Abstract
In the structure of photosynthetic reaction center (RC) of the purple bacterium Cereibacter sphaeroides the highly conserved amino acid residue Ile-M206 is located near the bacteriochlorophyll dimer P, which is the primary electron donor, and the monomeric bacteriochlorophyll BA, which is the nearest electron acceptor. Since Ile-M206 is close to the C2-acetyl group of bacteriochlorophyll PB, the hydroxyl group of Tyr-M210, and to the C9-keto group of bacteriochlorophyll BA, as well as to the water molecule near the latter group, this site can be used for introducing mutations in order to study mechanisms of primary photochemical processes in the RC. Previously it was shown that the Ile→Glu substitution at the M204 position (analog of M206 in the RC of C. sphaeroides) in the RC of the closely related purple non-sulfur bacterium Rhodobacter capsulatus significantly affected kinetics of the P+HA– state formation, whereas the M204 Ile→Gln substitution led to the loss of BChl BA molecule from the complex structure. In the present work, it is shown that the single I(M206)Q or double I(M206)Q + F(M208)A amino acid substitutions in the RC of C. sphaeroides do not change the pigment composition and do not markedly influence redox potential of the primary electron donor. However, substitution of Ile M206 by Gln affected positions and amplitudes of the absorption bands of bacteriochlorophylls, increased lifetime of the primary electron donor P* excited state from 3.1 ps to 22 ps, and decreased quantum yield of the P+QA– state formation to 60%. These data suggest significant changes in the pigment–protein interactions in the vicinity of the primary electron donor P and the nearest electron acceptor BA. A considerable decrease was also noticed in the resistance of the mutant RC to thermal denaturation, which was more pronounced in the RC with the double substitution I(M206)Q + F(M208)A. This was likely associated with the disruption of the dense packing of the protein near bacteriochlorophylls PB and BA. Possible reasons for different effects of identical mutations on the properties of two highly homologous RCs from closely related purple non-sulfur bacteria are discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
Photosynthetic reaction centers (RC) of purple bacteria are relatively simply organized structural analogs of photosystem-2 of higher plants, cyanobacteria, and algae, they have been used for many years as a convenient model to study mechanisms of light energy conversion into chemical energy of separated charges. By now RCs from Blastochloris viridis, Rhodobacter sphaeroides (further the new name Cereibacter sphaeroides will be used [1]), and Rhodobacter capsulatus are the most studied. Spatial structures of the first two RCs have been resolved with high resolution [2, 3]; for the RC of Rba. capsulatus the structure has not been yet elucidated in detail, but it is believed to be similar to that of the RC from C. sphaeroides [4]. The transmembrane pigment–protein complex of the C. sphaeroides RC consists of three units (L, M, and H) and ten cofactors integrated into the membrane: four molecules of bacteriochlorophyll (BChl), two bacteriopheophytin (BPheo) molecules HA and HB, two quinones QA and QB, a carotenoid molecule, and non-heme iron. The cofactors are organized as two transmembrane chains of electron transfer (A and B) located symmetrically relative to the 2-nd order symmetry axis [4] (Fig. 1a). It is known that in the RCs of purple bacteria only A-chain of the electron transfer is functionally active. Two BChls, PA and PB, on the periplasmic side of the membrane form the dimer P, which serves as a primary electron donor. After excitation by light, a series of rapid reactions of transmembrane electron transfer are induced in the RC accompanied by charge separation. Quantum yield of the photoinduced charge separation in the RCs of purple bacteria is close to 100% [5].
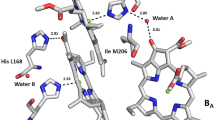
Structure of the reaction center of C. sphaeroides (PDB ID 3v3y). a) L is L-subunit; M is M-subunit; H is H-subunit; P is BChl dimer; BA and BB are monomeric BChl; HA and HB are monomeric BPheo; QA and QB are ubiquinones; car is carotenoid. b) Protein environment of bacteriochlorophylls PB and BA in the structure of the wild type RC (PDB ID 3v3y). Locations of Ile-M206, Phe-M208, Tyr-M210, water-A, fragments of D and E α-helices of the M-subunit are shown.
To date, it has been established that the protein of purple bacteria RC not only holds cofactors inside the membrane, but also controls their spectral and redox properties. Moreover, some amino acid residues play an important role in the photochemical charge separation [6]. This refers in particular to the Tyr-M210 located near the BChl P dimer and the BChl BA monomer. It was shown that changes in the conformation of the hydroxyl group of Tyr-M210 in the course of electron transfer from P* to BChl BA lead to decrease in the energy level of the P+BA– state and thus contribute to stabilization of this short-lived state [7, 8]. Water molecules also are essential components of the RC structure in the hydrophobic part of the complex. It has been shown that the water molecule detected by crystallography near the C9-keto group of BChl BA (in the literature it is often indicated as “water-A”) is necessary for the effective transfer of electron from P* to BA. In the absence of this water lifetime of the P* in the RC of C. sphaeroides increased eight-fold [9]. Establishment of a hydrogen bond between the water-A and C9-keto-group of BChl BA has been demonstrated in the course of the P+BA– state formation [9, 10]. It is assumed that the water-A is included in the chain of polar atoms involved in the electron transfer from the excited electron donor P* to the nearest acceptor BChl BA [11]. Despite a large body of available information about the influence of protein environment on the properties of the cofactors and efficiency of the electron and proton transfer into RC of purple bacteria, the detailed mechanisms of the most rapid initial stages of the charge separation is still remain a subject of discussion.
Introduction of site-directed amino acid substitutions in the environment of cofactors in order to change pigment–protein and protein–protein interactions is one of approaches for studying mechanisms of photosynthetic electron transfer in the RCs of purple bacteria [6]. In particular, in some works attention was focused on the highly conserved residue Ile-M206 located in the immediate vicinity of BChl PB, C9-keto group of the BChl BA monomer, Tyr-M210, and water-A (Fig. 1b). It was shown that the substitutions of Ile-M206 by His and Tyr in the RC of C. sphaeroides significantly affected optical properties of the complex in the absorption region of the dimer P and monomeric BChls [12, 13]. Moreover, a significant drop of the quantum yield of the P+QA– state formation was reported as well as decrease in the amount of the RC I(M206)H isolated from the membranes with the lauryldimethylaminoxide detergent (LDAO) [12, 14]. It was proposed that the introduction of a polar histidine residue into the M206 site in the area of the L- and M-subunit contact could influence the glycolipid binding [15] and thus could affect the RC structure stability [14]. In the RC of Rba. capsulatus the position of M204 corresponds to the M206 position in the RC of C. sphaeroides. In the work by Saggu et al. [16] it was shown that the I(M204)E mutation influenced mobility of the OH-group of Tyr-M210 and the I(M204)Q substitution resulted in the loss of BChl BA from the structure of the complex [17]. It is of interest to obtain stable mutant RCs of C. sphaeroides with similar properties in order to study mechanisms of the initial states of the photochemical process in this RC.
In the present work the mutant RCs of C. sphaeroides were obtained with a single I(M206)Q and double I(M206)Q + F(M208)A substitutions, and their spectral and photochemical properties, pigment composition, and thermal stability were studied. The M208 mutation Phe→Ala was introduced to increase homology of the protein environment of P and BA in the RCs of C. sphaeroides and Rba. capsulatus. Putative causes of different effects of identical mutations on the properties and pigment composition of the homologous RCs of these purple non-sulfur bacteria are discussed.
MATERIALS AND METHODS
Site-directed amino acid substitutions were introduced into RCs by PCR through oligonucleotide primers as described earlier [18]. Presence of mutations in the pufM gene encoding the M subunit of the RC was confirmed by DNA sequencing. The modified puf-operon was cloned into the shuttle-vector pRK-415 [19], the resulting plasmid was transferred by conjugation into the C. sphaeroides strain DD13 [20]. The cells of the resulting recombinant strains synthesized RCs with the desired mutations and did not contain light-harvesting complexes [18]. Pseudo-wild type (WT) RC isolated from the C. sphaeroides strain DD13 which contained pRK-415 derivative carrying non-modified copies of the puf-LMX genes, was used as a control reaction center [18]. C. sphaeroides were grown on a Hutner’s medium [21] in the presence of tetracycline (1 µg/ml) and kanamycin (5 µg/ml). Reaction centers were isolated using ion-exchange and affinity chromatography as described earlier [22, 23]. Complexes were solubilized from the membranes by treatment with LDAO detergent. After purification of the RCs, the detergent was replaced as described earlier [24]. The purified RCs were dissolved in 20 mM Tris-HCl buffer (pH 8.0) containing 0.2% sodium cholate or 0.6% n-octyl-glucoside. Thermostability of the RCs was studied at 48 and 55°C by monitoring thermo-dependent changes of the amplitude of the QY band for 60 min as described earlier [24]. Absorption spectra were recorded using Shimadzu UV-1800 spectrophotometer (Shimadzu, Japan) at room temperature. Sodium ascorbate (1 mM) was added to the samples to maintain the primary electron donor in the reduced state. The pigments were extracted and analyzed as described earlier [25], measurement error was determined by the standard deviation method. Value of the midpoint potential P/P+ was determined by electrochemical titration of the isolated RCs, using potassium ferricyanide as an oxidizer and sodium ascorbate as a reducer, as described earlier [25]. Transient absorption changes with femtosecond resolution were measured in the RCs using the earlier described experimental setup [26]. Light pulses with ~35 fs duration and repetition rate of 20 Hz were obtained using a titan-sapphire laser MaiTai SP (Spectra-Physics, USA) and a regenerative amplifier Spitfire Ace (Spectra-Physics). Energy of the Spitfire Ace output pulses was attenuated and used for pumping a parametric amplifier OPA800 CF (Spectra-Physics) to obtain excitation pulses at 865 nm (second harmonic of the idler beam). A small part of the pulse energy from the regenerative amplifier was used for generation of continuum in a 5 mm thickness cuvette with water as probing pulses. After the cuvette with a sample, spectra of the probing pulses were detected by a CCD-camera Pixis 400BR and a SpectraPro 2300i spectrograph (Princeton Instruments, USA) in the 750-1100 nm wavelength region 750-1100 nm. Transient absorption spectra were obtained by averaging 500 raw spectra for each time delay. Excitation beam with energy of ~1-5 µJ was focused onto the sample cuvette in order to obtain ~10% bleaching of the primary electron donor. The measurements were performed at room temperature. Relative polarization of the excitation pulses was set parallel to the measuring pulse. The reaction center structure was visualized and the amino acid substitutions were modeled with the PyMol program [27].
RESULTS
In the absorption spectra of the isolated WT RCs presented in Fig. 2 the QY P band with maximum at 865 nm belongs to absorption of the BChl dimer P, the absorption band with the maximum at 804 nm is ascribed to absorption of monomeric BChl (QY B), and to the high-energy exciton component of the QY P transition. The QY H band with the maximum at 760 nm corresponds to absorption of BPheo molecules. In the short wavelength region of the spectrum the band at 599 nm corresponds to QX-transitions in BChl molecules. Maximum of the absorption band attributed to BPheo molecules of the active and inactive chains, QX H, is located at 532 nm. Shoulder in the region of 500 nm demonstrates absorption of a carotenoid molecule. Soret band with maximum at 363 nm and shoulder at the long wavelength slope of the band at 390 nm corresponds to the absorption of all bacteriochlorins (Fig. 2).
In the absorption spectrum of the mutant RCs with single and double substitutions the observed changes are alike: a noticeable decrease in the amplitude of the QY P band and a 7-nm blue shift of this band. In addition, a decrease in the amplitude and a 4-nm red shift of the QY B band near 800 nm are observed (Fig. 2). Similarity between the absorption spectra of the two mutant RCs implies that the observed changes are associated with the mutation I(M206)Q, whereas the F(M208)A substitution does not have pronounced effect on the spectral properties of RCs.
The pigment analysis has shown that the ratio BChl/BPheo in the mutant RCs is the same as in the WT RC (table). These results indicate that the amino acid substitution M206 Ile→Gln in the RC of C. sphaeroides does not lead to removal of the monomeric BChl BA from the complex structure as was observed earlier for the RC of Rba. capsulatus with similar mutation I(M204)Q [17].
Value of the midpoint potential P/P+ and pigment analysis of the RC of C. sphaeroides
RC | Em P/P+, mV | BChl/BPheo |
|---|---|---|
Wild type | 490 ± 8 | 2.00 ± 0.1 |
I(M206)Q | 470 ± 5 | 1.98 ± 0.1 |
I(M206)Q + F(M208)A | 475 ± 5 | 1.98 ± 0.1 |
It was shown that the values of the midpoint potential, Em P/P+, were quite similar for the wild type and mutant RCs (table). Close values (within the measurement error) of the oxidative potential of P in the two mutant RCs suggest that the detected small decrease in Em of P/P+ is a result of introduction of the M206 substitution Ile→Gln. The decrease in the value of Em P/P+ also allows us to conclude that the mutation I(M206)Q did not result in formation of a hydrogen bond between the Gln-M206 and the C2-acetyl group of BChl PB conjugated to the π-electron system of the macrocycle. Based on the closeness of the Em P/P+ values of the wild type and mutant RCs, it can be assumed that the driving force of the photochemical reaction ΔG in the genetically modified complexes also did not change significantly. Considering that the F(M208)A substitution had no influence on the spectral properties and oxidative potential of P, we further estimated the lifetime of P* and quantum yield of the P+QA– state formation only for the RCs with the single substitution I(M206)Q.
Figures 3 and 4 present results of investigation of the charge separation in the wild type and I(M206)Q mutant RCs using pump-probe transient absorption spectroscopy. The purpose of this study was to elucidate how the Ile→Gln substitution at the M206 position affects the lifetime of the excited state of the primary electron donor and the yield of the P+QA– state.
Figure 3 presents kinetics of the stimulated emission decay from the excited state P* at 930 nm in the WT RC [open symbols (○)] and in the RC I(M206)Q [closed symbols (●)]. In WT RC, a fit of stimulated emission by an exponential yields a lifetime of 3.1 ± 0.03 ps, but in RC I(M206)Q, the lifetime increased up to 22 ± 1.4 ps. In the WT RC lifetime of the excited primary electron donor with blocked electron transfer is τ = 1 / Σki ≈ 300 ps (where ki are the rate constants of the excited state deactivation pathways) and decreases to value τ = 1 / (kp + Σki) ≈ 3 ps when the electron transfer channel is open. This allows us to estimate both the rate constant of the electron transfer from the excited primary donor to the active chain of cofactors, kp ≈ 3.3 ∙ 10−13 (1/s) and the charge separation yield, φp = kp / (kp + Σki) ≈ 0.99.
Increase of the lifetime of the excited primary electron donor to τ = 1 / (kp + Σki) ≈ 22 ps in the mutant reaction center is expected to result in the reduction of the rate constant of electron transfer from the excited primary donor to the active chain of cofactors to kp ≈ 4.2 ∙ 10−14 (1/s) and in slight decrease of charge separation yield to φp = kp / (kp + Σki) ≈ 0.92.
To estimate the yield of the charge separation state P+QA– in the RC I(M206)Q transient absorption spectra were examined (Fig. 4), which were recorded at certain delay times. Under pulse excitation the transient absorption spectrum of WT RC (Fig. 4a, curve 1) shows singlet excited state of the primary electron donor. At the delay time of ~20 ps the excited P* relaxes to the radical pair P+HA– with quantum yield 0.99 (curve 2). At the delay time of 1600 ps the P+QA– state can be observed (curve 3). In the mutant RC I(M206)Q (Fig. 4b) the same concentration of the excited primary electron donor P* (curve 1) relaxes until delay ~260 ps and yields only 60% of charge separated state P+HA– (curve 2) compared to that in the WT RC. The transient absorption spectrum of the RC I(M206)Q at the delay time of 1600 ps corresponds to formation of the P+QA– state (curve 3).
Difference between the expected value of quantum yield of the P+QA– state formation (92%) in the mutant RC I(M206)Q and that observed in experiment (60%) could be related to changes in the rate constants of both direct and reverse reactions of electron transfer, in particular, of the charge recombination of the P+BA– and P+HA– states.
Relatively high thermal stability of the RC of purple bacteria was noted previously [28]. It is also known that some amino acid substitutions can promote both weakening and strengthening of this complex structure [13, 29]. Effect of mutations introduced in this work or described earlier [12, 13] on stability of the RC structure was studied by monitoring changes in the amplitude of the QY B band at 48°C [24]. The buffer used to dissolve RCs contained sodium cholate detergent, which promoted stabilization of the complex [24].
It was shown that after incubation for 60 min under these conditions, the amplitude of the QY B band at 804 nm in the spectrum of the RC I(M206)Y decreased by less than 5%, similarly to the wild type RC, whereas in the RC I( M206)Q it decreased by ~10%. The RCs I(M206)Q + F(M208)A and I(M206)H exhibited the lowest resistance to thermal denaturation; their incubation at 48°C for 60 min resulted in a 20% decrease of the amplitude of the QY B band (Fig. 5). These results are in agreement with the previously obtained data on destabilizing effect of the I(M206)H substitution on the structure of the C. sphaeroides RC, which is more pronounced in the presence of the LDAO detergent [14]. Thus, it was shown that the double mutation I(M206)Q + F(M208)A that has a purpose to increase homology of the protein environment of the P and BA molecules in the RCs of C. sphaeroides and Rba. capsulatus, had a stronger destabilizing influence on the complex structure as compared to the single mutation I(M206)Q. The results shown in Fig. 5 allow us to suggest that the substitution Ile→Tyr at M206 did not affect thermal stability of the RC or even slightly stabilized the structure of the complex, as it was shown earlier for the RC with the M197 Phe→His substitution [29]. However, when heated in a buffer with the detergent n-octyl-glucoside at 48 and 55°C, the RC I(M206)Y displayed a significantly lower resistance to thermal denaturation than the WT RC (data not presented). Thus, a relatively high thermal stability of the mutant RC I(M206)Y structure could be explained by the stabilizing effect of the sodium cholate detergent shown previously [24].
DISCUSSION
As mentioned in the “Introduction” section, Ile-M206 is located close to molecules that presumably participate in the first stage of charge separation (Fig. 1b). Modeling of the amino acid substitution I(M206)Q using the PyMol program shows 16 possible positions of the side group of Gln-M206. Depending on this group position, the substitution of Ile M206 by Gln potentially could change interactions of any of the mentioned molecules with their environment that would affect quantum yield of the charge separation in the mutant RCs. In particular, it was shown that the mutation I(M204)E in the RC of Rba. capsulatus resulted in formation of a hydrogen bond between the side group of introduced Glu and the hydroxyl group of Tyr-M210 that affected kinetics of the P+HA– state formation [16].
It is shown in this study that the amino acid substitution I(M206)Q markedly influenced spectral properties of the dimer P and monomeric BChls. The available literature data demonstrate that various factors could affect position and amplitude of the QY P band. Among these factors, there are orientation of the C2-acetyl group of BChl, distance between the macrocycles in the dimer, electrostatic environment, etc. [30, 31]. According to the data obtained, oxidative potential of P and, obviously, orientation of the C2-acetyl group of the BChl PB in the mutant RCs were not changed noticeably. Gln, similarly to Ile, is an uncharged amino acid, and therefore, changes in the electrostatic environment of P as a result of the I(M206)Q mutation are also not expected. Therefore, the cause of spectral changes in the absorption region of the BChl dimer remains unclear. Due to overlapping of the QY B and QY P bands, decrease in the absorption in the QY B region near 800 nm in the mutant RCs seems to be associated with the decrease in the amplitude of the QY P band. Considering location of the mutation site, the long wavelength shift and decrease in the amplitude of the QY B band could indicate emergence of new interactions of BChl BA with its protein environment. We assume that the side group of Gln-M206 in the mutant RCs could form a hydrogen bond with the C9-keto group of BChl BA located near the mutation site. In such a case the above-mentioned electron transfer from P* to BA through the chain of polar bonds would be slowed down. As shown in the present work, lifetime of the excited state of the primary electron donor P* in the mutant RC I(M206)Q increased to ~22 ps demonstrating a significant deceleration of the electron transfer from P* to BChl BA, which is in agreement with this suggestion.
Earlier a similar 8-fold slowdown of the P* state decay was observed in the RC of C. sphaeroides with the G(M203)L substitution [9]. It was shown that in the structure of this RC the water-A became sterically displaced by the side group of Leu M203 introduced by the mutation. In the absorption spectrum of the RC G(M203)L a small blue shifts of the QY P and QY B bands were observed without decrease of their amplitudes [9]. Considering that the absorption spectra of the RCs I(M206)Q described in the present work and of the RC G(M203)L described earlier [9] are markedly different in the region of the QY B band (Fig. 2), it could be assumed that changes in the intermolecular interactions caused by the introduction of Gln-M206 should be different from the changes caused by the Gly-M203 substitution with Leu. To clarify the details of these interactions, information is required about the spatial structure of the RC with I(M206)Q substitution.
It is shown in this work that quantum yield of the P+QA– state formation decreased in the RC I(M206)Q to ~60% of that in the WT RC (Fig. 4). Alongside with the assumption that the given mutation could affect the rates of both direct and reverse reactions of the electron transfer (see the “Results” section), it can be also suggested that introduction of the amino acid substitution into the D-helix of the M subunit close to the active branch of cofactors could affect protein dynamics. In the recent work, Dods et al. [32] showed noticeable changes in the conformations of E α-helix of L-subunit and D α-helix of M-subunit occurring within photosynthetic reaction centre of purple bacterium during photochemical reaction and apparently associated with the effective course of this process.
Despite the fact that the genetic system for site-directed mutagenesis and structural-functional studies of the Rba. capsulatus RC was developed earlier than for other purple bacteria [33], attempts to crystallize this complex and resolve its 3D structure have been unsuccessful up to now. Foloppe et al. [34] presented a homologous model of the Rba. capsulatus RC structure based on known structures of the purple bacteria B. viridis and C. sphaeroides with the resolutions of 2.3 Å and 2.8 Å, respectively. A relatively low resolution of the C. sphaeroides RC structure used for the homologous modeling significantly limited possibilities of this approach. Nevertheless, small but reliable differences have been detected in the mutual orientation of monomeric BChl and BPheo in the B-branch between the C. sphaeroides RC and the model of the Rba. capsulatus RC [34]. No significant differences were found in the protein environment of the active branches of cofactors.
Until recently, the purple non-sulfur bacteria C. sphaeroides and Rba. capsulatus were assigned to the same Rhodobacter genus [1] and their RCs were considered highly homologous complexes [35]. We have shown that properties of the C. sphaeroides RC with I(M206)Q substitution are significantly different from the properties of the mutant RCs of Rba. capsulatus with similar substitution I(M204)Q in which this mutation led to the loss of BChl BA from the RC structure [17]. These results seem to be rather unexpected, if we take in consideration that Ile-M206 in the RCs of purple bacteria is a highly conserved residue, and there is a prominent similarity of the general structure of two pigment–protein complexes, same composition of the cofactors and specific features of the chromophores interaction with their protein environment [36], as well as high (77%) homology between the complete protein sequences of these RCs [35]. Especially high is homology of the protein environment of bacteriochlorophylls of the functionally active electron transfer chains reaching 96% for the D α-helix of M subunits of RCs.
Previously obtained experimental data of Wang et al. [37], allow us to speculate on the putative causes of the different effects of the Ile→Gln substitution in the similar positions M206 and M204 in the C. sphaeroides and Rba. capsulatus RCs, respectively. In that work properties of the isolated RCs of four purple bacteria were compared [37], including C. sphaeroides and Rba. capsulatus. It was shown that the positions of the maxima of the QY P bands in the absorption spectra, positions of the oxidized electron donor P+ band near 1250 nm as well as the dipole strength of these bands were different for these RCs. Moreover, ionic detergents were shown to have different effects on positions of the QY P band maxima in the two RCs [37]. Concurrently it was shown in the work by Rautter et al. [38] that the electron structure of P in the membrane-bound and isolated RCs of C. sphaeroides was identical, whereas in the Rba. capsulatus RCs it changed after isolation of these pigment–protein complexes from the membranes. Wang et al. [37] suggested that the RCs of C. sphaeroides and of Rba. capsulatus may differ in the type of binding and electrostatic interactions of the BChl dimer and its environment with membrane phospholipids. Because the site of the introduced mutation M206/M204 is situated in immediate protein environment of P, we assume that these differences, if they exist, could cause different effects of similar substitution on the properties of the C. sphaeroides and Rba. capsulatus RCs. Dissimilarities in the distances between the amino acid residue in the M206/M204 position and the nearest cofactors also cannot be ruled out; to elucidate molecular basis of these differences the structure of Rba. capsulatus should be examined.
It has been also shown in this study that the amino acid substitutions I(M206)Q and I(M206)Q + F(M208)A affected resistance of the RC structure to high temperatures. For the double mutant RC the decrease in thermal stability was more pronounced than for the RC with a single I(M206)Q substitution. Increase in the mobility of the transmembrane D α-helix of the M-subunit (Fig. 1b) due to decrease in the partial molecular volume of amino acid residues in the positions M206 and M208 as a result of introducing the mutations could likely be the cause of changes in the stability of the mutant RCs. In particular, substitution of the hydrophobic Ile residue by the polar Gln residue leads to a local decrease in the volume of the residue at the position M206 by 11.7 cm3/mol, and substitution Phe→Ala at the M208 position lowers partial volume in this region by 38.6 cm3/mol [39]. Thus, the combined decrease in the partial molecular volume of the M206 and M208 residues in the D-helix of the M-subunit due to the double mutation I(M206)Q + F(M208)A is 50.3 cm3/mol, which is comparable with the volume of a small amino acid residue. It can be expected that such change leads to disturbance in the dense packing of the transmembrane region of the protein near bacteriochlorophylls P and BA, weakening of Van-der-Waals and hydrophobic interactions in this region and, consequently, decrease in the stability of the mutant RCs. It should be noted also that of 28 amino acid residues of D-helices of M-subunits of the C. sphaeroides and Rba. capsulatus RCs only one residue at the M208 position is different [35], and in the C. sphaeroides RC this spot is the site of glycolipid binding [15]. We assume that there also could be differences in the interaction of two RCs with this lipid that could cause destabilization of the C. sphaeroides RC after introduction of the Phe→Ala substitution at M208.
CONCLUSIONS
A new mutant RC of C. sphaeroides with the amino acid substitution Ile→Gln at the M206 position near to PB and BA BChl molecules in the active chain of cofactors has been obtained and characterized in this work. It has been shown that this mutation does not lead to the change in the pigment composition of the complex, but markedly affects spectral properties of the cofactors close to the mutation site, lifetime of P*, as well as quantum yield of the charge separation that seems to be associated with formation of new intermolecular interactions in the active chain of cofactors. Properties of the C. sphaeroides and Rba. capsulatus RCs with the identical Ile→Gln substitution at the similar positions are significantly different that likely could be explained by the differences in the interactions of these complexes with lipids of the photosynthetic membrane.
Abbreviations
- BA and BB :
-
monomeric bacteriochlorophylls
- BChl:
-
bacteriochlorophyll
- BPheo:
-
bacteriopheophytin
- HA and HB :
-
monomeric bacteriopheophytins
- P:
-
bacteriochlorophyll dimer
- PA and PB :
-
P dimer bacteriochlorophylls
- QA and QB :
-
ubiquinones
- RC:
-
reaction center
- WT:
-
wild type
References
Hordt, A., Lopez, M. G., Meier-Kolthoff, J. P., Schleuning, M., Weinhold, L. M., et al. (2020) Analysis of 1,000+ type-strain genomes substantially improves taxonomic classification of Alphaproteobacteria, Front. Microbiol., 11, 468, https://doi.org/10.3389/fmicb.2020.00468.
Wöhri, A. B., Wahlgren, W. Y., Malmerberg, E., Johansson, L. C., Neutze, R., et al. (2009) Lipidic sponge phase crystal structure of a photosynthetic reaction center reveals lipids on the protein surface, Biochemistry, 48, 9831-9838, https://doi.org/10.1021/bi900545e.
Selikhanov, G., Fufina, T., Vasilieva, L., Betzel, C., and Gabdulkhakov, A. (2020) Novel approaches for the lipid sponge phase crystallization of the Rhodobacter sphaeroides photosynthetic reaction center, IUCrJ, 7, 1084-1091, https://doi.org/10.1107/S2052252520012142.
Komiya, H., Yeates, T. O., Rees, D. C., Allen, J. P., and Feher, G. (1988) Structure of the reaction center from Rhodobacter sphaeroides R-26 and 2.4.1: symmetry relations and sequence comparisons between different species, Proc. Natl. Acad. Sci. USA, 85, 9012-9016, https://doi.org/10.1073/pnas.85.23.9012.
Wraight, C. A., and Clayton, R. K. (1974) The absolute quantum efficiency of bacteriochlorophyll photooxidation in reaction centers of Rhodopseudomonas sphaeroides, Biochim. Biophys. Acta, 333, 246-260, https://doi.org/10.1016/0005-2728(74)90009-7.
Leonova, M. M., Fufina, T. Y., Shuvalov, V. A., and Vasilieva, L. G. (2014) Study of pigment-protein interactions in photosynthetic center of purple bacteria, in Current Problems of Photosynthesis (Allakhverdiev, S. I., Rubin, A. B., and Shuvalov, V. A., eds) vol. 1, Izhevsk Institute for Computer-Aided Studies, Moscow-Izhevsk, pp. 157-196.
Alden, R. G., Parson, W. W., Chu, Z. T., and Warshel, A. (1996) Orientation of the OH dipole of tyrosine (M)210 and its effect on electrostatic energies in photosynthetic bacterial reaction centers, J. Phys. Chem., 100, 16761-16770, https://doi.org/10.1021/jp961271s.
Yakovlev, A. G., Vasilieva, L. G., Shkuropatov, A. Ya., Bolgarina, T. I., Shkuropatova, V. A., et al. (2003) Mechanism of charge separation and stabilization of separated charges in reaction centers of Chloroflexus aurantiacus and of YM210W(L) mutants of Rhodobacter sphaeroides excited by 20 fs pulses at 90 K, J. Phys. Chem. A, 107, 8330-8338, https://doi.org/10.1021/jp0300647.
Potter, J. A., Fyfe, P. K., Frolov, D., Wakeham, M. C., van Grondelle, R. B. R., et al. (2005) Strong effects of an individual water molecule on the rate of light-driven charge separation in the Rhodobacter sphaeroides reaction center, J. Biol. Chem., 280, 27155-27164, https://doi.org/10.1074/jbc.M501961200.
Robert, B., and Lutz, M. (1988) Proteic events following charge separation in the bacterial reaction center: resonance Raman spectroscopy, Biochemistry, 27, 5108-5114, https://doi.org/10.1021/bi00414a024.
Yakovlev, A. G., Jones, M. R., Potter, J. A., Fyfe, P. K., Vasilieva, L. G., et al. (2005) Primary charge separation between P* and BA: electron-transfer pathways in native and mutant GM203L bacterial reaction centers, Chem. Phys., 319, 297-307, https://doi.org/10.1016/j.chemphys.2005.08.018.
Bolgarina, T. I., Khatypov, R. A., Vasilieva, L. G., Shkuropatov, A. V., and Shuvalov, V. A. (2004) Substitution of isoleucine M206 residue by histidine in the Rhodobacter sphaeroides reaction centers causes changes in the structure of the special bacteriochlorophyll pair molecule, Dokl. Biochem. Biophys., 394, 26-29, https://doi.org/10.1023/b:dobi.0000017147.33235.b4.
Fufina, T. Y., Selkhanov, G. K., Proskuryakov, I. I., Shuvalov, V. A., and Vasilieva, L. G. (2019) Properties of reaction centers of Rhodobacter sphaeroides with the Ile→Tyr substitution at positions L177 and M206, Biochemistry (Moscow), 84, 739-744, https://doi.org/10.1134/S0006297919050110.
Vasilieva, L. G., Fufina, T. Y., Gabdulkhakov, A. G., and Shuvalov, V. A. (2015) Different effects of identical symmetry-related mutations near the bacteriochlorophyll dimer in the reaction center of Rhodobacter sphaeroides, Biochemistry (Moscow), 80, 767-774, https://doi.org/10.1134/S0006297915060012.
Camara-Artigas, A., Brune, D., and Allen, J. P. (2002) Interactions between lipids and bacterial reaction centers determined by protein crystallography, Proc. Natl. Acad. Sci. USA, 99, 11055-11060, https://doi.org/10.1073/pnas.162368399.
Saggu, M., Carter, B., Zhou, X., Faries, K., Cegelski, L., et al. (2014) Putative hydrogen bond to tyrosine M208 in photosynthetic reaction centers from Rhodobacter capsulatus significantly slows primary charge separation, J. Phys. Chem. B, 118, 6721-6732, https://doi.org/10.1021/jp503422c.
Carter, B., Boxer, S. G., Holten, D., and Kirmaier, C. (2012) Photochemistry of a bacterial photosynthetic reaction center missing the initial bacteriochlorophyll electron acceptor, J. Phys. Chem. B, 116, 9971-9982, https://doi.org/10.1021/jp305276m.
Khatypov, R. A., Vasilieva, L. G., Bolgarina, T. I., and Shuvalov, V. A. (2005) Substitution of isoleucine L177 by histidine affects the pigment composition and properties of the reaction center of the purple bacterium Rhodobacter sphaeroides, Biochemistry (Moscow), 70, 1527-1533, https://doi.org/10.1007/s10541-005-0236-3.
Keen, N. T., Tamaki, S., Kobayashi, D., and Trollinger, D. (1988) Improved broad-host-range plasmids for DNA cloning in gram-negative bacteria, Gene, 70, 191-197, https://doi.org/10.1016/0378-1119(88)90117-5.
Jones, M. R., Visschers, R. W., van Grondelle, R., and Hunter, C. N. (1992) Construction and characterization of a mutant strain of Rhodobacter sphaeroides with the reaction center as the sole pigment–protein complex, Biochemistry, 31, 4458-4465, https://doi.org/10.1021/bi00133a011.
Cohen-Basire, G., Sistrom, W. R., and Stanier, R. Y. (1957) Kinetic studies of pigment synthesis by non-sulfur purple bacteria, J. Cell Comp. Physiol., 49, 25-68, https://doi.org/10.1002/jcp.1030490104.
Goldsmith, J. O., and Boxer, S. G. (1996) Rapid isolation of bacterial photosynthetic reaction centers with an engineered poly-histidine tag, Biochim. Biophys. Acta, 1276, 171-175, https://doi.org/10.1016/0005-2728(96)00091-6.
Fufina, T. Yu., Vasilieva, L. G., Khatypov, R. A., Shkuropatov, A. Ya., and Shuvalov, V. A. (2007) Substitution of isoleucine L177 by histidine in Rhodobacter sphaeroides reaction center results in the covalent binding of PA bacteriochlorophyll to the L subunit, FEBS Lett., 581, 57697-5773, https://doi.org/10.1016/j.febslet.2007.11.032.
Fufina, T. Y., and Vasilieva, L. G. (2021) Effects of detergents and osmolytes on thermal stability of native and mutant Rhodobacter sphaeroides reaction centers, Biochemistry (Moscow), 86, 517-524, https://doi.org/10.1134/S000629792104012X.
Vasilieva, L. G., Fufina, T. Yu., Gabdulkhakov, A. G., Leonova, M. M., Khatypov, R. A., et al. (2012) The site-directed mutation I(L177)H in Rhodobacter sphaeroides reaction center affects coordination of PA and BB bacteriochlorophylls, Biochim. Biophys. Acta, 1817, 1407-1417, https://doi.org/10.1016/j.bbabio.2012.02.008.
Khatypov, P. A., Khristin, A. M., Fufina, T. Y., and Shuvalov, V. A. (2017) An alternative pathway of light-induced transmembrane electron transfer in photosynthetic reaction centers of Rhodobacter sphaeroids, Biochemistry (Moscow), 82, 916-922, https://doi.org/10.1134/S0006297917060050.
DeLano, W. L. (2002) The PyMOL molecular graphics system, URL: http://www.Pymol.Org.
Jones, M. R. (2008) Structural plasticity of reaction centers from purple bacteria, in The Purple Phototrophic Bacteria (Hunter, C. N., Daldal, F., Thurnauer, M. C., and Beatty, N., eds) Springer, pp. 295-321, https://doi.org/10.1007/978-1-4020-8815-5_16.
Holden-Dye, K., Crouch, L. I., Williams, C. M., Bone, R. A., Cheng, J., et al. (2011) Opposing structural changes in two symmetrical polypeptides bring about opposing changes to the thermal stability of a complex integral membrane protein, Arch. Biochem. Biophys., 505, 160-170, https://doi.org/10.1016/j.abb.2010.09.029.
Spiedel, D., Roszak, A. W., McKendrick, R., McAuley, K. E., Fyfe, P. K., et al. (2002) Tuning of the optical and electrochemical properties of the primary donor bacteriochlorophylls in the reaction centre from Rhodobacter sphaeroides: spectroscopy and structure, Biochim. Biophys. Acta, 1554, 75-93, https://doi.org/10.1016/S0005-2728(02)00215-3.
Haffa, A. L. M., Lin, S., Katilius, E., Williams, J. C., Taguchi, A. K. W., et al. (2002) The dependence of the initial electron-transfer rate on driving force in Rhodobacter sphaeroides reaction centers, J. Phys. Chem. B, 106, 7376-7384, https://doi.org/10.1021/jp0257552.
Dods, R., Bath, P., Morozov, D., Gagner, V. A., Arnlund, D., et al. (2021) Ultrafast structural changes within a photosynthetic reaction centre, Nature, 589, 310-314, https://doi.org/10.1038/s41586-020-3000-7.
Youvan, D. C., Ismail, S., and Bylina, E. J. (1985) Chromosomal deletion and plasmid complementation of the photosynthetic reaction center and light-harvesting genes from Rhodopseudomonas capsulate, Gene, 38, 19-30, https://doi.org/10.1016/0378-1119(85)90199-4.
Foloppe, N., Ferrand, M., Breton, J., and Smith, J. C. (1995) Structural model of the photosynthetic reaction center of Rhodobacter capsulatus, Proteins, 22, 226-244, https://doi.org/10.1002/prot.340220304.
Michel, H., Epp, O., and Deisenhofer, J. (1986) Pigment-protein interactions in the photosynthetic reaction centre from Rhodopseudomonas viridis, EMBO J., 5, 2445-2451, https://doi.org/10.1002/j.1460-2075.1986.tb04520.x.
Lancaster, C. R. D., Ermler, U., and Michel, H. (1995) The structures of photosynthetic reaction centers from purple bacteria as revealed by X-ray crystallography. Anoxygenic Photosynthetic Bacteria, Advances in Photosynthesis and Respiration v. 2. (Blankenship, R. E., Madigan, M. T., Bauer, C. E., eds) Kluwer Academic Publishers, Netherlands, pp. 503-526, https://doi.org/10.1007/0-306-47954-0_23.
Wang, S., Lin, S., Lin, X., Woodbury, N. W., and Allen, J. P. (1994) Comparative study of reaction centers from purple photosynthetic bacteria: isolation and optical spectroscopy, Photosynth. Res., 42, 203-215, https://doi.org/10.1007/BF00018263.
Rautter, J., Lendzian, F., Lubitz, W., Wang, S., and Allen, J. P. (1994) Comparative study of reaction centers from photosynthetic purple bacteria: electron paramagnetic resonance and electron nuclear double resonance spectroscopy, Biochemistry, 33, 12077-12084, https://doi.org/10.1021/bi00206a010.
Nolding, B. (2005) The Latest Methods for Studying Biosystems, Tekhnosfera, Moscow.
Acknowledgments
Authors are grateful to M. M. Leonova for her contribution to the initial stage of the work.
Funding
This work was financially supported by the State Budget Project no. 122041100204-3.
Author information
Authors and Affiliations
Contributions
L. G. Vasilieva – concept of the study and supervision of the work; T. Yu. Fufina, O. A. Tretchikova, A. M. Khristin, R. A. Khatypov – performing experiments; L. G. Vasilieva, T. Yu. Fufina, O. A. Tretchikova, A. M. Khristin, R. A. Khatypov – discussion of the results of the study; L. G. Vasilieva, T. Yu. Fufina, A. M. Khristin, R. A. Khatypov – writing and editing text of the article.
Corresponding author
Ethics declarations
The authors declare no conflicts of interest in financial or any other sphere. This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Fufina, T.Y., Tretchikova, O.A., Khristin, A.M. et al. Properties of Mutant Photosynthetic Reaction Centers of Purple Non-Sulfur Bacteria Cereibacter sphaeroides with M206 Ile→Gln Substitution. Biochemistry Moscow 87, 1149–1158 (2022). https://doi.org/10.1134/S000629792210008X
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S000629792210008X