Abstract
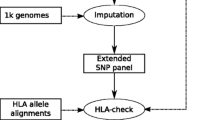
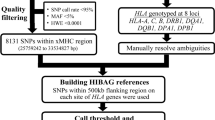
SNP-based imputation approaches for human leukocyte antigen (HLA) typing take advantage of the extended haplotype structure within the major histocompatibility complex (MHC) to predict classical HLA alleles using dense SNP genotypes, such as those available on chip panels of genome-wide association study (GWAS). These methods enable HLA analyses of classical alleles on existing SNP datasets genotyped in GWAS studies at no extra cost. Here, I describe the workflow of HIBAG, an imputation method with attribute bagging, for obtaining a sample’s HLA class I and II genotypes of two-field resolution using SNP data. Two examples are provided to illustrate with a publicly available HLA and SNP dataset: genotype imputation with pre-fit classifiers in GWAS, and model training to build a new classifier.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Shiina T, Hosomichi K, Inoko H, Kulski JK (2009) The HLA genomic loci map: expression, interaction, diversity and disease. J Hum Genet 54(1):15–39
Welter D, MacArthur J, Morales J, Burdett T, Hall P, Junkins H, Klemm A, Flicek P, Manolio T, Hindorff L et al (2014) The NHGRI GWAS catalog, a curated resource of SNP-trait associations. Nucleic Acids Res 42(Database issue):D1001–D1006
Bauer DC, Zadoorian A, Wilson LO, Thorne NP, Alliance MGH (2016) Evaluation of computational programs to predict HLA genotypes from genomic sequencing data. Brief Bioinform pii:bbw097
Erlich H (2012) HLA DNA typing: past, present, and future. Tissue Antigens 80(1):1–11
Meyer D, Nunes K (2017) HLA imputation, what is it good for? Hum Immunol 78(3):239–241
Zheng X, Shen J, Cox C, Wakefield JC, Ehm MG, Nelson MR, Weir BS (2014) HIBAG–HLA genotype imputation with attribute bagging. Pharmacogenomics J 14(2):192–200
Breiman L (1996) Bagging predictors. Mach Learn 24:123–140
Breiman L (2001) Random forests. Mach Learn 45(1):5–32
Khor SS, Yang W, Kawashima M, Kamitsuji S, Zheng X, Nishida N, Sawai H, Toyoda H, Miyagawa T, Honda M et al (2015) High-accuracy imputation for HLA class I and II genes based on high-resolution SNP data of population-specific references. Pharmacogenomics J 15(6):530–537
Levin AM, Adrianto I, Datta I, Iannuzzi MC, Trudeau S, McKeigue P, Montgomery CG, Rybicki BA (2014) Performance of HLA allele prediction methods in African Americans for class II genes HLA-DRB1, -DQB1, and -DPB1. BMC Genet 15:72
Nunes K, Zheng X, Torres M, Moraes ME, Piovezan BZ, Pontes GN, Kimura L, Carnavalli JE, Mingroni Netto RC, Meyer D (2016) HLA imputation in an admixed population: an assessment of the 1000 genomes data as a training set. Hum Immunol 77(3):307–312
Pappas DJ, Lizee A, Paunic V, Beutner KR, Motyer A, Vukcevic D, Leslie S, Biesiada J, Meller J, Taylor KD et al (2017) Significant variation between SNP-based HLA imputations in diverse populations: the last mile is the hardest. Pharmacogenomics J. https://doi.org/10.1038/tpj.2017.7
Gourraud PA, Khankhanian P, Cereb N, Yang SY, Feolo M, Maiers M, Rioux JD, Hauser S, Oksenberg J (2014) HLA diversity in the 1000 genomes dataset. PLoS One 9(7):e97282
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, Maller J, Sklar P, de Bakker PI, Daly MJ et al (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81(3):559–575
Zheng X, Levine D, Shen J, Gogarten SM, Laurie C, Weir BS (2012) A high-performance computing toolset for relatedness and principal component analysis of SNP data. Bioinformatics 28(24):3326–3328
Robinson J, Halliwell JA, Hayhurst JD, Flicek P, Parham P, Marsh SG (2015) The IPD and IMGT/HLA database: allele variant databases. Nucleic Acids Res 43(Database issue):D423–D431
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Zheng, X. (2018). Imputation-Based HLA Typing with SNPs in GWAS Studies. In: Boegel, S. (eds) HLA Typing. Methods in Molecular Biology, vol 1802. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-8546-3_11
Download citation
DOI: https://doi.org/10.1007/978-1-4939-8546-3_11
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-8545-6
Online ISBN: 978-1-4939-8546-3
eBook Packages: Springer Protocols