Abstract
MicroRNAs (miRNAs) control gene expression as key negative regulators at the post-transcriptional level. MiR160 plays a pivotal role in Arabidopsis growth and development through repressing expression of its target AUXIN RESPONSE FACTOR (ARF) genes; however, the function of miR160 in monocots remains elusive. In this study, we found that the mature rice miR160 (OsmiR160) was mainly derived from OsMIR160a and OsMIR160b genes. Among four potential OsmiR160 target OsARF genes, the OsARF18 transcript was cleaved at the OsmiR160 target site. Rice transgenic plants (named mOsARF18) expressing an OsmiR160-resistant version of OsARF18 exhibited pleiotropic defects in growth and development, including dwarf stature, rolled leaves and small seeds. mOsARF18 leaves were abnormal in bulliform cell differentiation and epidermal cell division. Starch accumulation in mOsARF18 seeds was also reduced. Moreover, auxin induced expression of OsMIR160a, OsMIR160b and OsARF18, whereas expression of OsMIR160a and OsMIR160b as well as genes involved in auxin signaling was altered in mOsARF18 plants. Our results show that negative regulation of OsARF18 expression by OsmiR160 is critical for rice growth and development via affecting auxin signaling, which will advance future studies on the molecular mechanism by which miR160 fine-tunes auxin signaling in plants.
Similar content being viewed by others
Introduction
MicroRNAs (miRNAs), which are small (~21 nucleotides) non-coding RNAs, act as critical negative regulators by binding to mRNA complementary sequences for mRNA destabilization and translational inhibition in both plants and animals1,2,3,4,5. In plants, primary miRNAs (pri-miRNA) are transcribed from MIRNA (MIR) genes. Stem-loop segments derived from pri-miRNAs are cleaved by RNase III-type endonucleases (also known as Dicers) to produce paired precursor miRNAs (pre-miRNA). After liberation of miRNA duplexes, mature miRNAs approximately 21-nucleotide long direct RNA-induced silencing complexes (RISC) to bind to their target mRNAs by complementary match, leading to cleavage or translational inhibition. In the model species Arabidopsis, the general roles of miRNAs are recognized by analysis of mutants that are impaired in miRNA biogenesis, while the functions of specific miRNAs have been established by either expressing miRNA-resistant versions of target genes or overexpressing miRNAs. The roles of miRNAs are also investigated by loss-of-function analyses of MIR genes or by interfering with mature miRNAs6,7,8. Despite identification of a large number of miRNAs from a variety of plant species, the functions of most individual miRNAs remain unclear, particularly in plant species other than Arabidopsis.
In rice (Oryza sativa L.), miRNAs play a remarkably wide number of roles in growth and development. Rice has six DICER-LIKE (OsDCL) genes with OsDCL1 being the major enzyme for producing mature miRNAs9,10,11. Strong OsDCL1 loss-of-function mutants are not viable, while weak lines exhibit pleiotropic defects. The rice AGO gene MEIOSIS ARRESTED AT LEPTOTENE1 (MEL1) is required for meiosis12. Loss-of-function studies of WAF1, an ortholog of Arabidopsis HEN1, have shown that WAF1 is responsible for shoot development via maintaining miRNAs and trans-acting-small interfering RNAs (ta-siRNAs)13. Characterization of the IPA1 (Ideal Plant Architecture 1) trait locus has revealed that the OsmiR156-controlled OsSPL14 is critical for grain yield14. In addition, OsmiR156 may target OsSPL16 to control grain size, shape and quality15. Furthermore, the gradually increased expression of OsmiR156 in leaves is important for leaf development16. Overexpression of OsmiR172 represses expression of AP2-like genes, which consequently causes abnormal floral meristem identity and defects in flower organ and seed development17,18. OsTIR1 and OsAFB2 are predicted targets of OsmiR393. Transgenic plants overexpressing OsmiR393a/b have defects similar to those observed in auxin signaling mutants19. Although various miRNAs have been found in rice20,21, it is imperative to study functions of individual OsmiRNAs in rice growth and development, particularly those of agricultural importance.
MiR160 is essential for plant growth and development6. In Arabidopsis, miR160 targets AUXIN RESPONSE FACTOR 10 (ARF10), ARF16 and ARF17. Expression of the miR160-resistant version of ARF16 (mARF16) results in reduced fertility and less lateral roots22. Plants expressing mARF17 exhibited pleiotropic defects in vegetative and reproductive development23. Analyses using similar approaches have revealed that the miR160-controlled ARF10 is essential for seed germination and many post-embryonic growth and developmental processes through the regulation of both auxin and ABA signaling24. Recent studies have shown the dormancy of mARF10 and mARF16 seeds is increased25. In tomato, SlymiR160 and the SlymiR160a target SlyARF10 are required for floral organ and early fruit development26,27. Moreover, in soybean, miR160 is involved in auxin and cytokinin signaling during nodulation28. Although miR160 is conserved in plants29,30, the role of miR160 in monocots is unknown. In this report, we examined expression of six OsMIR160 genes and found that the mature OsmiR160 was mainly derived from OsMIR160a and OsMIR160b genes. In addition, the OsARF18 transcript was cleaved at the OsmiR160 target site. We generated transgenic rice plants (named mOsARF18) that expressed an OsmiR160-resistant version of OsARF18. Phenotypic analyses revealed that mOsARF18 plants showed pleiotropic defects in growth and development. Furthermore, auxin treatment induced expression of OsMIR160a, OsMIR160b and OsARF18, whereas expression of OsMIR160a and OsMIR160b as well as other genes involved in auxin signaling was altered in mOsARF18 plants. Our results support the idea that deregulation of the OsmiR160 target gene OsARF18 leads to abnormal growth and development in rice through affecting the auxin signaling.
Results
Expression of OsMIR160 genes and OsmiR160 target OsARF genes in rice
Rice has six OsMIR160 (OsMIR160a to OsMIR160f) genes. Four of six mature OsmiR160s (OsmiR160a to OsmiR160d) have identical sequences, while the other two (OsmiR160e and OsmiR160f) each differs by a single nucleotide (Fig. S1a). Our RT-PCR results showed that expression of OsMIR160a and OsMIR160b genes was significantly higher than that of the other four OsMIR160 genes in leaf (L), young inflorescence (YI), mature inflorescence (MI) and stem (ST) (Fig. S1b). Moreover, quantitative real-time RT-PCR (qRT-PCR) revealed that OsMIR160a and OsMIR160b genes had higher expression levels in leaf and inflorescences than that in seedling and stem (Fig. 1a,b). Our studies suggest that the mature OsmiR160 might be mainly derived from OsMIR160a and OsMIR160b genes.
Expression analysis of OsMIR160a, OsMIR160b and OsmiR160 potential target OsARFs.
(a,b) Quantitative real time RT-PCR (qRT-PCR) results showing expression of OsMIR160a (a) and OsMIR160b (b). (c–f) qRT-PCR) results showing expression of OsmiR160 potential target genes: OsARF8 (c), OsARF10 (d), OsARF18 (e) and OsARF22 (f). S: seedling (7-day old), L: leaf, ST: stem, YI: young inflorescence and MI: mature inflorescence. Gene expression levels in other organs were normalized based on expression observed in seedling. Stars indicate significant difference (P < 0.01). (g) An unrooted phylogenetic tree constructed by the Maximum-Likelihood method showing miR160 target ARFs in Arabidopsis and rice. (h) The 5′ RACE result showing that OsARF18 is a target of OsmiR160. The arrow points to the OsmiR160-directed cleavage site at the OsARF18 transcript.
Our analyses using the psRNATarget web server31 showed that OsARF8 (Os02g41800), OsARF10 (Os04g43910), OsARF18 (Os06g47150) and OsARF22 (Os10g33940) were potential OsmiR160 target genes. Further phylogenetic analysis found that all four targets were similar to Arabidopsis ARF10 and ARF16, whereas OsARF18 had the highest similarity to the Arabidopsis ARF16 (Fig. 1g; Fig. S2). Our qRT-PCR results demonstrated that the expression of OsARF8 mainly occurred in young and mature inflorescences (Fig. 1c), while OsARF10 was primarily expressed in seedling and stem (Fig. 1d). OsARF18 was predominantly expressed in leaf as well as young and mature inflorescences (Fig. 1e). Expression levels of OsARF22 appeared similar in all examined organs except in leaf (Fig. 1f). A previous study using the “degradome sequencing” approach has shown that cleavage frequencies associated with OsARF18 and OsARF22 are the highest, but frequencies for OsARF8 and OsARF10 are extremely low32. We further identified that the OsARF18 transcript was cleaved at the OsmiR160 target site using a gene-specific 5′ RACE (Fig. 1h). Therefore, our results suggest that OsARF18 (Os06g47150) is a promising target gene of OsmiR160.
Negative regulation of OsARF18 by OsmiR160 is essential for rice growth and development
To investigate the function of OsmiR160 in rice, we chose its promising target OsARF18 to generate transgenic plants expressing an OsmiR160-resistant version of OsARF18 (resulting plants named mOsARF18) in the Oryza sativa Japonica cv. Nipponbare background (Fig. 2a). Our qRT-PCR results show that the expression of un-cleaved OsARF18 was significantly increased in leaf, stem and inflorescence of three examined mOsARF18 transgenic lines (Fig. 2b).
Generation of transgenic rice plants expressing an OsmiR160-resistant version of OsARF18 (mOsARF18).
(a) Domain structure of the OsARF18 protein. The B3 DNA binding domain (B3) and the C-terminal dimerization domain (CTD) are indicated in black and gray, respectively. The OsmiR160 complementary (or target) sequence in the OsARF18 mRNA and the corresponding region of amino acid sequence (GIQGAR) are shown. The silent mutations were created in mOsARF18 by introducing silent mutations (indicated by*). (b) qRT-PCR results showing that expression levels of un-cleaved OsARF18 were significantly (*P < 0.01) increased in leaf, stem and inflorescence of three examined independent mOsARF18 transgenic lines.
Eleven out of 19 mOsARF18 independent transgenic lines exhibited strong but similar defects in both growth and development, while the rest of lines showed mild phenotypes. Compared to wild-type plants, mOsARF18 lines were dwarf and formed less tillers (Fig. 3a,g,h). mOsARF18 plants produced short and rolled leaves (Fig. 3b; Fig. S3; Fig. 4a–c). mOsARF18 plants were also defective in reproduction, as indicated by abnormal flower and seed development (Fig. 3c–f). The lemma and palea did not enclose flowers (Fig. 3c). After fertilization, stamens remained attached to developing seeds, suggesting abnormal senescence of stamens (Fig. 3d). Moreover, mOsARF18 lines showed reduced seed setting (Fig. 3i). Although no change in seed length (Fig. 3e,f,j), mOsARF18 seeds had reduced width and weight when compared with wild type (Fig. 3e,f,k,l). Our results suggest that deregulation of OsARF18 results in abnormal growth and development in rice.
mOsARF18 transgenic plants exhibited pleiotropic defects in growth and development.
(a) Forty-day old wild-type (left), mOsARF18-3 (middle) and mOsARF18-5 (right) plants. Bar: 10 cm. (b) The fifth leaves of wild-type (left), mOsARF18–3 (middle) and mOsARF18-5 (right) plants. Bar: 1 mm. (c,d) Ten-DAP (Days After Pollination) wild-type (left), mOsARF18-3 (middle) and mOsARF18-5 (right) developing seeds. Bars: (c,d) 1 mm. Lemma and palea were removed in (d) to show attached stamens. (e,f) Wild-type (e) and mOsARF18-3 (f) mature seeds. Bars: (e,f) 1 mm. (g,h) Sixty-day old transgenic plants showed significantly (*P < 0.01) decreased pant height (g) and tiller numbers (h) than that of wild type plants. (i–l) Mature transgenic plant seeds showed no difference in seed length (j), but significantly (*P < 0.01) decreased seed setting (i), width (k) and weight (l).
mOsARF18 plants were abnormal in leaf cell division and differentiation as well as starch accumulation in seeds.
(a–c) Hand cutting of the fifth leaves of wild-type (a), mOsARF18-3 (b) and mOsARF18-5 (c) plants. Bars: (a) 500 μm; (b,c) 200 μm. (d–h) Semi-thin sections of the fifth leaves of wild-type, mOsARF18-3 and mOsARF18-5 plants: (d) wild type; (e,g) mOsARF18-3; and (f,h) mOsARF18-5. Arrows indicate bulliform cells. Arrowheads indicate the lack of typical bulliform cells. Red brackets indicate regions between two vascular bundles for epidermal cell counting. Bars: (d–h) 50 μm. (i–k) Numbers of bulliform cells (i), total vascular bundles (j) and abaxial epidermal cells between two vascular bundles (k) were significantly (*P < 0.01) decreased in leaves of mOsARF18-3, mOsARF18-5 and mOsARF18-8 transgenic lines in comparison with wild-type plants. (l) Endosperm in 10-DAP (Days After Pollination) wild-type developing seeds showing strongly stained starch granules by I-KI (iodine-potassium iodide). (m,n) Endosperm in 10-DAP mOsARF18-3 (b) and mOsARF18-5 (c) developing seeds showing small and weakly stained starch granules. Bars: (l–n) 20 μm.
Cell division and differentiation were abnormal in mOsARF18 leaves
To further examine leaf defects in mOsARF18 plants, we analyzed the structure of the fifth mature leaf via cross-section. Compared with wild type, mOsARF18 lines produced rolled leaves (Fig. 4a–c). Bulliform cells are specialized epidermal cells between two vascular bundles on the adaxial blade. Bulliform cells are large and highly vacuolated, which are important for leaf rolling through turgor pressure regulation33. Wild-type bulliform cells were arranged in groups of approximately 5 cells, among which the middle bulliform cell was larger than others along both sides (Fig. 4d,i). However, in mOsARF18 leaves, we observed reduced number and size of bulliform cells, or bulliform cells were absent (Fig. 4e,f). Further statistical analysis showed that bulliform cell numbers between two vascular bundles in mOsARF18 leaves were significantly reduced when compared with that of wild type (Fig. 4i). Numbers of total vascular bundles and abaxial epidermal cells between two vascular bundles were also decreased (Fig. 4d,g,h,j,k). In addition, with the exception of the bulliform cells, we did not observe significant size differences in other cells when comparing mOsARF18 leaves with the wild-type. Thus, our results suggest that repression of OsARF18 by OsmiR160 is important for epidermal cell division and bulliform cell differentiation during leaf development.
Starch accumulation during seed development was abnormal in mOsARF18 plants
To study why mOsARF18 plants produced small seeds, we examined starch accumulation in developing seeds. Our results demonstrated that endosperm in wild-type developing seeds contained many starch granules that were strongly stained by iodine (Fig. 4l), whereas endosperm in mOsARF18 developing seeds contained smaller, weakly stained starch granules (Fig. 4m,n). In conclusion, mOsARF18 plants were defective in starch accumulation during seed development.
The OsmiR160-regulated OsARF18 may control rice growth and development via affecting auxin signaling
ARFs play a primary role in auxin signaling. To test whether auxin signaling was affected in mOsARF18 transgenic plants, we examined expression of representative genes known to be involved in auxin signaling in rice, including the AUXIN RESPONSIVE FACTOR gene OsARF234, the auxin responsive gene OsGH3-135, the AUX1-LIKE gene OsLAX2 (OsAUX3 or OsRAU2; Os03g14080)36, the auxin efflux gene OsPIN1b37 and the auxin biosynthesis gene OsYUCCA238 via qRT-PCR. Expression levels of all tested genes were significantly decreased in mOsARF18 transgenic plant leaves (Fig. 5a–e). Thus, our results suggest that deregulation of OsARF18 affects auxin signaling, which might cause abnormal growth and development in rice.
Analysis of auxin signaling gene expression in mOsARF18 transgenic plants.
qRT-PCR results showing that expression of OsARF2 (a), OsGH3-1 (b), OsLAX2 (c), OsPIN1b (d) and OsYUCCA2 (e) was significantly decreased in leaves of mOsARF18 transgenic plants. Gene expression levels in mOsARF18-3, mOsARF18-5 and mOsARF18-8 were normalized based on expression observed in wild type. Stars indicate significant difference (P < 0.01).
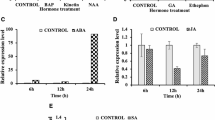
We further tested whether auxin affected expression of OsMIR160a, OsMIR160b and OsmiR160 target genes. Our qRT-PCR results showed that expression of OsMIR160a was significantly increased after 20 and 40-minute NAA treatment (Fig. 6a), while expression of OsMIR160b was significantly induced after 20-minute NAA treatment and then remained normal (Fig. 6b). NAA treatment also significantly induced expression of OsARF18 after 20 and 40 minutes (Fig. 6e) as well as expression of OsARF10 after 180 minutes (Fig. 6d). However, expression of OsARF8 (Fig. 6c) and OsARF22 (Fig. 6f) remained unchanged after NAA treatment.
Auxin up regulated expression of OsMIR160a, OsMIR160b and OsmiR160 target genes, whereas OsARF18 suppressed the expression of OsMIR160a.
(a–f) qRT-PCR results showing that expression changes of OsMIR160a, OsMIR160b and OsmiR160 target genes in seedlings (7-day old) treated with NAA for 20, 40 and 180 minutes. Expression levels without NAA treatment (0 minute) were used to normalize expression with treatments. Stars indicate significant difference (P < 0.01). Expression of OsMIR160a was significantly induced after 20 and 40-minute treatment (a), while expression of OsMIR160b was significantly increased only after 20-minute treatment (b). Expression of OsARF8 (c) and OsARF22 (f) remained unchanged. Expression of OsARF10 was significantly induced after 180-minute treatment (d), whereas expression of OsARF18 was significantly increased after 20 and 40-minute treatment (e). (g,h) qRT-PCR results showing that expression of OsMIR160a and OsMIR160b was significantly decreased in mOsARF18 transgenic lines in comparison with wild-type plants. Gene expression levels in #3 (mOsARF18-3), #5 (mOsARF18-5) and #8 (mOsARF18-8) were normalized based on expression observed in wild type. Stars indicate significant difference (P < 0.01).
Conversely, expression of OsMIR160a and OsMIR160b was found to be decreased in mOsARF18 plants (Fig. 6g,h), in which the expression of OsARF18 was highly increased (Fig. 2b). Our results suggest that auxin up regulates expression of OsMIR160a, OsMIR160b and OsARF18, whereas OsARF18 represses expression of OsMIR160a and OsMIR160b. Therefore, the positive regulation of OsMIR160a and OsMIR160b expression by auxin and the negative regulation of OsMIR160a and OsMIR160b by OsARF18 might be important for OsmiR160 to fine-tune auxin signaling in a negative feedback loop manner.
To test whether OsmARF18 seedlings are defective in auxin signaling, we treated OsmARF18 seedlings hydroponically with 1 μM of NAA for 7 days. We found that OsmARF18 seedlings produced significantly shorter primary roots than that of wild type (Fig. 7a,b,e). Primary root growth was significantly inhibited by NAA treatment in both wild type and OsmARF18 seedlings (Fig. 7c–e); however, the NAA inhibition effect on primary root length was similar in wild type and OsmARF18 seedlings, because there was no change in relative primary root length (percentage of root length between that treated and untreated) after the NAA treatment (Fig. 7f). Numbers of lateral roots were similar in wild type and OsmARF18 seedlings (Fig. 7a,b,g). NAA treatment significantly increased the lateral root number in wild type seedlings (Fig. 7a,c,g), conversely, the lateral root number in OsmARF18 seedlings was significantly decreased with the NAA treatment (Fig. 7b,d,g). Our results suggest that OsARF18 might be involved in auxin-regulated lateral root formation in rice.
Effect of NAA treatment on lateral root formation.
(a,c) Seven-day wild type (WT) seedlings without NAA (a) and with NAA (1 μM) (c) treatment (inset showing high magnification of roots in (c). (b,d) Seven -day OsmARF18 seedlings without NAA (b) and with NAA (1 μM) (d) treatment (inset showing high magnification of roots in (d). (e,f) Primary roots in WT 7-day seedlings were significantly (*indicating P < 0.01) longer than that of OsmARF18 seedlings. NAA treatment significantly (P < 0.01) inhibited primary root length of both WT and OsmARF18 seedlings, but the relative primary root length (percentage of root length between that treated and untreated) was similar. (g) Seven-day WT and OsmARF18 seedlings produced similar numbers of lateral roots. After NAA treatment, lateral root numbers in WT seedlings was significantly (P < 0.01) increased, whereas that in OsmARF18 seedlings was significantly (P < 0.01) decreased.
Discussion
Conserved and diverse roles of miR160 in plant growth and development
MiR160 is conserved throughout the plant kingdom from mosses to higher plants29,30. Sequence similarity of mature miR160s is more than 80% among different species. MiR160 target genes are also conserved, although their numbers vary with species. Arabidopsis contains MIR160a, MIR160b and MIR160c three genes which produce the same mature miR1606. MiR160 targets AUXIN RESPONSE FACTOR 10 (ARF10), ARF16 and ARF17 three genes and each target gene has conserved but some distinct functions in Arabidopsis. Plants expressing the miR160-resistant version of ARF10 (mARF10) exhibit pleiotropic defects that resemble phenotypes of some ABA and auxin defective mutants24, while mARF16 plants have reduced fertility and less lateral roots22. ARF10 and ARF16 are required for maintaining the expression of ABI3 gene25, suggesting that ARF10 and ARF16 are involved in both auxin and ABA signaling or cross-talk between them. ARF17 plays a general role in both vegetative and reproductive development via modulating expression of early auxin response genes23. SlymiR160 and the SlymiR160a target SlyARF10 control ovary patterning, early fruit development and floral organ abscission in tomato26,27, whereas the soybean miR160 regulates auxin and cytokinin signaling during nodulation28.
Six OsMIR160 (OsMIR160a to OsMIR160f) genes are found in rice. The mature OsmiR160 has OsARF8 (Os02g41800), OsARF10 (Os04g43910), OsARF18 (Os06g47150) and OsARF22 (Os10g33940) four potential target genes. In this study, we characterized the function of OsARF18 (Os06g47150), an orthologue of Arabidopsis ARF16. Arabidopsis mARF16 and rice mOsARF18 plants show some similar but also different defects in growth and reproductive development. In Arabidopsis, mARF16 plants produce curved leaves22. Our studies showed that the formation of rolled leaves in mOsARF18 plants was caused by alterations in shape, size and number of bulliform cells. Auxin plays a key role in controlling leaf shape39. Phenotypic analysis indicates that alteration of bulliform cells on the adaxial leaf blade surface is tightly linked to the formation of rolled leaves in rice. Disruption of the rice CONSTITUTIVELY WILTED1 (OsCOW1) gene, a member of OsYUCCA gene family, results in rupture of the largest bulliform cell and consequently rolled leaves40. In addition, alterations in number and size of bulliform cells lead to rolled leaves in the narrow leaf 7 (nal7) mutant, which is an oscow1 allele41. Therefore, our data support that the OsmiR160 target OsARF18 controls leaf shape via affecting auxin signaling. Besides rolled leaves, mOsARF18 plants produced less and smaller seeds than wild type. Moreover, starch accumulation in seeds from mOsARF18 plants was significantly decreased when compared to wild-type seeds. Collectively, miR160 plays conserved and diverse roles in plant growth and development. It will be interesting to test the loss-of-function of OsmiR160 and functions of other OsmiR160 target genes in rice growth and development.
Negative regulation of OsARF18 by OsmiR160 is important for its normal function
MiRNAs negatively regulate gene expression at the post-transcriptional level by binding to mRNA complementary sequences for mRNA destabilization and translational inhibition in both plants and animals1,2,3,4,5. One miRNA normally has multiple target genes. Over/ectopic-expression of normal miRNA-target genes usually does not cause a change in phenotype, because over/ectopic-expressed normal mRNAs can be still targeted by miRNAs for cleavage. Therefore, a primary approach for studying the function of miRNA and its target is to express the miRNA-resistant version of individual target gene. Employing 35S, Ubi and ACTIN promoters, previous studies identified functions of many miRNAs and their target genes via expressing miRNA-resistant versions of target genes, such as 35S::mTCP2, 35S::mTCP3, 35S::mTCP4, 35S::mCUP1 and 35S::mCUP2 (Arabidopsis miR164 target genes)42,43, 35S::mAP2 (Arabidopsis miR172 target gene)44, 35S::mSlyARF10 (tomato miR160 target gene)26, UBI:mGRF6 (rice miR396 traget gene)45, as well as ACTIN::mOSHB1, ACTIN::mOSHB3 and ACTIN:: mOSHB5 (rice miR166 target genes)46.
So far, no studies report that over/ectopic-expression of normal versions of miR160 target genes (ARF10, ARF16 and ARF17) causes phenotypes in leaf and flower development, but only plants expressing miR160-resistant versions of miR160 target genes (mARF10, mARF16 and mARF17) exhibit various phenotypes22,23,24,26. In this study, we used the Ubiquitin (Ubi) promoter to drive mOsARF18 expression. To further rule out the possibility that the phenotype of mOsARF18 was caused by ectopic activity of the Ubi promoter, we generated ARF16, mARF16, OsARF18 and mOsARF18 Arabidopsis transgenic plants under the control of the Ubi promoter. Of 30 ARF16 and 32 OsARF18 transgenic plants that we obtained, none of them showed detectable defects in growth and development (Fig. S5a–c,f–h; Fig. S6a–c,f–h,k–m,p); however, mARF16 (32 out of 40) and mOsARF18 (20 out of 29) transgenic plants formed narrow and curled leaves and had short stature (Fig. S5d,e,i,j). In addition, mARF16 and mOsARF18 plants produced abnormal flowers and a lower number of smaller seeds compared with wild-type plants (Fig. S6d,e,i,j,n,o,q). Our results showed that mARF16 and mOsARF18 caused defects in plant growth and development, but plants ectopically expressing normal ARF16 and OsARF18 were similar to the wild type. Taken together, the results from Arabidopsis and rice suggest that the negative regulation of OsARF18 and its Arabidopsis orthologous gene ARF16 by rice and Arabidopsis miR160, respectively, is critical for their function.
Molecular mechanism of miR160 in fine-tuning auxin signaling
MiRNAs play a pivotal role in auxin signaling by negative regulation of ARFs. In Arabidopsis, miRNAs are involved in expression regulation of 8 of a total 23 ARFs (ARF2, 3, 4, 6, 8, 10, 16 and 17). MiR390-derived trans-acting-small interfering RNAs (ta-siRNAs) target ARF2, ARF3 and ARF447,48. The tasiRNA gradient is important for establishing the normal patterning of ARF3 protein during leaf development49. During growth of lateral roots, miR390 affects production of tasiRNAs and thus inhibits ARF2, ARF3 and ARF450. Conversely, auxin activates ARF2, ARF3 and ARF4, which consequently influences the formation of miR390. Therefore, the regulatory network modulated by miR390 maintains normal expression of ARF2, ARF3 and ARF4. Negative regulation of ARF6 and ARF8 by miR167 is essential for anther and ovule development51. Moreover, miR160 negatively regulates expression of ARF10, ARF16 and ARF176,22,23,24. In rice, OsmiRNAs are predicted to regulate 11 of a total of 25 identified OsARFs52,53. Functional disruption of OsDCL4 causes increased expression of OsmiR165/166 and three OsARFs, which are orthologs of Arabidopsis ARF2, ARF3 and ARF454. OsmiR160 is predicted to have four OsARF targets, including OsARF18 that was characterized in this study. OsmiR167 potentially targets OsARF6 (Os02g06910), OsARF12 (Os04g57610), OsARF17 (Os06g46410) and OsARF25 (Os12g41950), which are highly similar to Arabidopsis ARF6 and ARF854. Together, miRNAs target a group of similar ARF genes in both monocots and dicots.
Auxin promotes the SCFTIR1/AFB E3 ligase-mediated degradation of Aux/IAA proteins, which sequester ARFs55,56. Upon the perception of auxin, the released ARFs activate or suppress expression of a large set of auxin-responsive genes by binding to auxin response elements (AuxREs) in their regulatory regions. Based on transient expression studies and protein structures, among 23 ARFs in Arabidopsis, ARF5-8 and 19 act as activators, while ARF1-4, 9-18 and 20-22 may function as repressors57,58,59.
Previous studies have implied that the feedback regulation between miRNA and its target ARF genes could provide a fine-tuning mechanism to regulate auxin signaling. During adventitious root development in Arabidopsis, expression of ARF6 and ARF8 is regulated by miR167 and miR160, whereas the abundance of ARF17 transcripts is controlled by miR16060. In addition, ARF6 and ARF8 activators as well as ARF17 repressors positively and negatively regulate the expression of each other. Thus, the delicate balance between miRNAs and ARFs is critical for auxin-regulated developmental processes. Our studies show that OsmiR160 negatively regulates the expression of OsARF18 by cleaving OsARF18 transcripts. Auxin induces expression of OsMIR160a, OsMIR160b and OsARF18, whereas expression of OsMIR160a and OsMIR160b was suppressed by OsARF18. Our analyses revealed that promoters and 3′ regions of OsMIR160a and OsMIR160b had clusters of AuxRE cores and AuxRR cis elements (Fig. S4), which may create OsARF binding sites by which OsARF activators or repressors could regulate expression of OsMIR160a and OsMIR160b.
In our hypothetical model, auxin promotes the release of OsARF activators and repressors via the SCFTIR1/AFB E3 ligase (Fig. 8). The balance between OsARF activators and repressors decides up or down expression of OsMIR160 genes, which consequently affects the abundance of mature OsmiR160. Conversely, OsmiR160 negatively modulates expression of its target OsARFs. OsARFs also mutually control their own expression. Future studies should focus on examining the feedback loop regulation between OsmiR160 and its target OsARFs, which might be important for fine-tuning highly dose-sensitive auxin signaling during rice growth and development.
A hypothetical working model for auxin signaling that is modulated by OsmiR160 during rice growth and development.
Auxin promotes the release of OsARF activators or repressors via the auxin receptor SCF E3 ligase. The balance between OsARF activators and repressors decides up or down expression of OsMIR160 genes. Conversely, the change in abundance of mature OsmiR160 negatively regulates expression of its target OsARFs. OsARF activators and repressors might positively or negatively regulate expression of each other.
Manipulating miRNAs and their target genes has demonstrated improvement of many important crop traits, including biomass yield, grain yield, fruit yield, nutritional quality, abiotic stress resistance (e.g. drought, salinity, cold, heat, oxidative stress, nutrient deprivation tolerance and heavy metal detoxification) and biotic stress resistance (e.g. virus, bacteria, fungus and nematode resistance)14,15,61,62. Our results showed that OsmiR160 played a pivotal role in rice growth and development via regulating auxin signaling. In particular, OsmiR160 is essential for leaf and seed development in rice. Leaf shape is important for photosynthesis, respiration and transpiration. Moderate leaf rolling can enhance photosynthesis and stress responses by inhibiting water loss and radiant heat absorption, which, therefore, increases crop yield. Future studies on the molecular mechanism by which OsmiR160 modulates auxin signaling will lead to potential applications for improving crop agronomic traits.
Methods
Plant materials and growth condition
Rice (Oryza sativa L. Japonica Nipponbare) plants were grown in Metro-Mix 360 soil (Sun-Gro Horticulture, Agawam, MA, USA) supplemented with sand and iron in a walk-in growth chamber under a 12-hour light (28 °C)/12-hour dark (22 °C) photoperiod regime. Transgenic rice plants were generated at the Plant Transformation Facility at Iowa State University. In total, 124 mOsARF18 plants were obtained from 19 independent transgenic lines. For expression studies, seven-day old wild-type seedlings were treated with 1 μm of NAA for 20, 40 and 180 min with untreated 7-day old seedlings as control. For studying the effect of exogenous auxin on root development, geminated seeds were hydroponically grown in the 1/2 Kimura B nutrient solution (pH 5.6) containing 1 μm of NAA for 7 days with continuous light at 25 °C.
Vector construction and rice transformation
PCR reactions (Primers are shown in Table S1) were performed using the Phusion High-Fidelity DNA Polymerase (New England Biolabs, Ipswich, MA, USA). The OsARF18 cDNA was amplified from rice leaf cDNAs and then cloned into the pCR2.1 vector (Invitrogen, Grand Island, NY, USA), resulting in pCR2.1-OsARF18. Point mutations of OsARF18 were created by overlapping PCR to generate pCR2.1-mOsARF18. mOsARF18 was then subcloned into the modified pCambia1301 binary vector harboring the Gateway cassette sequence and the maize Ubiquitin (Ubi) promoter using the Gateway LR recombinase II enzyme mix (Invitrogen, Grand Island, NY, USA).
For rice transformation, the mOsARF18 construct was introduced into the Agrobacterium strain EHA101. The callus induction (from mature embryos of Japonica cv. Nipponbare seeds), Agrobacterium infection, co-cultivation, selection of transformed calli and plant regeneration were performed essentially as described previously63.
RT-PCR and real time qRT-PCR
Total RNAs were isolated from different rice tissues using the RNeasy Plant Mini Kit (Qiagen, Valencia, CA, USA). After determining the RNA quantification by the NanoDrop 2000c (Thermo Scientific, Bannockburn, IL, USA), RNA reverse transcription was conducted using the QuantiTect Reverse Transcription Kit (Qiagen, Valencia, CA, USA). Real-time PCR (DNA Engine Opticon 2 system, Hercules, California, USA) and data analysis were performed as previously described6. Expression of OsMIR160a through OsMIR160f was examined by RT-PCR. Expression of OsMIR160a, OsMIR160b, OsARF8, OsARF10, OsARF18 and OsARF22, as well as other auxin signaling related genes was tested by real-time qRT-PCR. Three biological repeats were conducted and each value indicates the average with the standard error. All primer sequences are shown in Table S1.
5′ RACE
Using the GeneRacerTM kit (Invitrogen, Grand Island, NY, USA), a gene-specific 5′-rapid amplification of cDNA ends was conducted as described previously23. Gene-specific primers for OsARF18 (Os06g47150) are shown in Table S1.
Semi-thin section analysis
Semi-thin sectioning was performed as described previously64,65. The fifth leaves of 6-week-old rice were fixed in 2.5% of glutaraldehyde and post-fixed with 1% of OsO4 at room temperature. Samples were dehydrated through a graded acetone series (10% increments) for 60 minutes each. Infiltrated was started with 20% of Spurr’s resin and then 40%, 60% and 80% of Spurr’s resin every 3 hours. Following infiltration in three changes of 100% Spurr’s resin for 24 hours each, samples were finally embedded in 100% Spurr’s resin and polymerized at 60 °C overnight. Semi-thin sections (0.5 μm) were made using an RMC MT-7 ultramicrotome (Reichert-Jung, Depew, NY, USA) and were stained with 0.25% Toluidine Blue O. Images were photographed by an Olympus BX51 microscope equipped with an Olympus DP 70 digital camera (Olympus, Center Valley, PA, USA).
Histological detection of starch
Ten-DAP (Days After Pollination) seeds were fixed in FAA (50% ethanol, 10% formalin, 5% acetic acid). Following fixation, samples were dehydrated through an ethanol series, embedded in paraffin and sectioned at 8 μm with a Spencer 820 microtome. Sections were dewaxed and stained with Lugol’s iodine solution (6 mM iodine, 43 mM KI and 0.2 N HCl) for detection of starch granules. Images were photographed with an Olympus BX51 microscope equipped with an Olympus DP 70 digital camera (Olympus, Center Valley, PA, USA).
Additional Information
How to cite this article: Huang, J. et al. Deregulation of the OsmiR160 Target Gene OsARF18 Causes Growth and Developmental Defects with an Alteration of Auxin Signaling in Rice. Sci. Rep. 6, 29938; doi: 10.1038/srep29938 (2016).
References
Bartel, D. P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Carrington, J. C. & Ambros, V. Role of microRNAs in plant and animal development. Science 301, 336–338 (2003).
Jones-Rhoades, M. W., Bartel, D. P. & Bartel, B. MicroRNAs and their regulatory roles in plants. Annu. Rev. Plant Biol. 57, 19–53 (2006).
Voinnet, O. Origin, biogenesis and activity of plant microRNAs. Cell 136, 669–687 (2009).
Rogers, K. & Chen, X. Biogenesis, turnover and mode of action of plant microRNAs. Plant Cell 25, 2383–2399 (2013).
Liu, X. et al. The role of floral organs in carpels, an Arabidopsis loss-of-function mutation in MicroRNA160a, in organogenesis and the mechanism regulating its expression. Plant J. 62, 416–428 (2010).
Todesco, M., Rubio-Somoza, I., Paz-Ares, J. & Weigel, D. A collection of target mimics for comprehensive analysis of microRNA function in Arabidopsis thaliana. Plos Genet. 6, e1001031 (2010).
Yan, J. et al. Effective small RNA destruction by the expression of a short tandem target mimic in Arabidopsis. Plant Cell 24, 415–427 (2012).
Liu, B. et al. Loss of function of OsDCL1 affects microRNA accumulation and causes developmental defects in rice. Plant Physiol. 139, 296–305 (2005).
Song, X. et al. Roles of DCL4 and DCL3b in rice phased small RNA biogenesis. Plant J. 69, 462–474 (2012).
Liu, B. et al. Oryza sativa dicer-like4 reveals a key role for small interfering RNA silencing in plant development. Plant Cell 19, 2705–2718 (2007).
Nonomura, K. I. et al. A germ cell-specific gene of the ARGONAUTE family is essential for the progression of premeiotic mitosis and meiosis during sporogenesis in rice. Plant Cell 19, 2583–2594 (2007).
Abe, M. et al. WAVY LEAF1, an ortholog of Arabidopsis HEN1, regulates shoot development by maintaining MicroRNA and trans-acting small interfering RNA accumulation in rice. Plant Physiol. 154, 1335–1346 (2010).
Jiao, Y. et al. Regulation of OsSPL14 by OsmiR156 defines ideal plant architecture in rice. Nature Genet. 42, 541–544 (2010).
Wang, S. et al. Control of grain size, shape and quality by OsSPL16 in rice. Nature Genet. 44, 950–954 (2012).
Xie, K. et al. Gradual increase of miR156 regulates temporal expression changes of numerous genes during leaf development in rice. Plant Physiol. 158, 1382–1394 (2012).
Zhu, Q. H., Upadhyaya, N. M., Gubler, F. & Helliwell, C. A. Over-expression of miR172 causes loss of spikelet determinacy and floral organ abnormalities in rice (Oryza sativa). BMC Plant Biol. 9, 149 (2009).
Lee, D. Y. & An, G. Two AP2 family genes, supernumerary bract (SNB) and Osindeterminate spikelet 1 (OsIDS1), synergistically control inflorescence architecture and floral meristem establishment in rice. Plant J 69, 445–461 (2012).
Bian, H. et al. Distinctive expression patterns and roles of the miRNA393/TIR1 homolog module in regulating flag leaf inclination and primary and crown root growth in rice (Oryza sativa). New Phytol. 196, 149–161 (2012).
Sunkar, R., Girke, T., Jain, P. K. & Zhu, J. K. Cloning and characterization of microRNAs from rice. Plant Cell 17, 1397–1411 (2005).
Jeong, D. H. et al. Massive analysis of rice small RNAs: mechanistic implications of regulated microRNAs and variants for differential target RNA cleavage. Plant Cell 23, 4185–4207 (2011).
Wang, J. W. et al. Control of root cap formation by MicroRNA-targeted auxin response factors in Arabidopsis. Plant Cell 17, 2204–2216 (2005).
Mallory, A. C., Bartel, D. P. & Bartel, B. MicroRNA-directed regulation of Arabidopsis AUXIN RESPONSE FACTOR17 is essential for proper development and modulates expression of early auxin response genes. Plant Cell 17, 1360–1375 (2005).
Liu, P. P. et al. Repression of AUXIN RESPONSE FACTOR10 by microRNA160 is critical for seed germination and post-germination stages. Plant J. 52, 133–146 (2007).
Liu, X. et al. Auxin controls seed dormancy through stimulation of abscisic acid signaling by inducing ARF-mediated ABI3 activation in Arabidopsis. Proc. Natl. Acad. Sci. USA 110, 15485–15490 (2013).
Hendelman, A., Buxdorf, K., Stav, R., Kravchik, M. & Arazi, T. Inhibition of lamina outgrowth following Solanum lycopersicum AUXIN RESPONSE FACTOR 10 (SlARF10) derepression. Plant Mol. Biol. 78, 561–576 (2012).
Damodharan, S., Zhao, D. & Arazi, T. A common miRNA160-based mechanism regulates ovary patterning, floral organ abscission and lamina outgrowth in tomato. Plant J. doi: 10.1111/tpj.13127 (2016).
Turner, M. et al. Ectopic expression of miR160 results in auxin hypersensitivity, cytokinin hyposensitivity and inhibition of symbiotic nodule development in soybean. Plant Physiol. 162, 2042–2055 (2013).
Axtell, M. J., Snyder, J. A. & Bartel, D. P. Common functions for diverse small RNAs of land plants. Plant Cell 19, 1750–1769 (2007).
Luo, Y., Guo, Z. & Li, L. Evolutionary conservation of microRNA regulatory programs in plant flower development. Dev. Biol. 380, 133–144 (2013).
Dai, X. & Zhao, P. X. psRNATarget: a plant small RNA target analysis server. Nucleic Acids. Res. 39, W155–159 (2011).
Li, Y. F. et al. Transcriptome-wide identification of microRNA targets in rice. Plant J. 62, 742–759 (2010).
Zhang, G. H., Xu, Q., Zhu, X. D., Qian, Q. & Xue, H. W. SHALLOT-LIKE1 is a KANADI transcription factor that modulates rice leaf rolling by regulating leaf abaxial cell development. Plant Cell 21, 719–735 (2009).
Wang, D. et al. Genome-wide analysis of the auxin response factors (ARF) gene family in rice (Oryza sativa). Gene 394, 13–24 (2007).
Zhao, S. Q., Xiang, J. J. & Xue, H. W. Studies on the rice LEAF INCLINATION1 (LC1), an IAA-amido synthetase, reveal the effects of auxin in leaf inclination control. Mol. Plant 6, 174–187 (2013).
Chhun, T. et al. Saturated humidity accelerates lateral root development in rice (Oryza sativa L.) seedlings by increasing phloem-based auxin transport. J. Exp. Bot. 58, 1695–1704 (2007).
Li, G. et al. Rice actin-binding protein RMD is a key link in the auxin-actin regulatory loop that controls cell growth. Proc. Natl. Acad. Sci. USA 111, 10377–10382 (2014).
Yamamoto, Y., Kamiya, N., Morinaka, Y., Matsuoka, M. & Sazuka, T. Auxin biosynthesis by the YUCCA genes in rice. Plant Physiol. 143, 1362–1371 (2007).
Vanneste, S. & Friml, J. Auxin: a trigger for change in plant development. Cell 136, 1005–1016 (2009).
Woo, Y. M. et al. Constitutively wilted 1, a member of the rice YUCCA gene family, is required for maintaining water homeostasis and an appropriate root to shoot ratio. Plant Mol. Biol. 65, 125–136 (2007).
Fujino, K. et al. NARROW LEAF 7 controls leaf shape mediated by auxin in rice. Mol Genet. Genomics 279, 499–507 (2008).
Baker, C. C., Sieber, P., Wellmer, F. & Meyerowitz, E. M. The early extra petals1 mutant uncovers a role for MicroRNA miR164c in regulating petal number in Arabidopsis. Curr. Biol. 15, 303–315 (2005).
Koyama, T., Furutani, M., Tasaka, M. & Ohme-Takagi, M. TCP transcription factors control the morphology of shoot lateral organs via negative regulation of the expression of boundary-specific genes in Arabidopsis. Plant Cell 19, 473–484 (2007).
Chen, X. A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 303, 2022–2025 (2004).
Liu, H. et al. OsmiR396d-regulated OsGRFs function in floral organogenesis in rice through binding to their targets OsJMJ706 and OsCR4. Plant Physiol. 165, 160–174 (2014).
Itoh, J., Hibara, K., Sato, Y. & Nagato, Y. Developmental role and auxin responsiveness of Class III homeodomain leucine zipper gene family members in rice. Plant Physiol. 147, 1960–1975 (2008).
Williams, L., Carles, C. C., Osmont, K. S. & Fletcher, J. C. A database analysis method identifies an endogenous trans-acting short-interfering RNA that targets the Arabidopsis ARF2, ARF3 and ARF4 genes. Proc. Natl. Acad. Sci. USA 102, 9703–9708 (2005).
Fahlgren, N. et al. Regulation of AUXIN RESPONSE FACTOR3 by TAS3 ta-siRNA affects developmental timing and patterning in Arabidopsis. Curr. Biol. 16, 939–944 (2006).
Chitwood, D. H. et al. Pattern formation via small RNA mobility. Genes Dev. 23, 549–554 (2009).
Marin, E. et al. miR390, Arabidopsis TAS3 tasiRNAs and their AUXIN RESPONSE FACTOR targets define an autoregulatory network quantitatively regulating lateral root growth. Plant Cell 22, 1104–1117 (2010).
Wu, M. F., Tian, Q. & Reed, J. W. Arabidopsis microRNA167 controls patterns of ARF6 and ARF8 expression and regulates both female and male reproduction. Development 133, 4211–4218 (2006).
Guilfoyle, T. J. & Hagen, G. Auxin response factors. Curr. Opin. Plant Biol. 10, 453–460 (2007).
Shen, C. et al. Functional analysis of the structural domain of ARF proteins in rice (Oryza sativa L.). J. Exp. Bot. 61, 3971–3981 (2010).
Nagasaki, H. et al. The small interfering RNA production pathway is required for shoot meristem initiation in rice. Proc. Natl. Acad. Sci. USA 104, 14867–14871 (2007).
Dharmasiri, N., Dharmasiri, S. & Estelle, M. The F-box protein TIR1 is an auxin receptor. Nature 435, 441–445 (2005).
Tan, X. et al. Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature 446, 640–645 (2007).
Ulmasov, T., Hagen, G. & Guilfoyle, T. J. Activation and repression of transcription by auxin-response factors. Proc. Natl. Acad. Sci. USA 96, 5844–5849 (1999).
Wang, S., Tiwari, S. B., Hagen, G. & Guilfoyle, T. J. AUXIN RESPONSE FACTOR7 restores the expression of auxin-responsive genes in mutant Arabidopsis leaf mesophyll protoplasts. Plant Cell 17, 1979–1993 (2005).
Boer, D. R. et al. Structural basis for DNA binding specificity by the auxin-dependent ARF transcription factors. Cell 156, 577–589 (2014).
Gutierrez, L. et al. Phenotypic plasticity of adventitious rooting in Arabidopsis is controlled by complex regulation of AUXIN RESPONSE FACTOR transcripts and microRNA abundance. Plant Cell 21, 3119–3132 (2009).
Zhang, B. MicroRNA: a new target for improving plant tolerance to abiotic stress. J. Exp. Bot. 66, 1749–1761 (2015).
Zhou, M. & Luo, H. MicroRNA-mediated gene regulation: potential applications for plant genetic engineering. Plant Mol. Biol. 83, 59–75 (2013).
Hiei, Y., Ohta, S., Komari, T. & Kumashiro, T. Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J. 6, 271–282 (1994).
Jia, G., Liu, X., Owen, H. A. & Zhao, D. Signaling of cell fate determination by the TPD1 small protein and EMS1 receptor kinase. Proc. Natl. Acad. Sci. USA 105, 2220–2225 (2008).
Zhao, D. Z., Wang, G. F., Speal, B. & Ma, H. The EXCESS MICROSPOROCYTES1 gene encodes a putative leucine-rich repeat receptor protein kinase that controls somatic and reproductive cell fates in the Arabidopsis anther. Genes Dev. 16, 2021–2031 (2002).
Acknowledgements
We thank X. Li for his early involvement in this project, H.A. Owen, A. Smith, S. Malik and J. Skurski for critical reading of the manuscript. We are grateful for K. Wang, whose lab performed rice transformation. We also thank Z. Yang for generously offering us the binary vector for rice transformation, H.A. Owen for the phenotypic analysis and T. Schuck and J. Gonnering for the plant care. This work was in part supported by the National Science Foundation (NSF IOS-0721192 and IOS-1322796), the Research Growth Initiative (RGI) at the University of Wisconsin-Milwaukee and the UW-Madison/UW-Milwaukee Intercampus Research Incentive Grants Program. D. Zhao also thankfully acknowledges the support of the Shaw Scientist Award from the Greater Milwaukee Foundation.
Author information
Authors and Affiliations
Contributions
D.Z. and H.J. conceived and designed experiments. H.J. and Z.L. conducted experiments. D.Z. and H.J. wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Huang, J., Li, Z. & Zhao, D. Deregulation of the OsmiR160 Target Gene OsARF18 Causes Growth and Developmental Defects with an Alteration of Auxin Signaling in Rice. Sci Rep 6, 29938 (2016). https://doi.org/10.1038/srep29938
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep29938
- Springer Nature Limited
This article is cited by
-
Development of miRNA-SSR and target-SSR markers from yield-associate genes and their applicability in the assessment of genetic diversity and association mapping in rice (Oryza sativa L.)
Molecular Breeding (2024)
-
Negative regulators of grain yield and mineral contents in rice: potential targets for CRISPR-Cas9-mediated genome editing
Functional & Integrative Genomics (2023)
-
MicroRNA-mediated host defense mechanisms against pathogens and herbivores in rice: balancing gains from genetic resistance with trade-offs to productivity potential
BMC Plant Biology (2022)
-
Degradome sequencing reveals an integrative miRNA-mediated gene interaction network regulating rice seed vigor
BMC Plant Biology (2022)
-
Target-Mimicry-Based miR167 Diminution Confers Salt-Stress Tolerance During In Vitro Organogenesis of Tobacco (Nicotiana tabacum L.)
Journal of Plant Growth Regulation (2022)