there are emerging strategies to enhance performance of monobodies, such as through cross-linking with a particular focus on disulfide bonds [70].
Not every scaffold can tolerate mutations of the variable part, and this often leads to destabilization of the entire protein structure. Inconsistent success with heterologous expression points to inherent limitations in some scaffolds. However, there are recent examples of partial successes, e.g., an avimeric A-domain consisting of 35 a.a. with 12 conserved a.a. including six disulfides, despite such a small size has been produced in E. coli, and the required S-S bonds were formed by exposure to air, yielding a product of about 4 kDa with excellent stability.
It is possible to significantly change specificity of the high-affinity natural binding protein, which has been clearly demonstrated by the example of avidin transformed into the steroid-binding protein [71] and further development of the so-called antidines [72].
Grafting should be considered as a promising but not always fruitful strategy. There are numerous successful instances where binding loops from the effective antibodies have been grafted onto different proteins, such as green fluorescent proteins, to produce a new affinity tool that can bind a target [73, 74]. Such innovations typically pave the way for exciting, albeit non-therapeutic, applications like creating fluorescent sensors for proteins critical to oncogenesis, like MDM2 [75]. Notably, while grafting can occasionally eliminate stability reduction after selection, directed evolution targeting VEGFR2 resulted in the reduced thermostability and unchecked oligomerization. However, when loops were grafted onto the FN3Con monobody, an ultra-stable scaffold, the resultant construct (FN3Con-anti-VEGFR2) retained activity even after 2 years of storage at 36°C [51].
Problems of pharmacodynamics and pharmacokinetics of synthetic binding proteins. It should be noted that all small SBPs described here are easily excreted through kidneys, so the standard approaches of increasing total molecular weight, such as PEGylation, as well as introduction of the albumin-binding domain are used to prolong the circulation time [76]. A prevalent challenge with many other biologics introduced into the bloodstream is their excessive accumulation in the liver. It should be noted that accumulation and persistence of the targeted therapeutic drug in a tumor is influenced by a myriad of factors. These include the drug’s affinity for the target, molecular weight, half-life, propensity for extravasation, and more. DARPins, with their larger size, often have problems with adequate penetration even into the cultured spheroids [77].
On the contrary, tracers that clear faster from the non-target tissues have diagnostic advantages. They facilitate more accurate assessment of signals from various tissues, providing patients with quicker diagnoses, and allowing for a prompter commencement of anti-cancer treatments. For such diagnostic applications, the ideal choice would be smaller proteins that become visible in the target tissues within 4 hours post-administration
NEW APPROACHES TO IMPROVE SYNTHETIC BINDING PROTEINS FOR ONCOLOGY APPLICATIONS
Conjugation and chimerization. SBPs based on alternative scaffolds are increasingly regarded as promising modules for constructing multivalent affinity tools (review [78]). There are documented successes in conjugating these SBPs with small molecular weight drugs and toxins. Shipunova and Deyev provided an insightful review on the SBP conjugates with liposomes and nanoparticles [79]. Notably, they highlight the studies where the single-layer liposomes, sized between 80-90 nm in diameter and containing components like cytochrome c, fluorescent mCherry, and highly toxic exotoxin A from Pseudomonas aeruginosa with DARPin against HER2 specifically stain and kill the HER2+ cells [80]. Although trastuzumab has shown efficacy in the HER2-positive breast cancer therapies, specific antibodies typically fall short in treatment of solid tumors. This emphasizes significance of the developing protein-drug conjugates, especially given that the cytotoxic drugs can denature protein structures. This is important in the case of cytotoxic drugs that can denature protein structures, conjugation of small proteins needs to be tightly controlled in order to prevent aggregation. This important feature is crucial because most cytotoxic drugs are hydrophobic and can induce an uncontrolled aggregation of antibodies and other proteins used for conjugation.
Affitoxin (successfully produced in E. coli), consisting of a shortened diphtheria toxin and an anti-HER3 affibodies [81] coupled to the cytotoxic tubulin polymerization inhibitor DM1 exhibited strong cytotoxic effects on the BxPC-3 pancreatic carcinoma cells (IC50 7 nM) [76]. Advantage of monobodies is absence of cysteines, which permits introduction of one cysteine residue specifically intended for conjugation with small molecular weight drugs [82]. Conjugates of DARPin or recombinant antibody (scFvFc) against EGFR with monomethylauristatin E as a toxin showed subnanomolar cytotoxicity against the A431 human lung squamous cell carcinoma A431 cells in vitro but no specific anti-tumor activity in the mouse xenografts. The most interesting result was the comparison of penetration depths into cell spheroids: the monomer DARPin was superior over both antibody and divalent DARPin, whereas the DARPin-Fc was the least successful in this respect [83].
Extracellular delivery of synthetic binding proteins for cancer diagnosis, theranostics, and therapy. Binding of SBPs to the cell surface markers for the purposes of tumor growth diagnosis, therapy, or theranostics is at the forefront of the fight against cancer. Radioisotope imaging is covered in special reviews on antibodies and non-immunoglobulin scaffolds in [84, 85], a broad coverage of specific molecular tools can be found in [86, 87], and excellent reviews focused on tumor theranostics [79, 88] can be also recommended. We would like to highlight a valuable and very rare comparative work in which efficiency of binding of the full-length IgG, DARPin, and affibody covalently bound with the magnetic fluorescent nanoparticles directed against HER2 was examined. The affibody demonstrated superior specificity and selectivity for labeling cancer cells using nanoparticles [89].
VEGF and VEGFR2. When considering primary molecular targets for diagnostic and therapeutic agent development, the anti-angiogenic agents, such as repebodies targeting VEGF should be mentioned [29]. While VEGF is a secreted protein and not directly a tumor marker, its receptor, VEGFR2, serves as a prime target. For instance, the CT-322 monobody against VEGFR2 [90] advanced successfully through the phase I clinical trials for pancreatic cancer [51].
HER2 is an archetypal marker for numerous breast cancers, it has been the focal point for much developmental research, largely due to the success of trastuzumab and its SBP-based analogs (some of the works are mentioned above). Toxic DARPins targeting HER2, when conjugated with gold mini-nanorods, selectively accumulate in the HER2-positive xenograft tumors in mice. This accumulation promotes significant tumor reduction following photodynamic therapy [80]. The 68Ga-labelled affibody molecules allow accurate and specific measurement of HER2 expression in the breast cancer metastases via PET imaging [91].
EGFR. For EGFR, the radiolabeled affibodies are particularly valuable in theranostics [92]. Success of this theranostics approach is supported by the example of affiFAP, which consists of an affibody against EGFR and a fluorogene activating protein. This compact molecular recognition reagent can activate fluorescence upon binding, allowing tumor imaging with low levels of non-specific tissue staining [93]. Centyrines (from 10 kDa tenascin) and repebodies have performed well as a dye conjugate in the intraoperative fluorescence diagnosis of tumors overexpressing EGFR [94]. There is also an excellent work on therapeutic affibodies, where an anti-EGFR affibody was conjugated with IR700 for photodynamic therapy, reactive oxygen species (ROS) generated under IR irradiation induced immunogenic cell death (ICD) with subsequent dendritic cell maturation and in vivo therapeutic response was observed in the brain tumors shortly after IR irradiation [95].
Ecto-CRT (calreticulin that is exposed outside the cell surface) plays an important role in phagocytic clearance of apoptotic cells during immunotherapy. Two peptides (KLGFFKR from integrin-α and GQPMYGQPMY from Hep-I) specifically bind ecto-CRT during ICD induction. Monobodies grafted with both sequences showed good binding to ecto-CRT, effectively detecting pre-apoptotic cells upon treatment with doxorubicin but not gemcitabine, which does not induce ICD. Moreover, by using the CRT-specific monobodies, it is possible to detect induction of ecto-CRT in the cancer cells in response to drug exposure [66].
Using phage display to screen combinatorial libraries of knottins, researchers discovered the EDB-specific SBPs. These binding proteins were effective in marking EDB+ cells, which possess a tumor-specific fibronectin domain. Remarkably, they demonstrated this ability within the picomolar range when tested on the tissue sections from human glioblastoma U-87 MG mouse xenografts [96].
Synergies observed when two or more SBPs are used [97] are well illustrated by the example of the affitoxin consisting of a shortened form of diphtheria toxin and HER3-binding domains of an affibody [81], and also by a similar example with two DARPins [98]. The conjugates of Pseudomonas bacillus exotoxin A and barnase with DARPins were used in the mouse model of mammary carcinoma: in this case, the anti-HER2- or anti-EpCAM monotherapy was ineffective in contrast to the simultaneous action against HER2 and EpCAM [99]. Similar examples of strong enhancement of the SBP action on tumor cells have been reported in other studies [100, 101].
Oncolytic viruses (OVs). In the realm of oncolytic viruses (OVs), SBPs present an intriguing prospect for targeted therapy. They can be embedded into the OV capsid surface more effortlessly than traditional antibodies. There is a palpable optimism in the published data; for instance, the measles vaccine virus MeV was engineered to carry a DARPin targeting EGFR. Intriguingly, this configuration prompted activation by the tumor-linked matrix metalloproteases upon binding. Such a dual-targeted OV replicated in the EGFR+/MMP+ tumor cells but was safe for healthy cells (EGFR+ human keratinocytes). This virus destroyed glioblastoma cells and others [102]. Knottins were also used in a similar way; for example, CKP (cystine knot protein, binding αvβ3, αvβ5, and α5β1 integrins with nanomolar affinity); in this case the constructed OV (called MV-CKPint) infected, replicated in, and killed human glioblastoma, medulloblastoma, diffuse intrinsic pontine glioma, and melanoma cells in vitro. When administered intravenously, this OV was better able to reach glioblastoma cells, causing cytopathic effects similar to those of intratumoral virus injection [103].
Delivery of the replication-deficient viruses such as adeno-associated viruses (AAVs) used for vector-based cancer gene therapy can also be successfully enhanced by SBP, for example, by exposing the HER2-specific DARPin on the surface of AAVs [104].
Cell-based therapies. Similarly, SBPs can also be used to amplify cellular anti-tumor therapies based, for example on T-cells with chimeric antigen receptors (CAR-T) [105]. Here we should specifically mention the recent success of a large team of mainly Russian authors who improved CAR-T due to the high affinity of the bacterial toxin–antitoxin–barnase–barstar complex, where they used the DARPin-barnase conjugates to direct CAR T cells to solid tumors to kill the HER2+ carcinoma cells in vivo [106].
SYNTHETIC BINDING PROTEINS AGAINST INTRACELLULAR CANCER TARGETS
Inhibitors of interactions with MDM2. The product of p53 gene is one of the most important human oncosuppressors, restraining uncontrolled cancer cell division [107]. In tumors, p53 is often mutated or degraded at the protein level, largely due to the action of ubiquitin ligase MDM2, which in turn also undergoes post-translational modifications [108]. While there exists a number of low molecular weight inhibitors designed to disrupt the p53 and MDM2 interaction [109, 110], clinical trials for these inhibitors have yet to yield positive results. There have also been quite a few affinity tools, such as monobodies against the N-terminal domain of MDM2/X and against the α-helix present in the N-terminal transactivation domain of p53 [111], have demonstrated promising in vitro activity. However, these results have not yet been translated into in vivo success in clinical settings.
Attack on KRAS. GTPases of the Ras family, with KRAS being particularly noteworthy, are often mutated in a plethora of cancers. For instance, over half of the pancreatic ductal adenocarcinoma cells have KRAS mutations such as G12D, G12C, G12R, or G12V. These enzymes contain a pocket that is able to bind GTP and GDP with picomolar affinity, triggering intracellular signaling pathways that enhance cell proliferation. Since GTP and GDP are present at millimolar concentrations in the cell, any external small-molecule inhibitors face extremely stiff competition. The rest of the surface of the Ras protein is relatively flat, lacking any pronounced pockets for high-affinity interactions with small molecules.
In a remarkable discovery, the NS1 monobody was identified as an allosteric inhibitor of the Ras-mediated signaling. Surprisingly, NS1 binds to the surface that differs from the binding surfaces of other effector molecules that bind to Ras proteins. The mode of action of NS1 does not disrupt the Ras-Raf kinase interaction, but it inhibits Ras dimerization, which further inhibits Raf dimerization, leading to inhibition of the Ras-mediated signaling in its entirety [112]. Monobodies against the KRAS G12V mutant were first described as “RasIns” (or intrabodies). These monobodies are selective for the active state of Ras. At the same time, these monobodies bind both H- and K-Ras mutants in complex with GTP [67]. In this case, production in E. coli was successful. Recent progress in the development of monobodies against KRAS mutants has been substantial, for instance, the 12VC1 monobody (Fig. 3 and Table S1 in the Online Resource 1) recognizes the active state of KRAS with G12V and G12C mutations [113]; the R15 monobody reacts with all apo-Ras isoforms (those not bound to GTP or GDP), displaying a particular affinity for KRAS (G15C) in vivo [114]. Another promising candidate is the pan-Ras monobody JAM20, which binds to Ras independent on their GTP/GDP association [115].
Amino acid sequences of some SBPs aligned with their precursors. Identical amino acid residues are shown in grey, unique tyrosine and tryptophan residues in SBPs are shown in bold. a) Affibodies: alignment of an anti-HER2 affibody with the homologous region of S. aureus protein A sequence; b) DARPins: alignment a loopDARPin-type DARPin with the homologous region of human ankyrin (ANK2); c) alignment of monobodies (NS1 against Ras and 12VC1 against KRAS with G12V mutation) with the homologous region of the human fibronectin (FN).
The DARPin K27 predominantly interacts with the inactive form of Ras-GDP with Kd of 4 nM. Intracellular expression of K27 significantly reduces Ras activity, inhibits downstream signaling, particularly it reduces the level of phosphorylated ERK, and hampers growth of the HCT116 cells in soft agar [116]. Certain DARPins, which specifically inhibit the KRAS isoform by binding to the allosteric site spanning the region around the KRAS-specific histidine residue 95, specifically inhibit the KRAS/effector interaction, and signaling pathways dependent on it in the cancer cells [117].
The Ras-binding miniproteins, functioning as dimers, associate with the effector domain of Ras. In the specific Ras point mutants, these miniproteins stabilize the protein in an “open” conformation. The aPP miniprotein binds to the N-terminal type II polyproline helix linked by a short loop to the C-terminal α-helix, which is stabilized by the hydrophobic interactions. Notably, this miniprotein can be efficiently produced in E. coli [118].
Conformationally selective SBPs tailored for the mutant KRAS have also been designed based on Repebodies, which effectively block interaction between the active KRAS and the Ras-binding domain of BRAF, thereby inhibiting the KRAS-driven signaling [119].
The concept of forced degradation of intracellular protein targets using binding elements has attracted significant attention. In particular, monobodies can be engineered to degrade specific protein targets by selectively directing a degradation signal to an endogenous protein. This is accomplished either through conjugation or fusion with the E3 ubiquitin ligases (Fig. 4). The most popular ligases are MDM2 or oncoprotein suppressor VHL (Von-Hippel–Lindau). The strategy opens doors to myriad developmental opportunities, considering presence of approximately 600 distinct E3 ligases in the human cells. Notably, of all the SBPs, the fusion constructs of monobodies with E3 ligases seem to be the most promising [120, 121]. The challenges of delivery could be resolved using multiple potential solutions. A chimeric bacterial toxin consisting of a subunit of the shigatoxin B homologue (Stx2B) and translocation domain of the P. aeruginosa exotoxin A (ETA-II) fused to VHL and a monobody against endogenous tyrosine kinases could be mentioned as an example [120].
Synthetic binding proteins as transgenes of oncolytic viruses. Expression of the lipocalin-2 transgene can significantly enhance the action of oncolytic adenoviruses in some tumor models such as pancreatic adenocarcinoma and colorectal cancer cell lines [59, 60]. Clearly, OVs should be armed with the sequences that encode MDM2, VHL, or other ubiquitin ligase fusion proteins with monobodies against KRAS mutants and other cancer targets.
NEW DIRECTIONS FOR THE DEVELOPMENT OF SYNTHETIC BINDING PROTEINS FOR ONCOLOGY
Modular approach in the SBP design proves to be highly effective. In the case of miniproteins, solid-phase peptide synthesis is possible opening doors to a vast array of modifications including novel cross-links, structure of which may play a special role, as demonstrated with the successful example of knottin (33 a.a. polypeptide from the trypsin inhibitor from plant Ecballium elaterium that interacts to the αvβ3, αvβ5, and α5β1 integrins) dimerization [122], where a dimer of knottin helps to increase sensitivity of pancreatic cancer cells to gemcitabine.
There are various methods used for protein ligation, and it is best to refer the reader to the reviews that describe in detail pros and cons of the diverse methods like dockerin–cohesin interaction, SpyTag-SpyCatcher specific transglutaminase reaction system, sortase-mediated ligation and others [123], as well as formation of aldehyde groups on the protein surface by formyl glycine generating enzyme [124]. However, it is important to note that the use of cross-links should be expanded considering, for example, discovery of the lysine-cysteine bridges sensitive to redox potential (NOS and sulfur-oxygen-nitrogen-oxygen-sulfur, SONOS bridges) [125].
Catalytic synthetic binding proteins. Here we should mention catalytic antibodies, especially those with proteolytic activities. Such molecules could offer a dual advantage – binding to and subsequently degrading their targets without relying on the proteasome. Moreover, such SBPs would have to be extremely site-specific and personalized, potentially outperforming traditional proteases. So far, success in the development of proteolytic antibodies has been limited, because the already developed enzymatically active antibodies exhibit good Km but modest Vmax. However, it should be noted that future might bring innovations, and the non-immunoglobulin SBPs appear to be more promising candidates, since they may not share the aforementioned constraints like scaffold stiffness. Efforts continue, with some based on nanobodies: for example, a minimized L-asparaginase based on the nanobody with L-asparaginase amidohydrolase activity (this enzyme has been successfully used to treat acute lymphoblastic leukemia for over 50 years) was designed that successfully targeted the CD19-expressing cells [126].
Non-canonical amino acid residues in the synthetic binding proteins. It is clear that variability of the binding pockets in immunoglobulins or in the alternative SBPs seems to be nearing its natural chemical limits, although in a model system a quasi-specific binding can be achieved even with a quaternary [127, 128] or even a binary code in the complementarity-determining regions (CDR) of antibodies, for example with only two amino acid residues, namely tyrosine and serine like YXS [129]. High tyrosine content (see Fig. 2, this is shown for affibody and monobody but not DARPin) in many of these binding sequences [127] favor high affinity, but it is reasonable to assume that the appearance of oligotyrosine sites may reduce specificity to its target. Additionally, antibodies with non-specific, self-reactive properties tend to be rich in arginine and lysine [128], whereas pH-dependent ones are rich in histidine [130]. It is obvious from general considerations and also has been shown experimentally that 20 amino acid libraries surpass the 4 amino acid libraries [131], accordingly, further increase in diversity should also improve the properties of SBPs. This prompts the question: for how long should we limit ourselves to the traditional genetic code and its modest set of 20 basic amino acids? Recent advances in ribosomal synthesis involving non-canonical residues provide intriguing avenues to explore. Chemical (including enzyme-catalyzed) modification of binding pockets is feasible, but it is usually problematic in terms of desired (close to 100%) yield and specificity. Incorporating non-canonical amino acids in the regions such as CDRs, especially when they are well-tolerated, appears promising. Recent studies have integrated the yeast display screening with non-canonical amino acids to further assess proteins with “chemically extended” functions [124].
CONCLUSION
In the macroscopic world, crafting specific hooks tailored for individual fish species, adapting designs to the unique features of each mouth, is an intricate endeavor. It is even more incredible how much progress has been made in the development of various affinity reagents based on both Nature’s innovations and human ingenuity. It is also important to continue investigation of the well-known proteins such as immunoglobulins as has been demonstrated by the recent discovery of an unusual structure in the CDR of the bovine immunoglobulin [132] – a ‘stalk and knob’ elongation. Existence of such structure in nature could be very helpful for designing the SBPs based on non-immunoglobulin platforms. This paves the way for introducing extended loops, reminiscent of the earlier efforts to develop the so-called loopDARPins [133].
Abbreviations
- CDR:
-
complementarity-determining regions
- OV:
-
oncolytic viruses
- SBP:
-
synthetic binding protein
References
Weidle, U. H., Auer, J., Brinkmann, U., Georges, G., and Tiefenthaler, G. (2013) The emerging role of new protein scaffold-based agents for treatment of cancer, Cancer Genomics Proteomics, 10, 155-168.
Rabia, L. A., Desai, A. A., Jhajj, H. S., and Tessier, P. M. (2018) Understanding and overcoming trade-offs between antibody affinity, specificity, stability and solubility, Biochem. Eng. J., 137, 365-374, https://doi.org/10.1016/j.bej.2018.06.003.
Cunningham, O., Scott, M., Zhou, Z. S., and Finlay, W. J. J. (2021) Polyreactivity and polyspecificity in therapeutic antibody development: risk factors for failure in preclinical and clinical development campaigns, MAbs, 13, 1999195, https://doi.org/10.1080/19420862.2021.1999195.
Van Regenmortel, M. H. V. (2014) Specificity, polyspecificity, and heterospecificity of antibody-antigen recognition, J. Mol. Recognit., 27, 627-639, https://doi.org/10.1002/jmr.2394.
Pestov, N. B., Gusakova, T. V., Kostina, M. B., and Shakhparonov, M. I. (1996) Phage mimotopes of monoclonal antibodies against plasma membrane Ca2+-ATPase, Bioorg. Khim., 22, 664-670.
Rabia, L. A., Zhang, Y., Ludwig, S. D., Julian, M. C., and Tessier, P. M. (2018) Net charge of antibody complementarity-determining regions is a key predictor of specificity, Protein Eng. Des. Sel., 31, 409-418, https://doi.org/10.1093/protein/gzz002.
Gebauer, M., and Skerra, A. (2020) Engineered protein scaffolds as next-generation therapeutics, Annu. Rev. Pharmacol. Toxicol., 60, 391-415, https://doi.org/10.1146/annurev-pharmtox-010818-021118.
Aguilar, G., Vigano, M. A., Affolter, M., and Matsuda, S. (2019) Reflections on the use of protein binders to study protein function in developmental biology, Wiley Interdiscip. Rev. Dev. Biol., 8, e356, https://doi.org/10.1002/wdev.356.
Jenkins, T. P., Fryer, T., Dehli, R. I., Jürgensen, J. A., Fuglsang-Madsen, A., et al. (2019) Toxin neutralization using alternative binding proteins, Toxins (Basel), 11, E53, https://doi.org/10.3390/toxins11010053.
Olaleye, O., Govorukhina, N., van de Merbel, N. C., and Bischoff, R. (2021) Non-antibody-based binders for the enrichment of proteins for analysis by mass spectrometry, Biomolecules, 11, 1791, https://doi.org/10.3390/biom11121791.
Bondos, S. E., Dunker, A. K., and Uversky, V. N. (2022) Intrinsically disordered proteins play diverse roles in cell signaling, Cell Commun. Signal., 20, 20, https://doi.org/10.1186/s12964-022-00821-7.
Karlsson, O. A., Ramirez, J., Öberg, D., Malmqvist, T., Engström, Å., et al. (2015) Design of a PDZbody, a bivalent binder of the E6 protein from human papillomavirus, Sci. Rep., 5, 9382, https://doi.org/10.1038/srep09382.
Sha, F., Salzman, G., Gupta, A., and Koide, S. (2017) Monobodies and other synthetic binding proteins for expanding protein science, Protein Sci., 26, 910-924, https://doi.org/10.1002/pro.3148.
Hantschel, O., Biancalana, M., and Koide, S. (2020) Monobodies as enabling tools for structural and mechanistic biology, Curr. Opin. Struct. Biol., 60, 167-174, https://doi.org/10.1016/j.sbi.2020.01.015.
McConnell, S. J., and Hoess, R. H. (1995) Tendamistat as a scaffold for conformationally constrained phage peptide libraries, J. Mol. Biol., 250, 460-470, https://doi.org/10.1006/jmbi.1995.0390.
Ciesiołkiewicz, A., Lizandra Perez, J., and Berlicki, Ł. (2022) Miniproteins in medicinal chemistry, Bioorg. Med. Chem. Lett., 71, 128806, https://doi.org/10.1016/j.bmcl.2022.128806.
Kolmar, H. (2008) Alternative binding proteins: biological activity and therapeutic potential of cystine-knot miniproteins, FEBS J., 275, 2684-2690, https://doi.org/10.1111/j.1742-4658.2008.06440.x.
Silverman, J., Liu, Q., Bakker, A., To, W., Duguay, A., et al. (2005) Multivalent avimer proteins evolved by exon shuffling of a family of human receptor domains, Nat. Biotechnol., 23, 1556-1561, https://doi.org/10.1038/nbt1166.
Tiede, C., Tang, A. A. S., Deacon, S. E., Mandal, U., Nettleship, J. E., et al. (2014) Adhiron: a stable and versatile peptide display scaffold for molecular recognition applications, Protein Eng. Des. Sel., 27, 145-155, https://doi.org/10.1093/protein/gzu007.
Tiede, C., Bedford, R., Heseltine, S. J., Smith, G., Wijetunga, I., et al. (2017) Affimer proteins are versatile and renewable affinity reagents, Elife, 6, e24903, https://doi.org/10.7554/eLife.24903.
Shamsuddin, S. H., Jayne, D. G., Tomlinson, D. C., McPherson, M. J., and Millner, P. A. (2021) Selection and characterisation of Affimers specific for CEA recognition, Sci. Rep., 11, 744, https://doi.org/10.1038/s41598-020-80354-6.
Wicke, N., Bedford, M. R., and Howarth, M. (2021) Gastrobodies are engineered antibody mimetics resilient to pepsin and hydrochloric acid, Commun. Biol., 4, 960, https://doi.org/10.1038/s42003-021-02487-2.
Zoller, F., Markert, A., Barthe, P., Zhao, W., Weichert, W., et al. (2012) Combination of phage display and molecular grafting generates highly specific tumor-targeting miniproteins, Angew. Chem. Int. Ed. Engl., 51, 13136-13139, https://doi.org/10.1002/anie.201203857.
Cohen, I., Kayode, O., Hockla, A., Sankaran, B., Radisky, D. C., et al. (2016) Combinatorial protein engineering of proteolytically resistant mesotrypsin inhibitors as candidates for cancer therapy, Biochem. J., 473, 1329-1341, https://doi.org/10.1042/BJ20151410.
Sananes, A., Cohen, I., Shahar, A., Hockla, A., De Vita, E., et al. (2018) A potent, proteolysis-resistant inhibitor of kallikrein-related peptidase 6 (KLK6) for cancer therapy, developed by combinatorial engineering, J. Biol. Chem., 293, 12663-12680, https://doi.org/10.1074/jbc.RA117.000871.
Nishimiya, D., Kawaguchi, Y., Kodama, S., Nasu, H., Yano, H., et al. (2019) A protein scaffold, engineered SPINK2, for generation of inhibitors with high affinity and specificity against target proteases, Sci. Rep., 9, 11436, https://doi.org/10.1038/s41598-019-47615-5.
Jia, Z., Liu, Y., Ji, X., Zheng, Y., Li, Z., et al. (2021) DAKS1, a Kunitz scaffold peptide from the venom gland of Deinagkistrodon acutus prevents carotid-artery and middle-cerebral-artery thrombosis via targeting factor Xia, Pharmaceuticals (Basel), 14, 966, https://doi.org/10.3390/ph14100966.
Lee, S.-C., Park, K., Han, J., Lee, J., Kim, H. J., et al. (2012) Design of a binding scaffold based on variable lymphocyte receptors of jawless vertebrates by module engineering, Proc. Natl. Acad. Sci. USA, 109, 3299-3304, https://doi.org/10.1073/pnas.1113193109.
Hwang, D.-E., Ryou, J.-H., Oh, J. R., Han, J. W., Park, T. K., and Kim, H.-S. (2016) Anti-human VEGF repebody effectively suppresses choroidal neovascularization and vascular leakage, PLoS One, 11, e0152522, https://doi.org/10.1371/journal.pone.0152522.
Liu, H., Huang, H., Voss, C., Kaneko, T., Qin, W. T., et al. (2019) surface loops in a single SH2 domain are capable of encoding the spectrum of specificity of the SH2 family, Mol. Cell. Proteomics, 18, 372-382, https://doi.org/10.1074/mcp.RA118.001123.
Schlatter, D., Brack, S., Banner, D. W., Batey, S., Benz, J., et al. (2012) Generation, characterization and structural data of chymase binding proteins based on the human Fyn kinase SH3 domain, MAbs, 4, 497-508, https://doi.org/10.4161/mabs.20452.
Garlich, J., Cinier, M., Chevrel, A., Perrocheau, A., Eyerman, D. J., et al. (2022) Discovery of APL-1030, a novel, high-affinity nanofitin inhibitor of C3-mediated complement activation, Biomolecules, 12, 432, https://doi.org/10.3390/biom12030432.
Gocha, T., Rao, B. M., and DasGupta, R. (2017) Identification and characterization of a novel Sso7d scaffold-based binder against Notch1, Sci. Rep., 7, 12021, https://doi.org/10.1038/s41598-017-12246-1.
Kalichuk, V., Renodon-Cornière, A., Béhar, G., Carrión, F., Obal, G., Maillasson, M., et al. (2018) A novel, smaller scaffold for Affitins: showcase with binders specific for EpCAM, Biotechnol. Bioeng., 115, 290-299, https://doi.org/10.1002/bit.26463.
Steemson, J. D., Baake, M., Rakonjac, J., Arcus, V. L., and Liddament, M. T. (2014) Tracking molecular recognition at the atomic level with a new protein scaffold based on the OB-fold, PLoS One, 9, e86050, https://doi.org/10.1371/journal.pone.0086050.
Napolitano, E. W., Villar, H. O., Kauvar, L. M., Higgins, D. L., Roberts, D., et al. (1996) Glubodies: randomized libraries of glutathione transferase enzymes, Chem. Biol., 3, 359-367, https://doi.org/10.1016/S1074-5521(96)90119-2.
Lorey, S., Fiedler, E., Kunert, A., Nerkamp, J., Lange, C., et al. (2014) Novel ubiquitin-derived high affinity binding proteins with tumor targeting properties, J. Biol. Chem., 289, 8493-8507, https://doi.org/10.1074/jbc.M113.519884.
Pham, P. N., Huličiak, M., Biedermannová, L., Černý, J., Charnavets, T., et al. (2021) Protein Binder (ProBi) as a new class of structurally robust non-antibody protein scaffold for directed evolution, Viruses, 13, 190, https://doi.org/10.3390/v13020190.
Pan, X., Thompson, M. C., Zhang, Y., Liu, L., Fraser, J. S., et al. (2020) Expanding the space of protein geometries by computational design of de novo fold families, Science, 369, 1132-1136, https://doi.org/10.1126/science.abc0881.
Desmet, J., Verstraete, K., Bloch, Y., Lorent, E., Wen, Y., et al. (2014) Structural basis of IL-23 antagonism by an Alphabody protein scaffold, Nat. Commun., 5, 5237, https://doi.org/10.1038/ncomms6237.
Koide, A., Bailey, C. W., Huang, X., and Koide, S. (1998) The fibronectin type III domain as a scaffold for novel binding proteins, J. Mol. Biol., 284, 1141-1151, https://doi.org/10.1006/jmbi.1998.2238.
Lipovsek, D. (2011) Adnectins: engineered target-binding protein therapeutics, Protein Eng. Des. Sel., 24, 3-9, https://doi.org/10.1093/protein/gzq097.
Sullivan, M. A., Brooks, L. R., Weidenborner, P., Domm, W., Mattiacio, J., et al. (2013) Anti-idiotypic monobodies derived from a fibronectin scaffold, Biochemistry, 52, 1802-1813, https://doi.org/10.1021/bi3016668.
Chandler, P. G., and Buckle, A. M. (2020) Development and differentiation in monobodies based on the fibronectin type 3 domain, Cells, 9, E610, https://doi.org/10.3390/cells9030610.
Akkapeddi, P., Teng, K. W., and Koide, S. (2021) Monobodies as tool biologics for accelerating target validation and druggable site discovery, RSC Med. Chem., 12, 1839-1853, https://doi.org/10.1039/D1MD00188D.
Batori, V., Koide, A., and Koide, S. (2002) Exploring the potential of the monobody scaffold: effects of loop elongation on the stability of a fibronectin type III domain, Protein Eng., 15, 1015-1020, https://doi.org/10.1093/protein/15.12.1015.
Kükenshöner, T., Schmit, N. E., Bouda, E., Sha, F., Pojer, F., et al. (2017) Selective Targeting of SH2 domain-phosphotyrosine interactions of Src family tyrosine kinases with monobodies, J. Mol. Biol., 429, 1364-1380, https://doi.org/10.1016/j.jmb.2017.03.023.
Gross, G. G., Junge, J. A., Mora, R. J., Kwon, H.-B., Olson, C. A., et al. (2013) Recombinant probes for visualizing endogenous synaptic proteins in living neurons, Neuron, 78, 971-985, https://doi.org/10.1016/j.neuron.2013.04.017.
Porebski, B. T., Conroy, P. J., Drinkwater, N., Schofield, P., Vazquez-Lombardi, R., et al. (2016) Circumventing the stability-function trade-off in an engineered FN3 domain, Protein Eng. Des. Sel., 29, 541-550, https://doi.org/10.1093/protein/gzw046.
Porebski, B. T., Nickson, A. A., Hoke, D. E., Hunter, M. R., Zhu, L., et al. (2015) Structural and dynamic properties that govern the stability of an engineered fibronectin type III domain, Protein Eng. Des. Sel., 28, 67-78, https://doi.org/10.1093/protein/gzv002.
Chandler, P. G., Tan, L. L., Porebski, B. T., Green, J. S., Riley, B. T., et al. (2021) Mutational and biophysical robustness in a prestabilized monobody, J. Biol. Chem., 296, 100447, https://doi.org/10.1016/j.jbc.2021.100447.
Deuschle, F.-C., Ilyukhina, E., and Skerra, A. (2021) Anticalin® proteins: from bench to bedside, Expert Opin. Biol. Ther., 21, 509-518, https://doi.org/10.1080/14712598.2021.1839046.
Achatz, S., Jarasch, A., and Skerra, A. (2022) Structural plasticity in the loop region of engineered lipocalins with novel ligand specificities, so-called Anticalins, J. Struct. Biol., 6, 100054, https://doi.org/10.1016/j.yjsbx.2021.100054.
Gebauer, M., Schiefner, A., Matschiner, G., and Skerra, A. (2013) Combinatorial design of an anticalin directed against the extra-domain B for the specific targeting of oncofetal fibronectin, J. Mol. Biol., 425, 780-802, https://doi.org/10.1016/j.jmb.2012.12.004.
Chi, Y., Remsik, J., Kiseliovas, V., Derderian, C., Sener, U., et al. (2020) Cancer cells deploy lipocalin-2 to collect limiting iron in leptomeningeal metastasis, Science, 369, 276-282, https://doi.org/10.1126/science.aaz2193.
Yang, J., Bielenberg, D. R., Rodig, S. J., Doiron, R., Clifton, M. C., et al. (2009) Lipocalin 2 promotes breast cancer progression, Proc. Natl. Acad. Sci. USA, 106, 3913-3918, https://doi.org/10.1073/pnas.0810617106.
Chakraborty, S., Kaur, S., Guha, S., and Batra, S. K. (2012) The multifaceted roles of neutrophil gelatinase associated lipocalin (NGAL) in inflammation and cancer, Biochim. Biophys. Acta, 1826, 129-169, https://doi.org/10.1016/j.bbcan.2012.03.008.
Crescenzi, E., Leonardi, A., and Pacifico, F. (2021) NGAL as a potential target in tumor microenvironment, Int. J. Mol. Sci., 22, 12333, https://doi.org/10.3390/ijms222212333.
Xu, B., Zheng, W., Jin, D., Wang, D., Liu, X., and Qin, X. (2012) Treatment of pancreatic cancer using an oncolytic virus harboring the lipocalin-2 gene, Cancer, 118, 5217-5226, https://doi.org/10.1002/cncr.27535.
Xu, B., Zheng, W.-Y., Feng, J.-F., Huang, X.-Y., and Ge, H. (2013) One potential oncolytic adenovirus expressing Lipocalin-2 for colorectal cancer therapy, Cancer Biother. Radiopharm., 28, 415-422, https://doi.org/10.1089/cbr.2012.1352.
Nishimura, S., Yamamoto, Y., Sugimoto, A., Kushiyama, S., Togano, S., et al. (2022) Lipocalin-2 negatively regulates epithelial-mesenchymal transition through matrix metalloprotease-2 downregulation in gastric cancer, Gastric Cancer, 25, 850-861, https://doi.org/10.1007/s10120-022-01305-w.
Pinyopornpanish, K., Phrommintikul, A., Angkurawaranon, C., Kumfu, S., Angkurawaranon, S., et al. (2022) Circulating Lipocalin-2 level is positively associated with cognitive impairment in patients with metabolic syndrome, Sci. Rep., 12, 4635, https://doi.org/10.1038/s41598-022-08286-x.
Friedman, M., Lindström, S., Ekerljung, L., Andersson-Svahn, H., Carlsson, J., et al. (2009) Engineering and characterization of a bispecific HER2 x EGFR-binding affibody molecule, Biotechnol. Appl. Biochem., 54, 121-131, https://doi.org/10.1042/BA20090096.
Gorman, K., McGinnis, J., and Kay, B. (2018) Generating FN3-based affinity reagents through phage display, Curr. Protoc. Chem. Biol., 10, e39, https://doi.org/10.1002/cpch.39.
Davydova, E. K. (2022) Protein engineering: Advances in phage display for basic science and medical research, Biochemistry (Moscow), 87, S146-S110, https://doi.org/10.1134/S0006297922140127.
Zhang, Y., Thangam, R., You, S.-H., Sultonova, R. D., Venu, A., et al. (2021) Engineering calreticulin-targeting monobodies to detect immunogenic cell death in cancer chemotherapy, Cancers (Basel), 13, 2801, https://doi.org/10.3390/cancers13112801.
Cetin, M., Evenson, W. E., Gross, G. G., Jalali-Yazdi, F., Krieger, D., et al. (2017) RasIns: genetically encoded intrabodies of activated Ras proteins, J. Mol. Biol., 429, 562-573, https://doi.org/10.1016/j.jmb.2016.11.008.
Olson, C. A., and Roberts, R. W. (2007) Design, expression, and stability of a diverse protein library based on the human fibronectin type III domain, Protein Sci., 16, 476-484, https://doi.org/10.1110/ps.062498407.
Khramtsov, Y. V., Ulasov, A. V., Lupanova, T. N., Georgiev, G. P., and Sobolev, A. S. (2022) Delivery of antibody-like molecules, monobodies, capable of binding with SARS-CoV-2 virus nucleocapsid protein, into target cells, Dokl. Biochem. Biophys., 506, 220-222, https://doi.org/10.1134/S1607672922050088.
Lipovsek, D., Lippow, S. M., Hackel, B. J., Gregson, M. W., Cheng, P., et al. (2007) Evolution of an interloop disulfide bond in high-affinity antibody mimics based on fibronectin type III domain and selected by yeast surface display: molecular convergence with single-domain camelid and shark antibodies, J. Mol. Biol., 368, 1024-1041, https://doi.org/10.1016/j.jmb.2007.02.029.
Riihimäki, T. A., Hiltunen, S., Rangl, M., Nordlund, H. R., Määttä, J. A. E., et al. (2011) Modification of the loops in the ligand-binding site turns avidin into a steroid-binding protein, BMC Biotechnol., 11, 64, https://doi.org/10.1186/1472-6750-11-64.
Hytönen, V. P. (2020) (Strept)avidin as a template for ligands other than biotin: An overview, Methods Enzymol., 633, 21-28, https://doi.org/10.1016/bs.mie.2019.10.029.
Kiss, C., Fisher, H., Pesavento, E., Dai, M., Valero, R., et al. (2006) Antibody binding loop insertions as diversity elements, Nucleic Acids Res., 34, e132, https://doi.org/10.1093/nar/gkl681.
Kadonosono, T., Yabe, E., Furuta, T., Yamano, A., Tsubaki, T., et al. (2014) A fluorescent protein scaffold for presenting structurally constrained peptides provides an effective screening system to identify high affinity target-binding peptides, PLoS One, 9, e103397, https://doi.org/10.1371/journal.pone.0103397.
Chee, S. M. Q., Wongsantichon, J., Yi, L. S., Sana, B., Frosi, Y., et al. (2021) Functional display of bioactive peptides on the vGFP scaffold, Sci. Rep., 11, 10127, https://doi.org/10.1038/s41598-021-89421-y.
Rinne, S. S., Yin, W., Borras, A. M., Abouzayed, A., Leitao, C. D., et al. (2022) Targeting tumor cells overexpressing the human epidermal growth factor receptor 3 with potent drug conjugates based on affibody molecules, Biomedicines, 10, 1293, https://doi.org/10.3390/biomedicines10061293.
Sokolova, E., Kutova, O., Grishina, A., Pospelov, A., Guryev, E., et al. (2019) Penetration efficiency of antitumor agents in ovarian cancer spheroids: the case of recombinant targeted toxin DARPin-LoPE and the chemotherapy drug, doxorubicin, Pharmaceutics, 11, 219, https://doi.org/10.3390/pharmaceutics11050219.
Sachdev, S., Cabalteja, C. C., and Cheloha, R. W. (2021) Strategies for targeting cell surface proteins using multivalent conjugates and chemical biology, Methods Cell Biol., 166, 205-222, https://doi.org/10.1016/bs.mcb.2021.06.004.
Shipunova, V. O., and Deyev, S. M. (2022) Artificial scaffold polypeptides as an efficient tool for the targeted delivery of nanostructures in vitro and in vivo, Acta Naturae, 14, 54-72, https://doi.org/10.32607/actanaturae.11545.
Deyev, S., Proshkina, G., Baryshnikova, O., Ryabova, A., Avishai, G., et al. (2018) Selective staining and eradication of cancer cells by protein-carrying DARPin-functionalized liposomes, Eur. J. Pharm. Biopharm., 130, 296-305, https://doi.org/10.1016/j.ejpb.2018.06.026.
Nazari, M., Emamzadeh, R., Jahanpanah, M., Yazdani, E., and Radmanesh, R. (2022) A recombinant affitoxin derived from a HER3 affibody and diphteria-toxin has potent and selective antitumor activity, Int. J. Biol. Macromol., 219, 1122-1134, https://doi.org/10.1016/j.ijbiomac.2022.08.150.
Lipovšek, D., Carvajal, I., Allentoff, A. J., Barros, A., Brailsford, J., et al. (2018) Adnectin-drug conjugates for Glypican-3-specific delivery of a cytotoxic payload to tumors, Protein Eng. Des. Sel., 31, 159-171, https://doi.org/10.1093/protein/gzy013.
Karsten, L., Janson, N., Le Joncour, V., Alam, S., Müller, B., et al. (2022) Bivalent EGFR-targeting DARPin-MMAE conjugates, Int. J. Mol. Sci., 23, 2468, https://doi.org/10.3390/ijms23052468.
Sharma, R., Suman, S. K., and Mukherjee, A. (2022) Antibody-based radiopharmaceuticals as theranostic agents: an overview, Curr. Med. Chem., 29, 5979, https://doi.org/10.2174/0929867329666220607160559.
Luo, R., Liu, H., and Cheng, Z. (2022) Protein scaffolds: antibody alternatives for cancer diagnosis and therapy, RSC Chem. Biol., 3, 830-847, https://doi.org/10.1039/D2CB00094F.
Klont, F., Hadderingh, M., Horvatovich, P., Ten Hacken, N. H. T., and Bischoff, R. (2018) Affimers as an alternative to antibodies in an affinity LC-MS assay for quantification of the soluble receptor of advanced glycation end-products (sRAGE) in human serum, J. Proteome Res., 17, 2892-2899, https://doi.org/10.1021/acs.jproteome.8b00414.
Mayoral-Peña, K., González Peña, O. I., Orrantia Clark, A. M., Flores-Vallejo, R. D. C., Oza, G., et al. (2022) Biorecognition engineering technologies for cancer diagnosis: a systematic literature review of non-conventional and plausible sensor development methods, Cancers (Basel), 14, 1867, https://doi.org/10.3390/cancers14081867.
Shilova, O. N., and Deyev, S. M. (2019) DARPins: promising scaffolds for theranostics, Acta Naturae, 11, 42-53, https://doi.org/10.32607/20758251-2019-11-4-42-53.
Shipunova, V. O., Kolesnikova, O. A., Kotelnikova, P. A., Soloviev, V. D., Popov, A. A., et al. (2021) Comparative evaluation of engineered polypeptide scaffolds in HER2-targeting magnetic nanocarrier delivery, ACS Omega, 6, 16000, https://doi.org/10.1021/acsomega.1c01811.
Mamluk, R., Carvajal, I. M., Morse, B. A., Wong, H., Abramowitz, J., et al. (2010) Anti-tumor effect of CT-322 as an adnectin inhibitor of vascular endothelial growth factor receptor-2, MAbs, 2, 199-208, https://doi.org/10.4161/mabs.2.2.11304.
Tolmachev, V., and Orlova, A. (2020) Affibody molecules as targeting vectors for PET imaging, Cancers (Basel), 12, E651, https://doi.org/10.3390/cancers12030651.
Deyev, S., Vorobyeva, A., Schulga, A., Proshkina, G., Güler, R., et al. (2019) Comparative evaluation of two DARPin variants: effect of affinity, size, and label on tumor targeting properties, Mol. Pharm., 16, 995-1008, https://doi.org/10.1021/acs.molpharmaceut.8b00922.
Wang, Y., Ballou, B., Schmidt, B. F., Andreko, S., St Croix, C. M., et al. (2017) Affibody-targeted fluorogen activating protein for in vivo tumor imaging, Chem. Commun. (Camb.), 53, 2001, https://doi.org/10.1039/C6CC09137G.
Yun, M., Kim, D.-Y., Lee, J.-J., Kim, H.-S., Kim, H.-S., et al. (2017) A high-affinity repebody for molecular imaging of EGFR-expressing malignant tumors, Theranostics, 7, 2620-2633, https://doi.org/10.7150/thno.18096.
Mączyńska, J., Raes, F., Da Pieve, C., Turnock, S., Boult, J. K. R., et al. (2022) Triggering anti-GBM immune response with EGFR-mediated photoimmunotherapy, BMC Med., 20, 16, https://doi.org/10.1186/s12916-021-02213-z.
Lui, B. G., Salomon, N., Wüstehube-Lausch, J., Daneschdar, M., Schmoldt, H.-U., et al. (2020) Targeting the tumor vasculature with engineered cystine-knot miniproteins, Nat. Commun., 11, 295, https://doi.org/10.1038/s41467-019-13948-y.
Balalaeva, I. V., Krylova, L. V., Karpova, M. A., Shulga, A. A., Konovalova, E. V., Guryev, E. L., and Deyev, S. M. (2022) Synergistic effect of the combined action of targeted and photodynamic therapy on HER2-positive breast cancer, Dokl. Biochem. Biophys., 507, 330-333, https://doi.org/10.1134/S1607672922340038.
Proshkina, G. M., Shramova, E. I., Shilova, M. V., Zelepukin, I. V., Shipunova, V. O., et al. (2021) DARPin_9-29-targeted gold nanorods selectively suppress HER2-positive tumor growth in mice, Cancers (Basel), 13, 5235, https://doi.org/10.3390/cancers13205235.
Shramova, E., Proshkina, G., Shipunova, V., Ryabova, A., Kamyshinsky, R., et al. (2020) Dual targeting of cancer cells with DARPin-based toxins for overcoming tumor escape, Cancers (Basel), 12, E3014, https://doi.org/10.3390/cancers12103014.
Eijkenboom, L., Palacio-Castañeda, V., Groenman, F., Braat, D., Beerendonk, C., et al. (2021) Assessing the use of tumor-specific DARPin-toxin fusion proteins for ex vivo purging of cancer metastases from human ovarian cortex before autotransplantation, F. S. Sci., 2, 330-344, https://doi.org/10.1016/j.xfss.2021.09.004.
Xu, T., Schulga, A., Konovalova, E., Rinne, S. S., Zhang, H., et al. (2023) Feasibility of xo-targeting HER3 and EpCAM using seribantumab and DARPin-toxin fusion in a pancreatic cancer xenograft model, Int. J. Mol. Sci., 24, 2838, https://doi.org/10.3390/ijms24032838.
Hanauer, J. R. H., Koch, V., Lauer, U. M., and Mühlebach, M. D. (2019) High-affinity DARPin allows targeting of MeV to glioblastoma multiforme in combination with protease targeting without loss of potency, Mol. Ther. Oncolytics, 15, 186-200, https://doi.org/10.1016/j.omto.2019.10.004.
Lal, S., and Raffel, C. (2017) Using cystine knot proteins as a novel approach to retarget oncolytic measles virus, Mol. Ther. Oncolytics, 7, 57-66, https://doi.org/10.1016/j.omto.2017.09.005.
Strecker, M. I., Wlotzka, K., Strassheimer, F., Roller, B., Ludmirski, G., et al. (2022) AAV-mediated gene transfer of a checkpoint inhibitor in combination with HER2-targeted CAR-NK cells as experimental therapy for glioblastoma, Oncoimmunology, 11, 2127508, https://doi.org/10.1080/2162402X.2022.2127508.
Zajc, C. U., Salzer, B., Taft, J. M., Reddy, S. T., Lehner, M., and Traxlmayr, M. W. (2021) Driving CARs with alternative navigation tools – the potential of engineered binding scaffolds, FEBS J., 288, 2103-2118, https://doi.org/10.1111/febs.15523.
Stepanov, A. V., Kalinin, R. S., Shipunova, V. O., Zhang, D., Xie, J., et al. (2022) Switchable targeting of solid tumors by BsCAR T cells, Proc. Natl. Acad. Sci. USA, 119, e2210562119, https://doi.org/10.1073/pnas.2210562119.
Parfenyev, S., Singh, A., Fedorova, O., Daks, A., Kulshreshtha, R., and Barlev, N. A. (2021) Interplay between p53 and non-coding RNAs in the regulation of EMT in breast cancer, Cell Death Dis., 12, 17, https://doi.org/10.1038/s41419-020-03327-7.
Lezina, L., Aksenova, V., Fedorova, O., Malikova, D., Shuvalov, O., Antonov, A. V., et al. (2015) KMT Set7/9 affects genotoxic stress response via the Mdm2 axis, Oncotarget, 6, 25843-25855, https://doi.org/10.18632/oncotarget.4584.
Davidovich, P., Aksenova, V., Petrova, V., Tentler, D., Orlova, D., et al. (2015) Discovery of novel isatin-based p53 inducers, ACS Med. Chem. Lett., 6, 856-860, https://doi.org/10.1021/acsmedchemlett.5b00011.
Bulatov, E., Sayarova, R., Mingaleeva, R., Miftakhova, R., Gomzikova, M., et al. (2018) Isatin-Schiff base-copper (II) complex induces cell death in p53-positive tumors, Cell Death Discov., 4, 103, https://doi.org/10.1038/s41420-018-0120-z.
Lau, S.-Y., Siau, J. W., Sobota, R. M., Wang, C.-I., Zhong, P., et al. (2018) Synthetic 10FN3-based mono- and bivalent inhibitors of MDM2/X function, Protein Eng. Des. Sel., 31, 301-312, https://doi.org/10.1093/protein/gzy018.
Spencer-Smith, R., Koide, A., Zhou, Y., Eguchi, R. R., Sha, F., et al. (2017) Inhibition of RAS function through targeting an allosteric regulatory site, Nat. Chem. Biol., 13, 62-68, https://doi.org/10.1038/nchembio.2231.
Teng, K. W., Tsai, S. T., Hattori, T., Fedele, C., Koide, A., et al. (2021) Selective and noncovalent targeting of RAS mutants for inhibition and degradation, Nat. Commun., 12, 2656, https://doi.org/10.1038/s41467-021-22969-5.
Khan, I., Koide, A., Zuberi, M., Ketavarapu, G., Denbaum, E., et al. (2022) Identification of the nucleotide-free state as a therapeutic vulnerability for inhibition of selected oncogenic RAS mutants, Cell Rep., 38, 110322, https://doi.org/10.1016/j.celrep.2022.110322.
Wallon, L., Khan, I., Teng, K. W., Koide, A., Zuberi, M., et al. (2022) Inhibition of RAS-driven signaling and tumorigenesis with a pan-RAS monobody targeting the Switch I/II pocket, Proc. Natl. Acad. Sci. USA, 119, e2204481119, https://doi.org/10.1073/pnas.2204481119.
Guillard, S., Kolasinska-Zwierz, P., Debreczeni, J., Breed, J., Zhang, J., et al. (2017) Structural and functional characterization of a DARPin which inhibits Ras nucleotide exchange, Nat. Commun., 8, 16111, https://doi.org/10.1038/ncomms16111.
Bery, N., Legg, S., Debreczeni, J., Breed, J., Embrey, K., et al. (2019) KRAS-specific inhibition using a DARPin binding to a site in the allosteric lobe, Nat. Commun., 10, 2607, https://doi.org/10.1038/s41467-019-10419-2.
McGee, J. H., Shim, S. Y., Lee, S.-J., Swanson, P. K., Jiang, S. Y., et al. (2018) Exceptionally high-affinity Ras binders that remodel its effector domain, J. Biol. Chem., 293, 3265-3280, https://doi.org/10.1074/jbc.M117.816348.
Jung, Y. H., Choi, Y., Seo, H.-D., Seo, M.-H., and Kim, H.-S. (2023) A conformation-selective protein binder for a KRAS mutant inhibits the interaction between RAS and RAF, Biochem. Biophys. Res. Commun., 645, 110-117, https://doi.org/10.1016/j.bbrc.2023.01.019.
Schmit, N. E., Neopane, K., and Hantschel, O. (2019) Targeted protein degradation through cytosolic delivery of monobody binders using bacterial toxins, ACS Chem. Biol., 14, 916-924, https://doi.org/10.1021/acschembio.9b00113.
Röth, S., Macartney, T. J., Konopacka, A., Chan, K.-H., Zhou, H., et al. (2020) Targeting endogenous K-RAS for degradation through the affinity-directed protein missile system, Cell. Chem. Biol., 27, 1151-1163.e6, https://doi.org/10.1016/j.chembiol.2020.06.012.
Kim, J. W., Cochran, F. V., and Cochran, J. R. (2015) A chemically cross-linked knottin dimer binds integrins with picomolar affinity and inhibits tumor cell migration and proliferation, J. Am. Chem. Soc., 137, 6-9, https://doi.org/10.1021/ja508416e.
Gad, S., and Ayakar, S. (2021) Protein scaffolds: A tool for multi-enzyme assembly, Biotechnol. Rep. (Amst.), 32, e00670, https://doi.org/10.1016/j.btre.2021.e00670.
Islam, M., Kehoe, H. P., Lissoos, J. B., Huang, M., Ghadban, C. E., et al. (2021) Chemical diversification of simple synthetic antibodies, ACS Chem. Biol., 16, 344-359, https://doi.org/10.1021/acschembio.0c00865.
Rabe von Pappenheim, F., Wensien, M., Ye, J., Uranga, J., Irisarri, I., et al. (2022) Widespread occurrence of covalent lysine-cysteine redox switches in proteins, Nat. Chem. Biol., 18, 368-375, https://doi.org/10.1038/s41589-021-00966-5.
Maggi, M., Pessino, G., Guardamagna, I., Lonati, L., Pulimeno, C., and Scotti, C. (2021) A targeted catalytic nanobody (T-CAN) with asparaginolytic activity, Cancers (Basel), 13, 5637, https://doi.org/10.3390/cancers13225637.
Fellouse, F. A., Barthelemy, P. A., Kelley, R. F., and Sidhu, S. S. (2006) Tyrosine plays a dominant functional role in the paratope of a synthetic antibody derived from a four amino acid code, J. Mol. Biol., 357, 100-114, https://doi.org/10.1016/j.jmb.2005.11.092.
Birtalan, S., Zhang, Y., Fellouse, F. A., Shao, L., Schaefer, G., and Sidhu, S. S. (2008) The intrinsic contributions of tyrosine, serine, glycine and arginine to the affinity and specificity of antibodies, J. Mol. Biol., 377, 1518-1528, https://doi.org/10.1016/j.jmb.2008.01.093.
Gilbreth, R. N., Esaki, K., Koide, A., Sidhu, S. S., and Koide, S. (2008) A dominant conformational role for amino acid diversity in minimalist protein-protein interfaces, J. Mol. Biol., 381, 407-418, https://doi.org/10.1016/j.jmb.2008.06.014.
Bonvin, P., Venet, S., Fontaine, G., Ravn, U., Gueneau, F., Kosco-Vilbois, M., et al. (2015) De novo isolation of antibodies with pH-dependent binding properties, MAbs, 7, 294-302, https://doi.org/10.1080/19420862.2015.1006993.
Hackel, B. J., and Wittrup, K. D. (2010) The full amino acid repertoire is superior to serine/tyrosine for selection of high affinity immunoglobulin G binders from the fibronectin scaffold, Protein Eng. Des. Sel., 23, 211-219, https://doi.org/10.1093/protein/gzp083.
Svilenov, H. L., Sacherl, J., Protzer, U., Zacharias, M., and Buchner, J. (2021) Mechanistic principles of an ultra-long bovine CDR reveal strategies for antibody design, Nat. Commun., 12, 6737, https://doi.org/10.1038/s41467-021-27103-z.
Schilling, J., Schöppe, J., and Plückthun, A. (2014) From DARPins to LoopDARPins: novel LoopDARPin design allows the selection of low picomolar binders in a single round of ribosome display, J. Mol. Biol., 426, 691-721, https://doi.org/10.1016/j.jmb.2013.10.026.
Acknowledgments
The authors are grateful to Y. M. Rozenberg for thoughtful reading of the draft manuscript.
Funding
This work was financed by the Ministry of Science and Higher Education of the Russian Federation within the framework of state support for the creation and development of World-Class Research Centers ‘Digital Biodesign and Personalized Healthcare’ (no. 75-15-2022-305).
Author information
Authors and Affiliations
Contributions
N.A.B. – concept of the review and editing of the text; T.I.D., N.B.P., and T.V.K. – writing the text and creating illustrations.
Corresponding author
Ethics declarations
The authors declare no conflicts of interest in financial or any other sphere. This article does not contain description of studies with the involvement of humans or animal subjects performed by any of the authors.
Electronic supplementary material
Rights and permissions
Open access. This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution, and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit https://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
David, T.I., Pestov, N.B., Korneenko, T.V. et al. Non-Immunoglobulin Synthetic Binding Proteins for Oncology. Biochemistry Moscow 88, 1232–1247 (2023). https://doi.org/10.1134/S0006297923090043
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0006297923090043