Abstract
Organoids are simple tissue-engineered cell-based in vitro models that recapitulate many aspects of the complex structure and function of the corresponding in vivo tissue. They can be dissected and interrogated for fundamental mechanistic studies on development, regeneration and repair in human tissues, and can also be used in diagnostics, disease modelling, drug discovery and personalized medicine. Organoids are derived from either pluripotent or tissue-resident stem (embryonic or adult) or progenitor or differentiated cells from healthy or diseased tissues, such as tumours. To date, numerous organoid engineering strategies that support organoid culture and growth, proliferation, differentiation and maturation have been reported. This Primer highlights the rationale underlying the selection and development of these materials and methods to control the cellular/tissue niche; and therefore, the structure and function of the engineered organoid. We also discuss key considerations for generating robust organoids, such as those related to cell isolation and seeding, matrix and soluble factor selection, physical cues and integration. The general standards for data quality, reproducibility and deposition within the organoid community are also outlined. Lastly, we conclude by elaborating on the limitations of organoids in different applications, and the key priorities in organoid engineering for the coming years.
Similar content being viewed by others
Introduction
Stem cells are critical in maintaining organ size, structure and function through cellular renewal, migration, differentiation and apoptosis1. Stem cells reside in a defined microenvironment commonly referred to as the stem cell niche to regulate stem cell fate2. Given the importance of these environmental cues, there have been numerous tissue engineering attempts to mimic the stem cell niche in vitro to achieve high spatio-temporal control over cell–cell and cell–matrix interactions and reproduce mechano-chemical cues using engineered hydrogels and micro-devices3,4. In 1977, Matrigel, a basement membrane extracellular matrix (ECM) containing a unique mix of ECM components and growth factors, was extracted from mouse sarcoma tumours and used to support in vitro cell culture5. Matrigel was later shown to allow breast epithelial cells to grow in three dimensions and form lumens with milk protein secretion6, and adult intestinal stem cells embedded in Matrigel in the presence of a tissue-specific cocktail of growth factors were also shown to self-organize into 3D crypt–villus structures7. Organoid research intertwined with 3D cell culture, stem cell and tissue engineering for over a century, with various debates on the definition, standard and scope.
An organoid is a self-organized 3D tissue that is typically derived from stem cells (pluripotent, fetal or adult), and which mimics the key functional, structural and biological complexity of an organ8,9,10,11. Cells comprising organoids can be derived from induced pluripotent stem cells (iPSCs) or tissue-derived cells (TDCs), including normal stem/progenitor cells, differentiated cells and cancer cells12. Compared with conventional 2D cultures and animal models, organoid cultures enable patient specificity in the model while recapitulating in vivo tissue-like structures and functions in vitro. Organoid cultures are more accessible for manipulation and in-depth biological studies13 than animal models. As such, organoid cultures have been leveraged for a wide variety of applications including drug discovery14,15, personalized companion diagnostics15 and cell therapy13.
Organoid cultures exhibit significant heterogeneity and variable complexity in cellular composition, can undergo poorly controlled morphogenesis in the self-assembly process and often lack stromal, vascular and immunological components4,12. Hence, there is a great need to improve organoid culture by leveraging our understanding of organogenesis as well as how cells interact with their cellular and physical microenvironment in the form of the stem cell niche. Based on these insights, bioengineering strategies could be developed to precisely control stem cell decisions during organoid development. For example, from early embryogenesis studies, it is known that morphogen gradients regulate tissue patterning and development16,17. Microfluidics systems can be used to create the required concentration gradients of these by diffusing morphogens, giving rise to the desired cell types with spatial patterning16. Beyond biochemical cues, stem cells also experience active and passive forces from their external microenvironment and convert these physical stimuli into biochemical responses18. These physical cues arise from the matrix, external forces and/or cell–cell interactions. Rather than relying on a natural or biologically derived ECM such as Matrigel with limited stiffness tunability, synthetic hydrogels or other ECM combinations can be leveraged to control the physical properties of the matrix. Liquid friction against the cell membrane can also exert shear stress on cells19. The dynamic biofluidic environment has diverse effects on different cell types depending on the magnitude, direction and frequency19. Hence, microfluidic systems and bioreactors can be applied to provide perfusion at both the micro-scale and macro-scale20,21,22. Lastly, it is now known that cells interact with their neighbours and respond to external stimuli in a collective manner23; topographical cues, such as the curvature and shape of neighbouring cells, can affect stem cell decisions24. A recent neural tube model dissected the folding process and demonstrated that geometry constraints by micropatterning can control the final morphology of neural tube-like structures25.
It is debatable whether engineered cell-based in vitro models such as organoids need to faithfully recapitulate the structures and functions of the in vivo organ of origin. One trend is to recapitulate as much in vivo tissue architecture and function as possible in vitro in order to demonstrate the physiological relevance of models of increasing complexity. For bioengineers, the artificially created in vitro models only need to recapitulate specific features of the in vivo tissue, relevant to the physiological or diseased functions of interest. There is an optimism to creating highly complex models and expecting them to accurately mimic the in vivo organ of origin. For the majority of users, simpler models — such as a model with one or two cells in monolayer or 3D culture — are more robust for mechanistic studies and applications26,27,28 than more complex models, such as assembloids, or other multicellular models.
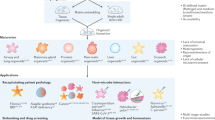
In this Primer, we focus on the rationale underlying the establishment of organoid cultures and provide guiding principles for the selection of suitable materials and methods for different applications. We first discuss the experimental considerations for setting up organoid-based cultures, categorized into four major components — cells, soluble factors, matrix and physical cues — and discuss approaches to integrate these components (Fig. 1). We also discuss key considerations for generating more complex yet robust organoids, such as those related to cell isolation and seeding, matrix and soluble factor selection, physical cues and integration. The general standards for data quality, reproducibility and deposition within the organoid community are also outlined. Lastly, we conclude by elaborating on the limitations of organoids in different applications, and the key priorities in organoid engineering for the coming years.
The establishment of organoid-based culture requires considerations about major components that make up organoid cultures — cells, soluble factors and matrix, physical cues — and the successful integration of these components. ASCs, adult stem cells; CSCs, cancer stem cells; ECM, extracellular matrix; FGF, fibroblast growth factor; iPSCs, induced pluripotent stem cells; OoC, organ-on-a-chip; TDCs, tissue-derived cells.
Experimentation
Cell source
Under defined physicochemical conditions, tissues such as small intestine7, colon29,30, stomach31,32, oesophagus29, tongue33, liver34,35,36,37, lung14, pancreas38,39,40, heart41, ear42 and skin43 have been obtained from iPSCs, adult or fetal cells and either stem/progenitor cells or differentiated cells. The starting cellular population for any given organoid is of prime importance, affecting not only the variability and heterogeneity in the structures obtained but also the function of the tissue they aim to model. To establish tissue-derived organoids or cancer organoids, tissue-resident stem/progenitor/differentiated cells or tumour cells, respectively, are obtained through an optimized tissue dissociation method. For iPSC-derived organoids, iPSC lines are established and fully characterized as the starting cells. Patient/tissue-derived stem cells are obtained through an optimized tissue dissociation method and then embedded into a 3D matrix mimicking stem cell niches. iPSCs can be maintained and expanded as undifferentiated clonal populations on feeder cells. To exemplify the generation of tissue-derived organoids we use intestinal organoids as an example (Fig. 2a), as this was the first tissue-derived organoid type established7. The small intestine and colon are opened longitudinally, washed and then cut into 2–4 mm fragments to increase the surface area for enzymatic digestion or further mechanical dissociation. EDTA treatment is used to chelate calcium, disrupting cell–cell adhesion and tissue integrity44. Larger tissue fragments and whole cells are removed from collected crypt fractions, and the harvested primary intestinal crypts are used for seeding and generation of intestinal organoid cultures.
Organoids can be generated from tissue-derived cells (TDCs) or induced pluripotent stem cells (iPSCs). a, To generate organoids from TDCs, tissue samples are obtained from humans or animals, such as the gut and stomach. The intestinal tissue samples are opened, washed and then cut into small fragments (2–4 mm) to increase the surface area for enzymatic digestion or further mechanical dissociation to isolate single intestinal stem cells or crypts. After several rounds of washing and purification, the harvested stem cells or crypts will be used for seeding and generation of organoid cultures for expansion. b, To generate organoids from iPSCs using genetic engineering, iPSCs are maintained and expanded as undifferentiated clonal populations on feeder cells or defined extracellular matrix (ECM) substrates to aggregate to embryoid bodies. Typically, iPSCs are harvested as cell aggregates, which preserve cell–cell contact and yield cell populations with higher viability. These aggregates are further induced through germ layer specification to form endodermal spheres, mesodermal domes and neuroectodermal matrix for additional applications.
The starting cellular populations for organoid cultures are generally obtained from adult or fetal tissue biopsy samples. The most commonly used tissue dissociation method is enzymatic digestion, which dissolves the ECM45. The composition of the enzymatic cocktail and the efficacy of the enzymatic dissociation process varies with tissue type46, and in certain cases DNase can be added to remove excessive DNA released from necrotic cells47. Depending on the tissue type, the tissue fragments can be further incubated with enzymes such as collagenase, elastase or dispase to generate single-cell suspensions and then seeded in Matrigel. The enzymatic dissociation method may affect the cell state of retrieved cells as it may require extended durations in the enzymatic mix to dissociate the majority of the tissue-resident stem cells. Tissue dissociation can also be achieved mechanically; although mechanical dissociation is much faster and less expensive, the cell yield and viability can be inconsistent46. Mechanical and enzymatic dissociation can be combined to generate better cell yield. After tissue dissociation, TDCs for organoid development are identified and collected using known biomarkers or physical characteristics45. Tissue-specific stem cell markers are typically used to identify and isolate the desired stem cells to generate organoids1,2. Fluorescence-activated cell sorting or magnetic-activated cell sorting isolates cells based on multiple parameters, including size, shape and cell-surface marker expression45,46. Other isolation techniques include laser capture microdissection and manual cell picking45.
iPSCs can be maintained and expanded as undifferentiated clonal populations over many generations. Undifferentiated human iPSCs are typically maintained on feeder cells or defined ECM substrates. As single iPSCs do not survive well in vitro, iPSCs are typically harvested as cell aggregates, which preserve cell–cell contact, yielding cell populations with higher viability. Physical scraping can also compensate for the lack of uniformity of cell aggregates. The dissociation enzymatic mixture should be chosen based on the level of cell sensitivity44 and whether the cultured cells secrete excessive ECM, making it difficult to detach the cells from the cell culture plate (Fig. 2b).
Tumour tissue, derived from either biopsy samples or surgical resections, is also typically processed akin to normal tissue to isolate tumour cells to grow as organoids15,48,49,50. Tumour cells isolated from liquid samples such as peripheral blood51, ascites49,52 and pleural effusions53 can be used as starting material to generate organoids. Patient-derived tumour organoids can be generated from samples obtained from minimally invasive Pap brush material54,55. Owing to their low numbers, tumour cells from biopsy samples or liquid samples can be first expanded in animal models as xenografts in order to obtain sufficient cells for organoid generation56. In the case of tumour tissue, it is preferable to limit tissue dissociation so that cell clusters rather than single cells are isolated. A critical factor that can influence the generation of tumour-derived organoids is the fact that isolated cells from tissue typically contain both cancer and normal cells. Although for some tumours it may be possible to enrich for tumour-forming cells by sorting a priori57, for the majority there is currently no robust method to separate normal and tumour cell populations prior to seeding into a matrix for culture. An approach to overcome this issue is to take advantage of culture conditions by using selective media that omits certain factors required for growth of normal organoids, as tumour cells gradually lose dependence on those factors during malignant transformation58. Blood contamination, particularly erythrocytes, can also affect organoid generation and matrix stability, and therefore standard approaches to eliminate these through lysis are typically used59.
Matrix
Following cell isolation, cells are typically seeded into biologically derived matrices such as Matrigel7,60 or a natural ECM such as collagen61, or into synthetic hydrogels3,62,63. Matrigel is mainly composed of laminin, collagen IV, entactin, perlecan and growth factors, and is similar in composition to the basement membrane61. As a continuation from the above example using intestinal organoids, we now briefly describe how cells are encapsulated into matrices. Isolated intestinal crypts are first re-suspended into cold Matrigel and pipetted into pre-warmed low-attachment well plates for culture, producing a flat semi-sphere gel of cell–matrix construct. The complete composition of intestinal organoid culture medium has been previously reported29. Organoids go through passaging, and the proliferation rate is measured by isolating organoids from the Matrigel, removing single cells and counting crypts. The expansion rate is calculated as the number of organoids from each well divided by the number of initial crypts seeded in Matrigel in that well.
Although Matrigel can support organoid culture, the inherently heterogeneous and poorly defined composition of this biologically derived matrix offers little control over the biochemical and biophysical spatio-temporal cues that are necessary for improving organoid culture. Therefore, other matrices with defined compositions63 have been explored as alternative matrices to Matrigel, such as recombinant human collagen61, fibrin64 or synthetic hydrogels3,62. Natural matrices can be recombinantly produced from proteins or polysaccharides to address the batch to batch variability of Matrigel60. On the other hand, synthetic hydrogels have emerged as powerful tools that enable independent manipulation of biochemical and biophysical matrix properties to control organoid features and enhance functionality. The ideal organoid matrix should, overall, be stress-relaxing and highly dynamic in biochemical and biophysical properties to accommodate or control changes in organoid structure during culture. For example, dynamic hydrogels based on polyethylene glycol (PEG) were recently shown to enable reproducible intestinal organoid formation and demonstrate how hydrogel properties could be tuned to control stemness and differentiation in cultured organoids3. The viscoelastic profile of hydrogels has also been shown to define the mechanical confinement of growing organoids65. In other examples, the activity of adult stem cells can be controlled using PEG hydrogels with photo-degradable moieties66 and biomimetic polymers can be modified to incorporate essential ECM signals to generate organoids with tailored features3. Tunable PEG hydrogels can promote intestinal crypt budding67, whereas dextran-based GMP-compatible hydrogels support expansion of cells for longer passages68. Lastly, microfabricated arrays were recently reported to enable the uniform production of crypt–villi-shaped epithelium69. Even with the recent advancements using the synthetic matrix to grow organoids, organoid growth by the synthetic matrix is still less efficient than Matrigel-cultured organoids. There is an unmet demand to develop a better matrix.
Organoids can arise either from round colonies generated by single cells7,34 or from initial multicellular structures such as intestinal crypts29,62, cell aggregates23 or micropatterned cells25. The aim of the latter approach is to establish a cellular niche which involves other cells of the same or different type from the beginning. To form cell aggregates, the simplest method is to use an ultra-low-attachment dish coated with hydrophilic hydrogel70 to prevent cell attachment; subsequent centrifugation can promote aggregate formation, enhancing cell–cell contacts71. The size and compaction of cell aggregates can be tuned and controlled, such as through the use of microwell arrays71,72. Another well-established method to form cell aggregates is the hanging drop method, where aggregates form at the bottom of the drop. In a recent example, droplet microfluidics was used to aggregate murine cholangiocytes to form complex organoids with liver mesenchymal cells73. Droplet microfluidics can print one organoid per well and enable the rapid generation of intra-organoid heterogeneity74. Droplet-based microfluidics has also been used to perform better single-cell RNA sequencing (scRNA-seq) analysis of intestinal organoid cell identities during various developmental stages, revealing extensive population heterogeneity75. Microwell structures or microfabricated pillar arrays69 have also been developed to enable enhanced uniformity in cell aggregation71,72,76. In one example, microfabricated patterns of Laminin-512 were shown to reproducibly support human pluripotent stem cells to form lumen structures in Matrigel and differentiate into human neural tube-like structures25. These examples, amongst others emerging in the field, illustrate how our knowledge of biomaterials and tissue engineering can be extrapolated to provide precise control over organoid structure and function.
Soluble factors
Organoid cultures are fundamentally based on our accumulated knowledge of developmental biology13, where soluble cues are presented to cells in a spatio-temporally controlled manner. Soluble factors used to differentiate TDCs and iPSCs into various tissue types are listed in Supplementary Table 1. In organoid culture, these soluble cues are recapitulated in vitro in the form of biologics, mainly as proteins such as growth factors77, or small-molecule drugs, which can activate or inhibit signalling pathways. Although growth factors could be costly and unstable, and many small-molecule drugs can affect multiple pathways resulting in poor reproducibility77, some organoid protocols have combined the use of both biologics and small-molecule drugs78. The use of conditioned medium from engineered cell lines producing biologically active growth factors, such as L-Wnt-3A, can replace commonly used growth factors, such as WNT3A ligands79. These conditioned media face a similar issue of batch to batch variation and require stringent tests to ensure reproducibility80. Thus, novel surrogate molecules are starting to arise as potential substitutes for conditioned medium79,80.
It is critical to consider how and when soluble cues are added to organoid cultures because soluble cues in vivo are typically presented to cells by the ECM or nearby cells, coordinated in time and space. This is the concept of spatio-temporal presentation. For example, it is now known that fibroblast growth factor (FGF) activity and specificity can be regulated by cell-surface heparan sulfate proteoglycans, suggesting that the addition of free FGF into cell culture medium may not recapitulate how FGF is available in tissue in vivo81. The importance of presenting soluble cues in a spatio-temporally relevant manner is especially important for growing human iPSCs into complex structures with multiple cell lineages, such as in the case of kidney organoids82. Based on previous studies83, it is known that the ureteric epithelium develops from early migrating presomitic mesoderm cells. To recapitulate this process, the effect of different durations of initial Wnt signalling before the addition of FGF was investigated82. The spatio-temporal presentation of soluble factors can be achieved using different tissue engineering approaches. In one strategy, these growth factors can be encapsulated within nanoparticles, and conjugated onto cell surfaces for controlled release84,85,86,87. To mimic how certain growth factors are bound to the ECM in vivo, researchers conjugated polymers with heparin, which can bind to growth factors, or conjugated these growth factors to the polymer itself88. Surface tethering can also be achieved with nanotechnologies, such as nano-imprint lithography, electron beam and electrospinning, and substrates with nanopillars, nanopits or nanochannels, mimicking the basement membrane for 3D organoid culture89. Lastly, microfluidic systems can be leveraged to create miniaturized niches with precise control over mechano-chemical properties90. With directed flow and gradients of gas or small molecules, these systems can finely control environmental parameters within organoids17. In one example, a microfluidic neural tube device was developed to present simultaneous opposing gradients of growth factors to direct neural tube patterning, enabling recapitulation of the in vivo structure91.
Physical cues
Beside biochemical cues, it is also important to consider when and whether it is necessary to provide appropriate physical cues to cultured organoids. Nutrient supply and waste removal, which are diffusion-dependent, become less efficient during organoid growth into larger tissue structures. This is the reason why intestinal organoids need to be fragmented into smaller cell clusters and re-seeded regularly7. Inadequate nutrient and waste diffusion is also problematic in brain organoid culture, where the resulting millimetre-sized constructs often exhibit necrosis within the inner core due to nutrient inaccessibility. This problem can be partially resolved using shaking cultures22,92, spinning bioreactors or suspension under continuous agitation22, or in continuously stirred bioreactors92. These bioreactors can monitor pH, temperature, oxygen and glucose levels to maximize mass transfer while minimizing shear stress. In this regard, perfusable microfluidic chips have also been developed to promote the long-term culture of organoids20,21,93. Lastly, it may be useful to consider providing topographical cues to control organoid culture in vitro; the topography of the substrate is known to modulate cell area, shape and cell–cell interactions, resulting in biochemical signals that can affect stem cell fate24. Topography-directed morphogenesis has been demonstrated using intestinal organoids grown on soft hydrogels94.
Integrating cues
In contrast to the original self-organizing organoid model, the above-described cues can be integrated to confer greater control over organoid morphogenesis. Integration of cues is a commonly employed strategy in the field of tissue engineering to construct tissues in vitro and in vivo95. Ideally, specific physical or chemical cues should be presented in a spatio-temporally, physiologically relevant manner using simple, reproducible and robust methods (Supplementary Table 2). We illustrate this using the example on intestinal organoids7. Following the identification and isolation of LGR+ intestinal stem cells residing in crypts, biologically derived Matrigel was leveraged to mimic the laminin-enriched crypt environment, supplemented with exogenously added growth factors in the cell culture medium. Most intestinal stem cells could survive, proliferate and form organoids7. However, given the stochastic nature of organoid development, the resulting organoids vary in size, bud number and function. In one example of how environmental cues (mechanical and biochemical) were integrated to exert control over organoid morphogenesis, the organoid maturation process was dissected into different stages, and biomaterials engineering was used to identify the optimal mechano-chemical environment at each stage3. It was shown that mechanically dynamic matrices that could switch from high to low stiffness over time enabled control over the morphogenesis process3. In another example, human neural tube morphogenesis25 was simulated using micropatterning, which created mechano-chemical gradients to regulate cell–cell/matrix interactions, and soluble factor presentation to orchestrate morphogenesis.
Besides these examples, other engineering methods have been used in organoid culture to control cell proliferation, differentiation and morphogenesis. A popular bioengineering approach for reconstructing tissues is bioprinting96 This technology uses bio-ink, comprising living organoids encapsulated within a biomaterial, to precisely create 3D biological geometries that mimic those of the native tissue in a layer by layer approach. In one recent example of how bioprinting was used to generate large tissue constructs, human small intestinal organoids were used to form self-evolving tissue constructs by a concept called bioprinting-assisted tissue emergence, where centimetre-scale intestinal tissues were printed sequentially to mimic boundaries in the gastrointestinal tract97. Another bioengineering approach to reconstruct tissues, especially across different tissues and organs, is the use of microfluidic platforms. Microfluidic systems have been used to create miniaturized cellular models comprising organoids that recapitulate critical aspects of organ physiology, known as the organ-on-a-chip98 (OoC). OoC devices can potentially incorporate other bioengineering techniques to imitate the key physiological and structural features of organs and tissues. For example, physical constraints were engineered into the organoid environment using synthetic scaffolds to provide physical boundaries99. OoC devices also support co-culture or multi-organ systems. The two-organ system has been simulated using connected chambers containing different tissue constructs to mimic liver injury (with intestinal epithelial cells and liver cells)100 and type 2 diabetes (with human pancreatic islets and liver spheroids)101. A two-organoid microfluidic device was recently fabricated with multiple read-outs using cardiac and hepatic organoids102. Moreover, OoC devices have improved the characteristics of human iPSC-derived pancreatic islets103, intestinal104, stomach105 and liver76 organoids by supporting physiological flow rates. Kidney organoids have been cultured in an OoC device that enables fluid flow to simulate shear stress106 and the generation of a vascular network107.
Results
It is important to evaluate whether cultured organoids recapitulate key aspects of the representative human tissue or organ. To achieve this, characterization of these organoids is typically performed by assessing organoid cellular composition, structure, functions and robustness of phenotype. In this section, we discuss commonly employed analytical methods using representative examples.
Organoid composition and structure
Organoid growth is a process of initial cell aggregation, proliferation, migration and differentiation. When assessing whether the organoids have been successfully established, it is critical to first determine whether the organoids contain the desired cell types, and the degree to which the organoids accurately mimic the functions of the corresponding tissue in vivo. To achieve this, both low-throughput gene expression validation and high-throughput whole-genome transcriptome analyses are typically performed. Real-time PCR is often first performed as an easy, fast and quantitative read-out on marker genes indicating cell identity, including key transcription factors and differentiation markers. Western blotting provides further quantitative information regarding protein abundance, protein degradation, protein–protein interactions and post-translational modifications, which represent the activities of a specific signalling pathway in a committed cell type. The most common tools used to evaluate organoid composition are immunofluorescence and immunohistochemical imaging using sections or whole mount, and the specific cell markers’ antibody staining further elucidates various cell types’ spatial distribution and proportion. A high-throughput analysis of scRNA-seq profiles all of the cell types in the organoids — undifferentiated and committed — at the whole-genome transcriptome level. These cell types are then compared with cells freshly isolated from the corresponding tissue or organ to evaluate the degree of similarity for each cell population. Given how it is expected that the in vitro cultured organoids contain different states of immature cells, scRNA-seq can be helpful for determining the extent of organoid heterogeneity in terms of cell differentiation status.
The assessment of organoid morphology is performed to determine whether there is structural similarity between the cultured organoids and corresponding in vivo tissue or organ108. For example, for secretory tissues such as pancreatic islets, the islet-like globular structure should be present within the organoids, and intra-cellular hormone vesicles109 are also expected to be observed. For branching epithelial organoids, such as breast organoids110, distinct branching structures are expected. For intestinal organoids, crypt-like structures99 should be observed. For more complex organoid culture systems comprising incorporated supportive cells, such as fibroblast111 and/or endothelial cells (ECs)106,109, it is expected that the incorporated ECs should form a vascular network106 that recapitulates the endothelial–epithelial interactions around the organoids.
Organoid functionality
Evaluation of organoid functionality includes, but is not limited to, the generation of mature cells, the formation of the vasculature or neuronal networks25,43, the accurate response to external stimuli and the effective secretion of cytokines or hormones, among other factors. Organoids should be compared with freshly isolated tissues as a positive control. Discrete physiological recapitulations are required for different tissues, such as acid secretion in gastric organoids and mucus secretion in intestinal organoids. Various methods of characterization to assess the structure and functions of organoids are listed in Table 1.
Pancreatic islet organoids
We use the example of mouse pancreatic islet organoids to discuss various characterization and validation methods (Fig. 3). The formation of four endocrine differentiated cell types (β-cell, α-cell, δ-cell and PP cell) is examined by the expressions of the corresponding hormones — insulin, glucagon, somatostatin and pancreatic polypeptide, respectively — by immunostaining112,113. To assess the extent of β-cell maturation, the ultrastructural morphometric analysis of insulin secretory granules is performed by immuno-gold transmission electron microscopy (TEM) imaging109. To evaluate the ability of the β-cells in organoid responsiveness to glucose, glucose-stimulated insulin secretion experiments are performed, for example using an insulin or a C-peptide ELISA assay with cyclic high and low glucose incubation114. As insulin release is associated with calcium dynamics, calcium signalling traces imaging rapidly increases in response to glucose and returns to baseline113,115. To test the full potential of the islet organoids upon transplantation, in vivo functional evaluation is needed to ameliorate the hyperglycaemic phenotypes of streptozotocin-induced type 1 diabetic mouse models109,113,116. The characterization parameters include normalized and stable blood glucose level, normal plasma insulin level and maintained body weight.
A, Representative view of pancreatic islet organoids at culture day 7. Aa, Organoids growing in Matrigel exhibit spherical shape. Ab, Viewed at high magnification, these spherical organoids are surrounded by endothelial cells (ECs) (arrows). Ac, A view of the organoids after they are released from Matrigel by dispase. B, Cell types and hormone secretion level validation by immunofluorescence or immunohistochemical staining. The expression of insulin (Ins; β-cells), glucagon (Gcg; α-cells), somatostatin (Sst; δ-cells) and pancreatic polypeptide (Ppy; PP cells) can be detected in mature organoids (shown by DAPI; blue). Scale bars, 20 μm. C, The morphology of mature organoids induced through prolonged culture for a total of 30 days: the organoids are surrounded by elaborate EC networks (parts Ca and Cb), shown by Cd31 (endothelium marker; red) and DAPI (organoid nuclei; blue) staining (part Cc). Scale bar, 50 μm. D, Real-time quantitative PCR analysis for some key transcription factors and differentiation markers. E, Pancreatic islet organoid function validation in vitro. Organoids are sequentially incubated with low (2 mM) and high (20 mM) concentration glucose for three rounds, followed by C-peptide ELISA and calcium imaging. F, Representative ELISA result of organoid secreting C-peptide measurement. The rise of C-peptide concentration in the media is sharply responsive to high glucose stimulation. **P < 0.01; ***P < 0.001. G, Representative curve of intensity of calcium signalling trace imaging, indicating the organoid’s capability of responding acutely to glucose, with high/low calcium activity consistent to high/low glucose stimulation. Potassium chloride incubation is used as a positive control. H, Islet organoid function validation in vivo. Organoids (or fresh islets as positive control) are transplanted into the kidney capsule of streptozotocin-induced diabetic mouse, followed by monitoring of certain physiological indexes, indicating the rescue of the diabetic phenotypes. Finally, the transplants are removed and the surging blood glucose levels in the mice are expected, further demonstrating that the transplants were responsible for the lowering glucose level effect. Parts A–C, F, G and H adapted from ref.113, Springer Nature Limited.
Intestinal organoids
Intestinal organoids are among the first organoid types successfully established in vitro7,29. Classical crypt–villus-like architecture is regarded as an important feature of successful organoid establishment, exhibiting proper branching morphology with robust multicellular composition and location reflecting the organoids’ differentiation and maturation level117,118. The expression of Lgr5 marker can analyse the maintenance of intestinal stem cells by real-time quantitative PCR or by imaging using a Lgr5-fluorescent reporter7,119. The differentiated cells are assessed by the staining of their markers, including lysozyme (for Paneth cells), villin (for enterocytes), Mucin 2 (for goblet cells) and chromogranin A (for enteroendocrine cells)7,99,120. These various cell types can also be distinguished by TEM according to their characteristic subcellular structure7,120. Functional intestinal organoids exhibit a relatively thick mucus layer due to mucus secretion from mature goblet cells121, which can be detected by Alcian blue and periodic acid–Schiff staining122,123.
Liver organoids
In the case of the liver, both cholangiocyte34,35 and hepatocyte organoids124,125 have been established, both from mouse and human tissue samples. Measuring the passaging time and proliferation of organoid cells is used to assess liver organoids. Regarding marker expression, specific cholangiocyte (for example KRT19, KRT7, SOX9) or hepatocyte (such as ALB, HNF4A, MRP4) markers are assessed both on the RNA level (real-time quantitative PCR, scRNA-seq) and on the protein level (immunofluorescence staining). Often, also markers of liver progenitors (such as LGR5) are used to assess the differentiation state of the cultured cells. For the assessment of organoid functionality, further markers such as CYP3A4 and CYP3A11 for hepatocytes are analysed on the RNA level. Furthermore, albumin secretion, low-density lipoprotein uptake, the presence of glycogen accumulation assessed with periodic acid–Schiff staining, bile acid production and activities of liver enzymes such as CYP3A4 are assessed using chemical assays to confirm hepatocyte function. The ultrastructure of the organoids is also often assessed for typical hepatocyte or cholangiocyte morphology by TEM. Finally, for an ultimate test of functionality, organoids are often transplanted in FRG immune-deficient mice, to look at integration and functionality of the graft in vivo34,124,125.
Validation of tumour organoids
Patient-derived tumour organoids must maintain the genomic, transcriptomic, morphologic and functional profiles of the tissue of origin. As such, it is important to validate them by comparison with the tissue they are derived from in terms of histology and immunohistochemistry profiles15, transcriptomics as well as genomics14,126. Cellular organization, tissue structure and protein expression patterns of tumour organoids can be easily compared with the cancer of origin15. Similarly, organoids are expected to have gene expression profiles similar to the tumour they are established from. This has been shown by RNA-seq on many tumour types including bladder cancer127 and oesophageal cancer128 amongst others. Lastly, it is important to confirm whether tumour organoids have diverged from the patient tumour at the genomic level. Several studies have shown concordance of organoids and parent tissue in terms of mutations and copy number variations14,126. A greater degree of divergence may be attributed to intra-tumour spatial heterogeneity and sampling bias: tumours can be spatially diverse, and organoids generated from a specific portion may lack the representativeness of distant regions129,130. In addition to this topographical issue, culture conditions can contribute to selecting specific clones, possibly altering or reducing clonality over time128.
Validation of tumour organoids is even more critical when culturing over a long period of time. Whereas some studies have reported that molecular characteristics can be maintained over long-term culture, others have observed tumour evolution after serial passaging128,131. For instance, exome sequencing of liver cancer organoids showed that mutation concordance decreased from 92% for organoids cultured for less than 2 months to 80% for those grown for more than 4 months131. Colorectal tumour organoids established from microsatellite instable tumours had higher de novo mutations after long-term culture than stable ones58. Some of the differences observed may be attributable to how different clones and subclones adapt and expand in culture. Using deep targeted sequencing of 500 cancer-associated genes, truncal mutations were shown to be retained whereas subclonal ones are gained or lost, with mutational changes at each passage126. Most mutations and copy number variations are preserved in the late passage of lung cancer organoids, whereas de novo mutations can be attributed to the subclonal expansion of small subsets of cells present in the tumour of origin14. Overall, these studies showed that genetic drift can occur in organoids cultured for extended time, influencing their reliability as models for functional and drug discovery studies132. This reiterates the importance of validation in the context of tumour organoid establishment, characterization and testing. As an example of analysis, crucial steps in characterization of liver cancer organoids are demonstrated (Fig. 4). Lastly, normal tissue contamination is an issue for establishing tumour organoids and should be addressed when validating patient-derived models. The presence of normal cells can be estimated by histological comparison and immunohistochemistry, as well as sequencing54,133.
a, Isolation of cells from patient samples and organoid culture, with schematic of tissue isolation and processing. b, Histological analysis of liver cancer samples. Top row shows tissue (scale bar: 50 μm), middle row shows organoid bright-field images (scale bar: 100 μm) and bottom two rows show histological haematoxylin and eosin (H&E) staining of organoids (scale bar: 50 μm). c, Analysis of specific marker gene expression, including immunofluorescence staining for AFP (hepatocyte/hepatocellular carcinoma (HCC) marker; red) and EpCAM (ductal/cholangiocarcinoma (CC) marker; green). DAPI staining shown in blue. Scale bar: 30 µm. d, Organoid passaging shown as growth and splitting curves, where dots represent a splitting time point and arrows represent continuous expansion. e, Transplantation of xenografts into immunodeficient mice. Xenograft and histopathology analysis, matching the patient’s original tissue sample. Scale bars: 125 μm, inset: 62.5 μm. f, Analysis of genetic changes in the cancer organoids and their concordance to the mutations in the original tumour sample. g, Half-maximal inhibitory concentration (IC50) curves for gemcitabine treatment showing organoid sensitivity to drugs. CHC, combined HCC/CC tumours; PLC, primary liver cancer; SC, subcutaneous. Adapted from ref.131, Springer Nature Limited.
Data analysis
Image processing and analyses
ImageJ/Fiji134, CellProfiler135 and other software are used for image processing. The number of organoids formed in a certain period, organoid size (measured as area or volume) and cell number can measure the cell proliferation rate. Cell proliferation can be quantified by the ratio of EdU-positive or Ki67-positive nuclei using available software plug-ins. Morphological evaluation — such as the bud formation ratio in the intestine organoid system — is also necessary when quantifying the success rate considering functional maturation. Mean and/or normalized fluorescence intensity of maturation markers for different tissue types can be recorded to measure the organoid maturation level. Long-term live imaging can be analysed using software to track the cell behaviour for lineage tracing purposes. Web resources are used for pathway analysis and visualization of multi-omics data to generate heat maps for understanding the differentiation stage of the organoid136,137. Other software, such as Profiler, GSEA, Cytoscape and EnrichmentMap, have been used for biomolecular interaction networks, pathway enrichment analysis and visualization138,139,140,141.
Data analytics classification of phenotypes
Miniaturization and automation using machine learning and artificial intelligence-based data analytics make high-throughput and multidimensional phenotyping of organoids more consistent. Various software for these purposes have been developed in recent years, including ilastik142, OrganoidTracer143 and Phindr3D (ref.144). Most of these have Fiji or CellProfiler plug-ins, which make handling large data sets in shorter time frames possible for users without substantial computational expertise. Those software usually contain functions for image segmentation, object classification, morphological characterization, fluorescence intensity quantification, cell counting and tracking for hundreds of images. Good image quality is a prerequisite for a robust result, so optimizing the image acquisition process is crucial. At different stages of the data processing, validation of the accuracy of the trained algorithm is recommended.
Applications
Organoids as engineered cell-based models to recapitulate relevant physiological structures and functions of interest have shown great potential in both basic research and clinical applications. Here, we summarize a few applications.
Tissue regeneration
Tissue-derived organoids can be a potential source of transplantable material for regenerative medicine. Organoids of murine intestines30, livers34,124,125 and pancreas68,145 have been successfully transplanted into mice with restoration of organ function. For example, liver organoids have been generated from hepatocytes isolated from murine livers125. The organoids were injected into mice with a deficiency in fumarylacetoacetate hydrolase (Fah), a defect causing liver injury resulting in short survival (40 days) after treatment with the hepato-protective nitisinone drug is withdrawn34. Organoid engraftment rates were as high as 80%, and the engrafted mice survived longer than 100 days after the interruption of nitisinone therapy125. Similarly, pancreatic islets cultured in an artificial ECM could be successfully engrafted into either streptozocin-induced or autoimmune-driven diabetic mouse models, rescuing insulin production and reversing hyperglycaemia146. Beyond murine organoids, human liver organoids derived from EpCAM+ ductal cells from liver samples have been engrafted into immunodeficient mice with acute liver damage induced34. Human pancreas organoids can be generated from ALDHhigh stem cells and have the potential to produce insulin145 without in vivo evidence yet to confirm engraftment. Intestinal organoids have been shown to fuse into tubular structures resembling in vivo intestines147,148. A more complicated intestinal organoid can also be generated in an artificial ECM with modulable biodegradability3. Although there is a long way to go for organoids to be used in regenerative medicine, improved protocols to generate normal tissue organoids, good manufacturing practices ensuring high reproducibility, grafting and function can already be established149.
Drug discovery and development studies
Technical challenges to rapid and high-throughput screening of 3D organoids have been addressed in recent years and surpassed by different means, including alternative plate design or seeding geometries15,150,151. A screening of 240 tyrosine kinase inhibitors on organoids derived from patients with ovarian or peritoneal tumours provided individualized drug sensitivity profiles within a week from surgery. The study included a carcinosarcoma of the ovary, a sporadic ovarian cancer for which no line therapy was established150. Tumour organoids can be screened to select effective drugs from structurally similar candidates152 and investigate the synergic effects of different drugs for combination therapy153. Non-tumour cells from adjacent normal tissue in biopsy samples or surgically resected samples can be used to develop control healthy organoids to confirm the tumour-specific efficacy of the tested drugs154. Organoids can be screened with libraries of immune cells engineered to express chimeric antigen receptors against different neo-antigens155. Checkpoint inhibitors and other immuno-oncology drugs can also be screened in tumour organoid models that include immune cells156.
Biomarker research
As organoids can maintain the genomic profile of the parent tissue157, drug screening can be used to provide connections between genetic mutations and drug responses36,153,158. By screening gastric cancer organoids from 7 patients against 37 drugs, tumours with ARID1A mutations were shown to be sensitive to ATR inhibitors158, supporting a previous study159 and functionally identifying an intervention for a class of mutations that were considered undruggable.
Precision medicine applications
There is growing interest in tailoring cancer therapy to each patient. Precision medicine is often synonymous with genomics160. However, as shown in several clinical trials160,161,162,163,164, only a small proportion of patients have an actionable alteration that can be coupled to a specific intervention160. In the NCI-MATCH trial that evaluates broad-scale implementation of sequencing-driven therapy, only approximately 37% of patients were found to have an actionable mutation165. This and other precision medicine trials have shown limited clinical benefits and modest response rates for many of the targeted interventions, uncovering a large degree of complexity165,166,167. For instance, the SHIVA trial that compared the outcome of genomic-guided interventions with conventional therapy selected by the oncologist for heavily pretreated patients with cancer showed no difference in progression-free survival160,161. Although the clinical efficacy of genomics-based precision medicine interventions can be debated, it is clear that most patients with cancer do not have a targetable genetic to guide therapy. Functional precision medicine — based on testing drug responses on patient tumours to identify treatment regimens — has been proposed as a more direct alternative, as it does not require any a priori knowledge of the tumour molecular profile or other characteristics160,168. Organoids are ideally suited for functional precision medicine because they are straightforward to establish, have high similarity with the tissue of origin, are tractable157,169 and have been deployed to identify therapeutic leads in numerous tumours170, including rare cancers that tend to be understudied and poorly characterized at the molecular level15,150,171,172. Recently, organoids established from patient-derived xenografts were used to identify eribulin as therapy for a patient with recurrent triple-negative breast cancer173. The patient’s progression-free survival was 3.5× longer on an eribulin regimen than on the previous therapy received173, indicating the potential of organoids in clinical care.
Source material for xenografts
Organoids are increasingly being used as seeding material to generate xenografts with engraftment rates. This process involves using intact organoids14,58 or individual cells after digestion54,174. Human intestinal organoids transformed by CRISPR–Cas9 to carry mutations in APC, SMAD4, TP53, KRAS and/or PIK3CA were seeded into the subcapsular kidney space of immunocompromised mice. Benign organoids carrying mutations in one or two genes were shown as unable to form tumours, whereas organoids with mutations in four or five genes were tumorigenic175.
Patient-derived organoids can also generate xenografts, confirm their tumorigenic capacity and recapitulate the histology of parental tumours14,54. In a study of colorectal cancer, benign tumour organoids showed no or minimal engraftment in mice. By contrast, colorectal cancer organoids derived from metastases were more invasive than those from primary tumours58. Comparison of the morphology of xenografts formed by glioblastoma multiforme cells cultured in vitro either as tumour spheres or organoids showed that whereas cells from tumorspheres displayed a solid growth pattern, organoids were more diffusive, similar to the tumour of origin174. Organoid lines derived from bladder tumours of lumenal origin transplanted in mice were shown to give rise to tumours with the same lumenal phenotype126.
Infectious diseases
Organoid models are now widely used in research on host–pathogen interactions. For instance, organoids derived from tissues and organs such as the intestine176,177, liver178, lung177,179, oral mucosa180 and stomach32,181 have been co-cultured with pathogens such as bacteria32,181,182, viruses176,178,179,180,183 and parasites177. In human intestinal organoids, differentiated intestinal cells are more susceptible to human rotavirus (HRV) infection than undifferentiated ones. Organoids infected with HRV or rotavirus enterotoxin displayed lumenal swelling, a characteristic of rotavirus-induced diarrhoea183. Human liver organoid was used as a drug screening platform to test anti-HRV drugs and antibodies, identifying mycophenolic acid, IFNγ and anti-rotavirus VP7 antibodies capable of blocking rotavirus replication ex vivo178. Organoids can also be used in studies of oncogenic pathogens. Infection of murine gastric organoids by Helicobacter pylori showed increased cytosolic β-catenin levels induced by bacterially originated CagA, which led to increased proliferation in infected organoids181.
Since the start of the COVID-19 pandemic, several laboratories have investigated the effects of the SARS-CoV-2 virus on organoids. SARS-CoV-2-infectable distal lung organoids were generated with ACE2-expressing lumenal cells184. SARS-CoV-2 was found to target goblet cells rather than ciliate ones, in contrast to a previous study in a 2D air–liquid interface model185. Studies in intestinal organoids show that SARS-CoV-2 can also infect enterocytes185,186, and CRISPR–Cas9 and drug screenings have shown that knock-outs of either TMPRSS2 or TMPRSS4, or treatment with Camostat, can reduce viral infection186.
Biology of common and rare diseases
Organoids have demonstrated significant utility in modelling and investigating both common as well as rare diseases arising from various different organs. Cystic fibrosis is a genetic disorder caused by mutations in the gene CFTR, which expresses a transmembrane chloride and bicarbonate transporter. More than 2,000 mutations in CFTR associated with cystic fibrosis have been reported, yet small-molecule therapeutics deployed clinically only target a subset of these187,188. Mutations in CFTR cause multi-organ dysfunctions affecting the lungs, pancreas, liver and intestine189. Cystic fibrosis organoid models of the colon190, liver and bile ducts191, rectum192,193, the respiratory system179,194 and the small intestine190,195 have been derived from adult stem cells of relevant organs of either mouse models or patients, with applications ranging from investigating the biology of disease to mirroring patients’ drug responses192. Forskolin, a CFTR activator, has been used to induce organoid swelling due to chloride and fluid flux into the lumen193, with reduced swelling of patient-derived rectal organoids correlated with clinical response parameters of patients, which could mimic individualized clinical responses to specific therapeutic regimens193. Cystic fibrosis organoids have also been used to investigate gene therapy interventions and are able to rescue the effect of CFTR mutations in intestinal organoids using CRISPR–Cas9 (ref.195).
Inflammatory bowel diseases such as ulcerative colitis and Crohn’s disease have also been studied with organoid models196. Organoids derived from intestinal epithelial cells of paediatric patients with inflammatory bowel diseases had inflammatory bowel disease-specific DNA methylation signatures despite retaining similar morphologies197. Liver organoids were generated to model liver diseases such as α1-antitrypsin deficiency35, Alagille syndrome35 and primary sclerosing cholangitis198. Lung organoids have been established to investigate diseases such as goblet cell metaplasia199.
Reproducibility and data deposition
Heterogeneity and reproducibility
Organoids exhibit heterogeneity and variability between cells forming the organoid (intra-organoid heterogeneity), between organoids in the same dish and between individual patients (inter-organoid heterogeneity). This heterogeneity is valuable to recapitulate the individual to individual differences, especially in the context of human diseases, and is valuable in cancer research to mimic patient-specific cancer development for personalized medicine126,127,128,131,154,158,169,200,201,202. Similarly, the intra-organoid variation reflects the complexity of cellular composition of the tissues. This is advantageous for modelling tissue development or regeneration, with cells forming an organoid encompassing different cellular states, from stem/progenitor cells to more differentiated cells35,124,203. Although these levels of variation and heterogeneity reflect the complexity of biological systems, they can jeopardize the reproducibility and robustness of the system when left uncontrolled.
Significant differences in morphology and functionality or even organoid formation efficiency depend on the different cells of origin of the culture or different cells in the organoid, as well as different medium composition119,204,205,206. Regarding the cells of origin, the batch and quality of the iPSCs/pluripotent stem cells used to generate the starting culture can influence the variability of the organoid line obtained. Similarly, the different primary cells isolated from different animals or patients also impact the subsequent cultures12. The media are usually optimized for cell proliferation whereas differentiation towards certain cell fates can be largely influenced by the culture conditions, thus increasing variability and heterogeneity. Refining of culture conditions has been demonstrated to increase the diversity of differentiated cell fates in human small intestinal organoids119. Undefined ECMs (such as Matrigel) have unknown composition and batch to batch variability, making reproducibility a significant concern12,207. Another important factor becomes apparent when looking into transcriptomic changes during organoid development208 or studying gene expression changes during organoid passages34,204. When organoids mimic development, these changes can lead to confounding effects, for example, the temporal heterogeneity. Therefore, recording the passage and the time in culture is essential to identify sources of variation that affect the results.
To better understand and control heterogeneity, the field has been moving from bulk analysis towards employing single-cell approaches, such as scRNA-seq, to help delineate the different populations of organoid-forming cells. Many single cell-resolved data sets are emerging, with several notable examples in more homogeneous organoid production3,69,99,209 and resolving organoid heterogeneity during growth progression206,210. Pooling together different cells and/or organoids for bulk analysis can lead to inaccurate results3,69,99,206,209,210. Therefore, good record-keeping about what was pooled in the experiment deems crucial to drawing robust conclusions.
Additionally, it is critical to reduce the sources of variability and heterogeneity as the field grows in complexity. Efforts to generate more complete organoid structures containing different cellular populations (reviewed elsewhere211,212) — such as the recently published endothelial and healthy human colon and human colorectal cancer organoids213, murine cholangiocyte and mesenchymal organoids73 and murine pancreatic islet and endothelial organoids109 — will require controlling sources of variation derived from incorporating additional cells to an already heterogeneous system. Exerting control over the variation while generating more functionally relevant structures represents a significant challenge for the field.
Finally, significant heterogeneity is also observed following organoid treatments. Although this represents a drawback when trying to use organoids as a tool for investigating molecular mechanism, it is beneficial when aiming to model patient responses. In particular, for cancer treatment, several different patient-derived organoid models tested (including, but not limited to, pancreas, colorectal and liver cancer) exhibit heterogeneous responses to known chemotherapeutic and anticancer agents48,131,214. This parallels the variability in patient outcomes observed in the clinic and has positioned cancer organoids as a potential predictive tool for drug testing (reviewed elsewhere215). This heterogeneity in treatment responses is now being exploited for the treatment of patients with cystic fibrosis193, with a large ongoing European multicentre clinical trial study, led by the HIT-Cystic Fibrosis consortium, to identify potential responders216. Similar initiatives are lagging behind for patients with cancer, although mostly because of disparities in establishment rates and lengthy timelines (reviewed elsewhere217). Organoids were recently used to inform the treatment of a patient suffering from a rare form of liver cancer, although unfortunately the treatment was discontinued due to deterioration of the patient’s general condition, preventing the authors from drawing conclusions regarding the predictive value of the organoid system for drug treatment218.
Minimum reporting standards
To improve the reproducibility and reliability of the results, it is essential to account for the variability between different cell isolations and intra-organoid heterogeneity. Although one could consider that at least three independent experiments with at least three different biological replicates are a minimum requirement, this may not always be sufficient. In fact, the number of organoids to study per biological replicate as well as the number of biological replicates depends on the research question, the specific experiment and the variability of the phenotype observed. Whenever possible, calculating the number of independent organoids per biological replicate, sample size calculations similar to those used for mouse experiments, taking into account statistical power and variance of the experiment, would be highly recommended. The data should be robust, meaning repeated in at least three independent biological replicates, which refers to independent isolations from different animals or patients. However, replicates and the research question for each experiment should be determined in advance to avoid hypothesis fishing and increasing replicates to get a significance between conditions219,220. Data should be reported correctly as a mean of each replicate, with each replicate being an average of a similar number of organoids. Recently, several publications have provided guidelines to improve reproducibility, including general biological reporting and accurate statistics221,222,223. SuperPlots is recommended for data visualization, as it shows both the mean of each independent biological replicate and the spread of the individual data points224,225. This type of plot is ideal for organoid reporting, as it allows immediate visualization of any potential artefact concerning batch variation. Journal collections that deepen all aspects of statistics used in biological sciences are valuable resources to facilitate accurate reporting226.
Organoids should be characterized in full, not only checked for the expression of markers but also benchmarked to the tissue of origin. It is crucial to not only determine whether cells have similar morphology and expression patterns as the tissue of origin and that they have a similar function (using, for example, ELISA for albumin, bile acid production by liver/hepatocyte organoids). Whenever possible, results between compared conditions should be normalized against DNA content (for genomic analysis) or structural proteins (such as actin, for western blotting), whereas functional data such as from ELISA should be normalized to the total number of cells or area of organoid structures in each well being compared. Improving reporting standards is the first step towards increasing reproducibility, thus enabling the field to move forward.
Data deposition
Standardized organoid data repositories are currently missing. A notable exception is a patient-derived repository for cancer organoids led by the National Cancer Institute (NCI) in the USA and the Human Cancer Models Initiative (HCMI), a combined international effort of several institutions (Cancer Research UK; NCI; Hubrecht Organoid Technology’s the HUB; and the US Office of Cancer Genomics’ OCG) (Supplementary Table 3). However, healthy patient-derived organoids and organoid models from non-cancer diseases, both human and murine, are yet to be documented in one database, indicating the apparent need to fully document, report and deposit all organoid lines generated. The recent launch of the Human Cell Atlas-Organoid initiative, documenting all human-derived organoids227, is the first step in that direction. For transcriptomics-related data, such as gene expression, non-coding RNA, ChIP, genome methylation, high-throughput PCR with reverse transcription, SNP arrays, SAGE and protein arrays, the Gene Expression Omnibus (GEO) is generally recommended228,229. For code used to analyse organoid data sets, GitHub is recommended. We envision that data and organoid depositories will arise shortly to fill in the presently missing gap in data repositories for organoid work. When depositing data from human organoids, it is important to consider all ethical regulations and the necessity of obtaining patient consent and keeping patient anonymity. Critical points have been extensively reviewed elsewhere230,231.
A consensus in organoid nomenclature is also missing. Aside from a few notable examples, where leaders in the intestine, liver and pancreas addressed the nomenclature of gut, hepatic, pancreatic and biliary organoids232,233, the field is waiting for a more clearly defined nomenclature system.
Limitations and optimizations
The reproducibility, both morphological and functional, of the obtained 3D organoid systems remains a major bottleneck. In this section, we elaborate on the limitations and recent developments in organoid research, providing a path towards a more optimal pipeline for developing, characterizing and benchmarking organoid systems.
Limited level of maturity and function
None of the present organoid model systems reproduces the entire physiological repertoire of cell types, maturation level and/or function of their respective organ; they, rather, exhibit certain functions of the tissue they predominantly form. The vast majority of tissue-derived organoid models are missing tissue-specific cell types, including niche-specific mesenchyme, immune cells, vascularization, innervation or microbiome. Recently, ductal cell–liver mesenchymal cell co-cultures have been shown to recapitulate part of the liver portal tract architecture73. Specifically challenging is that not all cell types have the same proliferation rate, growth factor requirements or even requirements for oxygen exposure (hypoxia for vasculature). Pluripotent stem cell-derived organoids are better in recapitulating different cell types and cellular interactions of the developing organ but fail on exhibiting adult-tissue structures and functions, as well as cell maturation. One strategy that helps is in vivo transplantation234. However, this is at the expense of giving up control over the formed tissue constructs. Meanwhile, optimizations on differentiation protocols are to enrich maturation and specific functions of interest.
Another factor contributing to limited maturity and function is the nutrient (in)accessibility and accumulation of dead cells in hollow lumens. This is particularly important for iPSC-derived organoids. As organoids grow in size, the nutrient supply to cells localized in the centre of the organoid gets restricted, resulting in cellular death. This is common with organoids that form a more compact structure, such as brain organoids. For tissue-derived organoids forming a hollow cyst (cholangiocyte, pancreas), dead cells will eventually start accumulating in their lumens, which cannot be avoided but can be resolved by mechanical fragmentation of organoids. Constant fragmentation of formed structures prevents carrying out the long-term studies. However, pluripotent stem cell-derived organoids cannot be fragmented and passaged; and new strategies to solve the nutrient accessibility problem are being developed, including the long-term maintenance of brain slices in vitro235.
Limited control of heterogeneity
Once cells form an organoid, we have minimal input in cellular behaviour within the organoid. Even in the same experimental settings, the result is often a plethora of phenotypic traits (shape, size, cell composition) rather than a stereotypic culture. Optimizing morphogenic gradients, tissue-specific cell–ECM interaction and local biochemical and biophysical properties are essential for minimizing batch to batch heterogeneity236.
To generate more complex multicellular mature and functional structures, the organoid field has started to create assembloids as demonstrated for human cortico-motor assembloids237. Such effort allows the creation of more complex structures, connecting multiple types of tissues with a defined interface such as connecting cerebral cortex, spine and skeletal muscle with neuro-muscular junctions, but at the cost of reproducibility. As recently discussed in another review focused on hepatic, biliary and pancreatic organoids233, reproducibility in multicellular and multi-tissue organoid systems decreases as it is challenging to coordinate proliferation and differentiation of multiple cell types.
The limited control of intra-organoid heterogeneity is detrimental for high-throughput screening applications and makes it difficult for studies requiring high spatio-temporal resolution imaging. Instead of creating more complex organoid systems, simpler models of reduced dimensions to recapitulate the essential tissue structures and functions of interest are gaining momentum. Variants of ECM combinations, micropatterned 2D mono-culture or co-culture238,239, cell sheets240, stacked 3D structures25 and micro-positioned ECM substrates69,241 allow the formation of reproducible tissue structures and functions with a high degree of spatio-temporal control such as stretching242 and osmotic forces243 (Fig. 5).
a–c, The organoid development process includes the formation of an initial 2D condition (part a), growth into a 3D structure (part b) and tissue morphogenesis (part c). d,e, Simpler models of reduced dimensions to recapitulate the essential tissue structures and functions of interest are gaining momentum. Micropatterned 2D mono-culture or co-culture allows the formation of a reproducible initial 2D condition which can further form the initial 3D structure. For example, crypt formation can be controlled by localized matrix softening and geometry can induce patterned differentiation with enterocytes (L-FABP stain) in the centre region and proliferating LGR5 cells in the buds69 (part d). Subsequently, a high degree of spatio-temporal control can be applied to direct certain tissue morphogenesis. For example, neural cell differentiation pattern can be modulated by stretch magnitude and timing242. Graph represents the stretch magnitude form 0 to 100% and scattered (S) and patterned (P) are the two configurations of differentiation patterning. f, Osmotic forces can control the crypt morphology in intestinal organoids243. Scale bars: 30 μm (part d), 50 μm (parts e and f ). Blebb, blebbistatin; hNTO, human neural tube organoid; PGE, prostaglandin E2. Part d adapted with permission from ref.69, AAAS. Part e adapted from ref.242, CC BY 4.0. Part f adapted from ref.243, Springer Nature Limited.
Optimizing ECM composition
Engineering methods have been implemented to optimize these limitations (Fig. 6). There are two main paths to overcome the use of non-specific ECM such as Matrigel: one is the use of synthetic matrices with more complete control over composition and stiffness, and the other is to take decellularized tissue and create tissue-specific matrices207,244. There are significant efforts to identify chemically defined, GMP-compatible ECMs that enable the growth and the long-term expansion of human organoids. In that regard, some advances were shown with human pancreas organoids, intestinal organoids and colorectal cancer organoids, which could grow in a dextran-based fully defined ECM; however, they would not expand long term3,68.
a, Accumulation of dead cells and cell debris inside cystic organoid lumina (left) has been overcome by designing perfusable open-end structures (right) that use inducible flow to wash out cell debris, which are more compatible with long-term experiments. b, Organoids grown in Matrigel domes display high variability of cell heterogeneity and morphology (left), which can be overcome by using grids with patterned synthetic extracellular matrix (ECM) (right) that provide cues for cell differentiation. These platforms are additionally compatible with high-throughput screenings. c, Single cell type-derived organoids do not recapitulate the cellular and physiological complexity of native tissue (left), but combining organoids with organ-on-a-chip (OoC) (right) as a novel technology would enable creating a controlled microenvironment, suitable for multiple cell types.
Organoids meet organs-on-a-chip
It has been shown that by maximizing the mass transfer and minimizing the shear stress in the perfusive soluble microenvironment, cells grown in OoC set-ups upregulate their functions, getting a step closer to a native tissue93,245,246. A more recent example shows how the presence of fluid flow enhances kidney organoids’ maturation and favours their vascularization in vitro106. Physical constraints were engineered into the organoid environment and intestinal cells when provided with boundaries through engineered scaffolds self-organized in crypts of the same size. At the same time, they overcame the inaccessibility of cystic organoids and clearance of cell debris by creating a perfusable culture of mini-intestines where cells are arranged to form tube-shaped epithelia and similar spatial arrangement to the in vivo tissue99.
Outlook
Moving forward, the trend is to develop more complex models that recapitulate in vivo structure and function as faithfully as possible, in terms of the recapitulating cell types over time, tissue architecture, measurable molecular events and phenotypic functions. Rather than focusing exclusively on the most prominent markers or functional assays, architectural benchmarking of native tissue should also be performed. In the example of hepatocyte organoids124, hepatocyte functions are preserved, yet the liver tissue architecture does not match the native tissue where hepatocytes are arranged in cords. Similarly, organoids such as pancreatic or colon cancer organoids grow isotropically, forming a cyst instead of the tubular structure they would form in their native tissue. To derive more complex functions, organoids with multicellular and multi-tissue structures will be important, especially in the context of studying cell–cell interactions247. Along this vein, assembloids and organs-on-chips are also becoming increasingly complex and more broadly adopted.
On the other hand, the engineer’s approach27,248 is to pursue simpler reductionist models defined by the minimal functional modules that drive a complex cellular or tissue function of interest, to study the mechano-biological causation in development or repair, or to develop a robust system for high-throughput screening. The basic premise is that a complex biological function is executed by coordinated operation of a limited number of functional modules, each described by a small set of molecules, and chemical reactions driving physical attribute changes of mesoscale (subcellular or intercellular tissue/multicellular) structures associated with functions of interest in the specific spatio-temporal stage/phase/step. For example, bile canaliculi in the liver exhibit hourly cycles of expansion and contraction. To study the causative contraction events with high resolution, only regions of adjacent hepatocytes forming the bile canaliculi are directly studied in the context of the entire regulatory machinery of adjacent hepatocytes26,249. One can choose to create a much larger structure involving cholangiocytes than the one operated by the minimal functional modules but the model will be noisy and costly. Each functional module is coupled to another and can be modelled together or independently at different length scale. Simple reductionist models have been useful for high-resolution mechanistic understanding of tissue morphogenetic events such as defects28,241,250.
Geometrically constraining the size of the initial 2D seeding pattern and 3D formation by micropatterning and supporting 3D cell growth using Matrigel enabled the induction of tissue-like neural tube morphogenesis and production of highly reproducible neural tubes. This also allowed identification of the mechanisms of neural tube folding and modelled neural tube defects25. In another example, symmetry breaking in a uniform sphere of cells and the emergence of a Paneth cell is a critical event in the early stage of intestinal organoid formation. The mechanism was recently elucidated: symmetry breaking is caused by transient activation of the mechanotransducer YAP1, which induces NOTCH-DLL1 lateral inhibition events206. YAP1 activation has since been controlled by applying geometrical constraints in hydrogel scaffolds and producing a uniform and reproducible intestinal microtissue3.
Organoids can be constrained by reducing the third dimension in a 2.5D culture. The 2.5D culture reduces the depth-driven variabilities of a typical organoid: diffusional constraints in the hypoxic core, limited accessibility for drugs/transfection agents and impeded imaging transparency251. Typical examples of restrictions in the third dimension include culturing cells on curved or patterned surfaces, a flattened or constrained cellular construct251 and overlaying the ECM on a flat cell monolayer at high confluency, which would pull the cells upward, forcing more cell–cell interactions to adopt 3D cell morphology. Hepatocytes in a collagen sandwich have sufficient contact area to attain polarity and form a bile canalicular lumen that contracts in the same periodic cycles as in vivo without the 3D network, wider and cholestatic compared with the native tissue. This cell-based model enables high-resolution dissection of the mechanism of bile canaliculi contraction into steps and an understanding of the molecular machinery regulating the phase transitions26,252. We would also envision more engineered organoid models based on CRISPR-edited cells for disease modelling, although these cells and models are synthetic.
Besides technological advances in creating more physiologically relevant, robust and easier to use organoid models, we envision seeing greater impact in applications. In the past two decades, although there have been discussions to replace animal testing, these efforts have not yet led to concrete actions. However, this is rapidly changing, with regulatory hard stops now established on multiple fronts253. Organoids that can recapitulate the complex physiological functions in vivo have also boosted confidence that the new alternative methods are now viable options. There will be more extrapolation of animal research findings in human organoids to better understand human biology and pathophysiology. We envision widespread adoption of organoids as cell sources for cell therapy, regenerative medicine, in vitro diagnostics and drug discovery.
References
Zakrzewski, W., Dobrzynski, M., Szymonowicz, M. & Rybak, Z. Stem cells: past, present, and future. Stem Cell Res. Ther. 10, 68 (2019).
Voog, J. & Jones, D. L. Stem cells and the niche: a dynamic duo. Cell Stem Cell 6, 103–115 (2010).
Gjorevski, N. et al. Designer matrices for intestinal stem cell and organoid culture. Nature 539, 560–564 (2016). This study introduces the concept of the stem cell niche, and describes that there are different requirements for mechanical cues at different stages of intestinal organoid formation.
Yi, S. A., Zhang, Y., Rathnam, C., Pongkulapa, T. & Lee, K. B. Bioengineering approaches for the advanced organoid research. Adv. Mater. 33, e2007949 (2021).
Orkin, R. et al. A murine tumor producing a matrix of basement membrane. J. Exp. Med. 145, 204–220 (1977).
Li, M. L. et al. Influence of a reconstituted basement membrane and its components on casein gene expression and secretion in mouse mammary epithelial cells. Proc. Natl Acad. Sci. USA 84, 136–140 (1987). This work demonstrates that Matrigel could support in vitro 3D culture and cell functions.
Sato, T. et al. Single Lgr5 stem cells build crypt–villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009). This work shows the stem cells’ potential to recapitulate native tissue-like structures and functions.
Huch, M. & Koo, B.-K. Modeling mouse and human development using organoid cultures. Development 142, 3113–3125 (2015).
Lancaster, M. A. & Knoblich, J. A. Generation of cerebral organoids from human pluripotent stem cells. Nat. Protoc. 9, 2329–2340 (2014).
Simian, M. & Bissell, M. J. Organoids: a historical perspective of thinking in three dimensions. J. Cell Biol. 216, 31–40 (2017).
Reis, R. L. 2nd Consensus conference on definitions on biomaterials science. J. Tissue Eng. Regen. Med. 14, 561–562 (2020).
Hofer, M. & Lutolf, M. P. Engineering organoids. Nat. Rev. Mater 6, 402–420 (2021). This review describes current limitations of organoid culture and proposed engineering approaches to address these limitations.
Clevers, H. Modeling development and disease with organoids. Cell 165, 1586–1597 (2016).
Kim, M. et al. Patient-derived lung cancer organoids as in vitro cancer models for therapeutic screening. Nat. Commun. 10, 3991 (2019). This work establishes personalized lung cancer organoids and normal bronchial organoids from patient tissues.
Al Shihabi, A. et al. Personalized chordoma organoids for drug discovery studies. Sci. Adv. 8, eabl3674 (2022). This publication is the first showing that patient-derived tumour organoids can be established for rare, indolent cancers and maintain the original tissue’s morphological and functional profiles.
Warmflash, A., Sorre, B., Etoc, F., Siggia, E. D. & Brivanlou, A. H. A method to recapitulate early embryonic spatial patterning in human embryonic stem cells. Nat. Methods 11, 847–854 (2014).
Wang, X., Liu, Z. & Pang, Y. Concentration gradient generation methods based on microfluidic systems. RSC Adv. 7, 29966–29984 (2017).
Kumar, A., Placone, J. K. & Engler, A. J. Understanding the extracellular forces that determine cell fate and maintenance. Development 144, 4261–4270 (2017).
White, C. R. & Frangos, J. A. The shear stress of it all: the cell membrane and mechanochemical transduction. Philos. Trans. R. Soc. Lond. B Biol. Sci. 362, 1459–1467 (2007).
Vatine, G. D. et al. Human iPSC-derived blood–brain barrier chips enable disease modeling and personalized medicine applications. Cell Stem Cell 24, 995–1005.e6 (2019).
Teng, Y., Zhao, Z., Tasnim, F., Huang, X. & Yu, H. A scalable and sensitive steatosis chip with long-term perfusion of in situ differentiated HepaRG organoids. Biomaterials 275, 120904 (2021).
Qian, X. et al. Brain-region-specific organoids using mini-bioreactors for modeling ZIKV exposure. Cell 165, 1238–1254 (2016).
Ladoux, B. & Mege, R. M. Mechanobiology of collective cell behaviours. Nat. Rev. Mol. Cell Biol. 18, 743–757 (2017).
Metavarayuth, K., Sitasuwan, P., Zhao, X., Lin, Y. & Wang, Q. Influence of surface topographical cues on the differentiation of mesenchymal stem cells in vitro. ACS Biomater. Sci. Eng. 2, 142–151 (2016).
Karzbrun, E. et al. Human neural tube morphogenesis in vitro by geometric constraints. Nature 599, 268–272 (2021). This work presents a chip-based culture system that enables self-organization of micropatterned stem cells into precise 3D cell fate patterns and organ shapes.
Gupta, K. et al. Bile canaliculi contract autonomously by releasing calcium into hepatocytes via mechanosensitive calcium channel. Biomaterials 259, 120283 (2020).
Sheetz, M. & Yu, H. The Cell as a Machine (Cambridge Univ. Press, 2018). This book explains the rationale of mechanobiology and the approaches to dissect complex biological functions.
Saw, T. B. et al. Topological defects in epithelia govern cell death and extrusion. Nature 544, 212–216 (2017).
Sato, T. et al. Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and Barrett’s epithelium. Gastroenterology 141, 1762–1772 (2011).
Yui, S. et al. Functional engraftment of colon epithelium expanded in vitro from a single adult Lgr5+ stem cell. Nat. Med. 18, 618–623 (2012).
Barker, N. et al. Lgr5+ve stem cells drive self-renewal in the stomach and build long-lived gastric units in vitro. Cell Stem Cell 6, 25–36 (2010).
Bartfeld, S. et al. In vitro expansion of human gastric epithelial stem cells and their responses to bacterial infection. Gastroenterology 148, 126–136.e6 (2015).
Aihara, E. et al. Characterization of stem/progenitor cell cycle using murine circumvallate papilla taste bud organoid. Sci. Rep. 5, 1–15 (2015).
Huch, M. et al. In vitro expansion of single Lgr5+ liver stem cells induced by Wnt-driven regeneration. Nature 494, 247–250 (2013). This study describes the first liver organoid cultures from adult mouse liver tissue, where cells can be expanded long term to form liver organoids even from a single cell.
Huch, M. et al. Long-term culture of genome-stable bipotent stem cells from adult human liver. Cell 160, 299–312 (2015).
Broutier, L. et al. Culture and establishment of self-renewing human and mouse adult liver and pancreas 3D organoids and their genetic manipulation. Nat. Protoc. 11, 1724–1743 (2016).
Takebe, T. et al. Vascularized and functional human liver from an iPSC-derived organ bud transplant. Nature 499, 481–484 (2013).
Greggio, C. et al. Artificial three-dimensional niches deconstruct pancreas development in vitro. Development 140, 4452–4462 (2013). This work presents the first fetal tissue organoid culture system that recapitulates pancreas progenitor development.
Huch, M. et al. Unlimited in vitro expansion of adult bi-potent pancreas progenitors through the Lgr5/R–spondin axis. EMBO J. 32, 2708–2721 (2013). This study is the first to produce pancreas organoids from adult tissue.
Manzar, G. S., Kim, E. M. & Zavazava, N. Demethylation of induced pluripotent stem cells from type 1 diabetic patients enhances differentiation into functional pancreatic β cells. J. Biol. Chem. 292, 14066–14079 (2017).
Drakhlis, L. et al. Human heart-forming organoids recapitulate early heart and foregut development. Nat. Biotechnol. 39, 737–746 (2021).
Koehler, K. R., Mikosz, A. M., Molosh, A. I., Patel, D. & Hashino, E. Generation of inner ear sensory epithelia from pluripotent stem cells in 3D culture. Nature 500, 217–221 (2013). This work presents the first inner ear organoids developed from iPSCs.
Lee, J. et al. Hair-bearing human skin generated entirely from pluripotent stem cells. Nature 582, 399–404 (2020).
Beers, J. et al. Passaging and colony expansion of human pluripotent stem cells by enzyme-free dissociation in chemically defined culture conditions. Nat. Protoc. 7, 2029–2040 (2012).
Hu, P., Zhang, W., Xin, H. & Deng, G. Single cell isolation and analysis. Front. Cell Dev. Biol. 4, 116 (2016).
Aronowitz, J. A., Lockhart, R. A. & Hakakian, C. S. Mechanical versus enzymatic isolation of stromal vascular fraction cells from adipose tissue. Springerplus 4, 713 (2015).
Gaipl, U. S. et al. Cooperation between C1q and DNase I in the clearance of necrotic cell-derived chromatin. Arthritis Rheum. 50, 640–649 (2004).
Vlachogiannis, G. et al. Patient-derived organoids model treatment response of metastatic gastrointestinal cancers. Science 359, 920–926 (2018).
Kopper, O. et al. An organoid platform for ovarian cancer captures intra- and interpatient heterogeneity. Nat. Med. 25, 838–849 (2019).
Tiriac, H. et al. Successful creation of pancreatic cancer organoids by means of EUS-guided fine-needle biopsy sampling for personalized cancer treatment. Gastrointest. Endosc. 87, 1474–1480 (2018).
Gao, D. et al. Organoid cultures derived from patients with advanced prostate cancer. Cell 159, 176–187 (2014).
Seino, T. et al. Human pancreatic tumor organoids reveal loss of stem cell niche factor dependence during disease progression. Cell Stem Cell 22, 454–467.e6 (2018).
Mazzocchi, A. et al. Pleural effusion aspirate for use in 3D lung cancer modeling and chemotherapy screening. ACS Biomater. Sci. Eng. 5, 1937–1943 (2019).
Lohmussaar, K. et al. Patient-derived organoids model cervical tissue dynamics and viral oncogenesis in cervical cancer. Cell Stem Cell 28, 1380–1396.e6 (2021).
Sartini, S. & Soragni, A. Cervical organoids go viral. Cell Stem Cell 28, 1337–1338 (2021).
De Angelis, M. L. et al. An organoid model of colorectal circulating tumor cells with stem cell features, hybrid EMT state and distinctive therapy response profile. J. Exp. Clin. Cancer Res. 41, 86 (2022).
Bergin, C. J. & Benoit, Y. D. Protocol for serial organoid formation assay using primary colorectal cancer tissues to evaluate cancer stem cell activity. STAR. Protoc. 3, 101218 (2022).
Fujii, M. et al. A colorectal tumor organoid library demonstrates progressive loss of niche factor requirements during tumorigenesis. Cell Stem Cell 18, 827–838 (2016).
Nguyen, H. T. L. & Soragni, A. Patient-derived tumor organoid rings for histologic characterization and high-throughput screening. STAR. Protoc. 1, 100056 (2020).
Aisenbrey, E. A. & Murphy, W. L. Synthetic alternatives to Matrigel. Nat. Rev. Mater. 5, 539–551 (2020).
Peng, H., Poovaiah, N., Forrester, M., Cochran, E. & Wang, Q. Ex vivo culture of primary intestinal stem cells in collagen gels and foams. ACS Biomater. Sci. Eng. 1, 37–42 (2015).
Cruz-Acuña, R. et al. Synthetic hydrogels for human intestinal organoid generation and colonic wound repair. Nat. Cell Biol. 19, 1326–1335 (2017).
Kratochvil, M. J. et al. Engineered materials for organoid systems. Nat. Rev. Mater. 4, 606–622 (2019).
Broguiere, N. et al. Growth of epithelial organoids in a defined hydrogel. Adv. Mater. 30, 1801621 (2018).
Chaudhuri, O., Cooper-White, J., Janmey, P. A., Mooney, D. J. & Shenoy, V. B. Effects of extracellular matrix viscoelasticity on cellular behaviour. Nature 584, 535–546 (2020).
Kloxin, A. M., Kasko, A. M., Salinas, C. N. & Anseth, K. S. Photodegradable hydrogels for dynamic tuning of physical and chemical properties. Science 324, 59–63 (2009).
Chrisnandy, A., Blondel, D., Rezakhani, S., Broguiere, N. & Lutolf, M. P. Synthetic dynamic hydrogels promote degradation-independent in vitro organogenesis. Nat. Mater. 21, 479–487 (2021).
Georgakopoulos, N. et al. Long-term expansion, genomic stability and in vivo safety of adult human pancreas organoids. BMC Dev. Biol. 20, 4 (2020).
Gjorevski, N. et al. Tissue geometry drives deterministic organoid patterning. Science 375, eaaw9021 (2022).
Valamehr, B. et al. Hydrophobic surfaces for enhanced differentiation of embryonic stem cell-derived embryoid bodies. Proc. Natl Acad. Sci. USA 105, 14459–14464 (2008).
Brandenberg, N. et al. High-throughput automated organoid culture via stem-cell aggregation in microcavity arrays. Nat. Biomed. Eng. 4, 863–874 (2020).
Czerniecki, S. M. et al. High-throughput screening enhances kidney organoid differentiation from human pluripotent stem cells and enables automated multidimensional phenotyping. Cell Stem Cell 22, 929–940.e4 (2018).
Cordero-Espinoza, L. et al. Dynamic cell contacts between periportal mesenchyme and ductal epithelium act as a rheostat for liver cell proliferation. Cell Stem Cell 28, 1907–1921.e8 (2021).
Jiang, S. et al. An automated organoid platform with inter-organoid homogeneity and inter-patient heterogeneity. Cell Rep. Med. 1, 100161 (2020).
Bues, J. et al. Deterministic scRNA-seq captures variation in intestinal crypt and organoid composition. Nat. Methods 19, 323–330 (2022).
Wang, Y. et al. In situ differentiation and generation of functional liver organoids from human iPSCs in a 3D perfusable chip system. Lab Chip 18, 3606–3616 (2018).
Goonoo, N. & Bhaw-Luximon, A. Mimicking growth factors: role of small molecule scaffold additives in promoting tissue regeneration and repair. RSC Adv. 9, 18124–18146 (2019).
Siller, R., Greenhough, S., Naumovska, E. & Sullivan, G. J. Small-molecule-driven hepatocyte differentiation of human pluripotent stem cells. Stem Cell Rep. 4, 939–952 (2015).
Janda, C. Y. et al. Surrogate Wnt agonists that phenocopy canonical Wnt and β-catenin signalling. Nature 545, 234–237 (2017).
Miao, Y. et al. Next-generation surrogate Wnts support organoid growth and deconvolute frizzled pleiotropy in vivo. Cell Stem Cell 27, 840–851.e6 (2020).
Ornitz, D. M. F. G. Fs heparan sulfate and FGFRs: complex interactions essential for development. Bioessays 22, 108–112 (2000).
Takasato, M. et al. Kidney organoids from human iPS cells contain multiple lineages and model human nephrogenesis. Nature 526, 564–568 (2015).
Mae, S. I. et al. Monitoring and robust induction of nephrogenic intermediate mesoderm from human pluripotent stem cells. Nat. Commun. 4, 1367 (2013).
Wang, Q. et al. Non-genetic engineering of cells for drug delivery and cell-based therapy. Adv. Drug Deliv. Rev. 91, 125–140 (2015).
Peng, H., Wang, C., Xu, X., Yu, C. & Wang, Q. An intestinal Trojan horse for gene delivery. Nanoscale 7, 4354–4360 (2015).
Davoudi, Z. et al. Intestinal organoids containing poly (lactic‐co‐glycolic acid) nanoparticles for the treatment of inflammatory bowel diseases. J. Biomed. Mater. Res. A 106, 876–886 (2018).
Davoudi, Z. et al. Gut organoid as a new platform to study alginate and chitosan mediated PLGA nanoparticles for drug delivery. Mar. Drugs 19, 282 (2021).
Gjorevski, N., Ranga, A. & Lutolf, M. P. Bioengineering approaches to guide stem cell-based organogenesis. Development 141, 1794–1804 (2014).
Dalby, M. J., Gadegaard, N. & Oreffo, R. O. Harnessing nanotopography and integrin–matrix interactions to influence stem cell fate. Nat. Mater. 13, 558–569 (2014).
Crane, G. M., Jeffery, E. & Morrison, S. J. Adult haematopoietic stem cell niches. Nat. Rev. Immunol. 17, 573–590 (2017).
Demers, C. J. et al. Development-on-chip: in vitro neural tube patterning with a microfluidic device. Development 143, 1884–1892 (2016).
Fluri, D. A. et al. Derivation, expansion and differentiation of induced pluripotent stem cells in continuous suspension cultures. Nat. Methods 9, 509–516 (2012).
Huh, D. et al. Reconstituting organ-level lung functions on a chip. Science 328, 1662–1668 (2010).
Perez-Gonzalez, C. et al. Mechanical compartmentalization of the intestinal organoid enables crypt folding and collective cell migration. Nat. Cell Biol. 23, 745–757 (2021).
Zhang, Z.-Z. et al. Orchestrated biomechanical, structural, and biochemical stimuli for engineering anisotropic meniscus. Sci. Transl. Med. 11, eaao0750 (2019).
Murphy, S. V. & Atala, A. 3D bioprinting of tissues and organs. Nat. Biotechnol. 32, 773–785 (2014).
Brassard, J. A., Nikolaev, M., Hübscher, T., Hofer, M. & Lutolf, M. P. Recapitulating macro-scale tissue self-organization through organoid bioprinting. Nat. Mater. 20, 22–29 (2021).
Park, S. E., Georgescu, A. & Huh, D. Organoids-on-a-chip. Science 364, 960–965 (2019).
Nikolaev, M. et al. Homeostatic mini-intestines through scaffold-guided organoid morphogenesis. Nature 585, 574–578 (2020). This study shows an intestinal organoid with a similar spatial arrangement of crypt-like and villus-like domains to that in vivo.
Esch, M. B., Mahler, G. J., Stokol, T. & Shuler, M. L. Body-on-a-chip simulation with gastrointestinal tract and liver tissues suggests that ingested nanoparticles have the potential to cause liver injury. Lab Chip 14, 3081–3092 (2014).
Bauer, S. et al. Functional coupling of human pancreatic islets and liver spheroids on-a-chip: towards a novel human ex vivo type 2 diabetes model. Sci. Rep. 7, 1–11 (2017).
Zhang, Y. S. et al. Multisensor-integrated organs-on-chips platform for automated and continual in situ monitoring of organoid behaviors. Proc. Natl Acad. Sci. USA 114, E2293–E2302 (2017).
Tao, T. et al. Engineering human islet organoids from iPSCs using an organ-on-chip platform. Lab Chip 19, 948–958 (2019).
Workman, M. J. et al. Enhanced utilization of induced pluripotent stem cell-derived human intestinal organoids using microengineered chips. Cell. Mol. Gastroenterol. Hepatol. 5, 669–677.e2 (2018).
Lee, K. K. et al. Human stomach-on-a-chip with luminal flow and peristaltic-like motility. Lab Chip 18, 3079–3085 (2018).
Homan, K. A. et al. Flow-enhanced vascularization and maturation of kidney organoids in vitro. Nat. Methods 16, 255–262 (2019). This study reports a microfluidic model to culture kidney organoids in vitro and shows flow-enhanced vascularization.
Grebenyuk, S. & Ranga, A. Engineering organoid vascularization. Front. Bioeng. Biotechnol. 7, 39 (2019).
Below, C. R. et al. A microenvironment-inspired synthetic three-dimensional model for pancreatic ductal adenocarcinoma organoids. Nat. Mater. 21, 110–119 (2022).
Wang, D. et al. Long-term expansion of pancreatic islet organoids from resident Procr+ progenitors. Cell 180, 1198–1211 e1119 (2020). This study identifies a new stem cell population with the potential to expand into organoids for long-term culture and transplantation.
Wang, J. et al. Endothelial Wnts control mammary epithelial patterning via fibroblast signaling. Cell Rep. 34, 108897 (2021).
Tuveson, D. & Clevers, H. Cancer modeling meets human organoid technology. Science 364, 952–955 (2019).
Friedlander, M. S. H., Nguyen, V. M., Kim, S. K. & Bevacqua, R. J. Pancreatic pseudoislets: an organoid archetype for metabolism research. Diabetes 70, 1051–1060 (2021).
Wang, J. et al. Isolation of mouse pancreatic islet Procr+ progenitors and long-term expansion of islet organoids in vitro. Nat. Protoc. https://doi.org/10.1038/s41596-022-00683-w (2022).
Ma, X. et al. Human expandable pancreatic progenitor-derived β cells ameliorate diabetes. Sci. Adv. 8, eabk1826 (2022).
Pagliuca, F. W. et al. Generation of functional human pancreatic β cells in vitro. Cell 159, 428–439 (2014).
Yoshihara, E. et al. Immune-evasive human islet-like organoids ameliorate diabetes. Nature 586, 606–611 (2020).
Date, S. & Sato, T. Mini-gut organoids: reconstitution of the stem cell niche. Annu. Rev. Cell Dev. Biol. 31, 269–289 (2015).
Rahmani, S., Breyner, N. M., Su, H. M., Verdu, E. F. & Didar, T. F. Intestinal organoids: a new paradigm for engineering intestinal epithelium in vitro. Biomaterials 194, 195–214 (2019).
Fujii, M. et al. Human intestinal organoids maintain self-renewal capacity and cellular diversity in niche-inspired culture condition. Cell Stem Cell 23, 787–793.e6 (2018).
Qu, M. et al. Establishment of intestinal organoid cultures modeling injury-associated epithelial regeneration. Cell Res. 31, 259–271 (2021).
Nakamura, T. Recent progress in organoid culture to model intestinal epithelial barrier functions. Int. Immunol. 31, 13–21 (2019).
Jung, K. B. et al. Interleukin-2 induces the in vitro maturation of human pluripotent stem cell-derived intestinal organoids. Nat. Commun. 9, 3039 (2018).
Son, Y. S. et al. Maturation of human intestinal organoids in vitro facilitates colonization by commensal Lactobacilli by reinforcing the mucus layer. FASEB J. 34, 9899–9910 (2020).
Hu, H. et al. Long-term expansion of functional mouse and human hepatocytes as 3D organoids. Cell 175, 1591–1606.e19 (2018). This study shows the establishment of hepatocyte organoid cultures from mouse and human fetal tissue.
Peng, W. C. et al. Inflammatory cytokine TNFα promotes the long-term expansion of primary hepatocytes in 3D culture. Cell 175, 1607–1619.e15 (2018). This study shows the establishment of hepatocyte organoid cultures for engraftment.
Lee, S. H. et al. Tumor evolution and drug response in patient-derived organoid models of bladder cancer. Cell 173, 515–528.e17 (2018).
Yu, L. et al. Patient-derived organoids of bladder cancer recapitulate antigen expression profiles and serve as a personal evaluation model for CAR-T cells in vitro. Clin. Transl. Immunol. 10, e1248 (2021).
Li, X. et al. Organoid cultures recapitulate esophageal adenocarcinoma heterogeneity providing a model for clonality studies and precision therapeutics. Nat. Commun. 9, 2983 (2018).
Honkala, A., Malhotra, S. V., Kummar, S. & Junttila, M. R. Harnessing the predictive power of preclinical models for oncology drug development. Nat. Rev. Drug Discov. 21, 99–114 (2022).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012).
Broutier, L. et al. Human primary liver cancer-derived organoid cultures for disease modeling and drug screening. Nat. Med. 23, 1424–1435 (2017). This study shows the first liver cancer organoid models from patient material.
Lo, Y. H., Karlsson, K. & Kuo, C. J. Applications of organoids for cancer biology and precision medicine. Nat. Cancer 1, 761–773 (2020).
Dijkstra, K. K. et al. Challenges in establishing pure lung cancer organoids limit their utility for personalized medicine. Cell Rep. 31, 107588 (2020).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
McQuin, C. et al. CellProfiler 3.0: next-generation image processing for biology. PLoS Biol. 16, e2005970 (2018).
Lindeboom, R. G. et al. Integrative multi-omics analysis of intestinal organoid differentiation. Mol. Syst. Biol. 14, e8227 (2018).
Hernández-de-Diego, R. et al. PaintOmics 3: a web resource for the pathway analysis and visualization of multi-omics data. Nucleic Acids Res. 46, W503–W509 (2018).
Raudvere, U. et al. g:Profiler: a web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 47, W191–W198 (2019).
Powers, R. K., Goodspeed, A., Pielke-Lombardo, H., Tan, A. C. & Costello, J. C. GSEA-InContext: identifying novel and common patterns in expression experiments. Bioinformatics 34, i555–i564 (2018).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Reimand, J. et al. Pathway enrichment analysis and visualization of omics data using g:Profiler, GSEA, Cytoscape and EnrichmentMap. Nat. Protoc. 14, 482–517 (2019).
Berg, S. et al. ilastik: interactive machine learning for (bio)image analysis. Nat. Methods 16, 1226–1232 (2019).
Kok, R. N. U. et al. OrganoidTracker: efficient cell tracking using machine learning and manual error correction. PLoS ONE 15, e0240802 (2020).
Mergenthaler, P. et al. Rapid 3D phenotypic analysis of neurons and organoids using data-driven cell segmentation-free machine learning. PLoS Comput. Biol. 17, e1008630 (2021).
Loomans, C. J. M. et al. Expansion of adult human pancreatic tissue yields organoids harboring progenitor cells with endocrine differentiation potential. Stem Cell Rep. 10, 712–724 (2018).
Elizondo, D. M. et al. Pancreatic islets seeded in a novel bioscaffold forms an organoid to rescue insulin production and reverse hyperglycemia in models of type 1 diabetes. Sci. Rep. 10, 4362 (2020).
Meran, L. et al. Engineering transplantable jejunal mucosal grafts using patient-derived organoids from children with intestinal failure. Nat. Med. 26, 1593–1601 (2020).
Sugimoto, S. et al. An organoid-based organ-repurposing approach to treat short bowel syndrome. Nature 592, 99–104 (2021).
Vives, J. & Batlle-Morera, L. The challenge of developing human 3D organoids into medicines. Stem Cell Res. Ther. 11, 72 (2020).
Phan, N. et al. A simple high-throughput approach identifies actionable drug sensitivities in patient-derived tumor organoids. Commun. Biol. 2, 78 (2019).
Tebon, P. J. et al. Drug screening at single-organoid resolution via bioprinting and interferometry. Preprint at bioRxiv https://doi.org/10.1101/2021.10.03.462896 (2021).
Chen, J. et al. An organoid-based drug screening identified a menin-MLL inhibitor for endometrial cancer through regulating the HIF pathway. Cancer Gene Ther. 28, 112–125 (2021).
Verissimo, C. S. et al. Targeting mutant RAS in patient-derived colorectal cancer organoids by combinatorial drug screening. eLife https://doi.org/10.7554/eLife.18489 (2016).
Nuciforo, S. et al. Organoid models of human liver cancers derived from tumor needle biopsies. Cell Rep. 24, 1363–1376 (2018).
Schnalzger, T. E. et al. 3D model for CAR-mediated cytotoxicity using patient-derived colorectal cancer organoids. EMBO J. 38, 100928 (2019).
Neal, J. T. et al. Organoid modeling of the tumor immune microenvironment. Cell 175, 1972–1988.e16 (2018).
Weeber, F. et al. Preserved genetic diversity in organoids cultured from biopsies of human colorectal cancer metastases. Proc. Natl Acad. Sci. USA 112, 13308–13311 (2015).
Yan, H. H. N. et al. A comprehensive human gastric cancer organoid biobank captures tumor subtype heterogeneity and enables therapeutic screening. Cell Stem Cell 23, 882–897.e11 (2018).
Williamson, C. T. et al. ATR inhibitors as a synthetic lethal therapy for tumours deficient in ARID1A. Nat. Commun. 7, 13837 (2016).
Letai, A. Functional precision cancer medicine — moving beyond pure genomics. Nat. Med. 23, 1028–1035 (2017). This review discusses the current stage of precision cancer and suggests future applications.
Le Tourneau, C. et al. Molecularly targeted therapy based on tumour molecular profiling versus conventional therapy for advanced cancer (SHIVA): a multicentre, open-label, proof-of-concept, randomised, controlled phase 2 trial. Lancet Oncol. 16, 1324–1334 (2015).
Meric-Bernstam, F. et al. Feasibility of large-scale genomic testing to facilitate enrollment onto genomically matched clinical trials. J. Clin. Oncol. 33, 2753–2762 (2015).
Sholl, L. M. et al. Institutional implementation of clinical tumor profiling on an unselected cancer population. JCI Insight https://doi.org/10.1172/jci.insight.87062 (2016).
Schwaederle, M. et al. On the road to precision cancer medicine: analysis of genomic biomarker actionability in 439 patients. Mol. Cancer Ther. 14, 1488–1494 (2015).
Flaherty, K. T. et al. Molecular landscape and actionable alterations in a genomically guided cancer clinical trial: national cancer institute molecular analysis for therapy choice (NCI-MATCH). J. Clin. Oncol. 38, 3883–3894 (2020).
Murciano-Goroff, Y. R., Taylor, B. S., Hyman, D. M. & Schram, A. M. Toward a more precise future for oncology. Cancer Cell 37, 431–442 (2020).
Murciano-Goroff, Y. R., Drilon, A. & Stadler, Z. K. The NCI-MATCH: a national, collaborative precision oncology trial for diverse tumor histologies. Cancer Cell 39, 22–24 (2021).
Friedman, A. A., Letai, A., Fisher, D. E. & Flaherty, K. T. Precision medicine for cancer with next-generation functional diagnostics. Nat. Rev. Cancer 15, 747–756 (2015).
van de Wetering, M. et al. Prospective derivation of a living organoid biobank of colorectal cancer patients. Cell 161, 933–945 (2015).
Xu, R., Zhou, X., Wang, S. & Trinkle, C. Tumor organoid models in precision medicine and investigating cancer–stromal interactions. Pharmacol. Ther. 218, 107668 (2021).
Kondo, T. Current status and perspectives of patient-derived rare cancer models. Hum. Cell 33, 919–929 (2020).
Kawasaki, K. et al. An organoid biobank of neuroendocrine neoplasms enables genotype-phenotype mapping. Cell 183, 1420–1435.e21 (2020).
Guillen, K. P. et al. A human breast cancer-derived xenograft and organoid platform for drug discovery and precision oncology. Nat. Cancer 3, 232–250 (2022).
Hubert, C. G. et al. A three-dimensional organoid culture system derived from human glioblastomas recapitulates the hypoxic gradients and cancer stem cell heterogeneity of tumors found in vivo. Cancer Res. 76, 2465–2477 (2016).
Matano, M. et al. Modeling colorectal cancer using CRISPR–Cas9-mediated engineering of human intestinal organoids. Nat. Med. 21, 256–262 (2015).
Ettayebi, K. et al. Replication of human noroviruses in stem cell-derived human enteroids. Science 353, 1387–1393 (2016).
Heo, I. et al. Modelling Cryptosporidium infection in human small intestinal and lung organoids. Nat. Microbiol. 3, 814–823 (2018).
Chen, S. et al. Rotavirus infection and cytopathogenesis in human biliary organoids potentially recapitulate biliary atresia development. mBio https://doi.org/10.1128/mBio.01968-20 (2020).
Sachs, N. et al. Long-term expanding human airway organoids for disease modeling. EMBO J. 38, 100300 (2019).
Driehuis, E. et al. Oral mucosal organoids as a potential platform for personalized cancer therapy. Cancer Discov. 9, 852–871 (2019).
Wroblewski, L. E. et al. Helicobacter pylori targets cancer-associated apical–junctional constituents in gastroids and gastric epithelial cells. Gut 64, 720–730 (2015).
Co, J. Y. et al. Controlling epithelial polarity: a human enteroid model for host-pathogen interactions. Cell Rep. 26, 2509–2520.e4 (2019).
Saxena, K. et al. Human intestinal enteroids: a new model to study human rotavirus infection, host restriction, and pathophysiology. J. Virol. 90, 43–56 (2016).
Salahudeen, A. A. et al. Progenitor identification and SARS-CoV-2 infection in human distal lung organoids. Nature 588, 670–675 (2020).
Lamers, M. M. et al. SARS-CoV-2 productively infects human gut enterocytes. Science 369, 50–54 (2020).
Zang, R. et al. TMPRSS2 and TMPRSS4 promote SARS-CoV-2 infection of human small intestinal enterocytes. Sci. Immunol. 5, 3582 (2020).
Van Goor, F. et al. Correction of the F508del-CFTR protein processing defect in vitro by the investigational drug VX-809. Proc. Natl Acad. Sci. USA 108, 18843–18848 (2011).
Van Goor, F. et al. Rescue of CF airway epithelial cell function in vitro by a CFTR potentiator, VX-770. Proc. Natl Acad. Sci. USA 106, 18825–18830 (2009).
Ratjen, F. et al. Cystic fibrosis. Nat. Rev. Dis. Primers 1, 15010 (2015).
Dekkers, J. F. et al. A functional CFTR assay using primary cystic fibrosis intestinal organoids. Nat. Med. 19, 939–945 (2013).
Verstegen, M. M. A. et al. Human extrahepatic and intrahepatic cholangiocyte organoids show region-specific differentiation potential and model cystic fibrosis-related bile duct disease. Sci. Rep. 10, 21900 (2020).
Dekkers, J. et al. Characterizing responses to CFTR-modulating drugs using rectal organoids derived from subjects with cystic fibrosis. Sci. Transl. Med. 8, 344ra384 (2016).
Berkers, G. et al. Rectal organoids enable personalized treatment of cystic fibrosis. Cell Rep. 26, 1701–1708.e3 (2019).
Brewington, J. J. et al. Detection of CFTR function and modulation in primary human nasal cell spheroids. J. Cyst. Fibros. 17, 26–33 (2018).
Schwank, G. et al. Functional repair of CFTR by CRISPR/Cas9 in intestinal stem cell organoids of cystic fibrosis patients. Cell Stem Cell 13, 653–658 (2013).
Boye, T., Steenholdt, C., Jensen, K. & Nielsen, O. Molecular manipulations and intestinal stem cell-derived organoids in inflammatory bowel disease. Stem Cells https://doi.org/10.1093/stmcls/sxac014 (2022).
Howell, K. et al. DNA methylation and transcription patterns in intestinal epithelial cells from pediatric patients with inflammatory bowel diseases differentiate disease subtypes and associate with outcome. Gastroenterology https://doi.org/10.1053/j.gastro.2017.10.007 (2017).
Soroka, C. et al. Bile‐derived organoids from patients with primary sclerosing cholangitis recapitulate their inflammatory immune profile. Hepatology https://doi.org/10.1002/hep.30470 (2018).
Danahay, H. et al. Notch2 is required for inflammatory cytokine-driven goblet cell metaplasia in the lung. Cell Rep. 10, 239–252 (2015).
Seidlitz, T. et al. Mouse models of human gastric cancer subtypes with stomach-specific CreERT2-mediated pathway alterations. Gastroenterology 157, 1599–1614.e2 (2019).
Sachs, N. et al. A living biobank of breast cancer organoids captures disease heterogeneity. Cell 172, 373–386.e10 (2018).
Boj, S. F. et al. Organoid models of human and mouse ductal pancreatic cancer. Cell 160, 324–338 (2015).
Rimland, C. A. et al. Regional differences in human biliary tissues and corresponding in vitro-derived organoids. Hepatology 73, 247–267 (2021).
Sampaziotis, F. et al. Reconstruction of the mouse extrahepatic biliary tree using primary human extrahepatic cholangiocyte organoids. Nat. Med. 23, 954–963 (2017).
Sampaziotis, F. et al. Cholangiocyte organoids can repair bile ducts after transplantation in the human liver. Science 371, 839–846 (2021).
Serra, D. et al. Self-organization and symmetry breaking in intestinal organoid development. Nature 569, 66–72 (2019).
Kozlowski, M. T., Crook, C. J. & Ku, H. T. Towards organoid culture without Matrigel. Commun. Biol. 4, 1387 (2021).
Aloia, L. et al. Epigenetic remodelling licences adult cholangiocytes for organoid formation and liver regeneration. Nat. Cell Biol. 21, 1321–1333 (2019).
Wang, Y. et al. A microengineered collagen scaffold for generating a polarized crypt–villus architecture of human small intestinal epithelium. Biomaterials 128, 44–55 (2017).
Lukonin, I. et al. Phenotypic landscape of intestinal organoid regeneration. Nature 586, 275–280 (2020).
Kretzschmar, K. & Clevers, H. Organoids: modeling development and the stem cell niche in a dish. Dev. Cell 38, 590–600 (2016).
Kim, J., Koo, B. K. & Knoblich, J. A. Human organoids: model systems for human biology and medicine. Nat. Rev. Mol. Cell Biol. 21, 571–584 (2020).
Palikuqi, B. et al. Adaptable haemodynamic endothelial cells for organogenesis and tumorigenesis. Nature 585, 426–432 (2020).
Tiriac, H. et al. Organoid profiling identifies common responders to chemotherapy in pancreatic cancerpancreatic cancer organoids parallel patient response. Cancer Discov. 8, 1112–1129 (2018).
Wensink, G. E. et al. Patient-derived organoids as a predictive biomarker for treatment response in cancer patients. NPJ Precis. Oncol. 5, 30 (2021).
van Mourik, P. et al. Rationale and design of the HIT-CF organoid study: stratifying cystic fibrosis patients based on intestinal organoid response to different CFTR-modulators. Transl. Med. Commun. https://doi.org/10.1186/s41231-020-00060-3 (2020).
Foo, M. A. et al. Clinical translation of patient-derived tumour organoids — bottlenecks and strategies. Biomark Res. 10, 10 (2022).
Meier, M. A. et al. Patient-derived tumor organoids for personalized medicine in a patient with rare hepatocellular carcinoma with neuroendocrine differentiation: a case report. Commun. Med. 2, 80 (2022).
Blainey, P., Krzywinski, M. & Altman, N. Points of significance: replication. Nat. Methods 11, 879–880 (2014).
Pollard, D. A., Pollard, T. D. & Pollard, K. S. Empowering statistical methods for cellular and molecular biologists. Mol. Biol. Cell 30, 1359–1368 (2019).
Krzywinski, M. & Altman, N. Significance, P values and t-tests. Nat. Methods 10, 1041–1042 (2013).
Krzywinski, M. & Altman, N. Points of significance: error bars. Nat. Methods 10, 921–922 (2013).
Nuzzo, R. Scientific method: statistical errors. Nature 506, 150–152 (2014).
Weissgerber, T. L., Milic, N. M., Winham, S. J. & Garovic, V. D. Beyond bar and line graphs: time for a new data presentation paradigm. PLoS Biol. 13, e1002128 (2015).
Lord, S. J., Velle, K. B., Mullins, R. D. & Fritz-Laylin, L. K. SuperPlots: communicating reproducibility and variability in cell biology. J. Cell Biol. https://doi.org/10.1083/jcb.202001064 (2020).
Nature. Statistics for biologists. Nature https://www.nature.com/collections/qghhqm/ (2017).
Bock, C. et al. The Organoid Cell Atlas. Nat. Biotechnol. 39, 13–17 (2021).
Barrett, T. et al. NCBI GEO: archive for functional genomics data sets — 10 years on. Nucleic Acids Res. 39, D1005–D1010 (2011).
Edgar, R., Domrachev, M. & Lash, A. E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002).
Bredenoord, A. L., Clevers, H. & Knoblich, J. A. Human tissues in a dish: the research and ethical implications of organoid technology. Science 355, eaaf9414 (2017).
Fuhr, A., Kurtz, A., Hiepen, C. & Müller, S. Organoids as miniature twins — challenges for comparability and need for data standardization and access. Organoids 1, 28–36 (2022).
Stelzner, M. et al. A nomenclature for intestinal in vitro cultures. Am. J. Physiol. Gastrointest. Liver Physiol 302, G1359–G1363 (2012).
Marsee, A. et al. Building consensus on definition and nomenclature of hepatic, pancreatic, and biliary organoids. Cell Stem Cell 28, 816–832 (2021).
Takebe, T. & Wells, J. M. Organoids by design. Science 364, 956–959 (2019).
Xiang, Z. et al. Long-term maintenance of mature hippocampal slices in vitro. J. Neurosci. Methods 98, 145–154 (2000).
Takebe, T., Zhang, B. & Radisic, M. Synergistic engineering: organoids meet organs-on-a-chip. Cell Stem Cell 21, 297–300 (2017).
Andersen, J. et al. Generation of functional human 3D cortico-motor assembloids. Cell 183, 1913–1929.e26 (2020).
Toh, Y. C., Xing, J. & Yu, H. Modulation of integrin and E-cadherin-mediated adhesions to spatially control heterogeneity in human pluripotent stem cell differentiation. Biomaterials 50, 87–97 (2015).
Toh, Y. C., Blagovic, K., Yu, H. & Voldman, J. Spatially organized in vitro models instruct asymmetric stem cell differentiation. Integr. Biol. 3, 1179–1187 (2011).
Efremov, A. K. et al. Application of piconewton forces to individual filopodia reveals mechanosensory role of L-type Ca2+ channels. Biomaterials 284, 121477 (2022).
Li, Q. et al. Extracellular matrix scaffolding guides lumen elongation by inducing anisotropic intercellular mechanical tension. Nat. Cell Biol. 18, 311–318 (2016).
Abdel Fattah, A. R. et al. Actuation enhances patterning in human neural tube organoids. Nat. Commun. 12, 3192 (2021).
Yang, Q. et al. Cell fate coordinates mechano-osmotic forces in intestinal crypt formation. Nat. Cell Biol. 23, 733–744 (2021).
Rezakhani, S., Gjorevski, N. & Lutolf, M. P. Extracellular matrix requirements for gastrointestinal organoid cultures. Biomaterials 276, 121020 (2021).
Zhang, C., Zhao, Z., Abdul Rahim, N. A., van Noort, D. & Yu, H. Towards a human-on-chip: culturing multiple cell types on a chip with compartmentalized microenvironments. Lab Chip 9, 3185–3192 (2009).
Toh, Y. C. et al. A novel 3D mammalian cell perfusion-culture system in microfluidic channels. Lab Chip 7, 302–309 (2007).
Koike, H. et al. Modelling human hepato-biliary-pancreatic organogenesis from the foregut–midgut boundary. Nature 574, 112–116 (2019).
Thomas, J. D., Lee, T. & Suh, N. P. A function-based framework for understanding biological systems. Annu. Rev. Biophys. Biomol. Struct. 33, 75–93 (2004).
Ghallab, A. et al. Bile microinfarcts in cholestasis are initiated by rupture of the apical hepatocyte membrane and cause shunting of bile to sinusoidal blood. Hepatology 69, 666–683 (2019).
Zhang, Y. et al. Biomimetic niches reveal the minimal cues to trigger apical lumen formation in single hepatocytes. Nat. Mater. 19, 1026–1035 (2020).
Baptista, D., Teixeira, L., van Blitterswijk, C., Giselbrecht, S. & Truckenmuller, R. Overlooked? Underestimated? Effects of substrate curvature on cell behavior. Trends Biotechnol. 37, 838–854 (2019).
Gupta, K. et al. Actomyosin contractility drives bile regurgitation as an early response during obstructive cholestasis. J. Hepatol. 66, 1231–1240 (2017).
Grimm, D. U.S. EPA to eliminate all mammal testing by 2035. Science https://www.science.org/content/article/us-epa-eliminate-all-mammal-testing-2035 (2019).
Acknowledgements
This work was supported in part by grants from CAS (XDA16020200 to Y.A.Z.), the National Key Research, Development Program of China (2020YFA0509002 to Y.A.Z.) and the National Institutes of Health (NIH) (R01CA240718, R01CA244729 and R01CA264248 to A.So.), and funding from the Institute of Bioengineering and Bioimaging, Biomedical Research Council, Agency for Science, Technology and Research (A*STAR) (Project Number IAF (H18/01/a0/017)), SMART CAMP, The Institute for Digital Medicine (WisDM) and Mechanobiology Institute of Singapore (R-714-106-004-135) to H.Y.
Author information
Authors and Affiliations
Contributions
Introduction (H.Y., Z.Z. and E.L.S.F.); Experimentation (Z.Z., K.B., E.L.S.F. and Q.W.); Results (X.C., A.M.D., A.Sl., L.L., Z.C., A.So., M.H. and Y.A.Z.); Applications (L.L. and A.So.); Reproducibility and data deposition (A.M.D., A.Sl. and M.H.); Limitations and optimizations (H.Y., A.M.D., A.Sl. and M.H.); Outlook (H.Y., Z.Z. and G.M.B.); Overview of the Primer (H.Y.).
Corresponding authors
Ethics declarations
Competing interests
M.H. is inventor on several patents on organoid technology. A.So. and L.L. are inventors on a patent on organoid technology. A.So. is founder and owner of Icona BioDx. H.Y. is inventor on several patents on cell-based models and founder of Ants Innovate and Invitrocue. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Methods Primers thanks Sebastian Merker, Toshiro Sato, Michael Schumacher and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Glossary
- 2.5D culture
-
3D culture with a restricted third dimension.
- Assembloids
-
Complex 3D structures combining several separately pre-generated cellular compartments and/or entities.
- Bioprinting
-
An additive manufacturing technique that enables direct deposition of stem cells, organoids and biomaterials to fabricate 3D organoid-based tissue structures with controlled cell–matrix structures.
- Engraftment rates
-
Degrees of organoid retention in a host tissue after transplantation.
- Organogenesis
-
The series of organized integrated processes that transform an amorphous mass of cells into a complete organ in the developing embryo.
- Passaging
-
The subculturing of organoids by creating either smaller organoid fragments or single cells using mechanical dissociation or enzymatic digestion.
- Stem cell niche
-
A specific tissue microenvironment where stem cells both reside and receive stimuli that regulate cell fate.
- Streptozotocin
-
An agent toxic to the insulin-producing β-cells of the pancreas in mammals.
- Temporal heterogeneity
-
Here, differences in the composition and changes during the time of passaging the organoids.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhao, Z., Chen, X., Dowbaj, A.M. et al. Organoids. Nat Rev Methods Primers 2, 94 (2022). https://doi.org/10.1038/s43586-022-00174-y
Accepted:
Published:
DOI: https://doi.org/10.1038/s43586-022-00174-y
- Springer Nature Limited
This article is cited by
-
Bispecific aptamer-decorated and light-triggered nanoparticles targeting tumor and stromal cells in breast cancer derived organoids: implications for precision phototherapies
Journal of Experimental & Clinical Cancer Research (2024)
-
Organoid as a promising tool for primary liver cancer research: a comprehensive review
Cell & Bioscience (2024)
-
Establishment of patient-derived organoids and a characterization based drug discovery platform for treatment of gastric cancer
Cancer Cell International (2024)
-
Organoids and tissue/organ chips
Stem Cell Research & Therapy (2024)
-
Tumoroids, a valid preclinical screening platform for monitoring cancer angiogenesis
Stem Cell Research & Therapy (2024)