Abstract
Centromere pairing is crucial for synapsis in meiosis. This study delves into the Skp1-Cullin1-F-box protein (SCF) E3 ubiquitin ligase complex, specifically focusing on F-box protein 47 (FBXO47), in mouse meiosis. Here, we revealed that FBXO47 is localized at the centromere and it regulates centromere pairing cooperatively with SKP1 to ensure proper synapsis in pachynema. The absence of FBXO47 causes defective centromeres, resulting in incomplete centromere pairing, which leads to corruption of SC at centromeric ends and along chromosome axes, triggering premature dissociation of chromosomes and pachytene arrest. FBXO47 deficient pachytene spermatocytes exhibited drastically reduced SKP1 expression at centromeres and chromosomes. Additionally, FBXO47 stabilizes SKP1 by down-regulating its ubiquitination in HEK293T cells. In essence, we propose that FBXO47 collaborates with SKP1 to facilitate centromeric SCF formation in spermatocytes. In summary, we posit that the centromeric SCF E3 ligase complex regulates centromere pairing for pachynema progression in mice.
Similar content being viewed by others
Introduction
During meiosis, homologous chromosomes generate DNA double-strand breaks (DSBs) by the topoisomerase SPO111,2, proceed homologous recombination to shuffle genetic materials through the process termed synapsis3, which is facilitated by synaptonemal complex (SC) assembly4. In mice, the SC is composed of three major components: two parallel lateral elements (LEs), transverse filaments connecting the two LEs, and a central element5. The transverse element, SYCP1, provides a core architecture to the SC6. The formation of the SC is governed by intricate molecular mechanisms during the zygotene to pachytene stages7. Ultimately, the recombination intermediates are resolved into crossover events that give rise to chiasmata8. These chiasmata physically link homologous chromosomes, ensuring their proper alignment and segregation during meiosis prophase I9.
Centromeres are sites where the kinetochore forms and attaches to spindle microtubules during mitosis and meiosis for chromosome segregation10,11. In mice, different from humans, centromeres are located at one of the telomeric ends on chromosomes12. Centromere protein C (CENP-C) is a conserved constitutive centromere protein in eukaryotes13,14,15, which is essential for centromere establishment and stability15.
Centromere clustering is an important event that occurs at the beginning of meiosis, where non-homologous centromeres cluster closely in groups of two or more to facilitate proper centromere pairing16,17,18. Centromere pairing can facilitate homologous chromosome pairing, a critical step in meiotic recombination that ensures the proper segregation of homologs during meiosis I17,19,20. During meiosis, not all chromosomes can undergo crossover to produce chiasmata that promote bivalents to attach to the meiotic spindle. For those chromosomes that fail to form crossovers, the only linkage to facilitate cell segregation is provided by the connection of centromeres16. Defects in chromosome segregation can lead to aneuploidy, which is a major cause of birth defects and cancer21,22.
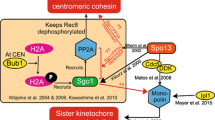
The molecular mechanisms governing centromere pairing are still being elucidated, yet several factors have been implicated in this process. CENP-C23 and centromeric SMC124 have been identified as essential for centromere clustering and pairing in Drosophila. Besides, SET domain bifurcated histone lysine methyltransferase 1 (SETDB1) plays a crucial role in centromere clustering and bouquet formation in mice25. Additionally, cohesin complex subunit ZmSMC3 participates in meiotic centromere pairing in maize26. Furthermore, components of SC act at the centromeres to promote the establishment and maintenance of centromere pairing, thereby enhancing the segregation fidelity of mammalian meiotic chromosomes27,28.
The S-phase kinase-associated protein 1 (Skp1)-Cullin1-F-box protein (SCF) E3 ubiquitin ligase complexes are integral to numerous biological processes29,30. The F-box motif of the F-box family proteins plays a vital role in these processes, as it mediates the recruitment of the SCF E3 ligase complex by directly interacting with the adapter protein SKP131. The specificity of substrate targeting for ubiquitination is determined by the particular F-box protein component within the SCF complex32,33. Research has elucidated that the SCF-Fbxo42 complex facilitates SC assembly during Drosophila female meiosis34. Furthermore, SKP1 has been identified as a crucial factor in maintaining DSB homeostasis and ensuring the successful progression of meiosis in mouse spermatocytes35,36.
Conflicting reports exist regarding the role of FBXO47 in mouse spermatocytes. One study reported that FBXO47 is a telomere-related protein that mediates bouquet formation and telomere-nuclear envelope (NE) attachment. This study suggests that knockout (KO) of Fbxo47 arrests at a late-zygotene stage and shows aberrations in telomere-protecting proteins37. However, another study asserts that FBXO47 has entirely different functions in mouse spermatocytes. It has been reported that Fbxo47 KO mice exhibit unrepaired DSBs, diplotene arrest, and bouquet formation is normal in these mice. It is suggested that FBXO47 is a stabilizer of SC in mouse spermatocytes through unknown underlying mechanisms38. Moreover, opposite conclusions were also drawn regarding whether FBXO47 interacts with SKP1.
Our initial motivation to experiment with Fbxo47 KO mice stemmed from our RNA-sequencing data, which indicated a notable expression of Fbxo47 in spermatocytes, suggesting a pivotal role for this protein in meiotic processes. The challenge of working with FBXO47 protein primarily stems from the difficulty in sourcing reliable antibodies, which can result in inconclusive outcomes both for our research and for the broader scientific community. Additionally, variations in knockout methodologies could account for the differing results observed across studies.
Through comprehensive analysis in this study, we have demonstrated that the deficiency of FBXO47 at the centromeres leads to centromere defects and destabilized centromere pairing, culminating in the collapse of the SC and premature disassembly, followed by (or synchronized with) the destruction of the telomere-NE attachment system at the pachytene stage. This novel finding addresses the discrepancies noted in prior studies and advances our understanding of the role of FBXO47. We have also delineated how FBXO47 contributes to stabilizing SKP1 expression and have confirmed the involvement of the centromeric SCF E3 ligase complex in the progression of meiosis.
Results
FBXO47 is located at centromeres in spermatocytes during meiosis prophase I
Firstly, we investigated the location of FBXO47 from prophase to metaphase I using chromosome spreads of wild-type (WT) mouse spermatocytes and indirect immunofluorescence (IF) staining. We observed that from zygotene to diplotene stages, FBXO47 was uniquely located on just one telomere of the autosomes (Fig. 1a–c, a′–c′, dotted lines, arrows), but not on the other telomere (Fig. 1a–c, a′–c′, dotted lines, arrowheads). At the diakinesis stage, FBXO47 signals were detected between SYCP3 and tubulin signals (Fig. 1d, d′, arrows) and disappeared at metaphase I (Fig. 1e, e′, arrowheads), indicating its functional role in prometaphase. With mild chromosome spreads where the nuclear membrane remained intact, we found that the FBXO47-expressing telomeric ends were consistently restricted to the brighter areas of DAPI staining, indicating the heterochromatic region of chromosomes from zygotene to diplotene stages (Supplementary Fig. S1a–c, arrows). This led us to speculate that FBXO47 might be localized at the centromeric ends of the chromosomes, especially since mouse centromeres are known to be located near the telomeres12.
a–e, a′–e′ Immunolocalization of SYCP3 (red) and FBXO47 (green) in mouse spermatocytes at the zygotene (a and a′), pachytene (b and b′), diplotene (c and c′), diakinesis (d and d′), and metaphase (e and e′) stages. f–i Immunolocalization of SYCP3 (blue), FBXO47 (green), and centromere (red) in Fbxo47+/+ (f) and Fbxo47−⁄− (i) spermatocytes. Detailed views of their location on autosomes (g) and the XY chromosomes (h) are provided. Chromosomes are outlined with dotted lines for clarity. Arrows indicate the FBXO47-positive ends of chromosomes, whereas arrowheads denote the FBXO47-negative ends. The PAR region is indicated for reference.
To further clarify the localization of FBXO47, we performed combined IF using anti-centromere antibody (ACA) along with FBXO47 and SYCP3 antibodies. We confirmed that FBXO47 was localized at the centromeres (Fig. 1f, dotted lines, arrows). No signal for either the centromere or FBXO47 was detected at the other ends of the same autosomes (Fig. 1f, dotted lines, arrowheads). Additionally, the FBXO47 signal co-localized with the centromere signals (Fig. 1g, arrows), but not at the non-centromeric end (Fig. 1g, arrowheads). Regarding the sex chromosomes, we also verified the co-localization of FBXO47 at the centromeric ends of X and Y chromosomes, which are known to separate during the pachytene stage of mouse spermatocytes (Fig. 1h, arrows)39, but not at the pseudoautosomal region (PAR) ends (Fig. 1h, arrowheads).
However, in Fbxo47−⁄− spermatocytes (KO strain will be mentioned later), no FBXO47 signals could be detected (Fig. 1i, dotted lines, arrows). Since non-specific binding could sometimes be observed in Fbxo47−⁄− spermatocytes by IF, we also conducted Immunohistochemistry (IHC) studies with this antibody on chromosome spreads. DAB staining revealed no non-specific binding (Supplementary Fig. S1d–g), indicating that this antibody is specific.
In summary, our findings demonstrated that FBXO47 is located at the centromeres of mouse spermatocytes during meiotic prophase I.
Delayed leptonema/zygonema and premature pachytene cell death in Fbxo47 −⁄− spermatocytes
To elucidate the impact of FBXO47 on meiosis, we created a Fbxo47−⁄− mouse strain by deleting exon 2 to exon 11 of Fbxo47 gene using CRISPR/Cas9 genome editing (Supplementary Fig. S2a, b). Male Fbxo47−⁄− mice exhibited infertility and reduced testicular size (Supplementary Fig. S2c), whereas Fbxo47+⁄− males remained fertile. Histological analyses showed that, compared to the normal PD30 testis with various differentiated spermatocytes (Supplementary Fig. S2d, left), the most advanced cells in Fbxo47−⁄− seminiferous tubules were likely pachytene cells (Supplementary Fig. S2d, right, red arrows), and no spermatids were observed. Our further analysis indicated that Fbxo47−⁄− spermatocytes displayed defective DSB repair, in agreement with one previous finding38, and a pachytene arrest without crossover formation (for details, see Supplementary Information and Fig. S2e–q).
Given the conflicting reports on the stage at which meiosis ceases in Fbxo47−⁄− spermatocytes, we conducted a synchronized meiosis analysis using a series of retinoic acid (RA) inhibitor (WIN18,446) injections to inhibit meiosis, followed by a single dose of RA injection to induce meiotic synchronization, letting the cells to develop with one accord. Testes were collected at various time points post-RA injection for chromosome spread analysis. Staining with SYCP3, SYCP1, and centromere antibodies revealed that, at 6.75 days post-RA injection, approximately 90% of the cells were pre-leptotene and 10% were leptotene in both Fbxo47+⁄− (Fig. 2a) and Fbxo47−⁄− (Fig. 2b) spermatocytes, indicating normal meiotic initiation in Fbxo47−⁄− mice. Signals of phosphorylated H2AX at Ser 139 (γH2AX), indicative of DSBs, began to appear on this day (Fig. 2a′, b′).
A synchronized analysis of meiotic progression was conducted in both Fbxo47−⁄− and Fbxo47+/− spermatocytes. Testicular chromosome spreads were collected at various time points following retinoic acid (RA) injection, and were stained for SYCP3 (red), SYCP1 (green), centromeres (blue), as well as with γH2AX antibodies. In the case of Fbxo47+/− testes: a, a′ At 6.75 days post-injection, cells are in the pre-leptotene stage; c, c′ By 7.75 days, cells have reached the zygotene stage, with the arrow denoting the synapsed region; e, e′ At 9 days, cells are in late-zygotene stage, with the arrow indicating the unsynapsed centromeric ends that remains; g, g′ By 10 days, cells have progressed to the pachytene stage; i, i′ At 11 days, cells continue to be observed in pachytene stage; k, k′ By 12 days, cells have entered early-diplotene stage, with the arrow pointing to an inflated chromosomal end; m, m′ At 14 days, diplotene cells are seen, with the arrow indicating partially desynapsed chromosomes; o, o′ Also at 14 days, the second wave of zygotene spermatocytes is present, with the arrow highlighting the synapsed region. In contrast, within Fbxo47−⁄− testes: b, b′ At 6.75 days post-injection, cells are also in the pre-leptotene stage; d, d′ By 7.75 days, cells are still at the leptotene stage; f, f′ At 9 days, cells are in early-zygotene stage, with the arrow indicating the synapsed region; h, h′ By 10 days, cells are in the zygotene stage, with arrows pointing out the synapsed chromosome segments; j, j′ At 11 days, cells resemble those in pachytene, with arrows pointing to unpaired centromeric ends; l, l′ By 12 days, cells are in a desynapsed pachytene (p)-like state, with the arrows indicating the SYCP1 signals; n, n′ Also at 12 days, the second wave of leptotene cells is observed; p, p′ At 14 days, the second wave of early-zygotene cells is present. All time points were repeated twice to ensure the accuracy and reproducibility of the observations. The data presented here underscore the critical role of Fbxo47 in the proper progression of meiosis during spermatogenesis.
At 7.75 days, Fbxo47+⁄− spermatocytes displayed SYCP1 signals along the chromosome axes, suggesting synapsis in zygotene cells (Fig. 2c, c′, arrow). However, Fbxo47−⁄− spermatocytes remained at the leptotene stage, with no SYCP1 signal observed (Fig. 2d, d′). By day 9, Fbxo47+⁄− spermatocytes progressed to the late-zygotene stage, with only some unpaired centromeres visible (Fig. 2e, arrow), and the γH2AX signal decreased as DSB repair occurred (Fig. 2e′). In contrast, Fbxo47−⁄− spermatocytes began to show SYCP1 signals at day 9, indicating early-zygotene stage (Fig. 2f, arrow), with strong γH2AX signals (Fig. 2f′). These findings suggest delays in leptotene and zygotene stages in Fbxo47−⁄− spermatocytes, likely due to extended repair duration based on increased RAD51, and RPA2 signals observed in these cells (Supplementary Information and Fig. S2e–j).
At 10 and 11 days, Fbxo47+⁄− spermatocytes reached the pachytene stage with well-paired autosomes (Fig. 2g, i), and γH2AX signals were observed only in the X-Y body region (Fig. 2g′, i′). However, Fbxo47−⁄− spermatocytes progressed to the zygotene stage with increased SYCP1 signals (Fig. 2h, h′, arrows), followed by a pachytene-like stage with unpaired centromeric ends (Fig. 2j, arrows) and reduced γH2AX signals (Fig. 2j′). We refer to these cells as “pachytene-like” because they contain highly condensed chromosomes, and over 80% of homologous chromosomes were synapsed in these cells. Additionally, the BRCT domain–containing protein TOPBP1 location (similar to BRCA1 reported previously38) largely confined to XY body, highly resembled those in wild-type pachytene stage (Supplementary Fig. S3c, g). For differentiation of each meiotic stages, please refer to Supplementary Information and Fig. S3. Notably, although meiotic progression was not perfectly synchronized, no separated interstitial chromosomal regions were observed at day 11 in Fbxo47−⁄− spermatocytes, indicating that chromosome dissociation had not yet occurred.
By day 12, Fbxo47+⁄− spermatocytes were at the early-diplotene stage with inflated chromosomal ends (Fig. 2k, arrow) and no γH2AX signal on autosomes (Fig. 2k′). However, Fbxo47−⁄− spermatocytes existed dissociated chromosome arms with rare or no SYCP1 signals (Fig. 2l, l′, arrows). These data indicate that FBXO47-deficient pachytene-like cells exist centromere pairing defects, fail to pass the pachytene checkpoint, which subsequently leads to premature desynapsis. We refer to these cells as “desynapsed pachytene-like” cells, as no TOPBP1 signal could be noticed at the desynapsed chromosomal regions (Supplementary Fig. S3h, i), resembling desynapsis in wild-type diplotene stage (Supplementary Fig. S3d). Additionally, the second wave of leptotene cells began to appear (Fig. 2n, n′).
By day 14, chromosomes in Fbxo47+⁄− spermatocytes had desynapsed, with SYCP1 signals only at synapsed regions (Fig. 2m, m′, arrow), and the second wave of zygotene cells could be observed (Fig. 2o, o′, arrow). However, in Fbxo47−⁄− spermatocytes, only zygotene-like cells were found, with no SYCP1 signals visible yet (Fig. 2p, p′). These findings indicate that the first wave of spermatocytes had already died by day 14, and the second wave of spermatocytes was also delayed in Fbxo47−⁄− mice.
In conclusion, the synchronized meiosis analysis revealed that Fbxo47−⁄− spermatocytes experienced delays at the leptotene and zygotene stages due to defective DSB repair. Fbxo47−⁄− spermatocytes were arrested at the pachytene-like stage with centromere pairing defects, exhibiting premature dissociation and cell death. These findings suggest that FBXO47 plays a critical role in meiotic DSB repair and pachynema progression in spermatocytes.
Defective centromere pairing in Fbxo47 −⁄− pachytene cells
To delve into the underlying causes of defects in Fbxo47−⁄− pachytene spermatocytes, we conducted IF staining on chromosome spreads from PD30 Fbxo47+/+ and Fbxo47−⁄− testes using antibodies against SYCP3, SYCP1, and centromeres. Enlarged images of the framed regions in Fig. 3 were included to show the details.
IF staining was conducted using antibodies against SYCP1 (green), SYCP3 (red), and centromeres (blue) to investigate the defective pachynema in Fbxo47−⁄− mice. a–f, a′–f′ depict the location of SYCP1, SYCP3, and centromeres in zygotene (a, a′, b, b′) and pachytene (e, e′, f, f′) cells of Fbxo47+/+ mice. c, c′, d, d′ show the location of SYCP1, SYCP3, and centromeres in zygotene cells of Fbxo47−⁄− mice; insets display enlarged sections. Arrows indicate centromeres, while arrowheads point to the SYCP1 signals. g–j, g′–j′ represent the expression of SYCP1, SYCP3, and centromeres in pachytene-like cells of Fbxo47−⁄− mice. Insets show magnified areas. Circled regions and red arrows highlight the unstable centromere pairing in Fbxo47−⁄− spermatocytes. k–m, k′–m′ demonstrate the expression of SYCP1, SYCP3, and centromeres in desynapsed pachytene-like cells of Fbxo47−⁄− mice. Insets provide enlarged views. Red arrowheads indicate desynapsed regions besides unpaired centromeres on chromosomes. The scale bar represents 5 µm. n presents the frequency of chromosomes with at least one unpaired centromere in Fbxo47+/+ and Fbxo47−⁄− mouse chromosome spreads (PD30) at the pachytene (like) stage. The inset shows the percentage of cells with varying numbers (1–2, 3–5, or over 5) of chromosomes with unpaired centromeres. “n” denotes the number of cells counted. Four to five mice were analyzed for each genotype. The data provided in this figure underscore the critical role of FBXO47 in ensuring proper centromere pairing and synaptic stability during meiosis.
For zygotene cells, we observed no significant differences in SC formation between Fbxo47+/+ (Fig. 3a, a′) and Fbxo47−⁄− (Fig. 3c, c′) spermatocytes. Partial SYCP1 (N-terminus) signals appeared in the midline of SYCP3 signals in both Fbxo47+/+ (Fig. 3b, b′, arrowhead) and Fbxo47−⁄− (Fig. 3d, d′, arrowhead) spermatocytes. Notably, in Fbxo47+/+ cells, the centromeric chromosomal ends were the last part to synapse and the 2 centromeres would pair at the last (Fig. 3b, b′, arrows), which was in agreement with previous studies that centromeric chromosomal ends are generally the last part to synapse in mice27,39. The 2 centromeric chromosomal ends in Fbxo47−⁄− spermatocytes were not synapsed either at the zygotene stage (Fig. 3d, d′, arrows).
For pachytene cells, Fbxo47+/+ cells exhibited complete synapsis, as indicated by the continuous SYCP3 and SYCP1 signals (Fig. 3e, e′). The SYCP1 signals were aligned along the chromosome axis and located in the midline of SYCP3 signals (Fig. 3f, f′, arrowheads), and centromeres were well-paired at the centromeric ends (Fig. 3f, f′, arrows). In contrast, in every observed pachytene-like Fbxo47−⁄− spermatocyte, we consistently observed failed centromere pairing (Fig. 3g, g′, circles and squares). This failed centromere pairing manifested as either two centromeres being separated with low SYCP1 signal (Fig. 3h, h′, red arrows), or widely separated homologous chromosomal ends with SYCP1 recruitment but without synapsis, suggesting a defective or unstable SC system in the absence of centromeric FBXO47 (Fig. 3i, i′, red arrows). Additionally, centromeric ends lacking SYCP1 signals were also noticeable (Fig. 3j, j′, red arrows). We believe that defective centromere pairing initiates with the corruption of SC at the centromere, subsequently affecting the entire chromosome arms. These data suggest that even if SYCP1 is loaded onto the homologous chromosomes, full synapsis cannot occur without properly paired centromeres.
Notably, in the observed desynapsed pachytene-like spermatocytes (Fig. 3k, k′, squares and red arrowheads), dissociation of chromosome axes occurred not only at the centromeric ends but also at non-centromeric ends or chromosome arms (Fig. 3l, l′, m, m′, red arrowheads). We hypothesize that the failed centromere pairing, and incomplete chromosome synapsis in Fbxo47−⁄− spermatocytes would trigger the dissociation of all the chromosomes in the cells.
We further quantified the incidence of unpaired centromeres in Fbxo47−⁄− pachytene spermatocytes. As summarized in Fig. 3n, 98.1% of Fbxo47−⁄− pachytene-like cells (with the remaining 1.9% of cells having at least one unpaired chromosome besides sex chromosomes) exhibited one or more pairs of unpaired centromeres (Fig. 3n, left). Among them, 19.1% of Fbxo47−⁄− pachytene cells had 1–2 pairs of unpaired centromeres, 57.9% had 3–5 pairs, and 22.9% had more than 5 pairs of unpaired centromeres (Fig. 3n, right). These data provide evidence suggesting that Fbxo47−⁄− spermatocytes exhibit unstable centromere pairing at the pachytene stage.
In summary, in Fbxo47−⁄− spermatocytes, the lack of FBXO47 leads to unstable centromere pairing, or some of the centromeres never pair, which prohibited their final synaptic pairing of chromosomes in Fbxo47−⁄− pachytene cells, resulting in premature dissociation of the chromosomes and cell death. In other words, FBXO47 plays a crucial role in centromere pairing, which is essential for achieving and maintaining full synaptic pairing as well as pachynema progression.
Fbxo47 −⁄− spermatocytes exhibit normal telomere-NE attachment in zygonema but failed telomere-NE tethering in pachynema
Since FBXO47 is located at centromeres, we also checked the other centromere association event, namely centromere clustering, which refers to centromeres cluster together in groups of two or more to facilitate proper centromere pairing, in Fbxo47−⁄− spermatocytes16,17. In mouse meiosis, centromere clustering concurrent to the bouquet formation40,41, and it remained unperturbed in Fbxo47−⁄− spermatocytes (For details, see Supplementary Information and Fig. S4).
During meiosis prophase I, telomere-NE attachment-related proteins (TRF142, KASH543 etc.) assist the force conduction from cytoskeleton to telomere to facilitate chromosome movements for homologous synapsis and recombination44,45. Our quest continued with an examination of telomere-NE attachment occurs during the bouquet stage, specifically at the zygotene stage46,47. Similar to their wild-type counterparts (Fig. 4a, a′ & Supplementary Fig. S5a, arrows), Fbxo47−⁄− spermatocytes exhibited normal telomere attachment to the NE during zygotene stage, as evidenced by TRF1 signals at the nuclear periphery (Fig. 4c, c′ & Supplementary Fig. S5c, arrows). However, at the pachytene-like stage, different from Fbxo47+/+ group (Fig. 4b, b′ & Supplementary Fig. S5b, arrows), TRF1 signals were not only diminished but also appeared detached from the NE (Fig. 4d, d′ & Supplementary Fig. S5d, arrows).
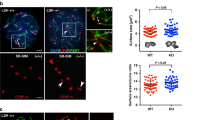
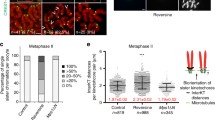
IF staining was employed to examine the tethering of telomeres to the NE in Fbxo47+/+ and Fbxo47−⁄− mice on chromosome spreads with an intact nuclear membrane. a, a′ In Fbxo47+/+ spermatocytes at the zygotene stage, TRF1 (green, arrows) and centromeres (red) are observed hanging on the NE (indicated by the circle), with SYCP3 (blue) highlighting the chromosome axes. b, b′ During the pachytene stage in Fbxo47+/+ spermatocytes, TRF1 (green) and centromeres (red) maintain their association with the NE (circle). c, c′ In Fbxo47−⁄− spermatocytes at the zygotene stage, TRF1 (green) and centromeres (red) are seen attaching to the NE (circle). d, d′ However, at the pachytene-like stage in Fbxo47−⁄− spermatocytes, the signal for TRF1 is weaker and appears detached from the NE. The scale bar represents 5 µm. e, f Representative section staining images of TRF1 (red, arrows) attached to NE at late-zygotene to pachytene stages of Fbxo47+/+ (e) and Fbxo47−⁄− (f) testes. DAPI staining (gray) delineate the nuclear periphery. g Quantification of TRF1 foci attached to NE at late-zygotene to pachytene stages. h The cell diameter of leptotene/zygotene and pachytene (like) cells from either Fbxo47+/+ or Fbxo47−⁄− mice. The data represent the mean ± SEM. “n” signifies the number of cells measured. i Immunoblotting analysis of TRF1 and KASH5 expression in leptotene/zygotene and pachytene cells isolated from the testes of Fbxo47+/+ or Fbxo47−⁄− mice at PD17-19. Three independent experiments were conducted, each time using a pool of at least two mice for each genotype. β-actin served as the loading control. The data provided in this figure emphasize the crucial role of FBXO47 in maintaining the proper tethering of telomeres to the NE at the pachytene stage.
Additionally, we conducted IF using antibodies against TRF1 and SYCP3 on paraffin-embedded testis sections from both genotypes, as depicted in Fig. 4e–e′′ and Fig. 4f–f′′. Given that both late-zygotene and pachytene cells exhibit highly condensed chromosomes (Fig. 4e, f, arrows), we randomly quantified the number of TRF1 foci in cells with highly condensed chromosomes and telomeres that were fully attached to the cell equator in both genotypes. After confocal scanning, 3 layers surrounding the cell equator were merged together for TRF1 foci number collection. DAPI staining was used to delineate the nuclear periphery. The proportion of cells with highly condensed chromosomes and telomeres fully attached to the NE was found to be 76.27% in wild-type mice and 50.72% in Fbxo47 KO mice (also noticed by Hua et al.37) (Supplementary Fig. S5e). Notably, the number of TRF1 foci attached to the NE in Fbxo47−⁄− cells was greater than in Fbxo47+/+ cells (Fig. 4g). This finding suggests that: (1) telomeres remain attached to the NE at least until the late-zygotene stage, and (2) the increased TRF1 signals on the NE in Fbxo47−⁄− cells are likely due to the previously observed unpaired centromeres, indicating that unpaired centromeres precede telomere detachment in Fbxo47-deficient spermatocytes.
Our findings suggest that centromeres and telomeres attach to the NE until late-zygotene stage in Fbxo47−⁄− spermatocytes, and then detach likely due to centromere pairing and synaptic instability.
To further elucidate this, we isolated leptotene/zygotene (diameter of 8–12 μm; purity: approximately 95%) and pachytene (diplotene cells are included due to limitation of method) or pachytene-like (diameter of 12-17 μm; purity: approximately 85%) spermatocytes using BSA gradient sedimentation48. The average diameter of leptotene/zygotene and pachytene cells from both Fbxo47-intact and deficient mice showed no discrepancy (Fig. 4h & Supplementary S5f–i), and this also contributes a reason why we called the defective Fbxo47−⁄− spermatocytes pachytene-like cells.
Western blotting (WB) analysis of telomeric proteins TRF1 and KASH5 revealed similar levels in leptotene/zygotene cells (Fig. 4i), facilitating normal centromere clustering (Supplementary Fig. S4). However, pachytene-like cells from Fbxo47-deficient testes exhibited significantly lower levels of these proteins compared to their wild-type counterparts (Fig. 4b, d, i & Supplementary Fig. S5b, d, j, k). These data suggest that telomere-NE detachment should be an indirect consequence of unstable centromeres and defective synapsis.
In summary, our research unveils that while centromere clustering remains unaffected in the absence of Fbxo47, telomere-NE detachment occurs later during the pachytene-like stage, most likely due to incomplete chromosome synapsis caused by unstable centromere pairing.
FBXO47 maintains centromere stability through sustaining centromeric SCF E3 ligase complex
To elucidate the impact of FBXO47 deficiency on centromere integrity and stability, we purified leptotene/zygotene and pachytene/pachytene-like spermatocytes from both Fbxo47+/+ and Fbxo47−⁄− testes. Through WB analysis, we assessed the expression levels of CENP-C, a crucial constitutive component of mammalian centromeres49. It exhibited normal expression levels in leptotene/zygotene cells, but a moderate reduction in Fbxo47−⁄− pachytene-like cells, compared to Fbxo47+/+ cells (Fig. 5a, a1, and Supplementary Fig. S8), suggesting a potential role of FBXO47 in centromere stabilization in mouse spermatocytes.
a, a1–3 WB analysis of CENP-C, SYCP1, and SKP1 expression in leptotene/zygotene cells and pachytene cells from both genotypes, as well as the quantitation of the results at pachytene (like) stage. b, b′ Location of SKP1 at synapsed regions of zygotene in Fbxo47+/+ spermatocytes. Arrows highlight the synapsed regions. c, c′ SKP1 location in pachytene spermatocytes of Fbxo47+/+ mice. Arrows indicate the synapsed regions. d, d′ Normal SKP1 signal observed in Fbxo47−⁄− zygotene cells. e, e′ Weakened SKP1 signal in pachytene-like cells of Fbxo47−⁄− mice. f, f′ SKP1 locates at centromeres on Fbxo47+/+ chromosomes. Arrows point to the centromeres, and insets show enlarged views. g, g′ Weak SKP1 signal in Fbxo47−⁄− pachytene-like cells. Scale bar, 5 µm. h Co-immunoprecipitation (Co-IP) study demonstrating that CENP-C1 can pull down overexpressed FBXO47 protein from HEK293T cell lysates. i SKP1 is also capable of pulling down overexpressed FBXO47 protein from HEK293T cell lysates. The arrow indicates Flag-SKP1. j Overexpression of FBXO47 leads to a decrease in the ubiquitination of SKP1 expressed in HEK293T cells. The arrow points to ubiquitinated Flag-SKP1. The Co-IP experiments were repeated three times with consistent outcomes. The data presented in this figure underscore the role of FBXO47 in maintaining centromere stability by ensuring the proper expression of SKP1 through the UPS pathway in spermatocytes.
We further investigated the interaction between FBXO47 and CENP-C by transfecting HEK293T cells with pcDNA3.1-MYC-Fbxo47 and pcDNA3.1-3xFLAG-Cenp-C1, followed by co-immunoprecipitation (Co-IP)-WB experiments. Our results indicated that MYC-FBXO47 could be co-immunoprecipitated with 3xFLAG-CENP-C1 (Fig. 5h and Supplementary Fig. S9a), implying a possible interaction between FBXO47 and CENP-C1 that may stabilize centromeres in mouse spermatocytes.
We also measured SYCP1 expression levels by WB in Fbxo47−⁄− spermatocytes. Normal expression was observed in leptotene/zygotene cells, consistent with normal SC formation at early prophase I (Fig. 3a, a′, b, b′), but reduced SYCP1 expression was noted in Fbxo47−⁄− pachytene-like cells (Fig. 5a, a2, and Supplementary Fig. S8), likely due to unstable synapsis and premature desynapsis.
Since F-box proteins determine the substrate specificity for ubiquitination in SCF E3 ligase complexes33, we hypothesized that FBXO47 might direct the functions of the SCF E3 ligase to centromeres, regulating specific substrates. Therefore, we measured the levels of SKP1 in Fbxo47−⁄− spermatocytes. The expression of SKP1 did not differ significantly in leptotene/zygotene cells but showed a moderate decrease in pachytene-like cells compared to Fbxo47+/+ spermatocytes (Fig. 5a, a3, and Supplementary Fig. S8).
To examine the localization of SKP1 on chromosomes, we conducted IF on chromosome spreads. In Fbxo47+/+ spermatocytes, SKP1 was observed at the synapsed parts of chromosomes during the zygotene stage (Fig. 5b, b′, arrows, & Supplementary Fig. S6a, a′) and displayed a robust signal along the lateral elements at the pachytene stage (Fig. 5c, c′, arrows, & Supplementary Fig. S6b–d, b′–d′), with distinct signals at the centromeric ends (Fig. 5f, f′, insets, arrows). Indeed, utilizing ImageJ software, we measured SPK1 signal intensity at chromosomal ends of 101 chromosomes from 7 cells (only those chromosomes with overlays were ignored), and found that the average ratio of SKP1 signal intensity at the centromeric end versus its counterpart on the same chromosome is 3.62 (Supplementary Fig. S6f). In Fbxo47−⁄− spermatocytes, SKP1 signal was present at synapsed regions of chromosome axes during the zygotene stage (Fig. 5d, d′, arrows), but were considerably weaker in pachytene-like cells (Fig. 5e, e′, red arrowheads, and Supplementary Fig. S6e). Moreover, the expression of SKP1 concentrated at the centromeric region was also dramatically reduced or disappeared in Fbxo47−⁄− spermatocytes (Fig. 5g, g′, red arrowheads), indicating degradation of SKP1 on centromeres and chromosomes without FBXO47 expression.
To confirm the interaction between SKP1 and FBXO47, which had been raised before37, we conducted transfection experiments in HEK293T cells by introducing pcDNA3.1-MYC-Fbxo47 and pcDNA3.1-3xFLAG-skp1. Immunoprecipitation of FLAG-SKP1 revealed the binding of MYC-FBXO47 to it (Fig. 5i, arrow, and Supplementary Fig. S9b), providing evidence for an interaction between FBXO47 and SKP1.
We then investigated the ubiquitination of SKP1 by co-transfecting HEK293T cells with pCMV-HA-Ubiquitin, pcDNA3.1-3xFLAG-Skp1, and with or without pcDNA3.1-MYC-Fbxo47. Our findings indicated that FBXO47 could reduce the ubiquitination of SKP1 (Fig. 5j, arrow, and Supplementary Fig. S10), suggesting that FBXO47 is involved in maintaining SKP1 levels by suppressing their ubiquitination.
Considering the depleted expression of SKP1 in Fbxo47−⁄− pachytene-like cells (Fig. 5a, a3) at centromeres (Fig. 5g) and chromosome arms (Fig. 5e), we propose that FBXO47 plays a crucial role in preserving SKP1 levels in spermatocytes.
Based on these comprehensive studies, we believed that the centromeric SFC complex is responsible for centromere pairing, ensuring the progression of spermatocytes through the pachytene stage.
FBXO47 probably regulates HORMAD1 to ensure proper chromosome synapsis
HORMAD1 is a component of unsynapsed chromosome axes, and it promotes DSB formation by recruiting pre-DSB complex (IHO1-REC114-MEI4) on unsynapsed regions to ensure the availability of sufficient DSBs for homology search50. It has been reported that the SCF complex prevents hyperactive DSB formation through proteasome-mediated degradation of HORMAD1 during early meiosis36, and HORMAD1 is required to direct DSB repair towards homologous chromosomes51. Notably, FBXO47 interact with HORMAD1 and promote its degradation through ubiquitin–proteasome system (UPS) in vitro36. Therefore, we conducted IF with HORMAD1 antibody on chromosome spreads.
We observed that in Fbxo47+/+ spermatocytes, HORMAD1 was located along the unsynapsed chromosomes at the leptotene stage and began to retreat at this stage (Supplementary Fig. S7a, a′, arrows). It rapidly disappeared from synapsed chromosomal parts at the zygotene stage (Supplementary Fig. S7b, b′, arrows). However, in Fbxo47−⁄− spermatocytes, no HORMAD1 retreat was observed at the leptotene stage (Supplementary Fig. S7c, c′), and unloading began at the zygotene stage (Supplementary Fig. S7d, d′, arrows). This was confirmed by the higher expression of HORMAD1 in leptotene/zygotene of Fbxo47−⁄− spermatocytes (Supplementary Figs. S7e and S11). The prolonged presence of HORMAD1 on chromosomes at leptotene/zygotene stages may contribute to the defective DSB repair in Fbxo47−⁄− spermatocytes.
At late-zygotene stage, HORMAD1 was exclusively localized to the unsynapsed centromeric ends of Fbxo47+/+ spermatocytes (Fig. 6a, a′, yellow arrows), and removed from the synapsed parts (Fig. 6a, a′, white arrows). In contrast, in Fbxo47−⁄− spermatocytes, although removal of HORMAD1 was observed in some of the chromosomes (Fig. 6d, d′, white arrows), unpaired centromeric ends without HORMAD1 assembly were often found (Fig. 6d, d′, yellow arrows), which also indicates defective centromeres as the cause of unpaired centromeres.
In Fbxo47+/+ mice, the location of HORMAD1 (green), SYCP3 (red), and centromeres (blue) is observed at different stages: a, a′ At late-zygotene stage, white arrows show the retreat of HORMAD1, while yellow arrows point to the remaining HORMAD1 expression at unpaired centromeric ends. b, b′ For pachytene stage, yellow arrows indicate the persistent HORMAD1 expression at the X-Y body. c, c′ In diplotene cells, yellow arrows demonstrate the recruitment of HORMAD1 expression on desynapsed regions. In Fbxo47−⁄− mice, the location of HORMAD1, SYCP3, and centromeres is altered: d, d′ Late-zygotene stage, where white arrows denote the retreat of HORMAD1, and yellow arrows indicate the lack of HORMAD1 expression at unpaired centromeres. e, e′ Pachytene-like stage, arrows point to the recruitment of HORMAD1 expression on chromosomes; yellow arrows indicate strong HORMAD1 at unpaired centromeric ends, while red arrows show slight HORMAD1 at paired parts. f, f′ Desynapsed pachytene-like stage, with yellow arrows showing the separation of non-centromeric ends and chromosome arms, and red arrows highlighting the weak expression of HORMAD1 on chromosome axes. The data from Fig. 6 underline the importance of HORMAD1 in participating proper chromosome pairing during meiosis.
HORMAD1 exclusively located to X-Y body in Fbxo47+/+ pachytene cells (Fig. 6b, b′, yellow arrows), then restored on desynapsed chromosomal regions at diplotene stage (Fig. 6c, c′, yellow arrows). In Fbxo47−⁄− spermatocytes at the pachytene-like stage, HORMAD1 was recovered on both synapsed (lower intensity, Fig. 6e, e′, red arrows) and unsynapsed (higher intensity, Fig. 6e, e′, yellow arrows) regions of all chromosomes. Notably, the unpaired centromeric ends exhibited higher intensity of HORMAD1 signals compared to synapsed regions (Fig. 6e, e′, yellow arrows versus red arrows). This recovery of HORMAD1 signals on the chromosomes could still be observed (Fig. 6f, f′, red arrows) in pachytene-like cells that just starting chromosome dissociation (Fig. 6f, f′, yellow arrows). We propose that the abnormal recruitment of HORMAD1 in Fbxo47−⁄− spermatocytes is related to the premature dissociation of chromosomes, ultimately resulting in cell death.
Discussion
Meiosis is a vital biological process responsible for the production of gametes through homologous recombination. During prophase I of meiosis, centromere interactions play a crucial role, including initial clustering at the onset of meiosis and subsequent pairing that coincides with the assembly of the SC52. Centromere pairing is an essential prerequisite for chromosome synapsis, a defining feature of meiosis47. This pairing ensures the proper segregation of bivalents, thereby preventing aneuploidy—a major cause of both infertility53 and cancer54.
In this study, we found that FBXO47 is located at centromeres. The knockout of Fbxo47 in mice resulted in developmental delay at the leptotene/zygotene stages due to defective DSB repair, a pachytene-like arrest with no crossovers, and premature degradation of spermatocytes. Increased RAD51 and RPA2 signals indicated disruption in the DSB repair process in Fbxo47-deficient spermatocytes, leading to a prolonged leptotene to zygotene stage. At pachytene stage, Fbxo47-deficient spermatocytes could not achieve complete centromere pairing even if the SC was formed due to defective centromeres without FBXO47, leading to a premature chromosome disassociation. Concurrently, there was a corruption of the telomere-NE attachment at the pachytene-like stage, and then cell death.
SKP1, the partner of F-box proteins in forming the SCF complex, is located on the LE and centromeric areas of chromosomes, as depicted in Fig. 5c, f, in addition to its cytoplasmic expression35. Guan et al. revealed that SKP1-deficient spermatocytes exhibit a “Y pachytene” phenotype characterized by unpaired centromeric ends at the pachytene stage, premature desynapsis, and precocious pachytene exit35. This finding aligns with our observations in Fbxo47−⁄− mice, where the centromeric ends failed to achieve full synapsis, leading to chromosome corruption.
The Co-IP study revealed that FBXO47 interacts with SKP1, and the introduction of FBXO47 to SKP1 resulted in a reduction of its ubiquitination, as shown in Fig. 5i, j. This suggests that the expression of FBXO47 is crucial for maintaining SKP1 expression. Based on our studies that FBXO47 deficiency resulted in the depletion of SKP1 on chromosomes, we believe that centromeric FBXO47 and SKP1 work cooperatively as the centromeric SCF complex to regulate centromere pairing and pachytene progression in mouse spermatocytes.
Besides, Guan et al. demonstrated that SKP1 plays crucial roles in maintaining DSB homeostasis. The SCF ubiquitin E3 ligase prevents hyperactive DSB formation through proteasome-mediated degradation of HORMAD1 during early meiosis36. Moreover, it has been reported that SKP1 deficiency induces HORMAD1 expression through the reduction of TRIP13 in mouse spermatocytes35.
Constitutive centromere protein CENP-C was reported to be involved in centromere pairing in Drosophila23. Degradation of CENP-C in pachytene-like cells may contribute to the failed centromere pairing in mouse spermatocytes. In fact, Guan et al. also observed severe down-regulation of CENP-C and BUB1 at late prophase I in SKP1 conditional knockout mice35. This information reinforces our thought of centromeric SCF complex functioning in meiosis progression.
Last but not least, we also analyzed HORMAD1 location, which is known to preferentially associate with unsynapsed chromosome axes throughout meiotic prophase51. In summary, in Fbxo47+/+ mice, the normal expression of FBXO47 and SKP1 from leptotene to early-zygotene stages facilitates the quick retreat of HORMAD1, ensuring normal DSB repair. By the late-zygotene stage, with the regression of HORMAD1, centromeric ends pair to complete chromosome synapsis, leading to the attainment of a pachytene stage where all chromosomes are well synapsed (Fig. 7).
Fbxo47+/+ Mice: By the late-zygotene stage, normal expression of FBXO47 and SKP1 ensures that the centromeric ends pair effectively, completing chromosome synapsis. This results in the attainment of the pachytene stage where all chromosomes are fully synapsed, with centromeres securely anchored at the NE. Fbxo47−/− Mice: In the absence of FBXO47 expression, some centromeric ends fail to pair even after the retraction of HORMAD1 at the late-zygotene stage. Subsequently, at the pachytene-like stage, abnormal recruitment of HORMAD1 back to the chromosome axes occurs (concurrently with SKP1 depletion). This leads to chromosomal instability and telomere detachment from the NE, ultimately culminating in cell death.
However, in Fbxo47−⁄− spermatocytes, the absence of FBXO47 expression during leptotene to early-zygotene stages results in a delayed HORMAD1 retreat, causing abnormal DSB homeostasis. In the late-zygotene to pachytene stages, some of the centromeric ends fail to pair even after HORMAD1 retreat, indicating defective centromeres as the cause of incomplete centromere pairing. Meanwhile, a recruitment of retreated HORMAD1 back to the chromosome axes was noticed at pachytene-like stage, which might be a response to the depletion to the SKP1 (Fig. 7).
In conclusion, we propose that centromeric SFC is involved in stabilizing centromere pairing in mouse spermatocytes. In Fbxo47−⁄− spermatocytes, the unstable centromere pairing leads to incomplete chromosome synapsis, triggers the collapse of the SC, chromosome dissociation, corruption of the telomere-NE attachment, and ultimately cell death, probably through the regulation of HORMAD1 (Fig. 7).
Recently, SKP1-related proteins (SKRs) have been confirmed as components of the SC in Caenorhabditis elegans. SKRs could repurpose their SCF-forming interfaces to dimerize and interact with SC proteins, driving synapsis independent of SCF activity55.
We believe that the desynapsed pachytene-like cells in this study are in line with diplotene-like cells (premature disassembly of chromosomes) observed by Tanno et al., because our TOPBP1 staining results were strikingly similar to their BRCA1 staining patterns38. Besides, there is a plausible role for centromeric SCF in stabilizing telomere-NE attachment proteins through the UPS. Our additional findings suggest that in HEK293T cells, FBXO47 could reduce the ubiquitination of SPDYA and CDK2, both of which are implicated in telomere-NE attachment56. Hua’s work also demonstrated that FBXO47 could stabilize shelterin protein TRF2 by reducing its ubiquitination in the same cell line37. It is likely that the depletion of SKP1 (SCF complex), along with the recruitment of HORMAD1 at the pachytene-like stage, could simultaneously induce both desynapsis of chromosomes and the detachment of telomeres from NE in Fbxo47−⁄− spermatocytes.
Although the location and quality of SYCP1 appear normal in zygotene cells in Fbxo47−⁄− mice, we hesitate to claim that SC formation is entirely correct due to the disturbed DSB homeostasis.
The molecular mechanisms that regulate centromere pairing during meiosis and their impact on this process are still not fully understood. For example, the exact relationship between centromere pairing and other cellular events, such as DNA repair, requires further investigation. Additionally, the roles of the SCF complex present at centromeres offer promising directions for future studies. Our subsequent research efforts will concentrate on uncovering the function of FBXO47 during prometaphase of spermatocytes.
Methods
Animals
C57BL/6 mice were housed under specific pathogen-free (SPF) conditions at 22 °C with a 12-h light/dark cycle, and they were provided with ample food and water. Fbxo47 knockout mice were generated by Cyagen Biosciences (Suzhou, China) and bred at the animal unit of Shenzhen PKU-HKUST Medical Center. The CRISPR/Cas9-mediated genome-editing system was employed to delete a genomic DNA fragment spanning exon 2 to exon 11, using gRNA1 (matching the forward strand of the gene): TAGGATGCCAAGAGGTATTTAGG and gRNA2 (matching the reverse strand of the gene): TATAGCCACTTTGACGTAAGTGG.
Founder mice were genotyped through PCR and DNA sequencing to confirm the deletion. A selected F0 founder mouse was bred with wild-type mice to obtain F1 heterozygous mutants (Fbxo47+/−). The Fbxo47+/− mice were then backcrossed with C57BL/6 wild-type mice to eliminate potential off-target mutations. Subsequently, heterozygous targeted mice were intercrossed to generate both wild-type (Fbxo47+/+) and homozygous Fbxo47-deficient (Fbxo47−⁄−) mice. Genotyping was performed using two pairs of PCR primers: F1: 5′-CATGCAGTTACATGTTTGATTTCTT-3′, R1:5′-CCTTCCATATTAAGTGTGTTTCTGCTATTG-3′, and R2: 5′-AAAGAGAACACGATAACAGTTGGG-3′ to select target mice. All experimental protocols were approved by the regional ethics committee of the University of Hong Kong-Shenzhen Hospital.
Antibodies
The primary antibodies utilized for IF were as follows: rabbit anti-SYCP3 (1:200 dilution; ab15093, Abcam), mouse anti-SYCP3 (1:200 dilution; ab97672, Abcam), rabbit anti-FBXO47 (1:100 dilution, Biorbyt, orb183644), mouse anti-phospho-histone H2AX (pSer139) (1:200 dilution; 05-636, Millipore), mouse- anti TRF1 (1:200 dilution; ab10579, Abcam), rat anti-Tubulin (1:200 dilution; ab6160, Abcam), human anti-centromere/kinetochore (1:200 dilution, Antibodies Incorporated), rabbit anti-SYCP1 (1:200 dilution; A12139, abclonal), rabbit anti-SKP1 (1:50 dilution; ab76502, Abcam), and rabbit anti-HORMAD1 (1:400 dilution; 13917-1-AP, Proteintech). Secondary antibodies for IF were as follows (All in 1:200 dilution, Abcam if not otherwise stated): Alexa Fluor goat anti-human 594 (ab96913), goat anti-rabbit 488 (ab96899)/594 (ab150084)/647 (ab150079), and goat anti-mouse 488 (ab150117)/594 (ab96881)/647 (ab150115), goat anti-rat 647 (ab150167), and goat anti-human 647 (A21445, Thermo Fisher Scientific).
Primary antibodies for WB included: β-actin (mouse, 1:5000 dilution; 4970, Cell Signaling Technology), TRF1 (1:1000 dilution; see above), CENP-C (rabbit, 1:500 dilution; NBP2-75438, Novusbio), SKP1 (1:1000 dilution; see above), SYCP1 (1:1000 dilution; see above), KASH5 (rabbit, 1:500 dilution, home-made). Secondary antibodies for WB were as follows: Goat anti-mouse HRP (1:10,000 dilution; ab6789, Abcam), Goat anti rabbit HRP (1:5000 dilution; AS014, ABclonal).
Immunofluorescent staining and imaging
The testes were prepared for chromosome spreading onto glass slides using a hypotonic solution (30 mM Tris-Cl pH 7.5, 50 mM sucrose, 17 mM sodium citrate, 5 mM EDTA) with a treatment duration of 30 min. Subsequently, harsh spreading was achieved with a 100 mM sucrose treatment for 5 min, while mild spreading utilized a 200 mM sucrose treatment for the same duration.
For IF studies, the samples underwent a 10-min wash with PBST (0.1% Triton X-100 in PBS) and two additional washes with PBS. The slides were then blocked with 5% bovine serum albumin (BSA) diluted in PBS for 1 h, followed by overnight incubation with the specified antibodies at 4 °C. Following PBS washing, a secondary antibody was applied for 1 h at room temperature.
Finally, the slides were washed in PBS and mounted with a DAPI-containing mounting medium (VECTASHIELD, H-1200). Visualization was performed using an LSM900 confocal microscope (Carl Zeiss AG) driven by ZEN software (Version 3.2.0.0000). Notably, For the FBXO47 antibody, the observation should be performed as soon as possible, as the signals would be extinguished.
Meiosis synchronization
To synchronize meiosis, male offspring aged 2 or 3 days (weighing over 1.5 g) were subcutaneously injected over the shoulders with N, N′-Octamethylenebis (2, 2-dichloroacetamide), also known as WIN18,446 (Santa Cruz Biotechnology, sc-295819), for 7 days, as outlined in a previous study57, to inhibit spermatogenesis. Subsequently, a single dose of retinoic acid (RA, Sigma Aldrich, R2625-50MG) was injected on the next day to synchronously restart spermatogenesis. The testes of Fbxo47−⁄− or Fbxo47+⁄− mice were collected at different days after RA injection to perform chromosome spreading, followed by staining with SYCP3, SYCP1, and centromere antibodies.
For the preparation and injection of WIN18,446, 5 mg of WIN18,446 was dissolved in 25 µL of DMSO. This solution was mixed with 475 µL corn oil (MedChemExpress, HY-Y1888) to make a 0.5 mL solution, and then shaken immediately to create a cloudy suspension. After aliquoting into small volumes, the solution was stored at −80 °C. Before each use, it was loaded into a 50 µL syringe with a 30G needle (Yeso med, China) after shaking to avoid clogging. Subcutaneous injections of 10 µL/g of body weight could be administered at any time of day, except for the first dose, which should be performed around noon. It was acceptable to skip a single dose, as long as injections were no more than 48 h apart.
For the preparation and injection of RA, a 100 mM RA stock (e.g., 50 mg RA + 1.664 mL DMSO) was first created. Subsequently, 1.917 mL of corn oil was mixed with 83 µL of RA stock and stored at −80 °C after aliquoting. After shaking well, 10 µL of RA suspension per gram of body weight was injected subcutaneously. It is important to note that the body weight of the mouse should be over 5 g for RA injection.
Purification of leptotene/zygotene and pachytene cells
The purification of distinct spermatocytes followed a previously outlined protocol58. Briefly, 2–3 postnatal day (PD) 17–19 mice were sacrificed for testes. After decapsulation, the seminiferous tubules were cut into pieces and incubated in 5 mL PBS containing 2 mg/mL collagenase (Gibco, 17100017) and 1 mg/mL DNase I (Applichem, A3778) at 37 °C for 5 min with gentle shaking. Subsequent centrifugation at 120 × g for 2 min was performed, and the pellet was re-suspended in 5 mL 0.25% Trypsin (Gibco, 15050057) with 1 mg/mL DNase I at 37 °C for 5 min with gentle shaking. After centrifugation at 300 × g for 5 min, cells were re-suspended in 20 mL DMEM (Gibco, 12100046) containing 0.5% BSA, followed by filtration through a 40 µm Nylon Cell Strainer (BD Falcon).
Subsequently, the recovered cells were re-suspended again in 20 mL DMEM containing 0.5% BSA and loaded into a cell separation apparatus (BOMEX Corporate) with a 2–4% BSA gradient in 600 mL DMEM. The cells were collected from the bottom of the separation apparatus at a rate of 10 mL/min into tubes (10 mL/tube) after 3 h of sedimentation. The cell type and purity in each fraction were assessed based on their diameters and morphological characteristics under a light microscope.
Western blotting
Purified cells from the mouse testes, as mentioned earlier, were collected in 1.5 mL tubes, lysed in a lysis buffer (50 mM Tris-HCl, pH 8.0, 120 mM NaCl, 1 mM EDTA, 6 mM EGTA, 1% NP-40, 1mM DTT, 10 mM NaF, 0.25 mM Na3VO4, 50 mM β-glycerolphosphate) supplemented with Complete Protease Inhibitor (Roche, 4693159001) on ice for 20 min, and subsequently centrifuged at 13,000 rpm for 20 min at 4 °C. The resulting supernatants were retained for western blotting studies, and 20 μg of proteins were used for each sample. β-actin served as the loading control, and a color-coded protein marker ranging from 10 to 250 kDa (Cell Signaling, #74124) was applied.
A 4–12% precast polyacrylamide gel (BIO-RAD, 4561094) was utilized and followed the provided instructions. Immunoreactive bands were detected and analyzed using the ChemiDoc MP Imaging System and Image Lab Software (Bio-Rad, USA). The experiments were conducted with a minimum of three repetitions to ensure consistency and reliability of results.
Quantitation of the WB results was performed by densitometric scanning using ImageJ, and the data were transformed as a ratio of the expression of respective targets in Fbxo47−⁄− samples over the Fbxo47+/+ samples after normalized with β-actin.
Paraffin embedding
Testes were dissected immediately after euthanasia and fixed in 4% PFA solution at 4 °C for around 20 h. After fixation, samples were washed with 70% ethanol, subjected to dehydration and then embedded in paraffin. Sections of 5 μm were prepared and mounted on glass slides. After deparaffinization, IF was performed as mentioned above.
Transfection and immunopreciptation
HEK293T cells were transfected with 1.5 μg of pcDNA3.1-3xFLAG-Cenp-C1 or pcDNA3.1-3xFLAG-skp1 along with 1.5 μg of pcDNA3.1-MYC-Fbxo47 using lipofectamine 3000. Cells were then harvested 24–48 h post-transfection, and protein extraction was carried out with lysis buffer (50 mM Tris-HCl, pH 8.0, 120 mM NaCl, 1 mM EDTA, 6 mM EGTA, 1% NP-40, 1 mM DTT, 10 mM NaF, 0.25 mM Na3VO4, 50 mM β-glycerolphosphate) supplemented with Complete Protease Inhibitor (Roche, 4693159001).
Twenty micrograms of protein lysate were reserved and used as input, while around 800–1000 μg of protein lysate were incubated with 5 μg of FLAG-tag antibody (Mouse, proteintech, 66008-3-Ig) overnight in rotation at 4 °C. The immuno-complexes were isolated using mixed protein A/G Sepharose beads (Roche, 647 11719416001 and 11719408001) for 1 h at 4 °C. After washing with a buffer containing 0.25 M NaCl for five times, the beads were denatured and centrifuged before loading the supernatants onto SDS-PAGE gels. After transferred onto the PVDF membrane, different protein tags were detected with the indicated antibodies.
Ubiquitination assay
For the ubiquitination assay, a co-transfection was performed with 1.3 μg each of pcDNA3.1-3xFLAG-skp1, pcDNA3.1-MYC-Fbxo47, and pCMV-HA-Ubiquitin into HEK293T cells. Cells were collected 24 h after transfection using the lysis buffer mentioned earlier for immunoprecipitation. Twenty micrograms of protein lysate were reserved as input, while approximately 1 mg of protein lysate was incubated with 5 μg of FLAG-tag antibody (Mouse, proteintech, 66008-3-Ig) overnight with rotation at 4 °C. Immuno-complexes were isolated using mixed protein A/G Sepharose beads for 1 h at 4 °C. Following washing, denaturing, and centrifuging, the supernatants were loaded onto SDS-PAGE gels.
The detection of ubiquitin bound to SKP1 was performed using the anti-HA antibody (Rabbit, Abcam, ab9110). Subsequently, anti-FLAG (Rabbit, proteintech, 20543-1-AP) or anti-MYC (Rabbit, proteintech,16286-1-AP) antibodies were used to detect SKP1 or FBXO47 after the immunoblotting process.
Statistical Analysis
The numerical data are expressed as means ± SEM. The statistical significance of the differences between the mean values for the different genotypes was measured by Student’s t test with an unpaired, two-tailed distribution. The data were considered significant when the P-value was less than 0.05, with * denoting P < 0.05, ** for P < 0.01, *** for P < 0.001, and **** for P < 0.0001.
Data availability
All relevant data are within the manuscript and its supplemental items, including original gel images in the Supplementary Information file and numerical data in the Supplementary Data file. All other data are available from the corresponding author on reasonable request.
References
Bouuaert, C. C., Tischfield, S. E., Pu, S., Mimitou, E. P. & Keeney, S. Structural and functional characterization of the Spo11 core complex. Nat. Struct. Mol. Biol. 28, 1–11 (2021).
Bolcun-Filas, E. & Handel, M. A. Meiosis: the chromosomal foundation of reproduction. Biol. Reprod. 99, 112–126 (2018).
Hinch, A. G., Becker, P. W., Li, T., Moralli, D. & Donnelly, P. The configuration of RPA, RAD51, and DMC1 binding in meiosis reveals the nature of critical recombination intermediates. Mol. Cell 79, 689–701 (2020).
Lake, C. M. & Hawley, R. S. Synaptonemal complex. Curr. Biol. 31, R225–r227 (2021).
Takeo, S. & Hawley, R. S. Rumors of its disassembly have been greatly exaggerated: the secret life of the synaptonemal complex at the centromeres. PLoS Genet. 8, e1002807 (2012).
Dunce, J. M. et al. Structural basis of meiotic chromosome synapsis through SYCP1 self-assembly. Nat. Struct. Mol. Biol. 25, 557–569 (2018).
Zhang, F. G., Zhang, R. R. & Gao, J. M. The organization, regulation, and biological functions of the synaptonemal complex. Asian J. Androl. 23, 580–589 (2021).
Baudat, F. & Massy, B. d. Regulating double-stranded DNA break repair towards crossover or non-crossover during mammalian meiosis. Chromosome Res. 15, 565–577 (2007).
Bloomfield, G. Atypical ploidy cycles, Spo11, and the evolution of meiosis. Semin. Cell Dev. Biol. 54, 158–164 (2016).
Talbert, P. B. & Henikoff, S. What makes a centromere? Exp. Cell Res. 389, 111895 (2020).
Mckinley, K. L. & Cheeseman, I. M. The molecular basis for centromere identity and function. Nat. Rev. Mol. Cell Biol. 17, 16–29 (2015).
Kipling, D., Wilson, H. E., Mitchell, A. R. & Taylor, B. Mouse centromere mapping using oligonucleotide probes that detect variants of the minor satellite. Chromosoma 103, 46–55 (1994).
Burkin, D. J., Jones, C., Burkin, H. R., McGrew, J. A. & Broad, T. E. Sheep CENPB and CENPC genes show a high level of sequence similarity and conserved synteny with their human homologs. Cytogenet Cell Genet. 74, 86–89 (1996).
McKay, S., Thomson, E. & Cooke, H. Sequence homologies and linkage group conservation of the human and mouse cenpc genes. Genomics 22, 36–40 (1994).
Falk, S. J. et al. CENP-C reshapes and stabilizes CENP-a nucleosomes at the centromere. Chromosomes 348, 699–703 (2015).
Kurdzo, E. L. & Dawson, D. S. Centromere pairing – tethering partner chromosomes in meiosis I. FEBS J. 282, 2445–2457 (2015).
Ines, O. D. & White, C. I. Centromere associations in meiotic chromosome pairing. Annu Rev. Genet. 49, 95–114 (2015).
Stewart, M. N. & Dawson, D. S. Changing partners: moving from non-homologous to homologous centromere pairing in meiosis. Trends Genet. 24, 564–573 (2008).
Kemp, B., Boumil, R. M., Stewart, M. N. & Dawson, D. S. A role for centromere pairing in meiotic chromosome segregation. Genes Dev. 18, 1946–1951 (2004).
Zhang, J. & Han, F. Centromere pairing precedes meiotic chromosome pairing in plants. Sci. China Life Sci. 60, 1197–1202 (2017).
Lakhani, A. A., Thompson, S. L. & Sheltzer, J. M. Aneuploidy in human cancer: new tools and perspectives. Trends Genet. 39, 968–980 (2023).
Cimini, D. & Degrassi, F. Aneuploidy: a matter of bad connections. Trends Cell Biol. 15, 442–451 (2005).
Unhavaithaya, Y. & Orr-Weaver, T. L. Centromere proteins CENP-C and CAL1 functionally interact in meiosis for centromere clustering, pairing, and chromosome segregation. Proc. Natl Acad. Sci. USA 110, 19878–19883 (2013).
Hatkevich, T., Boudreau, V., Rubin, T., Maddox, P. S. & Sekelsky, J. Centromeric SMC1 promotes centromer clustering and stabilizes meiotic homolog pairing. PLoS Genet. 15, e1008412 (2019).
Cheng, E. C., Hsieh, C. L., Liu, N., Wang, J. & Lin, H. The essential function of SETDB1 in homologous chromosome pairing and synapsis during meiosis. Cell Rep. 34, 108575 (2021).
Zhang, J. et al. The cohesin complex subunit ZmSMC3 participates in meiotic centromere pairing in maize. Plant Cell 32, 1323–1336 (2020).
Bisig, C. G. et al. Synaptonemal complex components persist at centromeres and are required for homologous centromere pairing in mouse spermatocytes. PLoS Genet. 8, e1002701 (2012).
Luciana, Pd. A. et al. Shugoshin protects centromere pairing and promotes segregation of nonexchange partner chromosomes in meiosis. Proc. Natl Acad. Sci. USA 116, 9417–9422 (2019).
Wang, Z., Liu, P., Inuzuka, H. & Wei, W. Roles of F-box proteins in cancer. Nat. Rev. Cancer 14, 233–247 (2014).
Bose, R., Manku, G., Culty, M. & Wing, S. S. Ubiquitin-proteasome system in spermatogenesis. Adv. Exp. Med. Biol. 759, 181–213 (2014).
Hindley, C. J., McDowell, G. S., Wise, H. & Philpott, A. Regulation of cell fate determination by Skp1-Cullin1-F-box (SCF) E3 ubiquitin ligases. Int J. Dev. Biol. 55, 249–260 (2011).
Richburg, J. H., Myers, J. L. & Bratton, S. B. The role of E3 ligases in the ubiquitin-dependent regulation of spermatogenesis. Semin. Cell Dev. Biol. 30, 27–35 (2014).
Cardozo, T. & Pagano, M. The SCF ubiquitin ligase: insights into a molecular machine. Nat. Rev. Mol. Cell Biol. 5, 739–751 (2004).
Barbosa, P., Zhaunova, L., Debilio, S., Anella, V. S. C. & Ohkura, H. SCF-Fbxo42 promotes synaptonemal complex assembly by downregulating PP2A-B56. J. Cell Biol. 220, e202009167 (2021).
Guan, Y., Leu, N. A., Ma, J., Chmátal, L. & Wang, P. J. SKP1 drives the prophase I to metaphase I transition during male meiosis. Sci. Adv. 6, eaaz2129 (2020).
Guan, Y. et al. SCF ubiquitin E3 ligase regulates DNA double-strand breaks in early meiotic recombination. Nucleic Acids Res. 50, 5129–5144 (2022).
Hua, R., Wei, H., Liu, C., Zhang, Y. & Liu, M. FBXO47 regulates telomere-inner nuclear envelope integration by stabilizing TRF2 during meiosis. Nucleic Acids Res. 47, 11755–11770 (2019).
Tanno, N. et al. FBXO47 is essential for preventing the synaptonemal complex from premature disassembly in mouse male meiosis. iScience 25, 104008 (2022).
Qiao, H., Chen, J. K., Reynolds, A., Höög, C. & Hunter, N. Interplay between synaptonemal complex, homologous recombination, and centromeres during mammalian meiosis. PLoS Genet. 8, e1002790 (2012).
Zhang, F. et al. The F-box protein ZYGO1 mediates bouquet formation to promote homologous pairing, synapsis, and recombination in rice meiosis. Plant Cell 29, 2597–2609 (2017).
Scherthan, H. et al. Centromere and telomere movements during early meiotic prophase of mouse and man are associated with the onset of chromosome pairing. J. Cell Biol. 134, 1109–1125 (1996).
Wang, L., Tu, Z., Chao, L., Liu, H. & Wei, L. Dual roles of TRF1 in tethering telomeres to the nuclear envelope and protecting them from fusion during meiosis. Cell Death Differ. 25, 1174–1188 (2018).
Morimoto, A. et al. A conserved KASH domain protein associates with telomeres, SUN1, and dynactin during mammalian meiosis. J. Cell Biol. 198, 165–172 (2012).
Lee, C. Y. et al. Mechanism and regulation of rapid telomere prophase movements in mouse meiotic chromosomes. Cell Rep. 11, 551–563 (2015).
Link et al. Structural and functional adaptations of the mammalian nuclear envelope to meet the meiotic requirements. Nucleus 6, 93–101 (2015).
Palmer, N., Talib, S. & Kaldis, P. Diverse roles for CDK-associated activity during spermatogenesis. FEBS Lett. 593, 2925–2949 (2019).
Mytlis, A. et al. Control of meiotic chromosomal bouquet and germ cell morphogenesis by the zygotene cilium. Science 376, eabh3104 (2022).
Bellvé, A. R., Cavicchia, J. C., Millette, C. F., O“Brien, D. A. & Dym, B. M. Spermatogenic cells of the prepuberal mouse. J. Cell Biol. 74, 68–85 (1977).
Saffery, R., Earle, E., Irvine, D. V., Kalitsis, P. & Choo, K. Conservation of centromere proteins in vertebrates. Chromosome Res. 7, 261–265 (1999).
Stanzione, M. et al. Meiotic DNA break formation requires the unsynapsed chromosome axis-binding protein IHO1 (CCDC36) in mice. Nat. Cell Biol. 18, 1208–1220 (2016).
Wojtasz, L. et al. Mouse HORMAD1 and HORMAD2, two conserved meiotic chromosomal proteins, are depleted from synapsed chromosome axes with the help of TRIP13 AAA-ATPase. PLoS Genet. 5, e1000702 (2009).
Obeso, D., Pezza, R. J. & Dawson, D. Couples, pairs, and clusters: mechanisms and implications of centromere associations in meiosis. Chromosoma 123, 43–55 (2014).
Charalambous, C., Webster, A. & Schuh, M. Aneuploidy in mammalian oocytes and the impact of maternal ageing. Nat. Rev. Mol. Cell Biol. 24, 27–44 (2023).
Ben-David, U. & Amon, A. Context is everything: aneuploidy in cancer. Nat. Rev. Genet. 21, 44–62 (2020).
Blundon, J. M. et al. Skp1 proteins are structural components of the synaptonemal complex in C. elegans. Sci. Adv. 10, eadl4876 (2024).
Tu, Z. et al. Speedy A–Cdk2 binding mediates initial telomere–nuclear envelope attachment during meiotic prophase I independent of Cdk2 activation. Proc. Natl Acad. Sci. USA 114, 592–597 (2017).
Romer, K. A., De Rooij, D. G., Kojima, M. L. & Page, D. C. Isolating mitotic and meiotic germ cells from male mice by developmental synchronization, staging, and sorting. Dev. Biol. 443, 19–34 (2019).
Li, M., Zheng, J., Li, G., Lin, Z. & Liu, K. The male germline-specific protein MAPS is indispensable for pachynema progression and fertility. Proc. Natl Acad. Sci. USA 118, e2025421118 (2021).
Acknowledgements
We sincerely acknowledge the financial support by Guangdong Basic and Applied Basic Research Foundation 2024A1515010059 and Shenzhen Science and Technology Program JCYJ 20220818101218040.
Author information
Authors and Affiliations
Contributions
A.N. Ma and J.V. Zhang conceived the initial study and designed the research. A.N. Ma wrote the paper; A.N. Ma, Y.L. Yang, L.B. Cao, and L.J. Chen performed the experiments and analyzed the data.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Damian Dudka and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Patrick Meraldi and Mengtan Xing.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ma, A., Yang, Y., Cao, L. et al. FBXO47 regulates centromere pairing as key component of centromeric SCF E3 ligase in mouse spermatocytes. Commun Biol 7, 1099 (2024). https://doi.org/10.1038/s42003-024-06782-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-024-06782-6
- Springer Nature Limited