Abstract
Astrocytes are the main homeostatic cells in the central nervous system (CNS) that provide mechanical, metabolic, and trophic support to neurons. Disruption of their physiological role or acquisition of senescence-associated phenotype can contribute to the CNS dysfunction and pathology. However, molecular mechanisms underlying the complex physiology of astrocytes are explored insufficiently. Recent studies have shown that miRNAs are involved in the regulation of astrocyte function through different mechanisms. Although miR-21 has been reported as an astrocytic miRNA with an important role in astrogliosis, no link between this miRNA and cellular senescence of astrocytes has been identified. To address the role of miR-21 in astrocytes, with special focus on cellular senescence, we used NT2/A (astrocytes derived from NT2/D1 cells). Downregulation of miR-21 expression in both immature and mature NT2/A by the antisense technology induced the arrest of cell growth and premature cellular senescence, as indicated by senescence hallmarks such as increased expression of cell cycle inhibitors p21 and p53 and augmented senescence-associated β-galactosidase activity. Additionally, in silico analysis predicted many of the genes, previously shown to be upregulated in astrocytes with the irradiation-induced senescence, as miR-21 targets. Taken together, our results point to miR-21 as a potential regulator of astrocyte senescence. To the best of our knowledge, these are the first data showing the link between miR-21 and cellular senescence of astrocytes. Since senescent astrocytes are associated with different CNS pathologies, development of novel therapeutic strategies based on miRNA manipulation could prevent senescence and may improve the physiological outcome.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
MicroRNAs (miRNAs) are small non-coding RNAs that negatively regulate gene expression at the post-transcriptional level. In recent years, miRNAs have been recognized as important players in the regulatory mechanism underlying neural differentiation [1]. They also regulate a diverse set of functions in the adult brain, both under physiological and pathological conditions [2].
MicroRNA-21 (miR-21) is one of the most intriguing miRNAs, due to its very broad roles. This miRNA acts as an oncogene and is upregulated in numerous malignancies, including glioblastoma [3, 4]. It is also involved in the regulation of different developmental processes. Thus, it was shown that miR-21 promotes neural stem cell proliferation and neural differentiation [5]. Additionally, it contributes to the maintenance of adult tissue homeostasis, since its upregulation was reported in reactivated astrocytes upon spinal cord injury (SCI), in which miR-21 mediates profound positive changes in the astrocyte responses [6, 7]. This miRNA has been also recognized as an immune system regulator able to modulate the “on/off switch” of the inflammatory response at the appropriate times by promoting or inhibiting the nuclear factor kappa B (NF-κB) and (NOD)-like receptor protein 3 (NLRP3) pathways that underlie chronic, low-level, systemic inflammation occurring during human aging [8].
Astrocytes are the main homeostatic cells in the CNS [9]. They are tightly integrated into the neural networks and participate in numerous aspects of CNS physiology [9] and pathology [10]. These cells have a unique ability to transform from quiescent to the reactive state in response to injury. This phenomenon is known as reactive astrogliosis and serves as a compensatory response that modulates tissue damage and recovery [11]. The loss of astrocytes function (or the gain of their neuroinflammatory function) as a result of cellular senescence could have profound implications in age-related neurodegenerative disorders [12]. In contrast to reactive astrogliosis, cellular senescence is a permanent state characterized by irreversible growth arrest coupled with other senescence hallmarks, such as increased activity of the senescence-associated beta-galactosidase (SA-β-gal) and expression of cell cycle inhibitors p21, p16, and p53 [12]. However, the molecular mechanisms underlying either of these processes are insufficiently explored. While miR-21 has been considered as a potential molecular target for manipulating astrogliosis and enhancing functional outcome after SCI [7], no link between this miRNA and cellular senescence of astrocytes has been reported.
To address the role of miR-21 in astrocytes, with special focus on cellular senescence, we used human pluripotent embryonal teratocarcinoma NT2/D1 cell line as a model system. NT2/D1 cells have the ability to differentiate along the neural lineage when treated with retinoic acid (RA), yielding both neuronal and glial cell populations [13, 14]. NT2/A (astrocytes derived from NT2/D1-cells) are similar to the primary astrocytes in their properties, such as the presence of intermediate filament proteins, growth arrest, and reactive response to injury [14]. Moreover, NT2/A have an active glutamate transport system and support the growth and survival of NT2/N (neurons derived from NT2/D1 cells) in mixed cultures [14]. We downregulated the expression of miR-21 in both immature and mature NT2/A and analyzed the effect of this downregulation on cells viability, proliferation, and induction of senescence. Our results showed that miR-21 downregulation induced growth arrest and premature cellular senescence of both immature and mature NT2/A.
MATERIALS AND METHODS
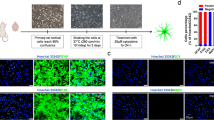
Cell culture and differentiation. Human pluripotent NT2/D1 embryonal carcinoma cells (ATCC® CRL-1973™) were maintained as previously described [15]. NT2/D1 cells were induced to differentiate in culture by adding 10 µM all-trans retinoic acid (Sigma-Aldrich, USA) for four weeks. A neuron-enriched population was isolated by mechanical detachment as previously described [13]. NT2-derived astrocytes were obtained from the remaining adherent cells according to the published protocol [14]. Briefly, after removal of NT2/N, the remaining cells were trypsinized and replated in fresh culture dish. The cells were maintained for additional four weeks in high-glucose DMEM with 10% FBS and subcultured when confluent (at 7-14 days). The morphology of cells was analyzed with a phase contrast DM IL LED Inverted Microscope (Leica Microsystems, Germany).
Lentiviral transduction of NT2/D1-derived astrocytes. Lentiviral particles were produced by transient cotransfection of HEK-293T cells (ATCC® CRL-1573™) using PEI MAX™ transfection reagent (Polysciences Inc., USA) according to the manufacturer’s instructions. Cotransfection of HEK-293T cells was performed in a 10-cm cell culture dish with 1.7 µg of pCMV-VSV-G (envelope plasmid), 1 µg of pRSV-REV and 2.5 µg of pMDLg/RRE (packaging plasmids), and 10 µg of either lentiviral antisense-miR-21 transduction construct (anti-miR-21) [16] or pGreenPuro empty vector (pGP; System Biosciences, USA). NT2/D1-derived astrocytes were incubated overnight in the medium containing lentiviral particles (harvested 48 h after transfection) and 8 µg/ml Polybrene (Sigma-Aldrich). Puromycin selection was started 48 h after transduction. The cells were grown for three days in the serum-containing medium supplemented with 0.3 µg/ml puromycin (selection medium). Non-transduced cells grown in the selection medium were used as puromycin efficiency control. The grown puromycin-resistant cells were used for further experimental analysis. GFP expression in the transduced cells was confirmed by fluorescence microscopy.
Immunocytochemistry (ICC). Four-week-old NT2/A were plated directly on coverslips; NT2/N and one-week old NT2/A were seeded on Geltrex®-coated coverslips. The cells were fixed in 4% paraformaldehyde (PFA) in phosphate buffered saline (PBS) for 20 min at room temperature (RT), permeabilized with 0.2% Triton X-100 for 20 min, and blocked with 1% BSA and 5% normal goat serum in PBS with 0.1% Triton X-100 (PBT) for 1 h at RT. Primary antibodies were diluted in PBT containing 1% BSA, and incubated overnight at 4°C as follows: mouse anti-MAP2, diluted 1 : 500 (ab11267, Abcam, UK); mouse anti-vimentin, diluted 1 : 300 (sc-6260, Santa Cruz Biotechnology, Germany); rabbit anti-Ki67, diluted 1 : 300 (ab15580, Abcam, UK); rabbit anti-GFAP, diluted 1 : 200 (Z 0334, Dako Cytomation, Denmark). The bound primary antibodies were labeled for 1 h at RT with appropriate secondary antibodies: goat anti-rabbit IgG (H+L) Alexa Fluor 488 (A-11034, Invitrogen, USA); goat anti-mouse IgG (H+L) Alexa Fluor 546 (A-11030, Invitrogen); or goat anti-mouse IgG (H+L) Alexa Fluor 488 (A-11029, Invitrogen) diluted 1 : 500 in PBT supplemented with 1% BSA. The anti-GFAP antibody was labeled first with biotinylated goat anti-rabbit IgG, diluted 1 : 500 (BA-1000, Vector Laboratories, USA) for 1 h at RT in PBS containing 1% BSA, followed by labeling for 1 h at RT with streptavidin conjugated with DyLight® 594 (SA-5594, Vector Laboratories, USA) diluted 1 : 500 in PBS. Nuclei were stained with 0.1 mg/ml diamino phenylindole (DAPI) (Sigma-Aldrich). The cells were observed and photographed with an Olympus BX51 fluorescence microscope using the Cytovision 3.1 software (Applied Imaging Corporation, USA) or Leica TCS SP8 confocal microscope using the Leica Microsystems LAS AF-TCS SP8 software (Leica Microsystems).
Reverse transcription (RT)-PCR analysis. Total RNA was isolated, treated with DNase I, and used for cDNA synthesis as previously described [17]. The cDNA was used as a template for the amplification of OCT4 and NEUROD1 using KAPA 2G Fast HotStart Ready Mix (Kapa Biosystems, USA) and for the amplification of p21Waf1/Cip1 and TP53 using Power SYBR Green PCR Master Mix (Applied Biosystems, USA) according to the manufacturer’s protocols. The experiment was done in at least three biological replicates. Amplification was performed in a 7500 Real Time PCR System (Applied Biosystems). All reactions were run in triplicate and the mean value was calculated. The relative levels of p21Waf1/Cip1 and TP53 expression were calculated using the ΔΔCt method and normalized to GAPDH as an endogenous control. The sequences of primers used in this study are: OCT4: 5′-GCTTTGAGGCTCTGCAGCTT-3′ (forward) and 5′-TCTCCAGGTTGCCTCTCACT-3′ (reverse), NEUROD1: 5′-GGAAACGAACCCACTGTGCT-3′ (forward) and 5′-GCCACACCAAATTCGTGGTG-3′ (reverse), GAPDH: 5′-GCCTCAAGATCATCAGCAATGC-3′ (forward) and 5′-CCACGATACCAAAGTTGTCATGG-3′ (reverse), p21Waf1/Cip1: 5′-GACACCACTGGAGGGTGACT-3′ (forward) and 5′-CAGGTCCACATGGTCTTCCT-3′ (reverse), TP53: 5′-CCCCTCCTGGCCCCTGTCATCTTC-3′ (forward) and 5′-GCAGCGCCTCACAACCTCCGTCAT-3′ (reverse).
TaqMan-miRNA assay and data analysis. TaqMan–miRNA assays (Applied Biosystems) were used for miR-21 (Cat. No. 4427975; Assay ID 000397), miR-30d (Cat. No. 4427975; Assay ID 000420), miR-30d* (Cat. No. 4427975; Assay ID 002305), miR-124a (Cat. No. 4427975; Assay ID 000446), miR-219-5p (Cat. No. 4427975; Assay ID 000522) and miR-941 (Cat. No. 4427975; Assay ID 002183). Reverse transcription was performed using MultiScribeTM reverse transcriptase (Applied Biosystems) as previously described [3]. Quantitative PCR was performed using TaqMan Universal PCR Master Mix in a 7500 Real Time cycler (Applied Biosystems) as previously described [3, 18]. The relative expression of specific miRNAs was evaluated using the ΔΔCt method [3] and normalized to RNU44 (Cat. No. 4427975; Assay ID 001095) as an endogenous control. All experiments were done in three biological replicates with at least two technical repeats.
Cell viability assay. Five days after transduction with anti-miR-21 or empty vector, the cells were seeded in 96-well plates at a density of 1.5 × 104 cells per well for one-week-old NT2/A and 2 × 104 cells per well for four-week-old NT2/A. Cell viability was tested with the MTT assay (Merck, USA) for the one-week-old NT2/A and the CellTiter 96® Aqueous One Solution Cell Proliferation Assay (Promega, USA) for the four-week-old NT2/A. The assays were performed according to the manufacturer’s instructions. Colorimetric measurements were performed with an Infinite 200 PRO microplate reader (Tecan, Switzerland). The assays were done in six technical replicates and repeated in three independent experiments.
Senescence analysis. For the assessment of the SA-β-gal activity, the cells were seeded in 24-well plates. Five days after transduction with the anti-miR-21 construct or empty vector, the cells were stained using Senescence β-galactosidase Staining Kit (Cell Signaling Technology) according to the manufacturer’s instructions. Briefly, the cells were washed with PBS, fixed, and stained for 12 h at 37°C for the β-galactosidase activity using a solution of X-gal. All cells were imaged under the same conditions with a phase contrast DM IL LED Inverted Microscope (Leica Microsystems); the intensity of SA-β-gal staining was quantified using the ImageJ software (National Institutes of Health, USA). The results were obtained from three independent biological replicates.
Target prediction data. The targets of miR-21 (hsa-miR-21-5p) were predicted using TargetScan Human Release 7.2 (http://www.targetscan.org/vert_72/) [19], DIANA TarBase v8 (https://carolina.imis.athena-innovation.gr/diana_tools/web/index.php?r=tarbasev8%2Findex) [20], and MirDB (http://mirdb.org/) [21] pre-computed predictions available online. Predicted or verified miR-21 targets obtained by each of these miRNA target prediction algorithms were overlapped with the genes upregulated in astrocytes upon irradiation-induced senescence published by Limbad et al. [22].
Statistical analyses. Statistical analyses were performed with the SPSS statistical software (version 20). The data represent mean ± SEM from at least three independent experiments; p-values were calculated using the Student’s t-test: * p ≤ 0.05, ** p ≤ 0.01, *** p ≤ 0.001.
RESULTS
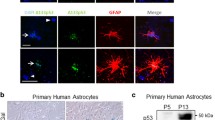
Characterization of in vitro neural differentiation of human pluripotent NT2/D1 cells. RA-induced in vitro neural differentiation of human pluripotent embryonal carcinoma NT2/D1 cells was performed as previously described [13]. The progression of neural differentiation was documented by changes in the cell morphology (Fig. 1a) and expression of specific markers (Fig. 1, b and c). Undifferentiated NT2/D1 cells, characterized by rapid cell division and small cell bodies, grew as a monolayer culture (Fig. 1a). These cells also expressed the pluripotency-associated stem cell marker OCT4 (octamer-binding transcription factor 4) (Fig. 1b). After four weeks of RA treatment, the cells formed an extremely dense multilayered culture with round cell clusters located above a very dense layer of adherent cells (Fig. 1a, +RA). This multilayered cell culture contained mixed populations of neuronal and glial progenitor cells as previously shown [13, 14]. Accordingly, at that phase of neural differentiation, we detected high expression levels of the neuronal progenitor marker NEUROD1 (neurogenic differentiation factor 1) (Fig. 1b). Next, we used these multi-layered cell cultures to isolate NT2/N and NT2/A (Fig. 1, a and c) using previously established protocols [13, 14]. The neuronal and astroglial phenotypes were confirmed by monitoring expression of specific cytoskeletal proteins. Thus, the NT2/N showed an intense staining for MAP2 (microtubule-associated protein 2), a neuron-specific cytoskeletal protein found throughout the neuronal cell bodies and processes (Fig. 1, c and c′, NT2/N). The antibodies against cytoplasmic filament proteins vimentin and GFAP (glial fibrillary acidic protein) were used for the detection of NT2/A. In line with the results reported by Sandhu et al. [14], all analyzed cells in the one-week-old culture of enriched NT2/A were vimentin-positive, while only a small subset of NT2/A displayed intense GFAP staining, a marker of mature astroglial cells (Fig. 1c, NT2/A).
a) Phase-contrast images showing morphological changes occurring during the RA-induced neural differentiation of NT2/D1 cells. NT2/D1, undifferentiated cells; +RA, NT2/D1 cells after four weeks of RA treatment; NT2/N, mature neurons; NT2/A, astrocytes (one-week-old); 10× magnification. b) Semiquantitative RT-PCR analysis of OCT4 and NEUROD1 expression. GAPDH was used as a loading control. c) Immunostaining of NT2/N with the anti-MAP2 antibody (red) and of NT2/A with the antibodies against GFAP (green) and vimentin (red). Cell nuclei were counterstained with DAPI (blue). The boxed region in (c) is enlarged in (c′). Scale bar, 100 µm.
Analysis of expression of selected miRNAs during RA-induced neural differentiation of NT2/D1 cells. Comprehensive miRNA expression analysis conducted by Smith et al. [23] has delineated miRNAs associated with specific phases of the RA-induced neural differentiation of NT2/D1 cells. To further characterize this model system with regard to the miRNAs expression patterns, we selected miR-219 and miR-30d, which are specific for the neural progenitors, as well as miR-124 and miR-21 characteristic for the NT2-derived neurons and astrocytes, respectively [23]. Additionally, we included miR-30d* (as another member of miR-30d family) and miR-941 in our analysis. miR-941 is particularly interesting, since it is human-specific, highly expressed in the brain, and affects genes involved in neurotransmitter signaling [24].
The highest expression levels of four out of six analyzed miRNAs (miR-30d, miR-30d*, miR-219, and miR-941) was detected in NT2/D1 cells within the four weeks of RA treatment (Fig. 2). Although expressed at considerably different levels, both miR-30 family members, miR-30d and miR-30d*, showed a similar expression pattern in the course of NT2/D1 neural differentiation (Fig. 2). miR-124 and miR-21 showed the highest level of expression in the NT2/N and NT2/A, respectively (Fig. 2). However, while miR-124 is well described as a brain-enriched miRNA known to mediate neurogenesis in both developing and adult brain [25, 26], the role of miR-21 in astrocytes has been less explored. Accordingly, we focused our studies on this miRNA and its role in NT2/A.
miRNA expression at different phases of RA-induced NT2/D1 neural differentiation: NT2/D1, undifferentiated cells; +RA, NT2/D1 cells after four weeks of RA treatment; NT2/N, mature neurons; NT2/A, astrocytes (one-week-old). Relative miRNA expression was calculated compared to the expression level in undifferentiated NT2/D1 cells. Data represent means ± SEM (n = 3), p-values were calculated using the Student’s t-test, * p ≤ 0.05, ** p ≤ 0.01, *** p ≤ 0.001.
Characterization of NT2/A at different stages of maturity. Maturation of NT2/A was documented based on changes in the cell morphology (Fig. 3a), decrease in the cell proliferation rate (Fig. 3, b and c), and expression of specific markers (Fig. 3a, lower panel). One-week-old NT2/A cultures consisted mostly of small star-like cells with numerous fiber-like processes (Fig. 3a, 1-week-old NT2/A). These fibrous or stellate astrocytes grew on the top of protoplasmic astrocytes during the first two weeks of cell culture. During further maturation of NT2/A, the proportion of fibrous astrocytes dramatically reduced and by the fourth week, the cultures were mainly composed of protoplasmic astrocytes, flat cells with short cytoplasmic extensions expanding from almost round cell bodies (Fig. 3a, 4-weeks-old NT2/A). While both fibrous and protoplasmic cells were vimentin-positive, intense GFAP staining was seen only in a subset of fibrous astrocytes, but not in large protoplasmic cells (Fig. 3a, lower panel).
a) Phase-contrast images showing morphological differences between the one-week-old and four-week-old NT2/A (upper panel); 10× magnification. Immunostaining with antibodies against vimentin (green) and GFAP (red) (lower panel). Cell nuclei were counterstained with DAPI (blue). Scale bar, 100 µm. b) Detection of Ki-67 expression (green) by immunofluorescence; 20× magnification. Boxed region in (b) (left panel) is enlarged in (b′). c) Statistical analysis of Ki-67 immunopositive cells in the one-week-old and four-week-old NT2/A cultures. The percentage of Ki-67 immunopositive cells was calculated relatively to the total number of DAPI-labeled cells (at least 100). Data represent means ± SEM (n = 3), p-values were calculated using the Student’s t-test, *** p ≤ 0.001. d) Relative miR-21 expression in the four-week-old NT2/A calculated compared to the expression level in the one-week-old NT2/A.
The proliferation rate of NT2/A at two different stages of maturation was determined by studying expression of Ki67, a nuclear protein associated with cellular proliferation (Fig. 3b). The number of NT2/A labeled with Ki67 decreased significantly during maturation (Fig. 3c). In particular, one-week-old NT2/A cultures contained 66% Ki67-labeled cells, while after four weeks of maturation, only 29% of mitotically active cells were observed (out of the total number of counted cells). Hence, maturation of NT2/A for additional three weeks was accompanied by an approximately 2-fold decrease in the NT2/A proliferation rate. Similar phenotypical changes during NT2/A maturation were described by Sandhu et al. [14]. We also examined whether the expression level of miR-21 changed during the NT2/A maturation. We detected a minor, non-significant decrease in the level of miR-21 expression in the four-week-old vs. one-week-old NT2/A (Fig. 3d), indicating that the level of miR-21 expression remains rather unchanged during the NT2/A maturation.
miR-21 downregulation affects NT2/A proliferation. To gain insight into the effect of miR-21 on the phenotypical characteristics of NT2/A, we downregulated its expression in both immature and mature NT2/A. We transduced one-week-old and four-week-old NT2/A with the construct expressing anti-miR-21 and analyzed the level of miR-21 expression five days after transduction. miR-21 expression was decreased to 0.39 ± 0.09 (in one-week-old NT2/A) and to 0.60 ± 0.03 (in four-week-old NT2/A) compared with the levels of miR-21 in the control cells transduced with the empty vector (Fig. 4a).
a) Relative miR-21 expression in one-week-old and four-week-old NT2/A after transduction with anti-miR-21 calculated in comparison to the expression level of miR-21 in control cells transduced with the empty vector. b) Relative viability of one-week-old and four-week-old NT2/A after transduction with anti-miR-21 calculated compared to the viability of cells transduced with the empty vector. Data presented in (a) and (b) represent means ± SEM (n = 3), p-values were calculated using the Student’s t-test, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001. c) Representative images of Ki-67 immunostaining (green) of one-week-old (left panel) and four-week-old (right panel) NT2/A transduced with anti-miR-21 or empty vector (control). Cell nuclei were counterstained with DAPI (blue); 20× magnification. d) Statistical analysis of Ki-67 immunopositive one-week-old and four-week-old NT2/A transduced with anti-miR-21 or empty vector. The percentage of Ki-67 immunopositive cells was calculated relatively to the total number of DAPI-labeled cells (at least 180). The results were obtained from two independent experiments.
First, we examined whether downregulation of miR-21 expression influenced the viability of NT2/A at both stages of maturity. The viability of NT2/A was analyzed five days after transduction by the MTT assay for the one-week-old NT2/A and by the MTS assay for mature (four-week-old) NT2/A. Downregulation of miR-21 led to a significant decrease in the cell viability – by 60 and 40% in one-week-old and four-week-old NT2/A, respectively (Fig. 4b). Next, we examined expression of Ki-67 by immunofluorescence to determine the effect of miR-21 downregulation on the cell proliferation rate of NT2/A (Fig. 4c). As presented in Fig. 4d, the number of Ki-67-positive cells decreased by approximately 30 and 20% in one-week-old and four-week-old NT2/A, respectively. Thus, the detected decline in the NT2/A cell viability was, at least in part, caused by reduction in the cell proliferation rate.
miR-21 downregulation induces NT2/A senescence. To evaluate if the growth arrest detected upon transduction of one-week-old and four-week-old NT2/A with anti-miR-21, was accompanied by changes in the expression levels of cell cycle inhibitors, we analyzed expression of p21Waf1/Cip1 and TP53 (Fig. 5a) and found that they were upregulated in both one-week-old and four-week-old NT2/A transduced with anti-miR-21 (Fig. 5a). Upregulated expression of the cell cycle inhibitors p21 and p53, as well as the cell growth arrest, could indicate induction of cellular senescence [12].
a) Quantitative RT-PCR analysis of p21Waf1/Cip1 and TP53 expression in the one-week-old and four-week-old NT2/A after transduction with anti-miR-21. Relative p21Waf1/Cip1 and TP53 expression levels in the transduced cells were calculated in comparison with the expression levels of p21Waf1/Cip1 and TP53 in the control cells transduced with the empty vector. Results were presented as the means ± SEM of three independent experiments, p-values were calculated using the Student’s t-test, * p ≤ 0.05. b) Representative images of SA-β-gal staining (green) of one-week-old and four-week-old NT2/A transduced with anti-miR-21 or empty vector. c) Relative increase of the SA-β-gal staining intensity in the one-week-old and four-week-old NT2/A transduced with anti-miR-21 was calculated in comparison with the SA-β-gal staining intensity in the cells transduced with the empty vector. Data represent means ± SEM (n = 3), p-values were calculated using the Student’s t-test, ** p ≤ 0.01.
To assess whether downregulation of miR-21 affects senescence, we analyzed the activity of SA-β-gal in both one-week-old and four-week-old NT2/A transduced with anti-miR-21. The number of senescent NT2/A increased during maturation (Fig. 5b, compare control/one-week-old and control/four-week-old NT2/A). In addition, intense SA-β-gal staining was detected in both one-week-old and four-week-old NT2/A transduced with anti-miR-21 (Fig. 5b). Quantification of the SA-β-gal staining intensity showed that miR-21 downregulation caused 4.6- and 3.8-fold increase in the SA-β-gal activity in the one-week-old and four-week-old NT2/A, respectively, compared to cells transduced with the empty vector (Fig. 5c).
Taken together, irreversible growth arrest coupled with the elevated expression of the cell cycle inhibitors p21 and p53 and increased SA-β-gal activity led us to the conclusion that miR-21 downregulation induced senescence of both one-week-old and four-week-old NT2/A.
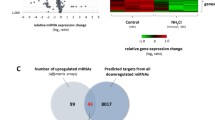
In silico analyses of senescence-associated miR-21 target genes. Comprehensive RNA-seq analysis performed by Limbad et al. [22] revealed vast transcriptome changes in astrocytes with the irradiation-induced senescence. Many genes crucial for astrocyte functions were downregulated, while numerous pro-inflammatory genes were upregulated [22]. To investigate whether potential miR-21 targets are among those identified upregulated genes, we conducted in silico analyses by applying three different online prediction software tools: TargetScan, TarBase, and MirDB. Our in silico study predicted many of the genes overexpressed in the irradiation-induced senescent astrocytes to be miR-21 targets (Table S1 in the Supplement). Among them, twenty-four genes, predicted by all three online tools, overlapped (Fig. 6). These findings further supported our results pointing to miR-21 as a potential regulator of cellular senescence of astrocytes.
Venn diagram showing predicted miR-21 target genes upregulated in senescent astrocytes [22] and shared between TargetScan, TarBase, and MirDB. Twenty-four genes common for all three algorithms are shown.
DISCUSSION
Large-scale expression profiling of miRNAs in human NT2/D1 cells during RA-induced transition from progenitors to fully differentiated neural phenotypes has revealed that this model system can be used for studying specific functions of miRNAs in human neural development [23]. In accordance with the comprehensive miRNAs expression analysis, we have shown that miR-219 and miR-30d are specific for the neural progenitor phases of RA-induced NT2/D1 neural differentiation. A similar conclusion can be drawn for the human-specific miR-941 (Fig. 2). The high level of miR-941 expression upon the four-week treatment of NT2/D1 cells with RA could be associated with cellular differentiation, since it was reported that this human-specific miRNA preferentially targets genes involved in signaling pathways essential for this process [24]. Detected high levels of miR-124 and miR-21 expression in NT2-derived neurons and astrocytes, respectively (Fig. 2), were also in line with the data published by Smith et al. [23], who showed elevated expression of these miRNAs in NT2-differentiated cells and their primary human counterparts [23].
NT2/A underwent phenotypical changes during their in vitro maturation (Fig. 3), consistent with previous study, including the presence of vimentin even in mature astrocytes [14]. The number of proliferating cells (Ki67-positive) decreased significantly with the age of NT2/A cultures (Fig. 3, b and c), which according to Sandhu et al. [14] is due to the cell cycle arrest in G0/G1 phase. Although these cells retained the capacity to resume proliferation after injury [14], our results indicated that the decreased growth rate in the aging NT2/A cultures was also due to the induction of cellular senescence, since we detected increased SA-β-gal activity in the four-week-old cultures compared to the one-week-old NT2/A cultures (Fig. 5b). This finding is in correlation with the fact that NT2/A cultures have a limited lifespan (six to ten passages) as reported by Sandhu et al. [14].
In the adult brain, the role of miR-21 has been mostly studied in the context of SCI. This miRNA has been proposed as the regulator of astrocytic function that can promote recovery after SCI both in vitro and in vivo [6, 7]. In particular, it was shown that the upregulated expression of miR-21 following SCI promotes proliferation and inhibits apoptosis in astrocytes [6]. In accordance with this data, we have shown that miR-21 down-regulation reduces proliferation of NT2/A. Interestingly, the effect was notable not only in the one-week-old NT2/A cultures, comprised mostly of mitotically active cells, but also in the four-week-old cultures, in which the majority of the cells had already reached the quiescence (Fig. 4, c and d). This pro-proliferative activity of miR-21 has been described in many other cellular contexts, mainly in different types of malignancies, including glioblastoma [27, 28]. Accordingly, it is surprising that our finding showed that miR-21 remained highly expressed in mature NT2/A cultures (Fig. 3d), even though the number of cycling cells decreased significantly. We can assume that astrocytes must maintain a certain level of miR-21 expression during maturation to prevent premature cellular senescence. Indeed, the growth arrest induced by the miR-21 downregulation was accompanied with an increased expression of p21Waf1/Cip1 and TP53 (Fig. 5a). Upregulation of these cell cycle inhibitors coupled with elevated SA-β-gal activity were clear indicators that miR-21 downregulation induced cellular senescence (Fig. 5, b and c). To the best of our knowledge, these are the first data showing the link between miR-21 and cellular senescence of astrocytes. Similar findings have been reported in other cell models [29, 30]. The changes in the basal miR-21 expression level often correlate with different CNS pathologies, most evidently, in glioblastoma [4]. Moreover, it has been reported that the level of miR-21 significantly changes in both tissues and bloodstream during aging and it is associated with a variety of inflammatory conditions and age-related diseases [8]. Taken together, we can assume that the maintenance of the basal level of miR-21 expression is important for normal physiological activity of astrocytes.
Senescence-associated dysfunction of astrocytes appears to have profound implication for the aging brain and neurodegenerative disorders [31]. The adverse effect of senescent astrocytes on the surrounding microenvironment is attributed mostly to their acquired pro-inflammatory secretory phenotype [32]. In addition, other alterations induced by senescence in astrocytes appear to be relevant for their neurotoxicity. Thus, the downregulation of transporters of potassium ions and glutamate, with the ensuing decrease of neuronal protection may lead to neurodegeneration, including Alzheimer’s disease and related dementias [22, 31, 33].
Our in silico study revealed twenty-four putative senescence-associated miR-21 target genes (Fig. 6). Several of these genes (TIMP3, TGFB1, IL12A, RECK, VCL, COL4A1, and EHD1) were also shown to be upregulated in human astrocytes in the oxidative stress-induced senescence [33]; some of them (TIMP3, RECK, SMAD7, SATB1, PITX2, MAP2K3, BTG2, CCL20, and ANKRD46) were experimentally validated as direct miR-21 targets in various cells [34-41]. These findings support our results pointing to miR-21 as a potential regulator of astrocyte senescence. Interestingly, putative miR-21 targets presented in Fig. 6 include the genes related to the senescence-associated secretory phenotype, such as genes for interleukin IL12A, cytokine CCL20, angiogenic factor TGFβ, and inhibitors of metalloproteases TIMP3 and RECK. Nevertheless, further direct functional analysis is needed to reveal miR-21-related mechanisms underlying astrocytes senescence. As discussed previously, senescent astrocytes are associated with different CNS pathologies. Therefore, discovering miRNA-related mechanisms underlying astrocytes senescence could be important also from the applicative point of view. New therapeutic strategies based on miRNA manipulation could prevent the senescent state and improve the physiological outcome.
Abbreviations
- anti-miR-21:
-
antisense-miR-21 transduction construct
- miR-21:
-
microRNA-21
- NT2/A:
-
astrocytes derived from NT2/D1-cells
- RA:
-
retinoic acid
- SA-β-gal:
-
senescence-associated beta-galactosidase
- SCI:
-
spinal cord injury
References
Rajman, M., and Schratt, G. (2017) MicroRNAs in neural development: from master regulators to fine-tuners, Development, 144, 2310-2322.
Qureshi, I. A., and Mehler, M. F. (2012) Emerging roles of non-coding RNAs in brain evolution, development, plasticity and disease, Nat. Rev. Neurosci., 13, 528-541.
Anastasov, N., Hofig, I., Vasconcellos, I. G., Rappl, K., Braselmann, H., et al. (2012) Radiation resistance due to high expression of miR-21 and G2/M checkpoint arrest in breast cancer cells, Radiat. Oncol., 7, 206.
Chan, J. A., Krichevsky, A. M., and Kosik, K. S. (2005) MicroRNA-21 is an antiapoptotic factor in human glioblastoma cells, Cancer Res., 65, 6029-6033.
Zhang, W. M., Zhang, Z. R., Yang, X. T., Zhang, Y. G., and Gao, Y. S. (2018) Overexpression of miR21 promotes neural stem cell proliferation and neural differentiation via the Wnt/betacatenin signaling pathway in vitro, Mol. Med. Rep., 17, 330-335.
Liu, R., Wang, W., Wang, S., Xie, W., Li, H., and Ning, B. (2018) microRNA-21 regulates astrocytic reaction post-acute phase of spinal cord injury through modulating TGF-beta signaling, Aging (Albany NY), 10, 1474-1488.
Bhalala, O. G., Pan, L., Sahni, V., McGuire, T. L., Gruner, K., et al. (2012) microRNA-21 regulates astrocytic response following spinal cord injury, J. Neurosci., 32, 17935-17947.
Olivieri, F., Prattichizzo, F., Giuliani, A., Matacchione, G., Rippo, M. R., et al. (2021) miR-21 and miR-146a: the microRNAs of inflammaging and age-related diseases, Ageing Res. Rev., 70, 101374.
Verkhratsky, A., and Nedergaard, M. (2018) Physiology of astroglia, Physiol. Rev., 98, 239-389.
Meldolesi, J. (2020) Astrocytes: news about brain health and diseases, Biomedicines, 8, 394, https://doi.org/10.3390/biomedicines8100394.
Barreto, G. E., Gonzalez, J., Torres, Y., and Morales, L. (2011) Astrocytic-neuronal crosstalk: implications for neuroprotection from brain injury, Neurosci. Res., 71, 107-113.
Cohen, J., and Torres, C. (2019) Astrocyte senescence: evidence and significance, Aging Cell, 18, e12937.
Pleasure, S. J., Page, C., and Lee, V. M. (1992) Pure, postmitotic, polarized human neurons derived from NTera 2 cells provide a system for expressing exogenous proteins in terminally differentiated neurons, J. Neurosci., 12, 1802-1815.
Sandhu, J. K., Sikorska, M., and Walker, P. R. (2002) Characterization of astrocytes derived from human NTera-2/D1 embryonal carcinoma cells, J. Neurosci. Res., 68, 604-614.
Andrews, P. W. (1984) Retinoic acid induces neuronal differentiation of a cloned human embryonal carcinoma cell line in vitro, Dev. Biol., 103, 285-293.
Radulovic, V., Heider, T., Richter, S., Moertl, S., Atkinson, M. J., and Anastasov, N. (2017) Differential response of normal and transformed mammary epithelial cells to combined treatment of anti-miR-21 and radiation, Int. J. Radiat. Biol., 93, 361-372.
Popovic, J., Stanisavljevic, D., Schwirtlich, M., Klajn, A., Marjanovic, J., and Stevanovic, M. (2014) Expression analysis of SOX14 during retinoic acid induced neural differentiation of embryonal carcinoma cells and assessment of the effect of its ectopic expression on SOXB members in HeLa cells, PLoS One, 9, e91852.
Kramer, M. F. (2011) Stem-loop RT-qPCR for miRNAs, Curr. Protoc. Mol. Biol., 95, 15.10.1-15.10.15, https://doi.org/10.1002/0471142727.mb1510s95.
Agarwal, V., Bell, G. W., Nam, J. W., and Bartel, D. P. (2015) Predicting effective microRNA target sites in mammalian mRNAs, Elife, 4, e05005, https://doi.org/10.7554/eLife.05005.
Karagkouni, D., Paraskevopoulou, M. D., Chatzopoulos, S., Vlachos, I. S., Tastsoglou, S., et al. (2018) DIANA-TarBase v8: a decade-long collection of experimentally supported miRNA-gene interactions, Nucleic Acids Res., 46, D239-D245.
Liu, W., and Wang, X. (2019) Prediction of functional microRNA targets by integrative modeling of microRNA binding and target expression data, Genome Biol., 20, 18.
Limbad, C., Oron, T. R., Alimirah, F., Davalos, A. R., Tracy, T. E., et al. (2020) Astrocyte senescence promotes glutamate toxicity in cortical neurons, PLoS One, 15, e0227887.
Smith, B., Treadwell, J., Zhang, D., Ly, D., McKinnell, I., et al. (2010) Large-scale expression analysis reveals distinct microRNA profiles at different stages of human neurodevelopment, PLoS One, 5, e11109.
Hu, H. Y., He, L., Fominykh, K., Yan, Z., Guo, S., et al. (2012) Evolution of the human-specific microRNA miR-941, Nat. Commun., 3, 1145.
Cheng, L. C., Pastrana, E., Tavazoie, M., and Doetsch, F. (2009) miR-124 regulates adult neurogenesis in the subventricular zone stem cell niche, Nat. Neurosci., 12, 399-408.
Kawahara, H., Imai, T., and Okano, H. (2012) MicroRNAs in neural stem cells and neurogenesis, Front. Neurosci., 6, 30.
Gaur, A. B., Holbeck, S. L., Colburn, N. H., and Israel, M. A. (2011) Downregulation of Pdcd4 by mir-21 facilitates glioblastoma proliferation in vivo, Neuro Oncol., 13, 580-590.
Lou, Y., Yang, X., Wang, F., Cui, Z., and Huang, Y. (2010) MicroRNA-21 promotes the cell proliferation, invasion and migration abilities in ovarian epithelial carcinomas through inhibiting the expression of PTEN protein, Int. J. Mol. Med., 26, 819-827.
Savickiene, J., Baronaite, S., Zentelyte, A., Treigyte, G., and Navakauskiene, R. (2016) Senescence-associated molecular and epigenetic alterations in mesenchymal stem cell cultures from amniotic fluid of normal and fetus-affected pregnancy, Stem Cells Int., 2016, 2019498.
Marasa, B. S., Srikantan, S., Martindale, J. L., Kim, M. M., Lee, E. K., et al. (2010) MicroRNA profiling in human diploid fibroblasts uncovers miR-519 role in replicative senescence, Aging (Albany NY), 2, 333-343.
Baker, D. J., and Petersen, R. C. (2018) Cellular senescence in brain aging and neurodegenerative diseases: evidence and perspectives, J. Clin. Invest., 128, 1208-1216.
Chinta, S. J., Woods, G., Rane, A., Demaria, M., Campisi, J., and Andersen, J. K. (2015) Cellular senescence and the aging brain, Exp. Gerontol., 68, 3-7.
Crowe, E. P., Tuzer, F., Gregory, B. D., Donahue, G., Gosai, S. J., et al. (2016) Changes in the transcriptome of human astrocytes accompanying oxidative stress-induced senescence, Front. Aging Neurosci., 8, 208.
Ahmed, M. I., Pickup, M. E., Rimmer, A. G., Alam, M., Mardaryev, A. N., et al. (2019) Interplay of microRNA-21 and SATB1 in epidermal keratinocytes during skin aging, J. Invest. Dermatol., 139, 2538-2542.e2.
Cui, M., Zhang, M., Liu, H. F., and Wang, J. P. (2017) Effects of microRNA-21 targeting PITX2 on proliferation and apoptosis of pituitary tumor cells, Eur. Rev. Med. Pharmacol. Sci., 21, 2995-3004.
Gabriely, G., Wurdinger, T., Kesari, S., Esau, C. C., Burchard, J., et al. (2008) MicroRNA 21 promotes glioma invasion by targeting matrix metalloproteinase regulators, Mol. Cell. Biol., 28, 5369-5380.
Liu, M., Wu, H., Liu, T., Li, Y., Wang, F., et al. (2009) Regulation of the cell cycle gene, BTG2, by miR-21 in human laryngeal carcinoma, Cell Res., 19, 828-837.
Marts, L. T., Green, D. E., Mills, S. T., Murphy, T., and Sueblinvong, V. (2017) MiR-21-mediated suppression of Smad7 induces TGFbeta1 and can be inhibited by activation of Nrf2 in alcohol-treated lung fibroblasts, Alcohol Clin. Exp. Res., 41, 1875-1885.
Yan, L. X., Wu, Q. N., Zhang, Y., Li, Y. Y., Liao, D. Z., et al. (2011) Knockdown of miR-21 in human breast cancer cell lines inhibits proliferation, in vitro migration and in vivo tumor growth, Breast Cancer Res., 13, R2.
Yao, T., and Lin, Z. (2012) MiR-21 is involved in cervical squamous cell tumorigenesis and regulates CCL20, Biochim. Biophys. Acta, 1822, 248-260.
Yao, X., Wang, Y., and Zhang, D. (2018) microRNA-21 Confers Neuroprotection against cerebral ischemia-reperfusion injury and alleviates blood-brain barrier disruption in rats via the MAPK signaling pathway, J. Mol. Neurosci., 65, 43-53.
Funding
This work was supported by the Ministry of Education, Science and Technological Development of the Republic of Serbia (projects nos. 451-03-68/2020-14/200042 and 451-03-9/2021-14/200042), IBRO/PERC InEurope Short Stay Grants, and Serbian Academy of Sciences and Arts (project no. 01-2021).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare no conflicts of interest. This article does not contain any studies involving human participants or animals performed by any of the authors.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Balint, V., Stanisavljevic Ninkovic, D., Anastasov, N. et al. Inhibition of miR-21 Promotes Cellular Senescence in NT2-Derived Astrocytes. Biochemistry Moscow 86, 1434–1445 (2021). https://doi.org/10.1134/S0006297921110079
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0006297921110079