Abstract
Four single nucleotide polymorphism (SNP)-based human leukocyte antigen (HLA) imputation methods (e-HLA, HIBAG, HLA*IMP:02 and MAGPrediction) were trained using 1000 Genomes SNP and HLA genotypes and assessed for their ability to accurately impute molecular HLA-A, -B, -C and –DRB1 genotypes in the Human Genome Diversity Project cell panel. Imputation concordance was high (>89%) across all methods for both HLA-A and HLA-C, but HLA-B and HLA-DRB1 proved generally difficult to impute. Overall, <27.8% of subjects were correctly imputed for all HLA loci by any method. Concordance across all loci was not enhanced via the application of confidence thresholds; reliance on confidence scores across methods only led to noticeable improvement (+3.2%) for HLA-DRB1. As the HLA complex is highly relevant to the study of human health and disease, a standardized assessment of SNP-based HLA imputation methods is crucial for advancing genomic research. Considerable room remains for the improvement of HLA-B and especially HLA-DRB1 imputation methods, and no imputation method is as accurate as molecular genotyping. The application of large, ancestrally diverse HLA and SNP reference data sets and multiple imputation methods has the potential to make SNP-based HLA imputation methods a tractable option for determining HLA genotypes.
Similar content being viewed by others
Introduction
Located on the short arm of chromosome 6p21, the human major histocompatibility complex (MHC) contains 226 genes with pivotal roles in the immune system. These include the human leukocyte antigen (HLA) genes, which have been extensively studied as central determinants of allogeneic transplantation success. More than 100 infectious, autoimmune, inflammatory diseases and cancers are associated with HLA variation.1 Furthermore, HLA genes have been associated with a number of immunologically mediated drug interactions. For example, HLA-B*57:01, DR7 and DQ3 are associated with hypersensitivity to the HIV/AIDS antiviral drug Abacavir,2, 3 HLA-B*58:01 is associated with adverse reactions to the chronic gout treatment allopurinol,4 and HLA-A*15:02, HLA-A*31:01 is associated with hypersensitivity to the epilepsy and neuropathic pain medication carbamazepine.5 Knowledge of patients’ HLA genotypes will help exclude those at risk of drug reactions that confer considerable morbidity and mortality.6 The HLA genes are highly polymorphic, with 15 635 allelic variants identified as of October 2016, and a variety of PCR-based HLA genotyping methods have been applied to identify specific HLA alleles.7
Although genome-wide association studies (GWAS) have identified genetic association signals for many common diseases,8, 9, 10 the structural complexity, high polymorphism and extensive linkage disequilibrium (LD) that characterize the MHC11, 12 have posed challenges for the interpretation of GWAS in this region. Although many of the strongest associations revealed to-date by GWAS with disease1, 13 and drug-induced hypersensitivity2, 3, 4, 5 are in the MHC, these associations have generally identified non-coding single nucleotide polymorphisms (SNPs), which are primarily related to gene function through LD.14 When association signals have been identified in the vicinity of HLA genes, the complexity of HLA polymorphism and the cost of molecular HLA genotyping have often limited efforts to fine-map causal HLA variants.7 The appreciation that individual SNPs, SNP haplotypes and other genetic markers are in strong LD with specific HLA alleles15, 16 has motivated the development of methods for the imputation of HLA genotypes from SNP genotypes, with the goal of interpreting associations identified within the MHC region17, 18, 19 in light of HLA allelic variation. These HLA imputation methods have also been applied to existing SNP data to confirm findings based on molecular HLA genotyping.5, 11
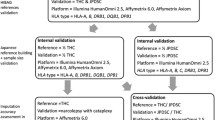
Although HLA imputation has primarily been evaluated in cohorts of European ancestry15 (and in non-Europeans to a lesser extent), no studies of multiple HLA imputation methods, applied to a worldwide range of populations, have been performed. Here, we describe the results from the ImmPute project, a consortium effort evaluating four HLA imputation methods (ensemble-based HLA prediction (e-HLA) (described in Supplementary Information), HLA Genotype Imputation with Attribute Bagging (HIBAG),17 HLA*IMP:02 (ref. 19) and Multi-allelic Gene Prediction (MAGPrediction)18). Each method was applied to impute HLA genotypes using SNP genotypes in the Human Genome Diversity Project (HGDP)20 cell panel after being trained on HLA and SNP genotypes in phase one 1000 Genomes (1000G) Project samples21 alone, and the results evaluated for accuracy and performance against HLA genotypes determined through standard molecular methods. The only variable in this approach is the applied imputation method, allowing the unobstructed comparison of method-specific variations in imputation outcome.
Materials and methods
MHC SNPS
A total of 12 352 extended MHC (xMHC; chr6: 26 000 000–36 000 000; genome build HG19/GRCh37) SNPs were obtained from two sources for 889 HGDP cell panel subjects. In total, 11 149 MHC SNPs were extracted from the UCLA Medical Center Illumina Immunochip22 HGDP Dataset 15 (ftp://ftp.cephb.fr/hgdp_supp15/), and additional 1203 MHC SNPs were extracted from the Stanford HGDP SNP Genotyping Dataset 2 (http://www.hagsc.org/hgdp/files.html). A total of 164 876 xMHC SNPs for the 1000G samples were extracted from whole-genome sequence data from the phase one 1000G Project repository21 using VCF tools.23 In total, 10 268 SNPs common to both data set were used for this study.
HLA genotyping
Sequence-based molecular HLA genotyping (SBT) was performed for the HLA class I (HLA-A, -B, -C) and class II (HLA-DRB1) genes in the 1000G samples as previously described (PMID: 24988075) HGDP HLA genotypes were generated using reverse-format sequence-specific oligonucleotide probe typing methods as previously described.24 The HLA-A, HLA-C, HLA-B and HLA-DRB1 loci were typed using Roche linear-array strips. In both methods, immobilized SSO probes, selected for maximum discriminating power between alleles in a given IMGT/HLA Database nomenclature epoch, are hybridized to locus-specific PCR products. Exons 2 and 3 were amplified and assessed for each of the HLA-A, HLA-C and HLA-B loci and exon 2 was amplified and assessed for HLA-DRB1. Historically, and in particular for transplantation, these are the four most commonly typed HLA loci.7 The HGDP and KG data sets were genotyped independently and at different, but overlapping loci. HLA-A, -C, -B, -DRB1, and -DPB1 data were available for the HGDP subjects, but DQB1 data were only available for African and European HGDP subjects. HLA-A, -C, -B, -DRB1 and -DQB1 data were available for the KG subjects.
Reference, testing and evaluation data sets
The ‘reference’, or training, data set consisted of genotypes for 10 268 xMHC SNPs and SBT molecular HLA genotypes data for 930 subjects in the phase one 1000G Project repository.21 These data are available online at immpute-project.immunogenomics.org. These HLA genotypes were recorded as G groups25 and represented only HLA-A, HLA-B and HLA-C exons 2 and 3 and HLA-DRB1 exon 2 nucleotide sequence variants. The ‘testing dataset’ consisted of genotypes for the same 10 268 xMHC SNPs for 889 HGDP subjects. The ‘evaluation dataset’ consisted of reverse-format sequence-specific oligonucleotide molecular HLA genotypes for the same 889 HGDP subjects. These HGDP subjects represent 27 distinct populations from five continental regions. For detailed subject ancestry, please refer to Supplementary Table 1.
Imputation methods
e-HLA uses an ensemble of classifiers to generate consensus predictions and confidence scores. HIBAG uses unphased SNP genotypes to predict HLA genes by averaging HLA posterior probabilities over an ensemble of classifiers constructed on K bootstrap samples with the same number of individuals.17 HLA*IMP:02 extends Browning and Browning’s method for SNP phasing and inference to predict HLA alleles from SNP genotypes using a graphical model of MHC haplotype structure.19 MAGPrediction uses a likelihood model for prediction of HLA genes from unphased SNP genotype data.18 For a detailed description of each method, see Supplementary Information.
The developers of the e-HLA, HIBAG, HLA*IMP:02, MAGPrediction and SNP2HLA imputation methods were supplied with the reference and testing data sets. HLA imputation was performed independently for each method. Detailed descriptions of each method are in the Supplementary Information. Following the initial submission of imputations, the performance of all methods was shared with all method developers, and each developer was given the opportunity to submit a second round of imputations reflecting algorithm improvement. The SNP2HLA developers withdrew from the study after the initial performance review. Data for this method were not included in the analyses presented here. The HIBAG and HLA:IMP*02 developers submitted second rounds of imputations. Only the most recently generated imputations performed with HIBAG version 1.3 and HLA:IMP*02 version 2.Fast (2.F) were used for the scoring and analyses presented here.
Scoring methods
Imputation accuracy (IA) was assessed by comparing concordance between the imputed genotypes and the evaluation data set at both 1-field and 2-field resolution.25 Accuracy included any imputation that (1) correctly imputed the known allele or (2) imputed an allele with identical nucleotide sequence (same G group, http://hla.alleles.org/alleles/g_groups.html) or identical encoded amino acid sequence (same P group, http://hla.alleles.org/alleles/p_groups.html) within exons 2 and 3 (HLA class I) or exon 2 (HLA class II).25 IA metrics reported at each locus included the total number of correctly imputed alleles, the total number of correctly imputed alleles per individual (zero, one or two matches) and the total number of correctly imputed four-locus genotypes (correct for all loci, in all alleles). Within each locus the IA was defined as the total number of correctly imputed alleles across all subjects (N) relative to the number of total chromosomes imputed (2N).

Score is a binary prediction accuracy value for each imputed allele at each locus, which was set to 1 or 0 for accurate and inaccurate predictions, respectively. The scores for each subject had a maximum of 2 and an overall combined locus maximum of 2N (Supplementary Table 1).
Confidence values between 0 and 1 (inclusive) were reported for each imputed allele at each locus (e-HLA, HLA*IMP:02) or for the entire genotype at each locus (HIBAG, MAGPrediction). Imputation performance was assessed by iteratively applying a confidence value threshold and recalculating the IA for the remaining imputed genotypes. The locus call rate was defined as the ratio of imputed genotypes remaining, relative to the number of total chromosomes (2N) remaining after each threshold reevaluation. Method and locus-specific thresholds were obtained from the unique list of confidence values reported with each imputed data set. Imputation performance was visualized by graphing the IA relative to the call rate. To aid in visualization, x and y axes were adjusted accordingly.
Results
Overall IA
Table 1 outlines the accuracy metrics for each method, including the 2-field IA, total count for correct imputations of zero, one or two alleles (Supplementary Table 2 for percentages), and number of subjects whose four-locus HLA genotypes were correctly imputed. We observe a statistically significant hierarchy of IA between loci, as illustrated in Figure 1. HLA-C ranks highest, with an IA range of 89.9–94.6% across methods, followed by HLA-A (IA 89.7–92.2%), HLA-B (IA 69–77%) and HLA-DRB1 (IA 62.4–70.1%) (all inter-method P<1e-07). As further illustrated in Figure 1, we observe fewer differences in IA across methods, with IA for HIBAG ranking higher (P=4.5e-9) than MAGPrediction and HLA*IMP:02, which ranks higher (P=0.037) than e-HLA. Supplementary Table 3 identifies those imputed alleles with IA >95% or <50% across all methods. Similar trends result from IA analyses restricted to European HGDP subjects (Supplementary Figure 1), and to sub-Saharan African or randomly selected subsets of HGDP subjects (data not shown). These variable levels of accuracy resulted in low performance overall for correctly imputed four-locus HLA genotypes, with HIBAG imputation demonstrating a marginal advantage (HIBAG=27.8% versus 20–17.2%, P=1.6 e-4).
Statistical significance of imputation accuracy across loci and methods. Statistical significance of imputation accuracy (IA) across loci and methods was assessed using a logistic regression model, as detailed in the Supplementary Information. Odds ratios and their confidence intervals are plotted relative to IA for a locus or method predictor, for all HGDP subjects, as informed by the model. HLA-A was selected as the predictor for locus comparisons, and e-HLA for method comparisons.
In addition to the imputed genotype, each method reported a per subject imputation confidence value (0–1), either for each individual allele (e-HLA, HLA*IMP:02) or for the genotype (HIBAG, MAGPrediction) at each locus. Figure 2 compares each method’s IA to the call rate (proportion of imputation results) at increasing confidence thresholds. As expected, removing lower confidence results increased accuracy at the expense of call rate, with the exception of MAGPrediction at HLA-C. IA increases in HLA-B and HLA-DRB1 were linear with respect to a wide range of call rates (50–80%), and confidence values for these loci failed to demonstrate robust correlations with correct imputations. In contrast, HLA-A and HLA-C exhibited a sharp increase in IA over a narrow range of call rates (80–100%), again with the exception of MAGPrediction at HLA-C. Variation in the 0.5 confidence threshold (diamonds, Figure 2), further illustrates the inconsistency of confidence values across methods; for HLA-B and HLA-DRB1, the 0.5 threshold is associated with a wide call rate range (60–90%) depending on method, whereas this threshold is restricted to 90–100% call rates in all methods for HLA-A and HLA-C.
Locus-level imputation performance. Imputation accuracy (IA) was assessed at different call rates by iterative application of a confidence value threshold and recalculation of the IA. Confidence value thresholds were derived from the unique list of confidence values reported for each allele or genotype. Line length was a function of the lowest reported confidence value (only non-zero call rates are graphed). Each panel corresponds to a different locus (HLA-A, -B, -C and -DRB1). Color corresponds to method: blue, e-HLA; orange, HIBAG; green, HLA*IMP:02; magenta, MAGprediction. For comparison, the 90% IA (gray dotted line) and the 0.5 confidence thresholds (diamonds) for each imputation are indicated. Only non-zero accuracies are graphed. For e-HLA and HLA*IMP:02, the distribution of confidence values was small compared with HIBAG and MAGPrediction and results in line termination at higher call rates. Different IA scales are presented for HLA-A and -C than for -B and -DRB1.
The number of subjects correctly imputed across all four loci is shown at the bottom of Table 1. Fewer than 27.8% of subjects were correctly imputed by any method. As illustrated in Supplementary Figure 2, only 77 (9.4%) subjects were correctly imputed by all four methods, and 51 (6.3%) subjects were correctly imputed by only one method. The call rates at which 50% of correctly imputed subjects remain for each method (21.2%, e-HLA; 25.5%, HIBAG; 12.1%, HLA*IMP:02; and 25%, MAGPrediction) decrease in step with the percentage of correctly imputed subjects, as illustrated in Figure 3, wherein the percentage of correctly imputed subjects decreases with call rate, as IA increases. HIBAG generated more correct imputations than the other methods, but over a larger range of confidence values. Regardless of the method applied, confidence values serve as unreliable predictors of correct four-locus imputations.
Subject-level imputation performance. For each method evaluated, two different imputation accuracy (IA) measures are plotted for each call rate (x axis) at the subject-level; only subjects for which the imputations at all four loci are correct are scored as accurate. 'Subset Accuracy', the percentage of correctly imputed subjects at each call rate threshold (dashed lines), as presented for individual loci in Figure 2, loci, is plotted alongside 'Global Accuracy', the percentage of subjects out of the total data set that are correctly imputed for each call rate threshold (solid lines). Color corresponds to method: blue, e-HLA; orange, HIBAG; green, HLA*IMP:02; magenta, MAGprediction.
To examine the extent to which variation in IA between loci results from the presence of HLA alleles in the evaluation data set that were absent from the reference data set (untrained alleles), subjects with untrained alleles were removed on a per locus basis and IA was recalculated. As illustrated in Figure 4, the locus-specific changes in IA (ΔIA) were smallest for HLA-C (max ΔIA 1.5%), followed by HLA-A (max ΔIA 1.6%), and were largest for HLA-DRB1 (max ΔIA 7%), followed by HLA-B (max ΔIA 5.7%). On average, the change in IA was 3.7% across all loci, suggesting that untrained alleles were not a major factor in the overall IA.
Imputation accuracy when masking untrained alleles. Imputation accuracy (IA) was assessed for each locus and method before and after removing carriers of untrained HLA alleles (that is, not present in the reference data set). HGDP subjects carrying one or two untrained HLA alleles were removed (masked) and IA recalculated on the remaining subjects, for which all alleles were present in the reference data set. The diagonal represents identical IA between masked and unmasked evaluation data sets. Changes in IA resulted in a shift from the diagonal. Shape corresponds to locus: circle, HLA-A; square, -B; diamond, -C; triangle, -DRB1. Color corresponds to method: blue, e-HLA; orange, HIBAG; green, HLA*IMP:02; magenta, MAGprediction.
IA within ancestries
The HGDP subjects were stratified into nine broad categories of continental origin (sub-Saharan Africa, North Africa, Europe, Southwest Asia, Southeast Asia, Oceania, Northeast Asia, North America and South America) to investigate variation in IA between samples from different world regions.26 Table 2 summarizes IA within these continental origin categories for each method and locus. Relative to the locus-specific median, IA values for sub-Saharan Africa and Oceania were consistently lower across all loci, whereas IA values for Northeast Asia were consistently higher. For individual loci, IA values for North America, Oceania, and South America were lowest across all methods for HLA-A (max IA 83.9%, 81.5%, 81.0%, 88.0%, respectively) and HLA-B (max IA 59.7%, 48.1%, 39.7%, respectively). IA values for Oceania were lowest for HLA-C (max IA 87%), whereas IA values for North America and South America were lowest for HLA-DRB1 (max IA 30.6 and 39.7%, respectively). Interestingly, despite the absence of North African and Southwest Asian individuals in the reference data set, IA for these regions was higher than the locus-specific median.
Application of multiple methods
The potential for imputation improvement through the application of multiple methods is illustrated in Figure 5. Although, the maximum possible IA for all combinations of methods is consistently higher than for any individual method (for example, 99.1% Max IA for HLA-C across all four methods), adjudicated IA values surpass individual method IAs by ~2% for all loci but HLA-DRB1, where with the maximum adjudicated improvement for HIBAG+HLA*IMP:02 is 3.2%.
Maximum, adjudicated and standardized imputation accuracies in method combinations. At each locus, and for each combination of two, three and four methods, the difference between the maximum imputation accuracy (IA), the adjudicated IA and the standardized IA is shown in comparison with the overall IA for each method. Maximum IA was calculated over all HGDP subjects by scoring the imputation for a given subject as correct if any of the predictions in a given combination of methods was accurate. Adjudicated IA was calculated over all HGDP subjects by choosing the prediction with the highest confidence score from among the predictions in a given combination of methods for each subject, and then comparing that prediction to the evaluation data set for accuracy. Standardized IA was calculated over all HGDP subjects by normalizing the confidence score distributions for each method and then choosing the highest confidence score as for Adjudicated IA. Ninety percent IA is indicated with the dotted line. The y axis for HLA-A and -C uses a different scale than the y axis for -B and -DRB1. Solid shapes correspond types of IA scores: circle, maximum IA score (Max); triangle, adjudicated IA score (Adj); asterisk, standardized IA (Std). Color corresponds to method: blue, e-HLA; orange, HIBAG; green, HLA*IMP:02; magenta, MAGprediction. Each panel corresponds to a different locus — HLA-A, -B, -C and -DRB1. For DRB1, the overall IA values for e-HLA and HLA*IMP:02 overlap.
The relationship between inter-method imputation agreement and the likelihood of a correct call at individual loci is illustrated in Supplementary Figure 3. Higher inter-method prediction agreement was associated with higher score (that is, number of correct imputations) and higher IA frequency. However, the frequency of agreement differed across loci (Supplementary Figure 3, red line). Agreement between all methods was less frequent for HLA-B and HLA-DRB1 (~40%), and in the case of HLA-DRB1, total agreement was associated with large variations in scores. Average IA within each method agreement category differed between loci, with IA for subjects with no inter-method agreement lowest for HLA-DRB1 (35%) and highest for HLA-C (47%). In cases where all methods agreed, IA is consistent with Figure 2 (HLA-A, 94.4%; HLA-C, 96.4%; HLA-B, 88.6%; HLA-DRB1, 76.5%), indicating greater agreement between methods at lower call rates.
Imputation using different developmental versions
The developers of HIBAG and HLA:IMP*02 opted to provide updated imputations, reflecting continued development of their methods. Supplementary Figure 4 details the imputation performance for both the legacy (initial submission) and the current versions of these methods. The updated imputation using HIBAG (v1.3) did not differ significantly from initial submission (P=0.89). However, of the two sets of updated HLA:IMP*02 imputations ('-v2 standard' and '-v2 fast'), only '-v2 fast' demonstrated an increase in performance over the legacy version (P=0.0013). For HLA*IMP:02-v2 fast, HLA-B demonstrated the greatest increase in performance relative to other loci, as illustrated in Supplementary Figure 5.
Discussion
Given the importance of the HLA genes in disease association and drug-induced hypersensitivity reactions,1, 2, 3, 4, 5, 13 and the abundance of SNP associations identified on chromosome 6p21 through GWAS, an in-depth investigation of HLA polymorphism is often warranted in disease association studies. Prediction of HLA genotypes through imputation from SNP data has been applied as an alternative to molecular HLA genotyping,27 especially in cohorts where chromosome 6 SNP data are already available. However, a detailed assessment of an imputation methods’ accuracy across a global selection of disparate populations has not been undertaken. In this study, the capacity of e-HLA, HIBAG, HLA*IMP:02 and MAGPrediction to correctly impute HLA genotypes at the HLA-A, HLA-B, HLA-C and HLA-DRB1 loci was assessed in the HGDP subjects, using the 1000G as a training data set. This is the first comprehensive comparison of multiple HLA genotype imputation methods across a wide range of populations, using large, well-characterized cohorts.
The accuracy of HLA allele imputation for the four most polymorphic and commonly investigated HLA loci (HLA-A, HLA-C, HLA-B and HLA-DRB1) varied more with respect to locus than with the method applied. When considering all predictions (100% call rate), imputation was most accurate for HLA-C, with IAs exceeding 89%, followed by HLA-A. HLA-DRB1 and HLA-B were the most difficult to impute across all methods, with IAs below 80%. That HLA-B proved difficult to impute is perhaps not surprising, as this is the most polymorphic HLA locus.28 However, HLA-DRB1 is less polymorphic than either HLA-A or HLA-C, suggesting that variation is not necessarily the primary obstacle to accurate imputation.
Studies involving three of the methods evaluated here have also indicated HLA-DRB1 as being difficult to impute.17, 18, 19 A recent comparison of sequence-based HLA genotyping with imputation of HLA-DRB1 alleles using HLA*IMP,29 HLA*IMP:02 and SNP2HLA8 (withdrawn from this study) in a small Finnish cohort also found accuracy rates to be very low (<30%) for this locus.30 HLA-DRB1 imputation also demonstrated the lowest concordance with sequence-based genotyping in an association study of Parkinson Disease and HLA polymorphism in the NeuroGenetics Research Consortium dataset.31 IA for this locus was also low in study of HIBAG imputation in the ethnically and racially diverse Women’s Interagency HIV Study cohort.32 It is possible that the SNPs in these studies did not sufficiently tag HLA-DRB1 allele or sequence variation.
As illustrated in Supplementary Figure 6, DRB1 IA in the ImmPute study was dependent on the DRB haplotype. Subjects with HLA-DRB1 alleles on the DR8 haplotype were most difficult to impute. This haplotype consists of the non-polymorphic HLA-DRA gene, HLA-DRB1 alleles in the HLA-DRB1*08 allele family and the HLA-DRB9 pseudogene, and may have been generated in a contraction of the DR52 haplotype resulting in the deletion of >60 KB of DNA between the HLA-DRB1 and HLA-DRB3 genes.33, 34 Traherne et al.35 have described a 'SNP desert' on the DR52 haplotype extending from HLA-DRB3 to HLA-DQB3. Gene content variation between DRB haplotypes may result in increased missing SNP rates and the systematic exclusion of DRB SNPs from panels during quality control evaluation, creating an effective SNP desert around DRB1.
Figure 6 illustrates the distribution of the SNPs included in this study relative to the HLA-A, HLA-B, HLA-C and HLA-DRB1 genes. Significantly fewer SNPs occur within 100 kb of the HLA-DRB1 locus relative to the class I loci. An effective SNP desert surrounding the HLA-DRB1 locus derives not from the absence of HLA-DRB1 SNPs in the genome, but from the absence of proximal HLA-DRB1 SNPs on the immunoChip and Illumina 650Y panels. As shown in Supplementary Figure 7, this SNP desert is also present on the Affymetrix Genome-Wide Human SNP Array 6.0 release 35 and the Illumina InfiniumOmniExpress-24 version 1.2 Array. This absence of informative SNPs contributes to lower HLA-DRB1 imputation performance, and suggests that the reassessment of SNP ascertainment in panel design, allowing the detection of structural variants in the DRB region, may improve HLA-DRB1 IA. Imputation concordance rates have been shown to be higher for SNP test data sets generated using genotyping platforms with higher SNP densities, as well as through increased numbers of reference SNPs.36, 37
SNP proximity and density for the HLA-A, -B, -C and DRB1 Loci. Primary Panel. The density of SNPs (ranging from 0–12) within 500 000 bases of the HLA-A, -B, -C and -DRB1 loci is shown. A distance of 0 indicates the location of each respective gene. Negative distances are telomeric of the gene in question; positive distances are centromeric. Bold line: Proximal subsets of the 164 876 SNPs present in the 1000G data set, prior to merging with the HGDP data set. Light shaded area: SNPs present after the merger of the 1000G and HGDP data sets (merged SNPs), prior to quality control (QC) evaluation. Dark shaded area: merged SNPs remaining after QC evaluation. Inset Panel: The cumulative number of SNPs, out of the 10 268 SNPs included in this study, within 200 000 bases of the HLA-A, -B, -C and –DRB1 loci is shown. Color corresponds to locus: green, HLA-A; orange, HLA-B, purple; HLA-C; magenta, HLA-DRB1.
Chromosomes with highly similar SNP patterns have been observed to carry different HLA alleles,38 so that SNP patterns across the HLA region may be generally difficult to distinguish. The SNPs included in the reference data set were extracted from genomic sequence data rather than determined using established SNP genotyping methods; however, many more genomic SNPs were identified than were detectable with the applied SNP-typing panels (Figure 6), and comparison of these extracted SNP data to HapMap39 data for a subset of the same cohort, revealed minimal discrepancies (see Supplementary Information).
HLA IA may also be diminished by the multi-population, multi-regional nature of the 1000G and HGDP collections; however, although the HGDP is a much more diverse sample than the 1000G, both capture the same variation (Supplementary Figure 8). In these cases, accuracy is challenged by the extent to which the reference data reflect the diversity and patterns of LD in the populations being tested.29 Such variation can affect performance and is a function of the underlying SNP framework, with its history of recombination, mutation, natural selection, genetic drift and gene flow.29 Individual HLA alleles have been observed on diverse SNP frameworks across populations,15 and multi-locus HLA haplotypes have been shown to be geographically restricted,40 posing challenges for imputation when there is low population-level correspondence between reference and testing data sets. These challenges are evident in Table 2, where sub-Saharan African and Oceanian IA was consistently below locus-specific median values; Oceanian populations were not represented in the training data set, whereas sub-Saharan African populations display the highest levels of genetic diversity in the human species, reducing the likelihood of correspondence between the training and testing data sets for these populations. These challenges can be addressed through the public availability of large reference data sets representing an ethnically diverse selection of populations.
Further to this point, SNP ascertainment has primarily been conducted in European cohorts, and most HLA imputation studies have been performed in cohorts of European ancestry as well. However, clinical use cases for HLA imputation (for example, patients seeking transplants from potential donors in unrelated donor registries) are likely more cosmopolitan. Of the methods evaluated in this study, only HIBAG and HLA*IMP:02 have been developed using multi-population data sets.17, 19 Hsieh et al.37 imputed HLA alleles in Han Chinese using MAGprediction, and found a generally high concordance between imputation and molecular HLA genotyping for HLA-A and HLA-C, but poor concordance with HLA-B and HLA-DRB1 using ancestry specific reference panels. Similarly, Kuniholm et al. (2016) found higher concordance between HIBAG imputation and molecular HLA genotyping for HLA-A and -C than for HLA-B and –DRB1 in the ethnically diverse WIHS cohort.32 Pillai et al.41 compared SNP2HLA predictions to molecular HLA genotyping for the Singapore Genome Variation Project in southern Han Chinese, Southeast Asian Malays and Tamil Indians. Using ethnic-specific reference panels, they reported similarly poor performance for HLA-B and HLA-DRB1. However, by combining the SGVP and International HapMap Project41 reference panels, they were able to markedly increase prediction performance for these two loci. Khor et al.42 developed specific custom classifiers for the Japanese population, and applied these in HIBAG to achieve high IA for high-risk class II haplotypes in Japenese narolepsy patients. Similarly, Levin et al.43 improved HIBAG IA, relative to that of HLA*IMP:02, in African Americans by applying models reflecting the African and European ancestry of this population.
As illustrated in Figure 5, the application of multiple imputation methods has the potential for large increases in IA, relative to individual methods. However, as they are currently generated, confidence scores cannot be effectively applied across methods to realize this potential. Only in the case of HLA-DRB1 did the application of confidence scores across methods result in a marked improvement in IA. Confidence thresholding may serve as an attractive option for increasing IA for an individual method, at the expense of call rates. However, the derivation of the confidence values is unique to the method and cannot be reliably compared across methods or HLA loci. Because they are calculated differently and thus have different meanings, no single threshold can be applied to obtain commensurate IAs, and normalization of confidence values across methods does not improve their utility. Moreover, confidence metrics did not reliably correlate with IA, especially for HLA-B and HLA-DRB1. Continued increases in the confidence threshold increased the likelihood of dropping correct imputations as demonstrated by the asymptotic nature of the performance curves, and combined four-locus confidence scores correlated poorly with correctly imputed subjects. Care should be exercised when considering where to set a confidence threshold for imputation of HLA genotypes, and the associated call rate should be reported for reliable comparison. Overall, this poor correlation between IA and confidence metrics stems from both the application of non-standard confidence values across methods, and the mechanisms by which HLA diversity is generated and maintained. Although LD is high across the MHC, recombination within HLA genes, recombination hotspots between HLA genes, selection for novel polymorphisms, and high HLA heterozygosity will degrade the utility of intergenic SNPs for imputing HLA genotypes.44, 45, 46 All these mechanisms have posed challenges for molecular HLA genotyping, and they suggest that the application of rare and tagging SNPs is not likely to improve IA,38 and that HLA imputation is unlikely to accurately predict the presence of rare HLA alleles. Rather than considering confidence scores, consensus predictions from multiple methods may ensure the most reliable, accurate imputation results, in particular for HLA-A, HLA-B, and HLA-C. To realize the full potential of HLA imputation, the burden may be placed on method developers to devise prediction confidence ratings that can be applied across methods.
The prediction accuracies reported in this study may be considered to be over-estimates when the total diversity of allelic HLA polymorphism is considered. The number of HLA alleles identified in the 1000G and HGDP data sets is a fraction of the number of alleles in the IMGT/HLA Database,28 a number that is likely to increase every 3 months for the foreseeable future,47 although most of these alleles have been reported only once.48 In addition, imputation scoring was generous in that matching was evaluated both for individual alleles and for the members of P and G groups (see Methods for definition). Perhaps most importantly, the imputation results reported here are based on restricted reference and testing data sets. Larger, multi-population, multi-ancestry reference data sets would be required to successfully predict a larger proportion of observed HLA alleles.19, 36 Klitz et al.47 have estimated that millions of distinct HLA alleles are maintained in the human population, with many combinations of alleles present in population-specific haplotypes.40 Suitable reference data sets appropriate for HLA imputation at these levels may prove elusive, as earlier studies have suggested that at least 10 copies of an allele may be required in a reference data set for accurate imputation.38
Finally, the improvement in performance for the second round of HLA*IMP:02 imputation underscores the importance of applying the most up-to-date version of a method for HLA imputation. Imputation method developers leverage programming innovations, larger, more comprehensive reference data sets and enhanced knowledge of the genomics of the HLA region to ensure a robust algorithm that maximizes IA.
Conclusions
Accurate determination of classical HLA allele genotypes is critical for clinical applications such as transplantation and important for enabling association studies to uncover the genetic risk of complex diseases. Although HLA-A and HLA-C imputation remains a tractable option for research, our results strongly suggest that further development will be necessary before such cost-effective methods should be considered suitable for all HLA loci in both the research and clinical settings.
References
Welter D, MacArthur J, Morales J, Burdett T, Hall P, Junkins H et al. The NHGRI GWAS Catalog, a curated resource of SNP-trait associations. Nucleic Acids Res 2014; 42: D1001–D1006.
Martin AM, Nolan D, Gaudieri S, Almeida CA, Nolan R, James I et al. Predisposition to abacavir hypersensitivity conferred by HLA-B*5701 and a haplotypic Hsp70-Hom variant. Proc Natl Acad Sci USA 2004; 101: 4180–4185.
Mallal S, Nolan D, Witt C, Masel G, Martin AM, Moore C et al. Association between presence of HLA-B*5701, HLA-DR7, and HLA-DQ3 and hypersensitivity to HIV-1 reverse-transcriptase inhibitor abacavir. Lancet 2002; 359: 727–732.
Hung SI, Chung WH, Liou LB, Chu CC, Lin M, Huang HP et al. HLA-B*5801 allele as a genetic marker for severe cutaneous adverse reactions caused by allopurinol. Proc Natl Acad Sci USA 2005; 102: 4134–4139.
McCormack M, Alfirevic A, Bourgeois S, Farrell JJ, Kasperaviciute D, Carrington M et al. HLA-A*3101 and carbamazepine-induced hypersensitivity reactions in Europeans. N Engl J Med 2011; 364: 1134–1143.
Pavlos R, Mallal S, Phillips E . HLA and pharmacogenetics of drug hypersensitivity. Pharmacogenomics 2012; 13: 1285–1306.
Erlich H . HLA DNA typing: past, present, and future. Tissue Antigens 2012; 80: 1–11.
Jia X, Han B, Onengut-Gumuscu S, Chen WM, Concannon PJ, Rich SS et al. Imputing amino acid polymorphisms in human leukocyte antigens. PLoS ONE 2013; 8: e64683.
Hirschhorn JN, Daly MJ . Genome-wide association studies for common diseases and complex traits. Nat Rev Genet 2005; 6: 95–108.
McCarthy MI, Abecasis GR, Cardon LR, Goldstein DB, Little J, Ioannidis JP et al. Genome-wide association studies for complex traits: consensus, uncertainty and challenges. Nat Rev Genet 2008; 9: 356–369.
de Bakker PI, Raychaudhuri S . Interrogating the major histocompatibility complex with high-throughput genomics. Hum Mol Genet 2012; 21: R29–R36.
Traherne JA . Human MHC architecture and evolution: implications for disease association studies. Int J Immunogenet 2008; 35: 179–192.
Fernando MM, Stevens CR, Walsh EC, De Jager PL, Goyette P, Plenge RM et al. Defining the role of the MHC in autoimmunity: a review and pooled analysis. PLoS Genet 2008; 4: e1000024.
Moore JH, Asselbergs FW, Williams SM . Bioinformatics challenges for genome-wide association studies. Bioinformatics 2010; 26: 445–455.
de Bakker PI, McVean G, Sabeti PC, Miretti MM, Green T, Marchini J et al. A high-resolution HLA and SNP haplotype map for disease association studies in the extended human MHC. Nat Genet 2006; 38: 1166–1172.
Malkki M, Single R, Carrington M, Thomson G, Petersdorf E . MHC microsatellite diversity and linkage disequilibrium among common HLA-A, HLA-B, DRB1 haplotypes: implications for unrelated donor hematopoietic transplantation and disease association studies. Tissue Antigens 2005; 66: 114–124.
Zheng X, Shen J, Cox C, Wakefield JC, Ehm MG, Nelson MR et al. HIBAG-HLA genotype imputation with attribute bagging. Pharmacogenomics J 2013; 14: 192–200.
Li SS, Wang H, Smith A, Zhang B, Zhang XC, Schoch G et al. Predicting multiallelic genes using unphased and flanking single nucleotide polymorphisms. Genet Epidemiol 2011; 35: 85–92.
Dilthey A, Leslie S, Moutsianas L, Shen J, Cox C, Nelson MR et al. Multi-population classical HLA type imputation. PLoS Comput Biol 2013; 9: e1002877.
Cann HM, de Toma C, Cazes L, Legrand MF, Morel V, Piouffre L et al. A human genome diversity cell line panel. Science 2002; 296: 261–262.
Abecasis GR, Auton A, Brooks LD, DePristo MA, Durbin RM, Handsaker RE et al. An integrated map of genetic variation from 1,092 human genomes. Nature 2012; 491: 56–65.
Parkes M, Cortes A, van Heel DA, Brown MA . Genetic insights into common pathways and complex relationships among immune-mediated diseases. Nat Rev Genet 2013; 14: 661–673.
Danecek P, Auton A, Abecasis G, Albers CA, Banks E, DePristo MA et al. The variant call format and VCFtools. Bioinformatics 2011; 27: 2156–2158.
Erlich H, Bugawan T, Begovich AB, Scharf S, Griffith R, Saiki R et al. HLA-DR, DQ and DP typing using PCR amplification and immobilized probes. Eur J Immunogenet 1991; 18: 33–55.
Marsh SG, Albert ED, Bodmer WF, Bontrop RE, Dupont B, Erlich HA et al. Nomenclature for factors of the HLA system, 2010. Tissue Antigens 2010; 75: 291–455.
Mack SJ, S MA, Meyer D,. Single RM, Tsai Y, Erlich HA . Methods used in the generation and preparation of data for analysis in the 13th International Histocompatibility Work- shop vol 1. IHWG Press: Seattle, WA, USA, 2007.
Frangoul H, Crowe D . Cost saving associated with implementing a stepwise approach to HLA typing of related donors before hematopoietic SCT. Bone Marrow Transplant 2014; 49: 850–851.
Robinson J, Halliwell JA, McWilliam H, Lopez R, Parham P, Marsh SG . The IMGT/HLA database. Nucleic Acids Res 2013; 41: D1222–D1227.
Dilthey AT, Moutsianas L, Leslie S, McVean G . HLA*IMP—an integrated framework for imputing classical HLA alleles from SNP genotypes. Bioinformatics 2011; 27: 968–972.
Vlachopoulou E, Lahtela E, Wennerstrom A, Havulinna AS, Salo P, Perola M et al. Evaluation of HLA-DRB1 imputation using a Finnish dataset. Tissue Antigens 2014; 83: 350–355.
Wissemann WT, Hill-Burns EM, Zabetian CP, Factor SA, Patsopoulos N, Hoglund B et al. Association of Parkinson disease with structural and regulatory variants in the HLA region. Am J Hum Genet 2013; 93: 984–993.
Kuniholm MH, Xie X, Anastos K, Xue X, Reimers L, French AL et al. Human leucocyte antigen class I and II imputation in a multiracial population. Int J Immunogenet 2016; 43: 369–375.
Andersson G . Evolution of the human HLA-DR region. Front Biosci 1998; 3: d739–d745.
Gorski J . The HLA-DRw8 lineage was generated by a deletion in the DR B region followed by first domain diversification. J Immunol 1989; 142: 4041–4045.
Traherne JA, Horton R, Roberts AN, Miretti MM, Hurles ME, Stewart CA et al. Genetic analysis of completely sequenced disease-associated MHC haplotypes identifies shuffling of segments in recent human history. PLoS Genet 2006; 2: e9.
Zhang XC, Li SS, Wang H, Hansen JA, Zhao LP . Empirical evaluations of analytical issues arising from predicting HLA alleles using multiple SNPs. BMC Genet 2011; 12: 39.
Hsieh AR, Chang SW, Chen PL, Chu CC, Hsiao CL, Yang WS et al. Predicting HLA genotypes using unphased and flanking single-nucleotide polymorphisms in Han Chinese population. BMC Genomics 2014; 15: 81.
Leslie S, Donnelly P, McVean G . A statistical method for predicting classical HLA alleles from SNP data. Am J Hum Genet 2008; 82: 48–56.
Thorisson GA, Smith AV, Krishnan L, Stein LD . The International HapMap Project Web site. Genome Res 2005; 15: 1592–1593.
Single RM, Meyer D, Mack SJ, Lancaster A, Nelson MP, Fernández-Viña M et al. Haplotype Frequencies and Linkage Disequilibrium among classical HLA genes vol. 1. IHWG Press: Seattle, WA, USA, 2007.
Pillai NE, Okada Y, Saw WY, Ong RT, Wang X, Tantoso E et al. Predicting HLA alleles from high-resolution SNP data in three Southeast Asian populations. Hum Mol Genet 2014; 23: 4443–4451.
Khor SS, Yang W, Kawashima M, Kamitsuji S, Zheng X, Nishida N et al. High-accuracy imputation for HLA class I and II genes based on high-resolution SNP data of population-specific references. Pharmacogenomics J 2015; 15: 530–537.
Levin AM, Adrianto I, Datta I, Iannuzzi MC, Trudeau S, McKeigue P et al. Performance of HLA allele prediction methods in African Americans for class II genes HLA-DRB1, -DQB1, and -DPB1. BMC Genet 2014; 15: 72.
Sasazuki T, Inoko H, Morishima S, Morishima Y . Gene map of the HLA region, Graves' disease and Hashimoto Thyroiditis, and hematopoietic stem cell transplantation. Adv Immunol 2016; 129: 175–249.
Begovich AB, McClure GR, Suraj VC, Helmuth RC, Fildes N, Bugawan TL et al. Polymorphism, recombination, and linkage disequilibrium within the HLA class II region. J Immunol 1992; 148: 249–258.
Solberg OD, Mack SJ, Lancaster AK, Single RM, Tsai Y, Sanchez-Mazas A et al. Balancing selection and heterogeneity across the classical human leukocyte antigen loci: a meta-analytic review of 497 population studies. Hum Immunol 2008; 69: 443–464.
Klitz W, Hedrick P, Louis EJ . New reservoirs of HLA alleles: pools of rare variants enhance immune defense. Trends Genet 2012; 28: 480–486.
Mack SJ, Cano P, Hollenbach JA, He J, Hurley CK, Middleton D et al. Common and well-documented HLA alleles: 2012 update to the CWD catalogue. Tissue Antigens 2013; 81: 194–203.
Acknowledgements
We thank Janelle Noble, Marc Salit and P Scott Pine for helpful discussions and Abeer Madbouly for assistance with the PCA plots. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH, NIAID, NINDS, NMSS, ONR or United States Government. This work was supported by Office of Naval Research (ONR) grant N00014-08-1-1207 (KB, DP, PAG, JAH, AL, SJM, MM and VP), National Institutes of Health (NIH) grants U01AI067068 (JAH and SJM) and U19AI067152 (ARRA administrative supplement) (PAG) awarded by the National Institute of Allergy and Infectious Diseases (NIAID), R01GM109030 (JAH, SJM and DJP) and P01GM099568 (XZ) awarded by the National Institute of General Medical Sciences (NIGMS), RO1NS076492 (PAG), RO1NS046297 (PAG) and R01NS049477 (PAG) awarded by the National Institute of Neurological Disorders and Stroke (NINDS), and National Multiple Sclerosis Society (NMSS) grant RG 2899-D11 (PAG). PAG is a recipient of the Race to Erase MS Junior Investigator Award and the European Federation for Immunogenetics Julia Bodmer Award. This work was supported by the Australian National Health and Medical Research Council (NHMRC), Career Development Fellowship ID 1053756 (S.L.); and by the Victorian Life Sciences Computation Initiative (VLSCI) grant number VR0240 on its Peak Computing Facility at the University of Melbourne, an initiative of the Victorian Government, Australia (S.L.). Research at the Murdoch Childrens Research Institute was supported by the Victorian Government’s Operational Infrastructure Support Program. We thank President Barack H. Obama for his support and appreciation of American science and basic research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
SL is a partner in Peptide Groove LLP. Peptide Groove has licensed HLA typing technology to Affymetrix Ltd.
Additional information
Supplementary Information accompanies the paper on the The Pharmacogenomics Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Pappas, D., Lizee, A., Paunic, V. et al. Significant variation between SNP-based HLA imputations in diverse populations: the last mile is the hardest. Pharmacogenomics J 18, 367–376 (2018). https://doi.org/10.1038/tpj.2017.7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tpj.2017.7
- Springer Nature Limited
This article is cited by
-
The lupus susceptibility allele DRB1*03:01 encodes a disease-driving epitope
Communications Biology (2022)
-
HLA-check: evaluating HLA data from SNP information
BMC Bioinformatics (2017)