Abstract
Irreversible neuron loss caused by central nervous system injuries usually leads to persistent neurological dysfunction. Reactive astrocytes, because of their high proliferative capacity, proximity to neuronal lineage, and significant involvement in glial scarring, are ideal starting cells for neuronal regeneration. Having previously identified several small molecules as important regulators of astrocyte-to-neuron reprogramming, we established herein that miR-124, ruxolitinib, SB203580, and forskolin could co-regulate rat cortical reactive astrocyte-to-neuron conversion. The induced cells had reduced astroglial properties, displayed typical neuronal morphologies, and expressed neuronal markers, reflecting 25.9% of cholinergic neurons and 22.3% of glutamatergic neurons. Gene analysis revealed that induced neuron gene expression patterns were more similar to that of primary neurons than of initial reactive astrocytes. On the molecular level, miR-124-driven neuronal differentiation of reactive astrocytes was via targeting of the SOX9-NFIA-HES1 axis to inhibit HES1 expression. In conclusion, we present a novel approach to inducing endogenous rat cortical reactive astrocytes into neurons through co-regulation involving miR-124 and three small molecules. Thus, our research has potential implications for inhibiting glial scar formation and promoting neuronal regeneration after central nervous system injury or disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
Functional neuron loss caused by central nervous system (CNS) injuries or diseases is usually irreversible and destructive [1, 2]. Upon injury stimulation, glial cells, especially astrocytes, proliferate, migrate and become reactive astrocytes (RAs) characterised by morphological changes (hypertrophy and process extension) and the upregulation of the RA markers GFAP and S100 [3, 4]. Moreover, the production of chondroitin sulphate proteoglycans and pro-inflammatory cytokine release further drives RAs to eventually form glial scars that impair neuronal regeneration [5, 6]. Since the regenerative capacity of the CNS is limited [7], it seems meaningful to eliminate the neurosuppressive effects of glial scars and restore neural function. Fortunately, cell reprogramming using somatic cells to produce functional neurons addresses these issues [8]. Astrocytes, the predominant glial cells in the mammalian CNS play vital roles in mediating neurotransmitter homeostasis and synaptic plasticity [9, 10]. Like neurons, they originate from neural stem cells (NSCs), and blocking astrocytic signals and activating neurophenotypic signals can promote transitions of astrocyte-to-neuron states without lineage switching [11]. As autogenous cells, astrocytes eliminate the possibility of immunorejection [12]; furthermore, neuronal conversion enables new local neural circuit formation and functional recovery. Thus, astrocytes are the most promising candidates for neuronal reprogramming.
Transcription factors (TFs) such as NEUROG2, ASCL1 and DLX2 can directly reprogram astrocytes into neurons [13]. However, ectopic viral TF expression in astrocytes can lead to gene mutations and chromosomal changes with tumorigenic risks. Moreover, the extremely complex injection process required to safely and effectively deliver viruses into the CNS hinders their applications [14]. Alternatively, the biological effects of small molecules are fine-tuneable and reversible; thus, their development into pharmaceuticals can provide broad application prospects for neuronal reprogramming without genetic modification [15]. The small molecule cocktail of VPA, CHIR99021, RepSox, forskolin, i-BET151 and ISX-9 reprogrammed adult astrocytes into mainly glutamatergic neurons [16], and foetal astrocyte-derived functional neurons could be obtained via mediation by a small molecule cocktail of SB431542, LDN193189, CHIR99021 and 2,4-diamino-5-phenylthiazole (DAPT) [17]. However, since the signals activated or inhibited by each small molecule differ, inconsistent chemical combinations lead to variable neuronal properties and conversion efficiencies. Additionally, the glutamatergic neurons produced in previous studies could not fully meet the needs of CNS injuries or diseases of other subtypes. Therefore, simplifying chemical compositions, improving conversion efficiency, inducing other subtype-specific neurons, and exploring more regulatory signals are critical for astrocyte-to-neuron reprogramming.
Combining small molecules with other agents to synergistically improve reprogramming effects has also garnered much attention. MicroRNAs (miRs) are small noncoding ribonucleic acids (RNAs) (20–22 nucleotides) that bind to the 3′ untranslated region of target messenger RNAs (mRNAs) to trigger translational repression or degradation [18] and can mediate direct neuronal reprogramming in vitro and in vivo [19, 20]. MiR-124, one of the most abundant characteristic miRs in adult mammalian brains [21], is predominantly expressed in neurons, and its expression gradually increases with neuronal maturation [22]. Moreover, miR-124 is a neuronal fate determinant for the subventricular zone, which mediates neurogenesis in adult mammalian brains. MiR-124 overexpression can induce neuron-specific differentiation and govern the dendritic plasticity of NSCs [23, 24]; in an ischaemic injury model, miR-124 administration could promote cortical neurogenesis [25]. Additionally, fibroblasts or HeLa cells with increasing miR-124 levels showed an expression profile shift toward that of a neuronal phenotype [26, 27]. This suggests that miR-124 not only regulates physiological and pathological neuronal differentiation of NSCs but also enables some non-nervous system cells to acquire neuronal identities. However, whether miR-124 has similar neuromodulative effects on RAs has not yet been assessed. Repressing the RNA-binding protein, PTBP1 has recently been reported to mediate astrocyte-to-dopaminergic neuron conversion in a Parkinson’s disease (PD) mouse model [28, 29]. We suspect that as one of the targets of miR-124 [30], Ptbp1 of RAs may be targeted by miR-124 to promote their neuronal differentiation.
Here, we present a novel RA-to-neuron reprogramming strategy using miR-124, ruxolitinib, SB203580 and forskolin, to inhibit RA glial scar formation and promote neuronal regeneration. Astrocytes were stimulated by exogenous TGF-β1 treatment to obtain in vitro RAs [31]. Among small molecules, in addition to forskolin, which was previously reported to reduce lipid peroxidation, change cell morphology and promote neuronal conversion efficiency [32, 33], we also used ruxolitinib and SB203580, inhibitors of the JAK/STAT and P38 MAPK pathways, respectively, which are involved in astrogliosis [34, 35], but have not been used before for neuronal reprogramming. Considering that miR-124 drives neuronal differentiation by inhibiting NOTCH signalling via the SOX9-NFIA-HES1 axis, we verified that simultaneous modulation of NOTCH, JAK/STAT and P38 MAPK signals is sufficient to convert rat cortex RAs into mainly cholinergic and glutamatergic neurons in vitro, opening new avenues for developing future CNS repair therapies.
Methods
Isolation and Culture of Primary Cortical RAs
All animals were supplied by the Experimental Animal Centre of the Jilin Yisi Company. All animal experiments were performed in accordance with the National Institute of Health Guidelines for the Care and Use of Laboratory Animals and were approved by the Ethics Committee of Jilin University.
Primary cortical astrocytes were isolated from neonatal Wistar rats (1–3 days old). Briefly, after removal of the blood vessels and pia mater, cerebral cortices were dissected and digested using papain (100 units, 40.4 mg/mL; Worthington, LS003127), L-cysteine (0.0036–0.0042 g/mL) and DNAse I (4 mg/mL; Worthington, LS002006) at 34°C; after 45 min, the reaction was terminated using 20% foetal bovine serum (FBS) (Sigma) and 10X Earle’s balanced salt solution (Sigma, E7510). Cells were passed through a 40 μm filter (Greiner, 542040) to obtain a single-cell suspension and plated at a density of 20×105 cells/T25 flask in RA-conditioned medium (AM) (Dulbecco’s modified Eagle’s medium/F12 [DMEM/F12] [Corning, 10-0920CVR] containing 10% FBS, 10 ng/mL TGF-β1 [PeproTech, 96-100-21-10] and 1% penicillin-streptomycin [Invitrogen, 15140122]) in a 5% CO2 humidified incubator at 37°C. Culture medium was refreshed once every 2 days. About 1 week later, when cultured cells had reached 90% confluence, flasks were moved to a shaker at 37°C (280 rpm/min, 10 h/day) for 2 days. Thereafter, 20 μM cytarabine (Sigma, C1768) was added for 24 h to obtain high purity RAs [31]. Subsequent experiments were performed when GFAP-positive cells exceeded 95% of the total cell number.
Isolation and Culture of Primary Rat Cortical Neurons
Single-cell suspensions of neonatal Wistar rat (1–3 days old) cortices were obtained by the method described for primary cortical astrocyte isolation. Subsequently, cells were plated (at a density of 20×105 cells/poly-D-lysine hydrobromide [Sigma, P6407] pre-coated T25 flask) in neuron-conditioned medium (NM) (Neurobasal [Gibco, 10888022] supplemented with 5% FBS, 1% B27 (Invitrogen, 17504044), 10 ng/mL BDNF, 15 ng/mL GDNF, 1% L-glutamine [Gibco, 25030081] and 1% penicillin-streptomycin) in a 5% CO2 humidified incubator at 37°C. After 12 h, new serum-free medium was added for another 12 h. Then, 10-μM cytarabine was added for 48 h. The culture medium was refreshed once every 2 days until the 10th day of culture to harvest high purity rat neurons [36, 37].
MiR-124 Transfection of Rat RAs
Purified RAs were seeded at a density of 5×105 cells/well in six-well plates for about 24 h before transfection. Thereafter, cells were transfected either with various concentrations of ron-miR-124-3p mimic, a double-stranded RNA synthesised by a chemical method that can effectively and significantly elevate the exogenous mature miR-124-3p content in cells to enhance its regulatory effects, or with the corresponding negative control (RiboBio, China) using lipofectamine RNAiMAX (Invitrogen, 13778075) [38]. The cells were harvested at different time points after transfection for further detection of miRs, mRNAs and proteins, or for the following differentiation experiments.
RNAi Transfection of Rat RAs
Cell preparation before RNAi transfection was identical to that described for miR-124 transfection of RAs. Cells were transfected with 50 nM RNAi oligonucleotides (si-r-Sox9, si-r-Nfia and si-r-Hes1) or negative control siRNA (RiboBio, China) using lipofectamine RNAiMAX. After 24 h, the medium was replaced by AM. Subsequently, cells were harvested at different time points for further experiments.
Neuronal Induction from Rat Cortical RAs
After ron-miR-124-3p mimic transfection of RAs for 24 h, the supernatant was switched to induced culture medium (DMEM/F12 medium supplemented with 2% FBS, 1% N2, 1 μM ruxolitinib [Selleck, S1378], 10 μM SB203580 [Selleck, S1076], 10 μM forskolin [Selleck, S2449], 10 ng/mL BDNF [PeproTech, 450-02-10], 15 ng/mL GDNF [PeproTech, 450-51-10] and 1% penicillin-streptomycin). The culture medium was refreshed once every 2 days for 1 week for further experiments or switched to NM until the third week [16, 17].
Immunofluorescence Staining
After harvesting, all culture samples were fixed in 4% paraformaldehyde for 20 min; permeabilised with 0.2% Triton X-100 in phosphate-buffered saline (PBS) for 20 min; and then incubated with 10% goat serum (Boster, AR0009) in PBS at room temperature for 1 h. After blocking, samples were incubated with primary antibodies (Online Resource 1) overnight at 4°C. After extensive washing, secondary antibodies (Online Resource 1) in PBS were added for 1 h in the dark, followed by nuclear labelling with Hoechst33342 for 15 min at room temperature [39]. Five randomly sampled images were captured for each culture using a laser scanning microscope (Olympus) and analysed using ImageJ software.
Quantitative Reverse Transcription-Polymerase Chain Reaction (qRT-PCR) Analysis
Total miR and mRNA were extracted using a miR extraction kit (QIAGEN, 217004) and Trizol (Invitrogen, 15596026). Reverse transcription of 1000 ng miRs from the total RNA was performed using the All-in-One miRNA first-strand cDNA Synthesis kit (GeneCopoeia, QP014); mRNA reverse transcription was performed using the TransScript-Uni One-Step gDNA Removal and cDNA Synthesis SuperMix (Transgen Biotech, AU311). MiR and mRNA PCR amplifications were performed on the 7300 Real-Time PCR System using the All-in-One miRNA qPCR Detection kit (GeneCopoeia, QP011) and NovoStart SYBR qPCR SuperMix Plus kit (Novoprotein, E096-01B), respectively. Expression levels of rat U6 and GAPDH were used as internal controls for miRs and mRNAs; values were calculated as the reverse log of the −ΔΔCT from controls [40]. All assays were performed at least thrice. The primers used for the experiments (Online Resource 2) were synthesised by Shanghai Sangon Biotech (Shanghai, China).
Western Blotting
Total protein (20 mg) was loaded onto 10% or 12% sodium dodecyl sulphate (SDS)-polyacrylamide gel electrophoresis (PAGE) gels and transferred to polyvinylidene fluoride membranes (Pall, BSP0161), which were blocked with 5% fat-free milk in Tris-buffered saline containing 0.1% Tween-20 (TBST) for 1 h at room temperature, followed by incubation with primary antibodies (Online Resource 3) at 4°C overnight. GAPDH was used as an internal control. After thorough washes with TBST, the membranes were incubated with peroxidase-conjugated secondary antibodies (Online Resource 3) at room temperature for 1 h. After washing, the signal was detected by chemiluminescence using the BeyoECL Plus (Beyotime, P0018) detection system and then analysed using ImageJ [41].
EdU Incorporation Assay
EdU incorporation and EdU staining were performed with slight modifications from a previous report [42]. Briefly, 24 h prior to fixation, 10 μM EdU were added on the 2nd, 4th and 6th day of induction. Cells were fixed by 4% paraformaldehyde for 30 min and permeabilised with 0.5% Triton X-100 in PBS at room temperature. The following EdU staining was performed following the manufacturer’s instructions of the EdU cell proliferation detection kit (RiboBio, C10310-1). Subsequently, cells were detected with DCX immunofluorescence staining. Images were captured and analysed with ImageJ software.
RNA-Sequencing (RNA-seq) Library Preparation and Data Analysis
Samples were sent to Sangon Biotech Co., Ltd. (Shanghai, China) for library construction, Illumina sequencing, data pre-processing and gene mapping. Total RNA extraction was performed using the Total RNA Extractor kit (Sangon Biotech, Shanghai). After RNA purity, concentration and integrity assessment, 2 μg high-quality RNA per sample were used as input material for RNA sample preparations. Sequencing libraries were generated using VAHTSTM mRNA-seq V2 Library Prep Kit for Illumina®, and index codes were added to attribute sequences to each sample. Subsequently, FastQC (version 0.11.2) was used for evaluating the quality of the sequenced data. Raw reads were filtered using Trimmomatic (version 0.36) and the obtained sequence reads were trimmed and mapped to the reference genome using HISAT2 (version 2.1.0), followed by RNA-seq assessment. StringTie (version 1.3.3b) was used to calculate the transcripts per million (TPM) of protein-coding genes in each sample. For samples with three biological replicates, DESeq (version 1.12.4) was used to obtain the significant differentially expressed genes (sDEGs). sDEGs between the two groups were identified using TPM and q-values (TPM ≥ 5 in at least one sample, |FoldChange|> 2, q-value < 0.05). Hierarchical cluster analysis was performed to show distinguishable mRNA expression profiles among the samples. Scatterplot analysis was conducted to depict the mRNA expression distribution. Gene ontology (GO) functional enrichment analysis, which provides all GO terms significantly enriched in differentially expressed genes (DEGs) comparing to the genome background, was performed using TopGO (version 2.24.0). Kyoto Encyclopedia of Genes and Genomes (KEGG) functional enrichment analysis, which identified significantly enriched metabolic pathways or signal transduction pathways in DEGs comparing with the whole genome background, was performed using clusterProfiler (version 3.0.5) [43].
Co-immunoprecipitation
An appropriate amount of immunoprecipitation lysis buffer (Beyotime, P0013) supplemented with protease inhibitor was added, and the resultant cell lysates were subject to immunoprecipitation with a specific antibody or IgG control (Beyotime, P7028) and protein A/G agarose beads (Beyotime, P2012) at 4°C overnight, followed by thorough washes with lysis buffer. Subsequently, the bound proteins were eluted by heating the beads in 2% SDS gel-loading buffer at 95°C for 5 min. Eluted proteins were resolved by SDS-PAGE and then detected by western blotting [44].
Statistical Analysis
GraphPad Prism 7.0 was used for all statistical analyses. The unpaired t-test was used for two-group comparisons, such as the GFAP and S100 expression after miR-124 overexpression in RAs, and SOX9, NFIA and HES1 mRNA/protein comparisons during the RA-to-neuron differentiation regulated by miR-124. The one-way ANOVA was used for multiple-group comparisons, such as the RA-to-neuron conversion efficiency of different sets of miR-124 and small molecules, and the HES1 protein comparisons of different groups of miR-124, small molecules and DAPT. The two-way ANOVA was performed to analyse the transfection efficiency of miR-124 in RAs. All experiments were repeated at least thrice. All data are represented as the mean ± standard deviation (SD). For the various analyses, *p< 0.05, **p< 0.01 and ***p< 0.001 were considered statistically significant.
Results
Identification and Characterisation of RAs from the Postnatal Rat Cortex
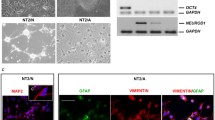
To avoid neural progenitor cell (NPC) contamination, only cerebral cortex regions where NPCs were not abundant were used. Cultured cells were expanded in medium containing TGF-β1, which stimulated astrocytes to become RAs [31]. For about 1 week, until the cells reached 90% confluence, cytarabine was added to remove proliferating oligodendrocytes and microglia. After purification, most cells were typically star-shaped or polygonal, producing branching processes from the cell bodies (Fig. 1a). More than 95% of the cells expressed the RA markers GFAP and S100 without DCX or MAP2 positive neuronal contamination (Fig. 1b–d). This suggested that the cultured cells were mainly RAs.
Morphological and biochemical characteristics of primary rat cortical RAs. a Purification processes of primary RAs and corresponding cell morphologies at different time points in vitro. Scale bars, 50 μm. b and c Immunostaining shows that the majority of cultured cells expressed the RA markers, GFAP and S100, but not the neuronal markers DCX and MAP2. Scale bars, 50 μm. d Statistical analysis shows that >95% of all cells were GFAP- and S100-positive RAs, but no DCX- or MAP2-positive cells were detected. Data are reported as mean ± SD; n = 5
Characterisation of RAs Overexpressing miR-124
Since we found that miR-124 expression in astrocytes, especially RAs, was much lower than that in neurons (Online Resource 4), if the RAs miR-124 expression artificially increased, it may regulate RAs neuronal differentiation. Subsequently, to create RAs overexpressing miR-124, we transfected RAs with a miR-124 mimic (miR-124) or a negative control (NC-124). After a 48 h transfection, cell bodies in the miR-124 group became smaller, and their branching processes slenderer compared with those in the NC-124 group (Fig. 2a). The qRT-PCR analysis demonstrated that miR-124 expression increased. The highest transfection efficiency was with 50 nM miR-124 for 48 h (about 23.4 times higher than in the control) (Fig. 2b and 2c). As expected, miR-124 overexpression reduced mRNA levels of the RA markers GFAP and S100 (Fig. 2d). Additionally, GFAP and S100 protein expression was also downregulated after miR-124 overexpression (Fig. 2e–h), which was consistent with the mRNA levels. These results suggested that we had successfully constructed miR-124 overexpressing RAs and that miR-124 overexpression suppressed the original astrocytic characteristics of RAs.
Overexpression of miR-124 in RAs attenuated astrocytic characteristics. a Phase-contrast images of RAs transfected with miR-124 mimic (miR-124) or negative control (NC-124) at 48 h. Notably, miR-124 group cells have smaller territories and more complex morphologies than NC-124 group cells. Scale bars, 50 μm. b and c qRT-PCR analysis after miR-124 overexpression at different concentrations and time points. The optimal transfection efficiency was obtained at 50 nM for 48 h. Data are reported as mean ± SD; n = 3; **P< 0.01, ***P< 0.001. d Expression of astrocytic markers in miR-124/NC-124 groups analysed using qRT-PCR. MiR-124 overexpression significantly downregulates Gfap and S100 mRNA expression in RAs. Data are reported as mean ± SD; n = 3; *P< 0.05. e and f Western blot (e) and densitometric quantitation (f) of GFAP and S100 in miR-124/NC-124 groups; signals were normalised to GAPDH. Data are reported as mean ± SD; n = 3; *P< 0.05, ***P< 0.001. g and h Immunostaining showing GFAP (g) and S100 (h) expression in the miR-124/NC-124 groups. Scale bar, 50 μm
RAs Synergistically Treated with miR-124, Ruxolitinib, SB203580 and Forskolin Acquired Neuronal Properties
Although miR-124 inhibited RA characteristics, the RA-to-neuron fate transition requires co-regulation of multiple signals [17]. We then treated RAs overexpressing miR-124 with ruxolitinib, SB203580 and forskolin (Fig. 3a). This synergistic combination dramatically converted RA morphology into neuron-like shapes after a 7-day treatment. The induced neuronal cell (iN) bodies became smaller and more compact and formed slender axon-like branching structures; no significant morphological changes were observed in the NC-124 control group (Fig. 3b). Moreover, miR-124 group cells had smaller territory sizes and longer branching processes (Fig. 3c and 3d). Immunostaining demonstrated that early neuronal markers DCX and TUJ1 and mature neuronal markers MAP2 and NEUN were detectable in the miR-124 group on the 7th day of induction (Fig. 3e–g); the neurosynaptic junction marker SYN1 was detectable on the 21st day (Fig. 3h). Based on neuronal marker expression and cell morphologies, the conversion efficiencies for DCX, TUJ1, MAP2, NEUN and SYN1 were approximately 41.5%, 64.4%, 38.7%, 87.3% and 0.7%, respectively (Fig. 3m). With regard to neuronal subtypes, our iNs were predominantly CHAT+ (25.9%) cholinergic and VGLUT1+ (22.3%) glutamatergic, occasionally TH+ (6.1%) dopaminergic, but rarely GAD67+ (1.4%) inhibitory neurons on the 7th day of induction (Fig. 3i–l and 3n). Additionally, qRT-PCR analysis also corroborated that gene transcripts corresponding to cholinergic neuron differentiation (e.g. Chat, Lhx8, Lhx6 and Tbr1) were significantly increased in these iNs (Online Resource 5). These data suggested that rat cortical RAs acquired a neuronal fate after synergistic miR-124 and small molecules treatment, mainly forming cholinergic and glutamatergic neurons.
Conversion of RAs into iNs by miR-124 and small molecules under defined conditions. a Schematic diagram showing the neuronal induction protocol. AM, RA conditioned medium; NM, neuron conditioned medium; R, ruxolitinib; S, SB203580; F, forskolin. b Phase-contrast images showing the change from astrocytic to neuronal morphology along the induction process. Bright-field images of RAs induced with NC-124/miR-124 and small molecules at day 7. Scale bars, 20 μm. c and d Statistical analysis of the total branch length (c) and total territory size (d) of iNs (miR-124 group) and RAs (NC-124 group). Territory size is defined as the two-dimensional area that is delineated by the border of the cell body. Data are reported as mean ± SD; n = 10; ***P< 0.001. e–h Immunostaining showing that iNs were positive for TUJ1, MAP2 (e), DCX (f) and NEUN (g) at day 7 and SYN1 (h) at day 21. Scale bars, 50 μm. i–l Immunostaining with different neuronal subtype markers reveals that iNs were mainly cholinergic (CHAT) (i), glutamatergic (VGLUT1) neurons (j) and rarely dopaminergic (TH) (k) or GABAergic (GAD67) (l) neurons. Scale bars, 50 μm. (m and n) Quantification of neuronal conversion efficiencies (m) and neuronal subtypes (n) was performed. Data are reported as mean ± SD; n = 5
Subsequently, to understand how the iNs converted from RAs, we performed an EdU incorporation experiment on our iNs on the 3rd, 5th and 7th day of induction. The results demonstrated that most of our iNs had no proliferative ability (Online Resource 6a and 6b), although some iNs with potential NPC proliferation characteristics could be occasionally observed during the conversion process, suggesting that in our neuronal environment, most iNs might not pass through the NPC stage, but the possibility was not completely ruled out that a few iNs might first have undergone the NPC stage and then differentiated and matured.
MiR-124, Ruxolitinib, SB203580 and Forskolin Were Indispensable for RA-to-Neuron Conversion
To explore whether the same induction effects could be achieved using fewer components, different sets of miR-124 and small molecules were used for induction. The small molecules alone and miR-124 combined with at least one small molecule promoted the generation of DCX-positive neurons at day 7, but conversion efficiency was the highest when miR-124 and the three small molecules were added synergistically (Fig. 4a–j). MAP2 immunostaining results were also consistent (Fig. 4k–t). As JAK/STAT3, P38 MAPK and ERK1/2 signalling is involved in RA migration, astrogliosis and proliferation [45,46,47], increased phosphorylation levels would enhance astrocyte activation. Therefore, we assessed the phosphorylation levels in the iNs. Western blot results at day 7 of induction revealed that phosphorylation levels were most downregulated when miR-124 and the three small molecules were added synergistically (Fig. 4u and v); this also reduced RA reactivity and benefited neuronal transdifferentiation the most.
Regulation of RA-to-neuron conversion by miR-124 and the three small molecules. a–j Representative profiles of immunostaining (a–i) at day 7 and the percentage (j) of DCX-positive cells after induction with different combinations of miR-124, ruxolitinib, SB203580 and forskolin. The conversion efficiency of miR-124 and the three small molecules treated group was higher than that of the other groups. Data are reported as mean ± SD; n = 5; **P< 0.01. Scale bars, 50 μm. n.d., not detectable. k–t Representative profiles of immunostaining (k–s) at day 7 and the percentage (t) of MAP2-positive cells after induction with different combinations. The conversion efficiency of miR-124 and the three small molecules treated group was higher than that of the other groups. Data are reported as mean ± SD; n = 5; *P< 0.05. Scale bars, 50 μm. n.d., not detectable. u and v Western blot (u) for measurement of phosphorylated STAT3 (p-STAT3), phosphorylated P38 MAPK (p-P38), phosphorylated ERK1/2 (p-ERK1/2), STAT3, P38 and ERK1/2 in RAs after induction with different combinations for 72 h. GAPDH was used as a loading control. Quantification (v) was done by normalisation of p-STAT3 (p-P38 or p-ERK1/2) over total STAT3 (total P38 or total ERK1/2). Data are reported as mean ± SD; n = 3; *P< 0.05
Interestingly, when we added SB431542, an inhibitor of the TGF-β pathway (whose inhibition is related to neuronal conversion), to the cocktail of miR-124 and three small molecules, no significant increase in DCX-positive conversion efficiency was observed (Online Resource 7a–c). These results suggested that the synergistic effects of miR-124 and the three small molecules best promoted RA-to-neuron conversion.
Gene Expression Profile of iNs Was Closer to That of Cortical Neurons Than RAs
To examine the differences between RAs, iNs and rat neurons at the genetic level, we first isolated rat cortical neurons and maintained them in a defined, serum-free NM. At day 10 after purification, the cells exhibited a typical neuronal morphology (Fig. 5a). Most cells were immune-positive for TUJ1 and MAP2 (Fig. 5b), suggesting that the isolated cells were mainly neurons. As miR-124 expression gradually increases during neuronal differentiation and maturation [48], we assessed miR-124 expression in iNs. At day 7 of induction, total miR-124 expression was 23.9 times higher than that in the control group (Fig. 5c). Considering that exogenous miR-124 decreases gradually (Fig. 2c), the total miR-124 overexpression was mainly caused by increased endogenous miR-124, further indicating RA-to-neuron conversion. To identify the gene expression profiles of RAs, iNs (day 7) and rat neurons (day 10), RNA-seq analyses were performed. The number of sDEGs between iNs and RAs (3065) was more than that between iNs and rat neurons (2695) (Fig. 5d), indicating fewer differences between the latter two. Consistently, hierarchical cluster analysis indicated that the iNs’ gene expression profile was closer to that of rat neurons than RAs (Fig. 5e), a similar finding to that of qRT-PCR (Fig. 5f). These results suggested that RAs silenced their astrocytic transcriptional programs and adopted a neuronal identity during induction.
iNs gene expression profile resembled that of rat cortical neurons. a Bright-field images of rat cortical neurons showing typical neuronal morphology at day 10. Scale bars, 50 μm. (b) Immunostaining showing that cultured rat cortical neurons expressed the neuronal markers TUJ1 and MAP2. Scale bars, 20 μm. (c) qRT-PCR analysis of miR-124 expression in iNs at day 7 after induction. Mainly endogenous miR-124 upregulation. Data are reported as mean ± SD; n = 3; ***P< 0.001. (d) Venn diagram indicating a bigger difference of sDEGs between iNs and RAs than between iNs and rat neurons. (e) Heatmap illustration showing more similar global expression profiles of iNs to rat neurons than to RAs. (f) qRT-PCR analysis of neuronal markers’ gene expression in RAs, iNs and rat neurons. iNs gene expression was closer to rat neurons. Data are reported as mean ± SD; n = 3; *P< 0.05, **P< 0.01, ***P< 0.001
As the induction efficiency was incomplete, we were interested in whether prolonged induction time or environmental factors would affect the fate of the remaining RAs. Therefore, we cultured the 21-day iNs in AM and NM. On the third day of continuous culturing, no morphological changes were observed. On the 11th day, cell morphology in NM was still unchanged, similar to that of rat neurons. However, cell numbers in AM increased, and the morphology was similar to that of RAs (Online Resource 8a), which might be because serum exposure allowed the remaining RAs to proliferate. These results indicated that prolonged exposure and a neuronal environment could not change the fate of non-neuronal transformed RAs. Subsequently, we assessed the gene expression of these cells, RAs and rat neurons. The genetic pattern of cells in AM resembled that of RAs, and that of cells in NM resembled that of rat neurons (Online Resource 8b).
Transcriptome Analysis of iNs and RAs
To explore the molecular and cellular changes of iNs compared with their initial cells, we further analysed the transcriptome of iNs and RAs. At day 7 of induction, the transcriptome scatterplot analysis revealed that, of the 3065 sDEGs, 1820 were upregulated and 1245 were downregulated (Fig. 6a). The expression of RA-enriched genes (e.g. Gfap, Aldh1l1, Vim, Nes, Slit2, Axin2 and Mmp2) was decreased, while that of neuron-enriched genes (e.g. Dcx, Tuj1, Syt1, Kcnc1, Map2, Syn1 and Ascl1) was significantly increased. Additionally, the expression of genes involved in brain development or cholinergic neuron differentiation (e.g. Foxg1, Bcl11b and Myt1l) was also significantly upregulated. Further, qRT-PCR results confirmed expression patterns of the selected genes, which were consistent with the RNA-seq results (Fig. 6b). The GO analysis (biological process, cellular component and molecular function) showed that the upregulated genes were significantly enriched in neuron projection morphogenesis, neuron differentiation, axonogenesis and synaptic vesicle (Fig. 6c), and the downregulated genes were significantly enriched in cell proliferation, cell migration, cell-substrate adhesion and proteoglycan binding (Fig. 6d), suggesting that the astrogliosis and scar formation properties of RAs had weakened, while neuronal characteristics had increased. Additionally, KEGG analysis revealed that upregulated cyclic guanosine monophosphate-dependent protein kinase G and cyclic adenosine monophosphate (cAMP) signalling, soluble NSF attachment protein receptor interactions in vesicular transport and other upregulated signals were mainly involved in neuron development, axon formation and neurotransmitter release (Fig. 6e), while downregulated MAPK signalling, cell adhesion molecules, PI3K-AKT signalling and other downregulated signals were related to astrogliosis and glial scarring of RAs (Fig. 6f), further pointing to RA-to-neuron conversion. Together, the transcriptional regulatory network of RAs was reprogrammed toward that of neuronal lineage under our induction.
Comparison of transcriptome analysis between iNs and RAs. a Scatter plots comparing gene expression levels between iNs (day 7) and RAs. Significantly upregulated genes are highlighted in red, significantly downregulated genes in green and non-differential genes in black. Dashed line indicates TPM ≥ 5 in at least one sample, |FoldChange|> 2 and q-value < 0.05. b qRT-PCR validation of the expression of representative RA- and neuron-enriched genes between RAs and iNs. Data are reported as mean ± SD; n = 3; *P< 0.05, **P< 0.01, ***P< 0.001. c and d GO enrichment analysis of upregulated/downregulated genes. The upregulated genes were significantly enriched in neuronal functions, and the downregulated genes were significantly involved in RA functions. e and f KEGG enrichment analysis of upregulated/downregulated genes. The upregulated genes were related to neuronal signals, and the downregulated genes were involved in RA signals
MiR-124 Regulated Neuronal Differentiation of RAs via the SOX9-NFIA-HES1 Axis
To clarify the molecular mechanism involved in the regulatory role of miR-124, we focused on the NOTCH downstream effectors SOX9 and NFIA, characteristic RA TFs [44], as well as HES1, whose oscillations are related to neurogenesis [49]. MiR-124 overexpression notably repressed SOX9, NFIA and HES1 expression (Fig. 7a and b), indicating the regulatory effect of miR-124 on them. Considering that Sox9 is a target of miR-124 [50], we explored the relationship between SOX9, NFIA and HES1 in RAs. After transfection with si-Sox9 (Online Resource 9a), Sox9 mRNA and protein levels were significantly reduced (Fig. 7c–e). Moreover, NFIA and HES1 expression was also downregulated with reductions in SOX9 levels (Fig. 7d and e), suggesting that SOX9 regulated NFIA and HES1. Additionally, a physical association between SOX9 and NFIA confirmed by co-immunoprecipitation further indicated that SOX9 directly downregulated NFIA (Fig. 7f). Subsequently, we transfected RAs with si-Nfia (Online Resource 9b), and Nfia mRNA level and protein expression were remarkably decreased (Fig. 7g–i). Moreover, HES1 also decreased upon reduced levels of NFIA (Fig. 7h and i), which reduces HES1 expression by binding to the Hes1 promoter region [51], suggesting the direct repression of NFIA on HES1. Altogether, miR-124 could downregulate HES1 expression by targeting the SOX9-NFIA-HES1 axis in RAs. To explore whether this axis was related to neurodifferentiation, we transfected RAs with si-Hes1 (Online Resource 9c), which significantly downregulated Hes1 mRNA expression (Fig. 7j–l). Furthermore, the mRNA expression of Ascl1, a gene related to neuronal differentiation, was significantly elevated by Hes1 silencing (Fig. 7j), revealing the negative regulation of Hes1 in neuronal differentiation. Simultaneously, we detected the effect of Sox9 or Nfia inhibition on Ascl1. The results showed that Sox9 or Nfia inhibition could also increase Ascl1 mRNA expression (Online Resource 9d), but not as much as Hes1 inhibition, further indicating the indispensable role of Hes1 in this regulatory axis. These results suggested that in RAs, miR-124 inhibited NOTCH signalling by targeting the SOX9-NFIA-HES1 axis, thereby promoting RA-to-neuron differentiation.
MiR-124 regulated HES1 expression via the SOX9-NFIA-HES1 axis in RAs. a and b Western blot (a) and densitometric quantitation (b) for SOX9, NFIA and HES1 expression in RAs after miR-124 overexpression at 96 h. Data are reported as mean ± SD; n = 3; *P< 0.05. c The expression of Sox9 mRNA was assessed using qRT-PCR in RAs transfected with si-Sox9 or si-control (Ctrl) for 48 h. Data are reported as mean ± SD; n = 3; ***P< 0.001. d and e The levels of SOX9, NFIA and HES1 in si-Sox9 or Ctrl transfected RAs were determined by western blot (d) at 96 h, and quantification (e) was performed. Data are reported as mean ± SD; n = 3; *P< 0.05, **P< 0.01. f Co-immunoprecipitation shows that SOX9 and NFIA physically interacted in RAs. g mRNA expression of Nfia in si-Nfia or Ctrl transfected RAs analysed by qRT-PCR at 48 h. Data are reported as mean ± SD; n = 3; ***P< 0.001. h and i Expression of NFIA and HES1 was assessed by western blot in RAs transfected with si-Nfia or Ctrl for 96 h. Data are reported as mean ± SD; n = 3; *P< 0.05. j Expression of Hes1 and Ascl1 at the mRNA level was analysed by qRT-PCR in RAs transfected with si-Hes1 or Ctrl for 48 h. Data are reported as mean ± SD; n = 3; **P< 0.01, ***P< 0.001. k and l Western blot analysis (k) and quantitation (l) by densitometry for HES1 expression of RAs transfected with si-Hes1 or Ctrl at 96 h was performed. All protein expression levels were normalised to GAPDH. Data are reported as mean ± SD; n = 3; *P< 0.05
HES1 Was Involved in the RA-to-Neuron Conversion Co-regulated by miR-124, Ruxolitinib, SB203580 and Forskolin
To decipher whether the SOX9-NFIA-HES1 axis operated during the induction after the small molecules were added, we first induced the aforementioned si-Hes1 RAs with the three small molecules for 3 days, and compared their HES1 expression with that in RAs, RAs treated with the three small molecules only, and RAs treated with both miR-124 and the three small molecules. Western blotting showed that HES1 expression was significantly suppressed in the miR-124 and small molecules group (Fig. 8a and b). Immunostaining for DCX demonstrated that conversion efficiency in the miR-124 and small molecules group was better than those in other groups (Fig. 8e–i). Given that the miR-124 and small molecules group had the lowest HES1 expression but the best conversion efficiency, we speculated whether HES1 expression determined differences in conversion efficiency. We then applied DAPT, a NOTCH pathway inhibitor, to further suppress HES1 expression. For better effects, we also prolonged induction time to 7 days. As expected, DAPT application further suppressed HES1 expression in the miR-124 and small molecules group (Fig. 8c and d), but conversion efficiency was still lower than that in the miR-124 and small molecules group without added DAPT (Fig. 8j–n), demonstrating that apart from suppressing HES1 expression, the combination might also regulate neuronal conversion by other means. These results suggested that HES1 expression was involved in RA-to-neuron conversion co-regulated by the miR-124 and small molecule cocktail, but this was not the only mechanism.
The RA-to-neuron conversion co-regulated by miR-124 and small molecules was associated with HES1 expression. a and b Western blot (a) and densitometric quantitation (b) of HES1 expression in RAs, RAs treated with small molecules for 3 days, RAs treated with si-Hes1 and small molecules for 3 days and RAs treated with miR-124 and small molecules for 3 days. Data are reported as mean ± SD; n = 3; *P< 0.05. c and d Western blot (c) and quantitation (d) by densitometry for HES1 expression in RAs treated with miR-124 and small molecules for 7 days, RAs treated with miR-124, small molecules and DAPT for 7 days, RAs treated with small molecules and DAPT for 7 days and RAs treated with miR-124 and DAPT for 7 days. All protein expression levels were normalised to GAPDH. Data are reported as mean ± SD; n = 3; *P< 0.05. e–n Immunostaining (e–h, j–m) and the percentage (i, n) of DCX-positive cells in RAs, RAs after induction with miR-124/si-Hes1/DAPT and/or small molecules for 3/7 days at different combinations. Data are reported as mean ± SD; n = 5; *P< 0.05, **P< 0.01. Scale bars, 50 μm. n.d., not detectable
Discussion
Recently, the use of diverse small molecule cocktails to reprogram astrocytes into neuronal cells has achieved considerable success [16, 17]. Yin et al. proved that using three to four small molecules (SB431542, CHIR99021, LDN193189 and DAPT) or their respective functional analogues to modulate the TGF-β/SMAD, BMP, NOTCH and WNT pathways was sufficient to reprogram human foetal astrocytes into neurons [17]. However, RA characteristics are also regulated by other signals, such as the JAK/STAT and P38 MAPK pathways and post-transcriptional levels of several noncoding RNAs. Here, we first developed and reported a new miR-124, ruxolitinib, SB203580 and forskolin cocktail, proving that this combination could significantly alter RA characteristics and promote RA-to-neuron conversion.
Studies have revealed the functional significance of miRs in regulating CNS development [52]. MiR-124, one of the most highly expressed miRs in the mammalian brain, is closely associated with the regulation of neurogenesis and neuronal differentiation [53]. Given that both astrocytes and neurons are derived from NSCs and that miR-124 is a neuronal fate determinant during neurodevelopment [54], we hypothesised that differential miR-124 expression might affect their distinct cell fates. Fortunately, we found that miR-124 expression in RAs was much lower than that in neurons. Moreover, GFAP, S100, SOX9 and NFIA expression was significantly downregulated after miR-124 overexpression, indicating that miR-124 upregulation could efficiently suppress inherent RA characteristics and might promote RA-to-neuron conversion. Additionally, as each miR regulates hundreds of target genes [55], and Smad4 (involved in both the TGF-β and BMP pathways), Dact1 and Axin1 (negative regulators of the WNT/β-catenin cascade) are all targets of miR-124 [56,57,58,59], we hypothesised that miR-124 might replace TGF-β and BMP pathway inhibitors as well as WNT/β-catenin pathway activators in RAs and implement functions similar to those of the SB431542, LDN193189 and CHIR99021 combination during RA-to-neuron reprogramming. As expected, when we randomly selected SB431542 to add to the cocktail of miR-124 and three small molecules, no increased DCX-positive iNs were obtained. Considering its numerous targets, miR-124 might promote RA-to-neuron conversion through other unknown mechanisms, and eventually, the effects of our cocktail were better than of the above three chemicals. However, miR-124 itself was not sufficient to reprogram RAs into neurons in vitro, suggesting that signals other than TGF-β, BMP and WNT, such as JAK/STAT [3] and P38 MAPK [46], that are crucial for maintaining RA characteristics, might be constantly activated and impair the transformation process. We thus hypothesised that additional inhibition of these signals might achieve more significant conversion efficiency.
JAK/STAT pathway activation is reported to be involved in CNS injuries, especially gliogenesis and glial scarring formation [45, 60]. Moreover, Jak3 silencing or Stat3 ablation promoted NPC neuronal differentiation and NSC neurogenesis, respectively [61, 62], revealing the neuronal promotion of JAK/STAT inhibition. Therefore, we speculated that JAK/STAT pathway suppression offers great possibilities for inhibiting inherent RA properties and promoting RA-to-neuron conversion. As expected, the conversion efficiency was significantly increased after treatment with ruxolitinib and miR-124. To the best of our knowledge, this is the first time a JAK/STAT pathway inhibitor has been used for neuronal induction.
P38 MAPK and ERK1/2 activation are also associated with astrogliosis [41]. During reactive gliosis, ASK1 phosphorylation is enhanced, and P38 MAPK relays the signal to astrocytes for activation and glial scar formation both in vitro and in vivo [35, 63]. Thus, suppressing P38 MAPK signalling was more likely to attenuate RA characteristics and favour neuronal induction. Here, we utilised SB203580, a common P38 inhibitor, to repress RA characteristics and promote neuronal conversion. Furthermore, forskolin is a well-known cAMP and PRKACA activator and promotes neuron survival and neuronal conversion efficiency [33, 64, 65]. As expected, the three small molecules promoted RA-to-neuron conversion, albeit inefficiently. However, when miR-124 was also added, an efficient conversion was obtained, further indicating the importance of combinatory functions.
Although the functional mechanisms of the three small molecules are clear, the function of miR-124 is more complicated. Here, we mainly focus on miR-124 and NOTCH signalling, a highly conserved morphogenic pathway that regulates NSC proliferation, self-renewal and neuronal differentiation [66]. Attenuated NOTCH1 signalling after stroke is necessary for striatal astrocyte neurogenesis in mice [67], indicating that RA-to-neuron conversion is related to NOTCH signalling. Here, we revealed that miR-124 promotes neuronal differentiation of RAs by targeting the SOX9-NFIA-HES1 axis and eventually elevates the expression of Ascl1 (Fig. 9), a proneural target gene of Hes1 that promotes neural differentiation [68]. This was consistent with previous studies showing that expression of the neural TF ASCL1 alone could reprogram astrocytes into neurons [69]. Nevertheless, persistent miR-124 overexpression alone did not produce DCX- or MAP2-positive RA-derived neurons. This might be because neurodifferentiation is promoted only when proneural activity is high enough [70]. Although Ascl1 was upregulated in RAs, the levels were not high enough for neuronal conversion. Moreover, HES1 expression disappears in differentiating neurons, leading to sustained ASCL1 expression, inducing neuronal differentiation [71]. In our study, miR-124 indirectly inhibited HES1 but failed to eliminate its expression. Thereafter, when the small molecules were added to regulate additional RA signals, we obtained DCX- and MAP2-positive iNs; however, when DAPT was added to further suppress HES1 expression in RAs, no more DCX-positive iNs were obtained. This might be because HES1 is a repressor-type basic helix-loop-helix protein, while ASCL1 is of the activator-type. When NOTCH signalling is inhibited, decreased HES1 expression upregulates ASCL1 to simultaneously promote neuronal fate determination and NOTCH ligands expression, activating NOTCH signalling and upregulating HES1 expression in neighbouring cells to maintain them in an undifferentiated state [72]. Therefore, HES1 and ASCL1 regulate each other, only allowing some cells to undergo neuronal differentiation. This may also explain why the remaining undifferentiated RAs in our study could not acquire a neuronal identity and increase the RA-to-neuron conversion efficiency even with prolonged induction. Of course, the synergy between miR-124 and the other three small molecules is extremely complex, and our analysis stems from the limited perspective of HES1 expression alone. However, from our RNA-seq results and previous reports, we speculate that an upregulated cAMP pathway, downregulated PI3K-AKT pathway, as well as the miR-124 target PTBP1 [28, 29] may also be involved in this conversion. Certainly, whether this synergism regarding RA-to-neuron conversion could operate through these mechanisms needs to be further explored.
Proposed mechanistic model for miR-124-driven neuronal differentiation of RAs. MiR-124 overexpression in RAs indirectly inhibited the expression of the NOTCH downstream target HES1 by targeting the SOX9-NFIA-HES1 axis, thereby reducing the inhibitory effect of HES1 on ASCL1 and ultimately promoting the neuronal differentiation of RAs
The neuronal subtypes after conversion from astrocytes depend on the combinations of inducing chemicals. Previous nine- and four-molecule cocktails mainly converted astrocytes into glutamatergic neurons through upregulation of NEUROD1 and NGN2 [17, 73]. Here, our RNA-seq results showed that Foxg1 and Myt1l mRNA expression was significantly upregulated. FOXG1 is one of the earliest TFs specifically induced in forebrain NPCs [74]. MYT1L, the only known TF specifically expressed in all neurons throughout life, is critical for maintaining neuronal identity [75, 76]. Thus, our induction modulated RAs to obtain a neuronal identity via upregulating Foxg1 and Myt1l. Interestingly, unlike with other protocols, many cholinergic neurons appeared in our experiments. The cholinergic hypothesis of Alzheimer’s disease (AD) proposes that brain acetylcholine depletion is a primary cause of cognitive decline in advanced age and AD [77, 78]. Moreover, cholinergic nuclei are differently affected by the pathogenetic mechanisms underlying major neurodegenerative diseases such as AD, PD and Huntington disease [77]. Given that astrocyte reactivity is a hallmark of the pathological CNS, RA-derived cholinergic neurons might be potential targets for the treatment of neurodegenerative diseases. Recently, Liang et al. reported that MYT1L could directly reprogram human brain vascular pericytes into cholinergic neurons [79], indicating the Myt1l expression in our experiments might be essential for converting and maintaining the cholinergic neuron phenotype. Although these mechanisms remain to be further explored, we believe that the different effects depended on our unique induction cocktail.
Glial scars formed by RAs after CNS injuries ultimately impair neuronal regeneration; therefore, ameliorating the cellular properties of glial scars is essential for neural repair. Although 41.5% RAs in this study were successfully converted into DCX-positive neurons, with continuous miR-124 and small molecules treatment, the remaining undifferentiated RAs would lose their reactivity to form glial scars, and further contact the iNs to support their maturation. Collectively, utilising our cocktail simultaneously achieved the two goals of reprogramming RAs into neurons and repressing RA cellular characteristics, which is significant for the treatment of CNS injuries.
Although the use of small molecules to reprogram cells has achieved remarkable success, they should be nontoxic and approved after clinical safety trials. Ruxolitinib, the first clinically effective, selective JAK1/2 inhibitor approved by the Food and Drug Administration, can penetrate the blood-brain barrier and reduce astrogliosis in mice with HIV encephalitis when systemically administered [80]. Additionally, losmapimod, a P38 MAPK α and β isoform inhibitor functionally analogous to SB203580, has entered stage I clinical trials for treating facioscapulohumeral muscular dystrophy. Forskolin is a natural compound used for centuries in traditional medicine. These chemicals selected for our formula or their functional analogues are closer to clinical application and more feasible for use.
Conclusions
In summary, if the in situ RAs within pathological lesions are artificially reprogrammed into neurons, RA’s harmful function, such as dense glial scar formation, will be effectively eliminated. Furthermore, RA-derived neurons will further connect with surrounding neurons to form circuits, thus promoting neuronal regeneration and functional recovery. To our knowledge, this is the first report showing that the combination of miR-124, ruxolitinib, SB203580 and forskolin could successfully reprogram RAs into neurons. Our induction cocktail has four advantages: (1) the chemical combination is novel and innovative; (2) the chemical composition includes only four elements; (3) our iNs are mainly cholinergic and glutamatergic neurons, providing more alternative neuronal subtypes; and (4) the selected small molecules are closer to clinical applications and more feasible to use. Although further investigations are required to prove whether or not this combination can effectively reprogram human RAs into neurons and have a similar conversion efficiency in vivo, herein, we explored a novel RA-to-neuron reprogramming approach and investigated its effectiveness and mechanisms in vitro, providing new clues for inhibiting glial scars and repairing CNS damage.
Data and Materials Availability
The datasets used and/or analysed during the current study are available from the corresponding author (Guangfan Chi, guangfan130@jlu.edu.cn) on reasonable request.
References
Ouyang W, Yan Q, Zhang Y, Fan Z (2017) Moderate injury in motor-sensory cortex causes behavioral deficits accompanied by electrophysiological changes in mice adulthood. PLoS One 12(2):e0171976. https://doi.org/10.1371/journal.pone.0171976
Bhowmick S, D'Mello V, Ponery N, Abdul-Muneer PM (2018) Neurodegeneration and sensorimotor deficits in the mouse model of traumatic brain injury. Brain Sci 8(1). https://doi.org/10.3390/brainsci8010011
Okada S, Hara M, Kobayakawa K, Matsumoto Y, Nakashima Y (2018) Astrocyte reactivity and astrogliosis after spinal cord injury. Neurosci Res 126:39–43. https://doi.org/10.1016/j.neures.2017.10.004
Burda JE, Sofroniew MV (2014) Reactive gliosis and the multicellular response to CNS damage and disease. Neuron 81(2):229–248. https://doi.org/10.1016/j.neuron.2013.12.034
Wang H, Song G, Chuang H, Chiu C, Abdelmaksoud A, Ye Y, Zhao L (2018) Portrait of glial scar in neurological diseases. Int J Immunopathol Pharmacol 31:2058738418801406. https://doi.org/10.1177/2058738418801406
Moeendarbary E, Weber IP, Sheridan GK, Koser DE, Soleman S, Haenzi B, Bradbury EJ, Fawcett J et al (2017) The soft mechanical signature of glial scars in the central nervous system. Nat Commun 8:14787. https://doi.org/10.1038/ncomms14787
McMurran CE, Kodali S, Young A, Franklin RJ (2018) Clinical implications of myelin regeneration in the central nervous system. Expert Rev Neurother 18(2):111–123. https://doi.org/10.1080/14737175.2018.1421458
Tsunemoto R, Lee S, Szucs A, Chubukov P, Sokolova I, Blanchard JW, Eade KT, Bruggemann J et al (2018) Diverse reprogramming codes for neuronal identity. Nature 557(7705):375–380. https://doi.org/10.1038/s41586-018-0103-5
Kanakov O, Gordleeva S, Ermolaeva A, Jalan S, Zaikin A (2019) Astrocyte-induced positive integrated information in neuron-astrocyte ensembles. Phys Rev E 99(1-1):012418. https://doi.org/10.1103/PhysRevE.99.012418
Farhy-Tselnicker I, Allen NJ (2018) Astrocytes, neurons, synapses: a tripartite view on cortical circuit development. Neural Dev 13(1):7. https://doi.org/10.1186/s13064-018-0104-y
Buganim Y, Faddah DA, Jaenisch R (2013) Mechanisms and models of somatic cell reprogramming. Nat Rev Genet 14(6):427–439. https://doi.org/10.1038/nrg3473
Li H, Chen G (2016) In Vivo Reprogramming for CNS repair: regenerating neurons from endogenous glial cells. Neuron 91(4):728–738. https://doi.org/10.1016/j.neuron.2016.08.004
Heinrich C, Gotz M, Berninger B (2012) Reprogramming of postnatal astroglia of the mouse neocortex into functional, synapse-forming neurons. Methods Mol Biol 814:485–498. https://doi.org/10.1007/978-1-61779-452-0_32
Su Z, Niu W, Liu ML, Zou Y, Zhang CL (2014) In vivo conversion of astrocytes to neurons in the injured adult spinal cord. Nat Commun 5:3338. https://doi.org/10.1038/ncomms4338
Lee C, Robinson M, Willerth SM (2018) Direct reprogramming of glioblastoma cells into neurons using small molecules. ACS Chem Neurosci 9(12):3175–3185. https://doi.org/10.1021/acschemneuro.8b00365
Gao L, Guan W, Wang M, Wang H, Yu J, Liu Q, Qiu B, Yu Y et al (2017) Direct generation of human neuronal cells from adult astrocytes by small molecules. Stem Cell Rep 8(3):538–547. https://doi.org/10.1016/j.stemcr.2017.01.014
Yin JC, Zhang L, Ma NX, Wang Y, Lee G, Hou XY, Lei ZF, Zhang FY et al (2019) Chemical conversion of human fetal astrocytes into neurons through modulation of multiple signaling pathways. Stem Cell Rep 12(3):488–501. https://doi.org/10.1016/j.stemcr.2019.01.003
Lagos-Quintana M, Rauhut R, Lendeckel W, Tuschl T (2001) Identification of novel genes coding for small expressed RNAs. Science 294(5543):853–858. https://doi.org/10.1126/science.1064921
Abernathy DG, Kim WK, McCoy MJ, Lake AM, Ouwenga R, Lee SW, Xing X, Li D et al (2017) MicroRNAs induce a permissive chromatin environment that enables neuronal subtype-specific reprogramming of adult human fibroblasts. Cell Stem Cell 21(3):332–348.e339. https://doi.org/10.1016/j.stem.2017.08.002
Ghasemi-Kasman M, Shojaei A, Gol M, Moghadamnia AA, Baharvand H, Javan M (2018) miR-302/367-induced neurons reduce behavioral impairment in an experimental model of Alzheimer's disease. Mol Cell Neurosci 86:50–57. https://doi.org/10.1016/j.mcn.2017.11.012
Mishima T, Mizuguchi Y, Kawahigashi Y, Takizawa T, Takizawa T (2007) RT-PCR-based analysis of microRNA (miR-1 and -124) expression in mouse CNS. Brain Res 1131 (1):37-43. doi:https://doi.org/10.1016/j.brainres.2006.11.035
Sun Y, Luo ZM, Guo XM, Su DF, Liu X (2015) An updated role of microRNA-124 in central nervous system disorders: a review. Front Cell Neurosci 9:193. https://doi.org/10.3389/fncel.2015.00193
Jiao S, Liu Y, Yao Y, Teng J (2018) miR-124 promotes proliferation and neural differentiation of neural stem cells through targeting DACT1 and activating Wnt/beta-catenin pathways. Mol Cell Biochem 449(1-2):305–314. https://doi.org/10.1007/s11010-018-3367-z
Xue Q, Yu C, Wang Y, Liu L, Zhang K, Fang C, Liu F, Bian G et al (2016) miR-9 and miR-124 synergistically affect regulation of dendritic branching via the AKT/GSK3beta pathway by targeting Rap2a. Sci Rep 6:26781. https://doi.org/10.1038/srep26781
Yang J, Zhang X, Chen X, Wang L, Yang G (2017) Exosome mediated delivery of miR-124 promotes neurogenesis after ischemia. Mol Ther Nucleic Acids 7:278–287. https://doi.org/10.1016/j.omtn.2017.04.010
Yoo AS, Sun AX, Li L, Shcheglovitov A, Portmann T, Li Y, Lee-Messer C, Dolmetsch RE et al (2011) MicroRNA-mediated conversion of human fibroblasts to neurons. Nature 476(7359):228–231. https://doi.org/10.1038/nature10323
Lim LP, Lau NC, Garrett-Engele P, Grimson A, Schelter JM, Castle J, Bartel DP, Linsley PS et al (2005) Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 433(7027):769–773. https://doi.org/10.1038/nature03315
Zhou H, Su J, Hu X, Zhou C, Li H, Chen Z, Xiao Q, Wang B et al (2020) Glia-to-Neuron Conversion by CRISPR-CasRx Alleviates symptoms of neurological disease in mice. Cell 181(3):590–603.e516. https://doi.org/10.1016/j.cell.2020.03.024
Qian H, Kang X, Hu J, Zhang D, Liang Z, Meng F, Zhang X, Xue Y et al (2020) Reversing a model of Parkinson’s disease with in situ converted nigral neurons. Nature 582(7813):550–556. https://doi.org/10.1038/s41586-020-2388-4
Mokabber H, Najafzadeh N, Mohammadzadeh Vardin M (2019) miR-124 promotes neural differentiation in mouse bulge stem cells by repressing Ptbp1 and Sox9. J Cell Physiol 234(6):8941–8950. https://doi.org/10.1002/jcp.27563
Lee HS, Lee SH, Cha JH, Seo JH, Ahn BJ, Kim KW (2015) Meteorin is upregulated in reactive astrocytes and functions as a negative feedback effector in reactive gliosis. Mol Med Rep 12(2):1817–1823. https://doi.org/10.3892/mmr.2015.3610
Gascón S, Murenu E, Masserdotti G, Ortega F, Russo GL, Petrik D, Deshpande A, Heinrich C et al (2016) Identification and successful negotiation of a metabolic checkpoint in direct neuronal reprogramming. Cell Stem Cell 18(3):396–409. https://doi.org/10.1016/j.stem.2015.12.003
Liu ML, Zang T, Zou Y, Chang JC, Gibson JR, Huber KM, Zhang CL (2013) Small molecules enable neurogenin 2 to efficiently convert human fibroblasts into cholinergic neurons. Nat Commun 4:2183. https://doi.org/10.1038/ncomms3183
You T, Bi Y, Li J, Zhang M, Chen X, Zhang K, Li J (2017) IL-17 induces reactive astrocytes and up-regulation of vascular endothelial growth factor (VEGF) through JAK/STAT signaling. Sci Rep 7:41779. https://doi.org/10.1038/srep41779
Li T, Xu R, Xia H, Hu X, Wang S, Li Y, Yan Y, Xia Y (2020) ASK1 phosphorylation regulates astrocytic reactive gliosis in vitro and in vivo. Neurosci Lett 716:134675. https://doi.org/10.1016/j.neulet.2019.134675
Koito H, Li J (2009) Preparation of rat brain aggregate cultures for neuron and glia development studies. J Vis Exp 31. https://doi.org/10.3791/1304
Xu SY, Wu YM, Ji Z, Gao XY, Pan SY (2012) A modified technique for culturing primary fetal rat cortical neurons. J Biomed Biotechnol 2012:803930–803937. https://doi.org/10.1155/2012/803930
Xu J, Lv S, Hou Y, Xu K, Sun D, Zheng Y, Zhang Z, Li X et al (2018) miR-27b promotes type II collagen expression by targetting peroxisome proliferator-activated receptor-γ2 during rat articular chondrocyte differentiation. Biosci Rep 38(1). https://doi.org/10.1042/bsr20171109
Ma K, Deng X, Xia X, Fan Z, Qi X, Wang Y, Li Y, Ma Y et al (2018) Direct conversion of mouse astrocytes into neural progenitor cells and specific lineages of neurons. Transl Neurodegener 7:29. https://doi.org/10.1186/s40035-018-0132-x
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 29(9):e45, 45e–e445. https://doi.org/10.1093/nar/29.9.e45
Wang CY, Yang SH, Tzeng SF (2015) MicroRNA-145 as one negative regulator of astrogliosis. Glia 63(2):194–205. https://doi.org/10.1002/glia.22743
Mao J, Huang S, Liu S, Feng XL, Yu M, Liu J, Sun YE, Chen G et al (2015) A herbal medicine for Alzheimer's disease and its active constituents promote neural progenitor proliferation. Aging Cell 14(5):784–796. https://doi.org/10.1111/acel.12356
Hernández-Montiel W, Collí-Dula RC, Ramón-Ugalde JP, Martínez-Núñez MA, Zamora-Bustillos R (2019) RNA-seq transcriptome analysis in ovarian tissue of Pelibuey breed to explore the regulation of prolificacy. Genes 10(5). https://doi.org/10.3390/genes10050358
Kang P, Lee HK, Glasgow SM, Finley M, Donti T, Gaber ZB, Graham BH, Foster AE et al (2012) Sox9 and NFIA coordinate a transcriptional regulatory cascade during the initiation of gliogenesis. Neuron 74(1):79–94. https://doi.org/10.1016/j.neuron.2012.01.024
Ben Haim L, Ceyzeriat K, Carrillo-de Sauvage MA, Aubry F, Auregan G, Guillermier M, Ruiz M, Petit F et al (2015) The JAK/STAT3 pathway is a common inducer of astrocyte reactivity in Alzheimer's and Huntington's diseases. J Neurosci 35(6):2817–2829. https://doi.org/10.1523/jneurosci.3516-14.2015
Roy Choudhury G, Ryou MG, Poteet E, Wen Y, He R, Sun F, Yuan F, Jin K et al (2014) Involvement of p38 MAPK in reactive astrogliosis induced by ischemic stroke. Brain Res 1551:45–58. https://doi.org/10.1016/j.brainres.2014.01.013
Jia M, Shi Z, Yan X, Xu L, Dong L, Li J, Wang Y, Yang S et al (2018) Insulin and heparin-binding epidermal growth factor-like growth factor synergistically promote astrocyte survival and proliferation in serum-free medium. J Neurosci Methods 307:240–247. https://doi.org/10.1016/j.jneumeth.2018.06.002
Smirnova L, Grafe A, Seiler A, Schumacher S, Nitsch R, Wulczyn FG (2005) Regulation of miRNA expression during neural cell specification. Eur J Neurosci 21(6):1469–1477. https://doi.org/10.1111/j.1460-9568.2005.03978.x
Ochi S, Imaizumi Y, Shimojo H, Miyachi H, Kageyama R (2020) Oscillatory expression of Hes1 regulates cell proliferation and neuronal differentiation in the embryonic brain. Development 147(4):dev182204. https://doi.org/10.1242/dev.182204
Zhou F, Zhang C, Guan Y, Chen Y, Lu Q, Jie L, Gao H, Du H et al (2018) Screening the expression characteristics of several miRNAs in G93A-SOD1 transgenic mouse: altered expression of miRNA-124 is associated with astrocyte differentiation by targeting Sox2 and Sox9. J Neurochem 145(1):51–67. https://doi.org/10.1111/jnc.14229
Piper M, Barry G, Hawkins J, Mason S, Lindwall C, Little E, Sarkar A, Smith AG et al (2010) NFIA controls telencephalic progenitor cell differentiation through repression of the Notch effector Hes1. J Neurosci 30(27):9127–9139. https://doi.org/10.1523/jneurosci.6167-09.2010
Cao DD, Li L, Chan WY (2016) MicroRNAs: key regulators in the central nervous system and their implication in neurological diseases. Int J Mol Sci 17(6). https://doi.org/10.3390/ijms17060842
Kutsche LK, Gysi DM, Fallmann J, Lenk K, Petri R, Swiersy A, Klapper SD, Pircs K et al (2018) Combined experimental and system-level analyses reveal the complex regulatory network of miR-124 during human neurogenesis. Cell Syst 7(4):438–452.e438. https://doi.org/10.1016/j.cels.2018.08.011
Akerblom M, Sachdeva R, Barde I, Verp S, Gentner B, Trono D, Jakobsson J (2012) MicroRNA-124 is a subventricular zone neuronal fate determinant. J Neurosci 32(26):8879–8889. https://doi.org/10.1523/jneurosci.0558-12.2012
Guarnieri DJ, DiLeone RJ (2008) MicroRNAs: a new class of gene regulators. Ann Med 40(3):197–208. https://doi.org/10.1080/07853890701771823
Zhang Z, Gong Q, Li M, Xu J, Zheng Y, Ge P, Chi G (2017) MicroRNA-124 inhibits the proliferation of C6 glioma cells by targeting Smad4. Int J Mol Med 40(4):1226–1234. https://doi.org/10.3892/ijmm.2017.3088
Zu L, Xue Y, Wang J, Fu Y, Wang X, Xiao G, Hao M, Sun X et al (2016) The feedback loop between miR-124 and TGF-beta pathway plays a significant role in non-small cell lung cancer metastasis. Carcinogenesis 37(3):333–343. https://doi.org/10.1093/carcin/bgw011
Jiao S, Liu Y, Yao Y, Teng J (2018) miR-124 promotes proliferation and neural differentiation of neural stem cells through targeting DACT1 and activating Wnt/β-catenin pathways. Mol Cell Biochem 449(1-2):305–314. https://doi.org/10.1007/s11010-018-3367-z
Lu Y, Zhang T, Shan S, Wang S, Bian W, Ren T, Yang D (2019) MiR-124 regulates transforming growth factor-beta1 induced differentiation of lung resident mesenchymal stem cells to myofibroblast by repressing Wnt/beta-catenin signaling. Dev Biol 449(2):115–121. https://doi.org/10.1016/j.ydbio.2019.02.010
Tapia VS, Herrera-Rojas M, Larrain J (2017) JAK-STAT pathway activation in response to spinal cord injury in regenerative and non-regenerative stages of Xenopus laevis. Regeneration (Oxf) 4(1):21–35. https://doi.org/10.1002/reg2.74
Kim YH, Chung JI, Woo HG, Jung YS, Lee SH, Moon CH, Suh-Kim H, Baik EJ (2010) Differential regulation of proliferation and differentiation in neural precursor cells by the Jak pathway. Stem Cells 28(10):1816–1828. https://doi.org/10.1002/stem.511
Cao F, Hata R, Zhu P, Nakashiro K, Sakanaka M (2010) Conditional deletion of Stat3 promotes neurogenesis and inhibits astrogliogenesis in neural stem cells. Biochem Biophys Res Commun 394(3):843–847. https://doi.org/10.1016/j.bbrc.2010.03.092
Cheon SY, Cho KJ, Song J, Kim GW (2016) Knockdown of apoptosis signal-regulating kinase 1 affects ischaemia-induced astrocyte activation and glial scar formation. Eur J Neurosci 43(7):912–922. https://doi.org/10.1111/ejn.13175
Smith DK, Yang J, Liu ML, Zhang CL (2016) Small molecules modulate chromatin accessibility to promote NEUROG2-mediated fibroblast-to-neuron reprogramming. Stem Cell Rep 7(5):955–969. https://doi.org/10.1016/j.stemcr.2016.09.013
Gascon S, Murenu E, Masserdotti G, Ortega F, Russo GL, Petrik D, Deshpande A, Heinrich C et al (2016) Identification and successful negotiation of a metabolic checkpoint in direct neuronal reprogramming. Cell Stem Cell 18(3):396–409. https://doi.org/10.1016/j.stem.2015.12.003
Roese-Koerner B, Stappert L, Brustle O (2017) Notch/Hes signaling and miR-9 engage in complex feedback interactions controlling neural progenitor cell proliferation and differentiation. Neurogenesis (Austin, Tex) 4(1):e1313647. https://doi.org/10.1080/23262133.2017.1313647
Magnusson JP, Goritz C, Tatarishvili J, Dias DO, Smith EM, Lindvall O, Kokaia Z, Frisen J (2014) A latent neurogenic program in astrocytes regulated by Notch signaling in the mouse. Science 346(6206):237–241. https://doi.org/10.1126/science.346.6206.237
Sueda R, Imayoshi I, Harima Y, Kageyama R (2019) High Hes1 expression and resultant Ascl1 suppression regulate quiescent vs. active neural stem cells in the adult mouse brain. Genes Dev 33(9-10):511–523. https://doi.org/10.1101/gad.323196.118
Liu Y, Miao Q, Yuan J, Han S, Zhang P, Li S, Rao Z, Zhao W et al (2015) Ascl1 converts dorsal midbrain astrocytes into functional neurons in vivo. J Neurosci 35(25):9336–9355. https://doi.org/10.1523/jneurosci.3975-14.2015
Boareto M, Iber D, Taylor V (2017) Differential interactions between Notch and ID factors control neurogenesis by modulating Hes factor autoregulation. Development 144(19):3465–3474. https://doi.org/10.1242/dev.152520
Imayoshi I, Kageyama R (2014) bHLH factors in self-renewal, multipotency, and fate choice of neural progenitor cells. Neuron 82(1):9–23. https://doi.org/10.1016/j.neuron.2014.03.018
Kageyama R, Ohtsuka T, Hatakeyama J, Ohsawa R (2005) Roles of bHLH genes in neural stem cell differentiation. Exp Cell Res 306(2):343–348. https://doi.org/10.1016/j.yexcr.2005.03.015
Zhang L, Yin JC, Yeh H, Ma NX, Lee G, Chen XA, Wang Y, Lin L et al (2015) Small molecules efficiently reprogram human astroglial cells into functional neurons. Cell Stem Cell 17(6):735–747. https://doi.org/10.1016/j.stem.2015.09.012
Cargnin F, Kwon JS, Katzman S, Chen B, Lee JW, Lee SK (2018) FOXG1 orchestrates neocortical organization and cortico-cortical connections. Neuron 100 (5):1083-1096.e1085. https://doi.org/10.1016/j.neuron.2018.10.016
Mall M, Kareta MS, Chanda S, Ahlenius H, Perotti N, Zhou B, Grieder SD, Ge X et al (2017) Myt1l safeguards neuronal identity by actively repressing many non-neuronal fates. Nature 544(7649):245–249. https://doi.org/10.1038/nature21722
Matsushita F, Kameyama T, Kadokawa Y, Marunouchi T (2014) Spatiotemporal expression pattern of Myt/NZF family zinc finger transcription factors during mouse nervous system development. Dev Dyn 243(4):588–600. https://doi.org/10.1002/dvdy.24091
Pepeu G, Grazia Giovannini M (2017) The fate of the brain cholinergic neurons in neurodegenerative diseases. Brain Res 1670:173–184. https://doi.org/10.1016/j.brainres.2017.06.023
Douchamps V, Mathis C (2017) A second wind for the cholinergic system in Alzheimer's therapy. Behav Pharmacol 28(2 and 3-Spec Issue):112–123. https://doi.org/10.1097/fbp.0000000000000300
Liang XG, Tan C, Wang CK, Tao RR, Huang YJ, Ma KF, Fukunaga K, Huang MZ et al (2018) Myt1l induced direct reprogramming of pericytes into cholinergic neurons. CNS Neurosci Ther 24(9):801–809. https://doi.org/10.1111/cns.12821
Haile WB, Gavegnano C, Tao S, Jiang Y, Schinazi RF, Tyor WR (2016) The Janus kinase inhibitor ruxolitinib reduces HIV replication in human macrophages and ameliorates HIV encephalitis in a murine model. Neurobiol Dis 92(Pt B):137–143. https://doi.org/10.1016/j.nbd.2016.02.007
Acknowledgements
We thank Shanghai Sangon Biotech Co., Ltd., for RNA-seq analysis and Guangzhou RiboBio Co., Ltd., for designing the miR mimics and si-RNA. We also thank Editage for English language editing.
Funding
This work was supported by the National Natural Science Foundation of China (Grant numbers 81571199 and 81870974) and the Fundamental Research Funds for the Central Universities, JLU. The funding body had no role in the design of the study, the collection, analysis and interpretation of data and in the writing of the manuscript.
Author information
Authors and Affiliations
Contributions
GC, CQ and YL conceived the study, designed the experiments, and together with YZ revised the manuscript. YZ performed the cellular and molecular experiments in vitro, with help from ZH and JX. KH, YY, SL and LC provided reagents and conducted the data analysis and interpretation. YZ wrote the manuscript. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Ethics Approval
The animal study has been reviewed and approved by the Ethics Committee of Jilin University (Ethical approval code: 2015-19).
Competing Interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zheng, Y., Huang, Z., Xu, J. et al. MiR-124 and Small Molecules Synergistically Regulate the Generation of Neuronal Cells from Rat Cortical Reactive Astrocytes. Mol Neurobiol 58, 2447–2464 (2021). https://doi.org/10.1007/s12035-021-02345-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-021-02345-6