Abstract
In the pharmaceutical industry, there is an abundance of regulatory documents used to understand the current regulatory landscape and proactively make project decisions. Due to the size of these documents, it is helpful for project teams to have informative summaries. We propose a novel solution, MedicoVerse, to summarize such documents using advanced machine learning techniques. MedicoVerse uses a multi-stage approach, combining word embeddings using the SapBERT model on regulatory documents. These embeddings are put through a critical hierarchical agglomerative clustering step, and the clusters are organized through a custom data structure. Each cluster is summarized using the bart-large-cnn-samsum model, and each summary is merged to create a comprehensive summary of the original document. We compare MedicoVerse results with established models T5, Google Pegasus, Facebook BART, and large language models such as Mixtral 8\(\times \)7b instruct, GPT 3.5, and Llama-2-70b by introducing a scoring system that considers four factors: ROUGE score, BERTScore, business entities and the Flesch Reading Ease. Our results show that MedicoVerse outperforms the compared models, thus producing informative summaries of large regulatory documents.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Introduction
The pharmaceutical industry has witnessed a remarkable surge in published literature, encompassing a diverse range of topics within the life sciences. This valuable repository of knowledge includes journals, academic publications, and research papers in fields such as medicine, genetics, epidemiology, and more. Pharmaceutical science literature serves as a rich source of information, providing comprehensive insights into the latest advancements and discoveries in various life sciences domains. Researchers, medical professionals, and others rely on this extensive corpus of data to gain knowledge, enhance patient care, drug development, influence public health policies, etc., thereby playing a vital role in advancing scientific understanding and promoting evidence-based decision-making. Despite the invaluable knowledge in biomedical science literature, researchers often face significant challenges in staying current and publications pose hurdles in efficiently analyzing vast volumes of text. Given this scenario, there is a need for an effective approach to streamline the information extraction process and locate the key contents of the text. Text summarization which emerges as a promising solution in natural language processing (NLP) addresses the challenges posed by the extensive biomedical science literature.
Text summarization1 is the process of condensing a large amount of text into a concise, informative summary without compromising the underlying meaning of the original text. There are two primary approaches to text summarization, extractive summarization2,3 and abstractive summarization4. In extractive summarization, key phrases and words are extracted from the raw text and then merged to generate a summary. Abstractive summarization works by creating new sentences that resonate with the meaning of the original text. This method involves a two-step approach of first selecting important phrases and then paraphrasing them. Both techniques are considered to be supervised machine learning problems.
Research has demonstrated that neural network based abstractive summarization have been shown to achieve state-of-the art performance5,6. These methods often employ encoder-decoder architectures7, which are typical in sequence-to-sequence models. The addition of the attention mechanism in Transformers4,8,9,10,11,12 has significantly enhanced these models. Currently, abstractive text summarization is commonly achieved through Transformer-based models and their variations, as they offer reduced computational requirements and enable concurrent training13,14,15,16,17. Early Transformer-based models for summarizing text were assessed using ROUGE scores, whereas newer models utilize BERTScore18 for tasks like text simplification19 and correcting grammatical errors20. Lately, there has been a shift towards using reinforcement learning to optimize rewards based on various evaluation criteria21,22, including the ROUGE-L score23. The BERTSUM model14, which utilizes the BERT model24 for the Transformer encoder-decoder, has attained the highest performance on various datasets. Motivated by these developments, we have created an innovative method that merges unsupervised hierarchical agglomerative clustering25,26 with Transformer models for the purpose of abstractive text summarization.
Our work focuses on providing an abstractive text summarization tool designed to extract key insights from medical and scientific research papers. In this paper, we introduce our solution MedicoVerse, a novel biomedical text summarizer developed to address the pressing need for effective information synthesis in the biomedical domain. Our approach harnesses NLP techniques and models to construct abstractive summaries that encapsulate the core knowledge embedded within the original text. Our approach has a multi-stage methodology, encompassing techniques such as word embeddings27,28, hierarchical agglomerative clustering29,30, abstractive summarization, and disease-chemical keyword annotation. We also introduce a novel scoring technique to evaluate the effectiveness of our approach, integrating four key metrics: ROUGE31, BERTScore18, Business Entities and the Flesch Reading Ease32. This scoring technique ensures a comprehensive assessment on the quality of the summaries.
Results
Dataset
Our main dataset is in the domain of regulatory requirements, consisting of 227 parts of texts from publically available 38 regulatory documents33, and parts of texts from 23 PubMed documents34. These documents are primarily used for understanding the current regulatory landscape and proactively making project decisions based on recent regulatory activity. For example, this may be new U.S. Food and Drug Administration guidelines to include data from studies. Due to the size of these regulatory documents, it is helpful for regulatory teams to have their informative summaries. These informative summaries are also valuable for biologics license application reviewers because those reviewers can in turn provide better inputs if they know about the current regulatory trends. One benefit of informative summaries of regulatory documents is that they can reduce project timelines by decreasing the number of regulatory queries received from regulators, as each query is potentially a delay to the final approval date. There is also a cost-saving benefit as it could mean less work responding to regulators. The 227 parts of texts from the 38 regulatory documents is preprocessed and fed through the MedicoVerse system architecture. Finally, the summaries are evaluated and compared along with results from other models.
System architecture
Our approach addresses the challenge of condensing extensive regulatory documents into concise and informative abstractive summaries. The proposed summarizer, MedicoVerse, uses advanced NLP models, hierarchical clustering, summarization techniques, and disease-chemical keyword annotation. A comprehensive breakdown of the MedicoVerse architecture is shown in Fig. 1.
Data collection and pre-processing
Our main dataset consists of 227 parts of texts from publically available 38 regulatory documents, and parts of texts from 23 PubMed documents. All documents are preprocessed through the elimination of unnecessary characters and newlines, text normalization, as well as ensuring a consistent format. Moreover, the text is segmented into smaller units using a customized delimiter. This segmentation enhances the feasibility of subsequent analysis and processing.
sapBERT word embeddings
To capture the intricacies of the terminology and context, the preprocessed data is inputted into the cambridgeltl/SapBERT-from-PubMedBERT-fulltext24,35 model to generate BERT embeddings. Instead of providing a fixed embedding for each word, BERT produces contextualized word embeddings, so that the representation of each word is influenced by its surrounding words in a given sentence. This model has been pre-trained on an extensive corpus of full-text biomedical articles from PubMed36,37 and clinical text data38, enabling it to capture intricate language patterns and domain-specific knowledge present in biomedical literature. This ensures that our summarizer gains the capability to comprehend the specialized language inherent in documents. The resulting embeddings are extracted from the last hidden state of the model which generates the contextualized information for each token in the sequence. The last hidden state of the model refers to the final internal representation generated by the encoder in a sequence-to-sequence model. It encapsulates the key information from the entire input sequence and serves as a condensed representation. This last hidden state is crucial in summarization tasks, as it is used as the initial context for the decoder to generate a concise and contextually informed summary of the input text. This process effectively provides a good representation of the original text, allowing our summarization tool to create comprehensive summaries that capture the essence of the content.
Hierarchical clustering with agglomerative clustering
The resulting sentence embeddings from the cambridgeltl/SapBERT-from-PubMedBERT-fulltext24,35 model are put through a hierarchical agglomerative clustering algorithm29,30 that works in a bottom-up approach. This process begins by treating each sentence as an individual cluster. It uses the cosine similarity metric to measure the similarity between sentences and employs the average linkage method to create a hierarchical structure. The output is a distance matrix that represents how sentences cluster together based on their similarity. Iteratively, a pair of closest clusters are merged, reducing the number of clusters by half. This merging process continues until all of the data points belong to one cluster, forming a hierarchical structure of clusters. Leveraging this technique, we are able to group semantically related sentences into clusters. The resulting dendrogram, shown in Fig. 2, can be cut at different levels to obtain various clusters. The number of clusters will be highly dependent on the complexity of the document.
We arrived at the optimal number of clusters empirically through experimentation. Our refined approach on the number of clusters is an integral part of \(\frac{n+1}{3}\), where n is the total number of sentences in the document. This formulation allowed us to achieve clusters each with coherent set of sentences. The heuristic \(\frac{n+1}{3}\) is designed to balance over-segmentation and under-segmentation of text data.
Each resulting cluster is ranked based on the smallest index among the sentences within that cluster. The rankings of these clusters will be used to order the summaries coming from each cluster, yielding a final cohesive summary.
Hierarchical clustering of a piece of regulatory document39. Top left: piece of regulatory document, top right: colored sentences corresponding to clusters, bottom left: dendrogram showing cluster heirarchy of sentences, and bottom right: key for colors corresponding to cluster ids.
Summarization with bart-large-cnn-samsum
The next step of the MedicoVerse solution focuses on the generation of concise summaries for a set of clustered texts, employing the large language model philschmid/bart-large-cnn-samsum40. This is a pre-trained sequence-to-sequence model that adopts the Transformer model architecture12, leveraging self-attention mechanisms to capture comprehensive dependencies between words within a sequence. The bart-large-cnn-samsum model is a Transformer-based text summarization model fine-tuned on the SamSum dataset40, which includes general conversational text and their corresponding summaries. This dataset features interactions that also include conversations between doctors and patients, discussing diseases, symptoms, and healthcare. Models trained on the SamSum dataset are also known to perform well on MTSamples, a collection of transcribed medical sample reports, and the Medical Information Mart for Intensive Care (MIMIC) dataset41.
Each cluster resulting from the agglomerative clustering technique is subsequently fed into the model. Summarizing each of these clusters ensures our ability to capture the context present across the entire document. Finally, MedicoVerse merges the summaries of the clusters with respect to the order of the clusters based on the original sentence indices.
Disease and chemical keyword annotation with SciSpacy
Within the scope of the current task, one notable application of the SciSpaCy42,43 model en_ner_bc5cdr_md lies in its capability to perform entity annotation by color-coding disease and chemical entities within each text. The value of this functionality is in the identification of relevant entities within the summarized text. By color-coding these entities, our summarization approach offers a visual cue, empowering users to swiftly discern and extract pertinent information concerning diseases and chemicals.
MedicoVerse summary of a piece of regulatory document from Fig. 2 highlighting diseases and chemical entities.
Quality of clusters and sumarries
The top left of Fig. 2 shows an example of a piece of regulatory document. Its color-coded form at the top right in Fig. 2 depicts the cluster of the corresponding sentences. The hierarchical clustering bins the sentences based on their semantic content. The color coding scheme and corresponding cluster number of Fig. 2 can be considered more carefully to understand what each cluster comprises.
-
Cluster 0 sentences discuss the water deficit in DKA and HHS, showing a thematic focus on the conditions impact on hydration levels.
-
Cluster 1 sentences cover the initial fluid therapy, its direction, and factors influencing the choice of fluid, indicating a focus on treatment steps.
-
Cluster 2 sentences elaborate on the specifics of fluid therapy, such as the type of saline and rate of infusion.
-
Cluster 3 sentences are the largest group and seems to discuss the management of the patients condition in a more comprehensive manner, covering various aspects of fluid therapy, insulin administration, and their effects.
-
Cluster 4 has a single sentence that emphasizes the importance of considering urinary losses, a unique aspect not covered in other clusters.
-
Cluster 5 contains a defining sentence about DKA and HHS, which is likely quite distinct from the operational treatment discussions in the other clusters.
Figure 3 shows the summary of a part of a regulatory document39 from Fig. 2. This summary indicates that key themes such as volume depletion in DKA and HHS, fluid therapy protocols, and the consideration of electrolytes and hydration status have been maintained. Specific details such as the rate and type of saline administration, the goal of therapy, and precautions with insulin administration in hypotensive patients have also been preserved. The summary has reduced the text by almost 46%. This suggests that the clustering managed to eliminate redundant information while retaining the essence of the text. Over the 227 pieces of texts from 38 regulatory documents, an average of 37% reduction was seen in their summaries. The ability to condense content without sacrificing the quality or missing vital information showcases the effectiveness of the clustering approach.
The summarizer also retains a high level of readability and presents the information in a manner that is accessible to both medical professionals and readers less familiar with the domain. It demonstrates the potential of advanced natural language processing techniques to support researchers in distilling and communicating complex datasets effectively.
Evaluation metrics
A multifaceted approach was employed for a comprehensive evaluation of MedicoVerse. We utilized the ROUGE metrics (ROUGE-1, ROUGE-2, ROUGE-L) to assess the quality of the generated summaries. ROUGE44,45,46 (Recall-Oriented Understudy for Gisting Evaluation) is a set of metrics used for the automatic evaluation of text summarization and machine translation systems. The ROUGE metrics measure the quality of the generated summary or translation by comparing it to one or more reference summaries or translations. The scoring ranges from 0 to 1, with 1 being the best score. There are 3 main variants under ROUGE metrics: ROUGE-1, ROUGE-2, and ROUGE-L. The ROUGE-1 metric compares the unigrams between the generated summary and the reference summary while ROUGE-2 compares the bigrams. The ROUGE-L metric is used to compute the longest common subsequence between the machine-generated summary and the reference summary. By treating each summary as a sequence of words, ROUGE-L considers sequence matches of sentences that reflect sentence-level word order. The ROUGE metrics precision, recall, and F1 score are assessed by comparing the model-generated summaries in reference to the raw text, which serves as the ground truth for evaluation.
In our work, we employed the ROUGE F1 score as the primary evaluation metric for assessing the quality of summaries generated by multiple models, given the absence of ground truth references. We also incorporated BERTScore to evaluate the similarity between the generated summary and the reference summary by utilizing contextual embeddings from models such as BERT. BERTScore computes precision, recall, and F1 scores based on token-level matches within the embedding space. While the average BERT F1 score and ROUGE F1 score provides a balanced assessment, it is important to acknowledge its sensitivity to the choice of evaluation metric and the potential for bias inherent in automated evaluation methods. To mitigate these limitations and broaden the scope of the evaluation on our models, we used business key performance indicators (KPIs) where subject matter experts identified business entities that were used to measure the models capacity to capture domain-specific entities. As a last measure, we considered the readability index that quantifies the ease with which a piece of text can be understood by its readers. This index provides a numerical value that reflects the complexity of the texts’ vocabulary, sentence structure, and overall readability. We employed the Flesch Reading Ease32 readability metric to assess the text’s readability, which gauges how easily a document can be comprehended. This metric relies on both the average sentence length and the average number of syllables per word in order to calculate readability. It evaluates the ease of comprehending a summary after the summarization process has been carried out.
Under these four metrics, we constructed a final score in Eq. (1), where Eq. (2) shows how to compute the ROUGE F1 Score Weighted Average term.
This final score of Eq. (1) was used to compare multiple summarization models such as bart-large-cnn-samsum, Facebook BART, Google Pegasus47, T5, Mixtral 8\(\times \)7b instruct, GPT 3.5, and Llama-2-70b in addition to our own approach. The weights of Eqs. (1) and (2) were determined empirically.
Evaluation
This paper’s primary objective is to assess the effectiveness of various summarization approaches and their ability to generate concise and coherent summaries for regulatory and PubMed documents. We introduced a novel scoring technique to evaluate the effectiveness of our approach, integrating four key metrics: ROUGE, BERTScore, Unique Business KPIs, and Flesch Reading Ease. A detailed analysis of the results from our evaluation is illustrated through performance Tables 1 and 2, with discussions of key findings, and comparisons to existing approaches. We also delve into the implications of our findings and their potential applications in pharmaceutical and biomedical sciences.
According to Table 1, the MedicoVerse model demonstrates robust performance across all ROUGE metrics, consistently achieving scores exceeding 0.4. The MedicoVerse model displays recall and precision scores of 0.57, 0.46, 0.56, 0.86, 0.73, 0.85, exhibiting a balance between precision and recall scores compared to the other models. The ability to strike this balance ensures that the generated summaries contain essential information while avoiding excessive inclusion of irrelevant terms.
On the contrary, models such as philschmid/bart-large-cnn-samsum, Facebook/bart-large-cnn, Google/pegasus-xsum, and T5 models without clustering demonstrate low recall scores and high precision, consistently surpassing 0.6. The high precision and low recall scores of these models suggest that the generated summaries fall short of capturing the most relevant words, phrases, or context from the reference text. Furthermore, the F1-scores associated with these models are notably low, residing within the range of 0.05–0.2, indicating a substantially lower overall performance compared to that of MedicoVerse.
To broaden the scope of our analysis and ensure a more exhaustive comparison, we explore the capabilities of MedicoVerse alongside large language models such as Mixtral 8\(\times \)7b instruct, GPT 3.5, and Llama-2-70b, as illustrated in Table 1. We utilized MedicoVerse’s architecture but substituted the bart-large-cnn-samsum model with Large Language Models (LLM). The Mixtral 8\(\times \)7b instruct exhibits notable F1-score performance, with values of (0.66, 0.51, 0.66) closely resembling those of MedicoVerse. Additionally, its recall scores (0.52, 0.39, 0.52) and precision scores (0.92, 0.76, 0.92), surpassing the 0.4 threshold, position it as the second-best performing model in our evaluation.
On the other hand, models such as GPT 3.5 and Llama-2-70b demonstrate noteworthy F1-score performances with values of (0.60, 0.31, 0.54) and (0.63, 0.42, 0.60), respectively. Furthermore, the recall and precision scores for both models fall within the range of 0.23 to 0.54, with precision values consistently exceeding 0.4. This finding highlights a favorable balance between recall and precision, indicating that the generated summaries effectively capture only the relevant information from the reference text.
Table 1 shows that MedicoVerse performs better than other compared models consistently under the evaluation metrics. MedicoVerse, a unique composition of hierarchical clustering combined with the philschmid/bart-large-cnn-samsum model, demonstrates significant advantages. However, a limitation inherent in the philschmid/bart-large-cnn-samsum model is the persistent inclusion of 1–2% noise in the generated summaries. This constraint is effectively mitigated by the Mixtral 8\(\times \)7b instruct, where the summaries are concise and domain-specific. Notably, models like GPT 3.5 and Llama-2-70b yield summaries that are lengthy, easy to comprehend, and tailored for a broader audience.
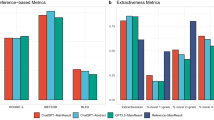
To assess the significance of the hierarchical clustering in the summarization process, we conducted evaluations using Mixtral 8\(\times \)7b instruct, GPT 3.5, and Llama-2-70b models as shown in the Table 1 to generate summaries without the inclusion of clustering as an additional layer. Comparing the Mixtral 8\(\times \)7b instruct without and with clustering, we observe an increase in ROUGE-1 scores from 0.15 to 0.52, ROUGE-2 from 0.04 to 0.39, and ROUGE-L from 0.14 to 0.52. This demonstrates the effectiveness of the supplementary clustering layer in capturing essential information and relationships within the raw text. Extending the analysis to the other models such as GPT 3.5 and Llama-2-70b, when going from not using hierarchical clustering to using it, the F1-scores have increased from (0.52, 0.28, 0.51) to (0.6, 0.31, 0.54) and (0.57, 0.37, 0.53) to (0.63, 0.42, 0.6), respectively. The application of hierarchical clustering consistently results in higher ROUGE scores across the models, reinforcing the general applicability of clustering techniques in the context of text summarization. The summaries of ten sampled data of regulatory documents across all these models are further reviewed by subject matter experts (SME’s), as depicted in Fig. 4. Notably, the x-axis in the graph represents the different models, while the y-axis indicates the score given by SME’s and computational scores. The graph illustrates a clear distinction in the scores assigned by SME’s between summaries generated with and without clustering. SME’s reviewed both the summaries and original text, assigned scores based on their expertise. Specifically, SME’s consistently rated the summaries generated by models with clustering higher compared to those without. This trend suggests that incorporating clustering techniques in the summarization process contributes to a deeper understanding of the input data, enabling the models to generate summaries of greater accuracy.
Table 2 provides an overview of the performance of eleven distinct models, with each metric averaged across ten sampled data of regulatory documents. We are using Eq. (1) for calculating the final score.
Table 2 shows that MedicoVerse emerges as the standout model with the highest final score of 0.64, demonstrating its overall performance. A more granular analysis of each metric starting with ROUGE F1 score weighted average illustrates that MedicoVerse attained a score of 0.60, while Mixtral 8\(\times \)7b instruct outperformed with a score of 0.63, followed closely by Llama-2-70b with a score of 0.61. Delving deeper into the ROUGE F1 score weighted average across ten sampled data of regulatory documents, it is evident that Mixtral 8\(\times \)7b instruct surpasses MedicoVerse and Llama-2-70b, showcasing its ability in effectively capturing the most relevant context from the reference text. This highlights that using hierarchical clustering with Mixtral 8\(\times \)7b instruct is a compelling choice for regulatory document text summarization.
In addition to the ROUGE scores, BERT F1 scores provide another layer of evaluation for contextual understanding. Here, Hierarchical Clustering + Mixtral 8\(\times \)7b instruct again leads with a score of 0.77, slightly ahead of both MedicoVerse and Hierarchical Clustering + Llama-2-70b, each scoring 0.76. The Hierarchical Clustering + GPT 3.5 Model also performs well with a score of 0.76, indicating strong contextual relevance across these models.
With regards to the average Flesch Reading Ease scores, MedicoVerse achieves the second highest score of 0.43, suggesting moderately easy-to-understand summaries suitable for a broader audience. However, it’s worth noting that the Flesch Reading ease scores for all the large language models such as Mixtral 8\(\times \)7b instruct, Llama-2-70b, and GPT 3.5, with and without clustering fall within the range of 0.18 to 0.29, reflecting highly concise and domain-specific summaries. In contrast, philschmid/bart-large-cnn-samsum, Facebook/bart-large-cnn, Google/Pegasus-xsum, and T5 models without clustering exhibit comparatively higher values of 0.35, 0.39, 0.33, and 0.44 and average ROUGE scores ranging from 0.1 to 0.32. These scores indicate that while the summaries generated by these models are easy to understand, they lack capturing the key elements from the reference text.
Lastly, MedicoVerse attains a Business KPI score of 0.70 while Llama-2-70b with hierarchical clustering stands out as the top performer among all other models with a score of 0.72. Following closely, with hierarchical clustering, GPT 3.5 achieves a score of 0.63, and Mixtral 8\(\times \)7b instruct achieves a score of 0.56. Conversely, models such as philschmid/bart-large-cnn-samsum, Facebook/BART-large-cnn, Google/Pegasus-xsum, and T5 models without clustering received significantly lower scores ranging from 0.06 to 0.31, indicating limited alignment with business objectives.
Tables 1 and 2 provide a clear analysis, indicating that MedicoVerse is the most effective model that meets all the business requirements. Its comprehensive performance, coupled with the advantage of being free-to-use, generating summaries with relevant context, positions it as a promising option. Among the other models evaluated, Llama-2-70b outperforms most of the benchmarks demonstrating competitive performance. Despite its scores, the summaries generated by Llama-2-70b with clustering lack conciseness, thereby leading to lengthy summaries. As a result of this drawback, Mixtral 8\(\times \)7b instruct with clustering emerges as a compelling option, positioned as the second-best performer.
Methods
Number of clusters
The determination of the optimal number of clusters in text clustering is a critical task that directly impacts the quality of results. Traditional methods such as the Elbow Method and Silhouette Score often fall short when applied to textual data which tends to be high-dimensional and sparse49,50,51. In this study, we propose a novel heuristic, \(\frac{n+1}{3}\), where n represents the number of sentences, to determine the number of clusters in hierarchical agglomerative clustering. This heuristic emerged from extensive experimentation and was found to provide consistent and meaningful clustering results across the datasets. The heuristic \(\frac{n+1}{3}\) is designed to balance over-segmentation and under-segmentation of text data. By incorporating a constant adjustment factor, this formula adapts to varying corpus sizes while maintaining a manageable number of clusters for interpretability. Our comparative analysis, illustrated in Table 3, demonstrates that the Silhouette Score tends to under-segment, the Elbow Method tends to over-segment, and the heuristic \(\frac{n+1}{3}\) offers an intermediate number of clusters, thereby achieving a more balanced segmentation.
Metrics
The performance of MedicoVerse is measured through metrics falling under four categories. The first is the ROUGE metric, which is commonly used for measuring text summarization models. The second is BERTScore, utilized to capture the semantic and contextual information of texts in both candidate and reference summaries. The third category is in business entity identification, where the capacity of MedicoVerse to accurately capture domain-specific business entities is measured. The final category is around measuring the readability of the summaries produced by MedicoVerse. After obtaining these four metrics, we constructed a weighted score.
ROUGE metrics
ROUGE31,52 (Recall-Oriented Understudy for Gisting Evaluation) is a set of metrics used for the automatic evaluation of text summarization and machine translation systems. ROUGE measures the quality of the generated summary by comparing it to one or more reference summaries. It ranges from 0-1, with 1 being a highest score.
ROUGE assesses the output summary of a model based on the reference summaries crafted by human evaluators. Since the ground truth is unavailable in our study, we consider the raw text itself as the designated reference summary for evaluation purposes. There are 3 main variants of ROUGE on which models have been evaluated, ROUGE-1, ROUGE-2, and ROUGE-L.
The ROUGE-1 score measures the overlap of unigrams (single words) between the generated summary and the reference summary. It calculates the precision and recall of unigrams in the generated summary compared to the reference summary as
In general, ROUGE-1 F1-score around 0.3 to 0.4 may be considered low and ROUGE-1 F1-score around 0.4 to 0.5 may be considered moderate, while ROUGE-1 F1-score above 0.5 is considered good.
The ROUGE-2 score evaluates the overlap of bigrams (pairs of adjacent words) between the generated summary and the reference summary. It calculates the precision and recall of bigrams in the generated summary compared to the reference summary as
In general, ROUGE-2 F1-score around 0.1 to 0.2 may be considered low and ROUGE-2 F1-score around 0.2 to 0.4 may be considered moderate, while ROUGE-2 F1-score above 0.4 is considered good.
ROUGE-L measures the longest common subsequence between the generated summary and the reference summary. A subsequence is a sequence of words that appear in the same order, but not necessarily consecutively. ROUGE-L calculates the precision and recall of the longest common subsequence as
In general, ROUGE-L F1-score around 0.3 to 0.4 may be considered low and ROUGE-L F1-score around 0.4 to 0.5 may be considered moderate, while ROUGE-L F1-score above 0.5 is generally considered good. ROUGE metrics assess the effectiveness of summarization models by considering both precision (the ability to produce relevant words) and recall (the ability to capture all relevant words). However, depending on the specific use case and the complexity of the text being summarized, it is possible to establish rough guidelines for evaluating the individual models performance.
BERTScore
Introduced in53,54, BERTScore is a metric used in NLP workflows to measure the similarity between candidate texts and reference texts using contextual embeddings. It leverages pre-trained BERT embeddings to capture the semantic and contextual information of words and phrases in both the candidate and reference texts.
Given a reference sentence of words \( \langle x_1, x_2, \ldots , x_n \rangle \), and candidate sentence of words \( \langle {\hat{x}}_1, {\hat{x}}_2, \ldots , {\hat{x}}_m \rangle \), we first use BERT to generate a sequence of word embeddings for both reference and candidate sentences as
Note that each \({\textbf{x}}_i\) and \({\hat{\textbf{x}}}_j\) are vectors since they are word embeddings.
To measure the similarity between two individual embeddings, we will use the cosine similarity which is defined as
which simply reduces to \({\textbf{x}}_i^\top \cdot {\hat{\textbf{x}}}_j\) since both \({\textbf{x}}_i\) and \({\hat{\textbf{x}}}_j\) are normalized. With these definitions, we can now calculate the BERT-Precision, BERT-Recall, and BERT-F1.
BERTScore comprises of three metrics: BERT-Precision, BERT-Recall, and BERT-F1. BERT-Precision assesses how well the candidate texts avoid introducing irrelevant content, BERT-Recall evaluates how well the candidate texts retain relevant content, and BERT-F1 combines both precision and recall to measure the overall effectiveness of the candidate texts in capturing and retaining relevant information from the reference texts. We recall the formulas found in53,54 for BERT-Precision (\(P_{BERT}\)), BERT-Recall (\(R_{BERT}\)) and BERT-F1 (\(F_{BERT}\)) as
where \({\textbf{X}} = \varvec{\langle {\textbf{x}}_1, {\textbf{x}}_2, \ldots , {\textbf{x}}_n \rangle }\) and \({\hat{\textbf{X}}} = \langle {\hat{\textbf{x}}}_1, {\hat{\textbf{x}}}_2, \ldots , {\hat{\textbf{x}}}_m \rangle \).
We employed the BERT-F1, \(F_{BERT}\), for a weighted average calculation, aiming to assess how well the candidate texts captured and retained relevant information from the reference texts in context. We assigned a weight of 0.2 to BERT Score, 0.2 to ROUGE, 0.4 to business KPIs, and 0.2 to the readability index.
Business entity identification
In this research, subject matter experts play a crucial role in identifying essential business entities. Businesses provide a list of keywords, critical context, units, and other important details to ensure they are not missed in the text summaries. After generating a summary, we conduct an analysis to determine the occurrence of important business terminology in both the original text and the summary. We evaluate the algorithm’s performance by measuring the percentage of these significant business terms captured in the summary. This analysis is a key aspect of our research findings, offering valuable insights into the algorithm’s effectiveness in retaining critical business information.
Readability index
Readability index is a formula that quantifies the ease in which a piece of text can be understood by its readers. It provides a numerical value that reflects the complexity of the texts vocabulary, sentence structure, and overall readability.
To compute the readability of the summaries, the Flesch Reading Ease index is considered. The Flesch Reading Ease32,55,56 is a readability metric that measures how easy it is to read a given piece of text. It uses the average sentence length and the average number of syllables per word to calculate readability as
The Reading Ease score of Eq. (3) ranges from 0 to 100, with high values indicating easier readability.
Conclusion
The MedicoVerse text summarizer stands out as a pioneering solution in the field of pharmaceutical sciences, leveraging hierarchical agglomerative clustering with SapBERT embeddings and the philschmid/BART-large-cnn-samsum model. Notably, it captures key business entities while maintaining a balance between precision and recall across various ROUGE metrics. Moreover, the summaries produced by MedicoVerse demonstrate a high readability index, falling within the moderate-to-easy range. This ensures that the conveyed information is accessible and understandable to a broad audience. An advantage of MedicoVerse is in its performance, coupled with the accessibility of free-to-use, easy-to-understand summaries enriched with relevant context. Such attributes position MedicoVerse as a promising option for both researchers and practitioners in pharmaceutical sciences. In comparison to other models evaluated in our analysis, hierarchical clustering with Mixtral 8\(\times \)7b emerges as the second-best performer, offering concise and domain-specific summaries. However, it is important to note that models such as GPT 3.5 with hierarchical clustering produces text summaries which will be useful for a broader audience. The model Llama-2-70b produces lengthy summaries that may lack the precision and relevance required within the pharmaceutical science domain. Therefore, by leveraging MedicoVerse or similar state-of-the-art models integrated with hierarchical clustering, researchers and practitioners can efficiently distill vast amounts of information into concise and insightful summaries.
As the landscape of large language models is rapidly evolving, there is a need to consider more recent models. One of the advantages of the architecture of MedicoVerse is that the model for the text summarization can be interchanged. One possible area of research is to see how the architecture of MedicoVerse would behave with the recent large language models such as GPT and Google Gemini models. One can also consider the architecture change by varying the models for embeddings. Another possible area of research is to vary the number of clusters for the hierarchical clustering over newer models of choice. Finally, we find that the architecture of MedicoVerse around a large language model performs better than just using the same large language model for text summarization. This may have to do with the embedding model or more so with the hierarchical clustering. It would be interesting to see if this can be shown to be true in general or to find a large language model that can perform better than with the MedicoVerse architecture for text summarization tasks.
Data availability
The data analyzed in this study may be made available upon request. Contact contributing author Sumit Ranjan for requests around the data.
Code availability
The code used in this study may be made available upon request. Contact contributing author Sumit Ranjan for requests around the code.
References
Bui, D., Del Fiol, G., Hurdle, J. & Jonnalagadda, S. Extractive text summarization system to aid data extraction from full text in systematic review development. J. Biomed. Inform. 64, 265–272 (2016).
Alguliev, R. & Aliguliyev, R. Evolutionary algorithm for extractive text summarization. Intell. Inf. Manag. 1, 128–138 (2009).
Sinha, A., Yadav, A. & Gahlot, A. Extractive Text Summarization Using Neural Networks (2018). Preprint at https://arxiv.org/abs/1802.10137.
Nallapati, R., Zhou, B., Gulcehre, C., Xiang, B. & Dos santos, C. N. Abstractive Text Summarization Using Sequence-to-Sequence rnns and Beyond. (2016). Preprint at https://arxiv.org/abs/1602.06023.
Lin, H. & Ng, V. Abstractive summarization: A survey of the state of the art. Proc. AAAI Conf. Artif. Intell. 33, 9815–9822 (2019).
Gupta, S. & Gupta, S. K. Abstractive summarization: An overview of the state of the art. Expert Syst. Appl. 121, 49–65 (2019).
Sutskever, I., Vinyals, O. & Le, Q. V. Sequence to sequence learning with neural networks. Advances in neural information processing systems27 (2014).
Bahdanau, D., Cho, K. & Bengio, Y. Neural Machine Translation by Jointly Learning to Align and Translate. arXiv preprint arXiv:1409.0473 (2014).
Luong, M.-T., Pham, H. & Manning, C. D. Effective Approaches to Attention-Based Neural Machine Translation. arXiv preprint arXiv:1508.04025 (2015).
See, A., Liu, P. J. & Manning, C. D. Get to the Point: Summarization with Pointer-Generator Networks. arXiv preprint arXiv:1704.04368 (2017).
Cohan, A. et al. A Discourse-Aware Attention Model for Abstractive Summarization of Long Documents. arXiv preprint arXiv:1804.05685 (2018).
Vaswani, A. et al. Attention is All you Need. (2017). Preprint at https://arxiv.org/abs/1706.03762.
Zhang, H., Xu, J. & Wang, J. Pretraining-Based Natural Language Generation for Text Summarization. arXiv preprint arXiv:1902.09243 (2019).
Liu, Y. & Lapata, M. Text Summarization with Pretrained Encoders. arXiv preprint arXiv:1908.08345 (2019).
You, Y., Jia, W., Liu, T. & Yang, W. Improving abstractive document summarization with salient information modeling. Proceedings of the 57th Annual Meeting of the Association for Computational Linguistics 2132–2141 (2019).
Xu, S. et al. Self-attention guided copy mechanism for abstractive summarization. Proceedings of the 58th Annual Meeting of the Association for Computational Linguistics 1355–1362 (2020).
Pilault, J., Li, R., Subramanian, S. & Pal, C. On extractive and abstractive neural document summarization with transformer language models. Proceedings of the 2020 Conference on Empirical Methods in Natural Language Processing (EMNLP) 9308–9319 (2020).
Zhang, M. et al. Bertscore: Evaluating Text Generation with Bert (2020). Preprint at https://arxiv.org/abs/1904.09675.
Alva-Manchego, F., Scarton, C. & Specia, L. The (un)suitability of automatic evaluation metrics for text simplification. Comput. Linguist. 47, 861–889. https://doi.org/10.1162/coli_a_00418 (2021).
Bryant, C. et al. Grammatical error correction: A survey of the state of the art. Comput. Linguist. 49, 643–701. https://doi.org/10.1162/coli_a_00478 (2023).
Li, Y. Deep Reinforcement Learning: An Overview. arXiv preprint arXiv:1701.07274 (2017).
Keneshloo, Y., Shi, T., Ramakrishnan, N. & Reddy, C. K. Deep reinforcement learning for sequence-to-sequence models. IEEE Transact. Neural Netw. Learn. Syst. 31, 2469–2489 (2019).
Paulus, R., Xiong, C. & Socher, R. A Deep Reinforced Model for Abstractive Summarization. arXiv preprint arXiv:1705.04304 (2017).
Devlin, J., Chang, M., Lee, K. & Toutanova, K. Bert: Pre-training of Deep Bidirectional Transformers for Language Understanding. (2018). Preprint at https://arxiv.org/abs/1810.04805.
Ward, J. H. Hierarchical grouping to optimize an objective function. J Am. Stat. Assoc. 58, 236–244 (1963).
Gowda, K. C. & Krishna, G. Agglomerative clustering using the concept of mutual nearest neighbourhood. Pattern Recognit. 10, 105–112. https://doi.org/10.1016/0031-3203(78)90018-3 (1978).
Patel, K., Patel, D., Golakiya, M., Bhattacharyya, P. & Birari, N. Cohen, K., Demner-Fushman, D., Ananiadou, S. & Tsujii, J. (eds) Adapting Pre-trained Word Embeddings for Use in Medical Coding. (eds Cohen, K., Demner-Fushman, D., Ananiadou, S. & Tsujii, J.) BioNLP 2017 (Association for Computational Linguistics, Vancouver, Canada, 2017).
Wang, Y. et al. A comparison of word embeddings for biomedical natural language processing. J. Biomed. Inform. 87, 12–20 (2018).
Ushioda, A. Scott, D. (ed.) Hierarchical clustering of words and application to NLP tasks. (ed.Scott, D.) Fourth Workshop on Very Large Corpora (Association for Computational Linguistics, Herstmonceux Castle, Sussex, UK, 1996).
Murtagh, F. & Contreras, P. Methods of Hierarchical Clustering. (2011). Preprint at https://arxiv.org/abs/1105.0121.
Lin, C.-Y. Looking for a few good metrics : Rouge and its evaluation. Proc. of the 4th NTCIR Workshops, Tokyo, Japan (2004).
Crossley, S. et al. A large-scaled corpus for assessing text readability. Behav. Res. Methods 55, 491–507 (2022).
Food, U. & Administration., D. Establishment Registration and Device Listing. https://www.accessdata.fda.gov/scripts/cdrh/cfdocs/cfrl/rl.cfm.
of Health National Library of Medicine., N. I. Pubmed. https://pubmed.ncbi.nlm.nih.gov/.
Liu, F., Shareghi, E., Meng, Z., Basaldella, M. & Collier, N. Self-alignment pretraining for biomedical entity representations. (2020). Preprint at https://arxiv.org/abs/2010.11784.
Fiorini, N., Lipman, D. & Lu, Z. Cutting edge: Towards pubmed 2.0. eLife (2017).
Williamson, P. & Minter, C. Exploring pubmed as a reliable resource for scholarly communications services. J. Med. Libr. Assoc. 107, 16–29 (2019).
Spasic, I. & Nenadic, G. Clinical text data in machine learning: systematic review. JMIR Med. Inform.8 (2020).
Gosmanov, A., E.O., G. & A.E., K. Hyperglycemic crises: Diabetic ketoacidosis and hyperglycemic hyperosmolar state. Endotext [Internet] (2021).
Gliwa, B., Mochol, I., Biesek, M. & Wawer, A. Samsum corpus: A human-annotated dialogue dataset for abstractive summarization. (2019). Preprint at https://arxiv.org/abs/1911.12237.
Labs., J. S. Summarize Clinical Notes (augmented). https://nlp.johnsnowlabs.com/2023/03/30/summarizer_clinical_jsl_augmented_en.htmluno/abcde.html.
Neumann, M., King, D., Beltagy, I. & Ammar, W. Scispacy: Fast and Robust Models for Biomedical Natural Language Processing. (2019). Preprint at https://arxiv.org/abs/1902.07669.
Tarcar, A. et al. Healthcare Ner Models Using Language Model Pretraining. (2019). Preprint at https://arxiv.org/abs/1910.11241.
Lin, C.-Y. Zaimis, E. (ed.) Rouge: A package for automatic evaluation of summaries. (ed.Zaimis, E.) Text Summarization Branches Out, 74–81 (Association for Computational Linguistics, Barcelona, Spain, 2004).
Ganesan, K. Rouge 2.0: Updated and improved measures for evaluation of summarization tasks. (2018). Preprint at https://arxiv.org/abs/1803.01937.
Cohan, A. & Goharian, N. Revisiting Summarization Evaluation for Scientific Articles. (2016). Preprint at https://arxiv.org/abs/1604.00400 .
Zhang, Y., J.and Zhao, Saleh, M. & Liu, P. Pegasus: Pre-training with Extracted Gap-Sentences for Abstractive Summarization. (2019). Preprint at https://arxiv.org/abs/1912.08777.
Janssen. Rybrevant amivantamab-vmjw, injection bla/nda number: 761210 product quality review. (2021). https://www.accessdata.fda.gov/drugsatfda_docs/nda/2021/761210Orig1s000ChemR.pdf.
Humaira, H. & Rasyidah, R. Determining the Appropiate Cluster Number Using Elbow Method for k-Means Algorithm (2020).
Rousseeuw, P. Silhouettes: A graphical aid to the interpretation and validation of cluster analysis. J. Comput. Appl. Math. 20, 53–65 (1987).
Januzaj, Y., Beqiri, E. & Luma, A. Determining the optimal number of clusters using silhouette score as a data mining technique. Int. J. Online Biomed. Eng. (iJOE) 19, 174–182 (2023).
Barbella, M. & Tortora, G. Rouge Metric Evaluation for Text Summarization Techniques. (2022). Preprint at https://papers.ssrn.com/sol3/papers.cfm?abstract_id=4120317 .
Zhang, T., Kishore, V., Wu, F., Weinberger, K. Q. & Artzi, Y. Bertscore: Evaluating Text Generation with Bert. arXiv preprint arXiv:1904.09675 (2019).
Zhang, T., Kishore, V., Wu, F., Weinberger, K. & Artzi, Y. Bertscore: Evaluating Text Generation with BERT (2020). https://openreview.net/forum?id=SkeHuCVFDr.
Stajner, S., Evans, R., Orasan, C. & Mitkov, R. Rello, L. & Saggion, H. (eds) What can readability measures really tell us about text complexity? (eds Rello, L. & Saggion, H.) Proceedings of the LREC’12 Workshop: Natural Language Processing for Improving Textual Accessibility (NLP4ITA) (European Language Resources Association (ELRA), Istanbul, Turkey, 2012).
Wrigley Kelly, N., Murray, K., McCarthy, C. & O’Shea, D. An objective analysis of quality and readability of online information on Covid-19. Heal. Technol. 11, 1093–1099 (2021).
Acknowledgements
We would like to acknowledge Allison Lo and Valerie Carothers from Pfizer INC., Biotherapeutics & Pharmaceutical Sciences - Transformational Technology Digital Sciences for contributions to edits. We would also like to acknowledge the scientists from Pfizer INC., Biotherapeutics & Pharmaceutical Sciences for their involvment as subject matter experts for reviewing summaries of regulatory documents.
Author information
Authors and Affiliations
Contributions
All authors are contributed to manuscript in equal ways.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Dalal, A., Ranjan, S., Bopaiah, Y. et al. Text summarization for pharmaceutical sciences using hierarchical clustering with a weighted evaluation methodology. Sci Rep 14, 20149 (2024). https://doi.org/10.1038/s41598-024-70618-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-70618-w
- Springer Nature Limited