Abstract
Group B Streptococcus (GBS) is an encapsulated, gram-positive pathogen that is an important cause of neonatal invasive infections, including sepsis and meningitis. There are ten known GBS serotypes based on distinct capsule compositions (Ia, Ib, II-IX), and current candidate capsular polysaccharide conjugate vaccines target only a subset of these. Serotyping of GBS isolates is important for understanding local epidemiology and for monitoring for serotype replacement or capsular switching. However, serotyping generally requires either latex agglutination, multiplex PCR with analysis of band sizes, or analysis of whole genome sequences–all techniques that are either expensive or not widely available. Here we report the development of a robust real-time PCR assay for determining GBS serotypes. Using both a diverse reference set of strains encompassing all ten serotypes and a collection of clinical isolates, we demonstrate concordance between real-time PCR serotyping and latex agglutination. We propose that real-time PCR serotyping represents an attractive alternative to current serotyping methods and may allow for improved acquisition of GBS serotype data.
Similar content being viewed by others
Introduction
Streptococcus agalactiae (Group B Streptococcus [GBS]) is a major cause of neonatal morbidity and mortality worldwide1,2. While the incidence of GBS early-onset sepsis has decreased substantially in the United States following the implementation of universal screening and intrapartum antimicrobial prophylaxis3,4, these represent resource-intensive policies that are subject to missed opportunities for prevention5,6. Because of these limitations, improved methods for prevention of GBS disease are urgently needed. Immunization with capsular polysaccharide (CPS)-protein conjugates can induce serotype-specific immunity7, and candidate GBS conjugate vaccines are currently being tested in clinical trials8.
The serotype of a GBS strain is determined by the genes in the cps locus, and 10 distinct serotypes (Ia, Ib, II-IX) have been described to date9,10. Understanding the serotype distribution of GBS in both disease and colonization states is crucial to inform rational vaccine policies and to detect events such as capsular switching11,12 and serotype replacement. Capsular typing of GBS is generally performed using latex agglutination with type-specific antibodies, though both multiplex PCR13,14,15 and flow cytometry16 strategies have been described. More recently, whole-genome sequences have been used to determine GBS serotypes17. Here we describe a novel TaqMan-based real-time PCR strategy for serotyping GBS based on detection of specific cps locus sequences and demonstrate its applicability using both a defined set of strains from a reference laboratory and a set of primary clinical isolates.
Materials and Methods
Bacterial strains and growth conditions
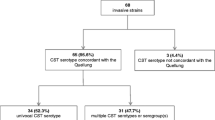
A total of 68 clinical GBS isolates were included in this study. Of these, 47 S. agalactiae serotype reference stains were provided by the Streptococcus laboratory at the Respiratory Diseases Branch of the Centers for Disease Control and Prevention (CDC). For some isolates, multilocus sequence type (MLST) determination was determined by the Streptococcus reference laboratory using established protocols (http://pubmlst.org/sagalactiae/). The remaining 21 isolates were deidentified sterile site isolates collected between 2010 and 2015 in New York City. All specimens were cultured on trypticase soy (TS) or 5% sheep blood agar plates at 37 °C and then grown overnight in TS media at 37 °C.
Serotyping by latex agglutination
All GBS isolates were serotyped by latex agglutination with the Immulex Latex Agglutination Streptococcus B kit (Staten Serum Institute; Copenhagen, Denmark) according to the manufacturer’s instructions. Briefly, 10 μl of the latex reagent was added to a single colony of GBS suspended in 10 μl of saline. The reaction was rotated and interpreted as positive if agglutination was visible after 30 seconds.
TaqMan real-time PCR serotyping
Primers and probes were designed to amplify unique regions of the polysaccharide capsular genes of each of the serotypes of S. agalactiae (Table 1). In order to generate these primer/probe sets, DNA sequences from specific capsular polysaccharide (cps) operons were obtained from National Center for Biotechnology Information databases (Table 1). Trimmed predicted cpsE-cpsL regions were aligned with using the CLUSTALW algorithm (Supplemental File 1). Specific primer and probe sequences and target regions were hand-selected from this multiple alignment of all 10 cps operons and are listed in Table 1. Oligonucleotides and probes were obtained from Integrated DNA Technologies (Coralville, IA). Oligonucleotides were unmodified and desalted, and probes were labeled with a fluorescent probe (5′ 6-carboxyfluorescein (6-FAM)) and two quenchers (internal ZENTM, and 3′ Iowa Black® FQ) and purified by high-pressure liquid chromatography.
Genomic DNA was extracted from overnight cultures of GBS using the Qiagen DNeasy Blood and Tissue Kit according to the manufacturer’s instructions. PCR reactions were performed in a final volume of 20 μl and consisted of 10 μl Taqman Universal Mastermix (Applied Biosystems), 7.4 μl sterile water, 0.2 μl forward primer (100 μM stock), 0.2 μl reverse primer (100 μM stock), 0.2 μl probe (100 μM stock), and 2 μl GBS DNA (25 ng/μl, unless otherwise indicated in the text.) Triplicate reactions were performed on a StepOne Plus thermal cycler (Applied Biosystems) and analyzed using StepOne software. Reaction parameters were as follows: initial incubation at 50 °C for 2 minutes; initial denaturation at 95 °C for 10 minutes; 35 cycles of PCR at 95 °C for 15 seconds and 60 °C for 1 min. Positive reactions were defined as a cycle threshold (CT) < 30 for 50 ng DNA template/reaction. Negative control reactions (no DNA template) were included with every run. Results were compared to latex agglutination.
Reaction sensitivity and interference by non-targeted strains
Sensitivity was evaluated by determining the cycle threshold on 10-fold serial dilutions of bacterial DNA from 5 ng to 50 pg per reaction. Potential interference was assessed by real-time PCR using each primer/probe set under the conditions above with a mixture of chromosomal DNA from all ten GBS serotypes (strains AR959 - AR968) at a final amount of 25 ng for each strain (250 ng total DNA) per reaction as template.
Results
Accuracy of real-time PCR serotyping using a validation set
We used a set of 47 validation strains (≥3 per serotype) from the CDC Streptococcus laboratory to determine whether the primer/probe combinations listed in Table 1 could be used for accurate serotyping. In all cases, there was detection of the predicted sequence using the real-time PCR protocol (Table 2) and no positive reactions with any of the other serotype primer/probe combinations (not shown). We confirmed the serotype of each of the 47 strains by latex agglutination, and there was 100% concordance between the PCR-based and latex agglutination methods for the validation set.
Serotyping of clinical isolates
We used the real-time PCR serotyping assay on a collection of 21 clinical sterile-site GBS isolates (Table 3). In 21/21 (100%) of cases, a single serotype was detected at CT < 30, and in all of those, the result was confirmed by latex agglutination.
Sensitivity of real-time PCR serotyping and potential interference
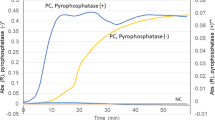
In order to determine the sensitivity of the real-time PCR protocol, we performed serial dilutions of the extracted DNA from 20 of the validation strains, from 5 ng to 50 pg per reaction. As predicted, CT increased as the total amount of bacterial DNA template in the reaction decreased (Fig. 1), though the majority of strains remained under the 30 cycle threshold, even at 50 pg per reaction. For all primer sets, detection of target sequences (CT < 30) was robust to the inclusion of an excess of non-targeted GBS DNA.
Serial dilution of GBS genomic DNA template demonstrates sensitivity of real-time PCR serotyping.
Serotyping reactions were performed on 20 validation strains using 5 ng, 500 pg, and 50 pg of template per reaction. Cycle threshold increased with decreasing template amounts but remained detectable for all strains even at the lowest concentration tested.
Discussion
Clinical microbiology laboratories routinely identify GBS but do not typically report serotype data. However, as the development of vaccines that target a subset of the ten known GBS serotypes proceeds, it will become increasingly important to understand the distribution of vaccine and non-vaccine serotypes and to monitor for capsular switching and serotype replacement. Non-nucleic acid-based laboratory methods for GBS serotyping techniques can be labor intensive and expensive, may require high-titer serotype-specific antisera and can create an added barrier for performing GBS serotype prevalence studies. Latex agglutination kits may be used for GBS serotyping in reference laboratories but are costly and not routinely available.
Molecular capsular typing techniques, including multiplex PCR13,14 and targeted analysis of whole genome sequences17, may be advantageous because of adaptability to newly available sequence information and relative ease of performance. In one prior study, real-time PCR was reported to distinguish among serotypes Ia, Ib, and III only18. We designed custom primer-probe sets for detection of all ten GBS serotypes. Unlike many prior methods, serotyping by real-time PCR is not based on operator interpretation of biochemical reactions and allows for specific identification of a GBS serotype despite interference from other bacterial DNA. Although the current assay is likely too technically demanding for routine laboratory adoption, it may be useful in the context of epidemiologic studies as an alternative to existing multiplex PCR assays. With further development, this novel method may allow for detection of GBS serotypes directly from clinical specimens without the need for culture. Limitations of this real-time PCR-based serotyping include an inability to identify new serotypes directly (as specific primer-probe sets are required for detection) and dependence on conservation of the targeted genetic regions within a particular serotype. Further validation of these primer-probe sets on a globally representative set of isolates is underway, as all of the tested strains originated in North America. Studies of larger, more diverse strain collections will be required to ensure that the primer-probe sets are sensitive and specific across an appropriate range of isolates.
Additional Information
How to cite this article: Breeding, K. M. et al. Real-time PCR-based serotyping of Streptococcus agalactiae. Sci. Rep. 6, 38523; doi: 10.1038/srep38523 (2016).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Edmond, K. M. et al. Group B streptococcal disease in infants aged younger than 3 months: systematic review and meta-analysis. Lancet 379, 547–556 (2012).
Le Doare, K. & Heath, P. T. An overview of global GBS epidemiology. Vaccine 31, D7–D12 (2013).
Schrag, S. J. et al. Group B streptococcal disease in the era of intrapartum antibiotic prophylaxis. N Engl J Med 342, 15–20 (2000).
Verani, J. R., McGee, L. & Schrag, S. J. Division of Bacterial Diseases National Center for Immunization and Respiratory Diseases Centers for Disease Control and Prevention CDC. Prevention of perinatal group B streptococcal disease–revised guidelines from CDC, 2010. MMWR Recomm Rep 59, 1–36 (2010).
Stoll, B. J. et al. Early Onset Neonatal Sepsis: The Burden of Group B Streptococcal and E. coli Disease Continues. Pediatrics 127, 817–826 (2011).
Verani, J. R. et al. Early-onset group B streptococcal disease in the United States: potential for further reduction. Obstetrics and gynecology 123, 828–837 (2014).
Kasper, D. L. et al. Immune response to type III group B streptococcal polysaccharide-tetanus toxoid conjugate vaccine. J Clin Invest 98, 2308–2314 (1996).
Madhi, S. A. et al. Safety and immunogenicity of an investigational maternal trivalent group B streptococcus vaccine in healthy women and their infants: a randomised phase 1b/2 trial. Lancet Infect Dis. doi: 10.1016/S1473-3099(16)00152-3 (2016).
Berti, F. et al. Structure of the type IX group B Streptococcus capsular polysaccharide and its evolutionary relationship with types V and VII. J Biol Chem 289, 23437–23448 (2014).
Cieslewicz, M. J. et al. Structural and genetic diversity of group B streptococcus capsular polysaccharides. Infect Immun 73, 3096–3103 (2005).
Bellais, S. et al. Capsular switching in group B Streptococcus CC17 hypervirulent clone: a future challenge for polysaccharide vaccine development. Journal of Infectious Diseases 206, 1745–1752 (2012).
Martins, E. R., Melo-Cristino, J. & Ramirez, M. Evidence for rare capsular switching in Streptococcus agalactiae. J Bacteriol 192, 1361–1369 (2010).
Poyart, C. et al. Multiplex PCR assay for rapid and accurate capsular typing of group B streptococci. J Clin Microbiol 45, 1985–1988 (2007).
Imperi, M. et al. A multiplex PCR assay for the direct identification of the capsular type (Ia to IX) of Streptococcus agalactiae. J. Microbiol. Methods 80, 212–214 (2010).
Yao, K. et al. Capsular gene typing of Streptococcus agalactiae compared to serotyping by latex agglutination. J Clin Microbiol 51, 503–507 (2013).
Rosini, R. et al. Genomic Analysis Reveals the Molecular Basis for Capsule Loss in the Group B Streptococcus Population. PLoS ONE 10, e0125985 (2015).
Sheppard, A. E. et al. Capsular Typing Method for Streptococcus agalactiae Using Whole-Genome Sequence Data. J Clin Microbiol 54, 1388–1390 (2016).
Morozumi, M. et al. Direct identification of Streptococcus agalactiae and capsular type by real-time PCR in vaginal swabs from pregnant women. J. Infect. Chemother. 21, 34–38 (2014).
Acknowledgements
The authors are grateful to Dr. Bernard Beall (CDC Streptococcus Laboratory) for providing characterized GBS isolates and helpful advice and to the Active Bacterial Core Surveillance (ABCs) program. This work was supported by the Doris Duke Charitable Foundation (#2014084 to A.J.R.).
Author information
Authors and Affiliations
Contributions
A.J.R. performed bioinformatic analyses and designed primers and probes. K.M.B., B.R., K.D.L., M.M., and T.M.R. performed serotyping and PCR assays. K.M.B., T.M.R., and A.J.R. drafted the main manuscript text. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Breeding, K., Ragipani, B., Lee, KU. et al. Real-time PCR-based serotyping of Streptococcus agalactiae. Sci Rep 6, 38523 (2016). https://doi.org/10.1038/srep38523
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep38523
- Springer Nature Limited
This article is cited by
-
Prevalence of group B streptococcus colonization in pregnant women in Jiangsu, East China
BMC Infectious Diseases (2021)
-
Molecular characterization of Streptococcus agalactiae isolated from pregnant women and newborns at the University of Gondar Comprehensive Specialized Hospital, Northwest Ethiopia
BMC Infectious Diseases (2020)
-
Development of a droplet digital PCR method for detection of Streptococcus agalactiae
BMC Microbiology (2020)
-
Development and analytical validation of real-time PCR for the detection of Streptococcus agalactiae in pregnant women
BMC Pregnancy and Childbirth (2020)
-
Comparison of molecular serotyping approaches of Streptococcus agalactiae from genomic sequences
BMC Genomics (2017)