Abstract
Intrahepatic cholangiocarcinoma (iCCA) presents a challenging diagnosis due to its nonspecific early clinical manifestations, often resulting in late-stage detection and high mortality. Diagnosing iCCA is further complicated by its limited accuracy, often necessitating multiple invasive procedures for precise identification. Despite carbohydrate antigen 19–9 (CA19-9) having been investigated and employed for iCCA diagnosis, it demonstrates modest diagnostic performance. Consequently, the identification of novel biomarkers with improved sensitivity and specificity remains an imperative yet formidable task. Autoantibodies, as early indicators of the immune response against cancer, offer a promising avenue for enhancing diagnostic accuracy. Our study aimed to identify non-invasive blood-based autoantibody biomarkers capable of distinguishing iCCA patients from healthy individuals (CTRs). We profiled autoantibodies in 26 serum samples (16 iCCAs and 10 CTRs) using protein microarrays containing 1622 functional proteins. Leveraging machine learning techniques, we identified a signature composed of three autoantibody biomarkers (NDE1, PYCR1, and VIM) in conjunction with CA19-9 for iCCA detection. This combined signature demonstrated superior diagnostic performance with an AUC of 96.9%, outperforming CA19-9 alone (AUC: 83.8%). These results suggest the potential of autoantibody biomarkers to develop a complementary non-invasive diagnostic utility for routine iCCA screening.
Similar content being viewed by others
Introduction
Intrahepatic cholangiocarcinoma (iCCA) constitutes 10–15% of newly diagnosed liver cancers worldwide1. For Thailand, the annual incidence rate is 14.6 per 100,000 people2. While the curative treatment for iCCA is early surgical resection, the nonspecific early clinical presentations often lead to delayed diagnosis. This delay limits treatment options and results in reduced survival rates1. Therefore, 60–70% of patients are diagnosed at advanced stages, with a median survival of 12–15 months3. According to the American Cancer Society, the 5-year relative survival rate is 34.2% for early localized cases but decreases to 2.5% for advanced cases with distant metastases4. These alarming statistics emphasize the need for early detection of iCCA.
Currently, iCCA diagnosis relies heavily on imaging modalities, including computerized tomography (CT), magnetic resonance imaging (MRI), and positron emission tomography (PET) scans, followed by pathological examination of resected lesions5. However, these approaches exhibit moderate sensitivity, ranging from 18 to over 90%, potentially leading to false-negative results, and the requirement for repeated biopsies further complicates iCCA diagnosis5,6. Carbohydrate antigen 19–9 (CA19-9), a tetrasaccharide typically attached to O-glycans on the cell surface and a well-known serological tumor marker, has been investigated and used for diagnostic and monitoring purposes in iCCA. However, it demonstrates limited diagnostic performance, with 72% sensitivity and 84% specificity7. Consequently, the identification of non-invasive blood-based biomarkers with higher sensitivity and specificity has become a research priority to address the challenges of iCCA diagnosis and early detection8,9,10.
Tumor-associated autoantibodies (AAbs) offer a promising avenue for early cancer diagnosis as they are part of the initial humoral immune response against tumor-associated antigens (TAAs). These elevated autoantibody levels can be detected in the peripheral blood of patients up to five years before clinical symptoms manifest, enabling the potential for early blood-based diagnostic biomarkers11. It is suggested that autoantibodies could be generated through the over-expression, mutation, release of TAAs from damaged tissues, or misfolding of proteins, leading to their recognition by the immune system12,13,14. While the exact mechanisms of autoantibody production in cancer remain elusive, their high specificity, stability, and non-invasive nature make them attractive candidates for enhancing the accuracy of iCCA diagnosis12,13,14. Previous reports showed that autoantibodies possess predictive abilities for cancer recurrence, metastasis, and overall survival in various cancer types, improving timely interventions and patient outcomes11. Hence, the selection of autoantibodies using multiple machine learning approaches could provide new biomarkers for iCCA diagnosis. However, detecting and enriching viable autoantibodies within complex biological matrices remain challenging tasks.
To address these complexities, this study utilized high-density protein microarrays consisting of various correctly folded human proteins, including TAAs, cytokines, interleukins, and transcription factors15,16, to comprehensively profile sera of iCCA patients and healthy controls (CTRs) from the Thai population17. Through statistical analyses and a machine learning-based feature selection pipeline, we identified an optimal biomarker signature, which includes three autoantibodies and CA19-9. This signature demonstrated superior diagnostic performance compared to CA19-9 alone. Furthermore, this study aimed to investigate potential pathways of autoantibody-targeted antigens involved in iCCA pathogenesis, which might lead to the development of better treatment and clinical management for iCCA patients.
Materials and methods
Study population and sample selection
We selected 16 serum samples from iCCA patients diagnosed through combinations of imaging studies and histological investigations, along with 10 serum samples from age-, sex-, and location-matched CTRs with no history of cancer. These samples were obtained from the Thailand Initiative in Genomics and Expression Research for Liver Cancer (TIGER-LC) biorepository. Additionally, we integrated mRNA levels of autoantibody-targeted antigens from 76 iCCA subjects, sourced from our previous TIGER-LC study of Chaisaingmongkol et al.17, to confirm the validity of our candidate autoantibodies. Details of the sample size calculation (power analysis) are available in the Supplementary Data 1 online.
Written informed consent was obtained from all participants included in this study in accordance with the Declaration of Helsinki and Good Clinical Practice guidelines. The study protocols were approved by the Institution Review Boards of National Cancer Institute (NCI protocol number 13CN089), Chulabhorn Research Institute (CRI protocol number 18/2555), Chulabhorn Hospital (Chulabhorn Hospital protocol number 11/2553), Thai National Cancer Institute (Thai NCI protocol number EC163/2010), Chiang Mai University (Chiang Mai University protocol number TIGER-LC), and Khon Kaen University (Khon Kaen University protocol number HE541099).
Protein microarray profiling
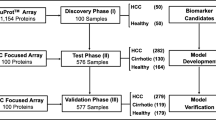
The serum autoantibody profiling process is summarized in Fig. 1a. We utilized the protein microarray platform (Immunome Arrays, Sengenics), which consists of quadruplicate spots of 1622 human proteins, tagged with biotin carboxyl carrier protein (BCCP)-Myc folding and printed onto a streptavidin-conjugated hydrogel slide (as detailed in Supplementary Fig. S1 online)15,16. A comprehensive list of these proteins and their corresponding UniProt IDs can be found as Supplementary Data 2 online. The steps and procedures for the protein microarray profiling were performed as previously described18. In brief, sera aliquots from iCCA and/or CTR samples, diluted to 1:400 in a buffer containing 0.1% v/v Triton X100, 0.1% w/v BSA in 1X PBS, were incubated on the slides for 2 h. Detection of the reaction was carried out using Cy3-conjugated anti-human IgG (Dako Cytomation) and scanned using a microarray laser scanner (Agilent Technologies) at a resolution of 10 µm.
Experimental strategy for iCCA biomarker and signature identification. (a) Immunome arrays analyzed autoantibody profiles in 26 serum samples. After data preprocessing and filtering, differentially expressed autoantibodies were identified through various criteria and pathway analysis was performed. Candidate autoantibody biomarker expressions were validated against their targeted-antigen mRNA expressions. (b) The selected candidate biomarker data was utilized to train the models. All possible combinations of biomarkers were generated and individually trained using Random Forest algorithms. The model with the optimal diagnostic performance formed a promising iCCA biomarker signature. Graphic was created with BioRender.com.
Data pre-processing
Scanned images underwent pre-processing and quality control (QC) procedures using GenePix Pro 7 (Molecular Devices), following established methods19, with a summary of the QC results detailed in Supplementary Table S1 online. The signal intensities reflecting autoantibody binding to specific proteins were quantified as relative fluorescent intensities (RFUs). Subsequently, the raw RFUs underwent composite normalization employing both quantile-based and intensity-based modules on Cy3-labelled biotinylated BSA-positive control probes, as previously outlined by Duarte et al.20. We further conducted a filtering process, excluding proteins with a mean expression below 450 RFUs to enhance confidence in the candidate proteins. This threshold was determined by systematically testing different threshold values and evaluated their impact on the number of proteins with a false discovery rate-adjusted p-value (FDR) ≤ 0.1 using Welch's unequal variance t-test21. The threshold that yielded the maximum number of proteins was selected, as illustrated in Supplementary Fig. S2 online. The data were then log2 transformed for all downstream analysis.
Selection of differentially expressed autoantibodies
To identify differentially expressed autoantibodies between the iCCA and CTR groups, a Welch’s t-test21 with Benjamini–Hochberg procedure (FDR correction) was performed on the filtered data, using an FDR threshold of ≤ 0.1. A higher cutoff of 0.1 was employed to retain potential candidates in these early stages of discriminating cancer and non-cancer samples22. Log2 fold change (log2 FC) was then performed to confirm the validity of the highly expressed autoantibodies derived from the t-test. Candidates with absolute log2 FC greater than 1 were selected for further analysis. A univariate receiver operating characteristic (ROC) analysis for binary classification (iCCA vs CTR) was conducted on the selected hits using generalized linear models (GLMs) with leave-one-out cross-validation (LOOCV). Autoantibodies with corresponding AUC scores of at least 75% were chosen as an initial set of most relevant candidate biomarkers. All analyses were performed using R Statistical Software version 4.3.123. The heatmaps presented in this paper were generated using the ‘ComplexHeatmap’ package version 2.12.124, with hierarchical clustering of columns and rows based on ‘canberra’ distances and the ‘ward.D2’ linkage. GLMs were performed on individual candidates using the ‘caret’ package version 6.0–9325, while the assessment of AUC was carried out using the ‘pROC’ package version 1.18.026.
Comparative analysis of candidate autoantibody and targeted antigen gene expression profiles
The mRNA expression profiles for iCCA tumor (T) and adjacent non-tumor (NT) tissue pairs were obtained from the gene expression omnibus (GEO) database (accession number GSE76297)17. A total of 76 pairs of tissues from iCCA patients were included. Autoantibody-targeted antigen genes were selected based on previous analysis. Autoantibody and mRNA expression profiles were z-score standardized. The p-values were computed using log2-transformed data and the Mann–Whitney U test with FDR correction. Autoantibody and gene expression log2 FC values were calculated, and candidates with opposite regulatory directions were excluded. The remaining candidates were assessed for multicollinearity using the ‘corrplot’ function in R, employing Spearman’s rank correlation. Principal component analysis (PCA) was also performed on the final hits using ‘prcomp’ function in R.
Biomarker signature selection for iCCA diagnosis
To enhance predictive performance, we created biomarker signatures from autoantibody candidates plus CA19-9. We assessed various combinations using random forest models27. The pipeline is outlined in Fig. 1b. We opted for the random forest algorithm due to its superior average median training accuracy, outperforming seven other machine learning methods, including GLMs, Naïve Bayes28, regularized discriminant analysis29, support vector machine30, k-nearest neighbors31, distance weighted discrimination32, and Bayesian GLMs33. A detailed comparison is provided in Supplementary Table S2 online.
To identify the optimal iCCA diagnostic signature using the random forest classifier, we generated sets of candidate biomarkers using the ‘combn’ function in R. This process produced 120 unique signature combinations. Each signature was individually trained using the ‘caret’ package25 in R, employing ‘method = rf’, ‘tuneLength = 50’, and LOOCV. Performance metrics, including AUC, accuracy, optimal threshold(s), sensitivity(ies), and specificity(ies) were assessed through ROC curve analysis using the ‘pROC’ package26. The ‘best’ threshold(s) were determined using the ‘Youden’ method (Sensitivity + Specificity 1). Ninety-five percent confidence intervals (95% CIs) for these metrics were calculated based on 2000 stratified bootstrap replicates. The panel with an optimal performance was identified as a potential signature for iCCA detection. PCA analysis was also performed on the optimal signature using ‘prcomp’ function in R to confirm the result. Of note, the autoantibody expression and CA19-9 concentration data were preprocessed through a sequence of steps, including ‘medianImpute’ imputation, log2 transformation, upsampling, centering, and scaling before modeling. To ensure result reproducibility, a fixed random seed of ‘123’ was consistently used throughout the study’s analyses, including processes involving randomness such as imputation, resampling, and modeling.
Functional enrichment analysis of iCCA associated autoantibody-targeted antigens
In this study, the functional enrichment analysis is focused on the interactions among autoantibody-targeted antigens themselves, rather than the interaction between the autoantibodies. Our hypothesis driving this investigation is that autoantibodies arise because of immune responses triggered by the aberrant expression of their targeted antigens. Therefore, our aim is to gain insights into the functions of these autoantibody-targeted antigens and their involvement in the statistically significant pathways associated with iCCA development.
To assess the functional relevance of the targeted antigens linked to iCCA associated autoantibodies, we first performed the Over-Representation Analysis (ORA) of differentially expressed proteins derived from the Welch's t-test21 with FDR ≤ 0.1 by using their UniProt accession numbers with the STRING database version 1234. Protein–protein interactions (PPI) were examined with medium confidence (a combined score > 0.4), an FDR of 5%, and 1622 proteins in the array as a statistical background. The Markov clustering (MCL)35 was applied to the network using an inflation parameter of 2. Additionally, we applied Gene Set Enrichment Analysis (GSEA) to our immunome data using the ‘clusterProfiler’ package version 4.8.3 in R36 to identify enriched biological pathways or functional annotations associated with iCCA, utilizing the gene ontology (GO) database. This analysis included all 1622 proteins with their corresponding log2 fold changes, and statistically significant terms were defined with FDR ≤ 0.05.
Results
Analysis of clinicopathological parameters
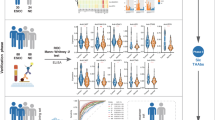
Demographic, clinicopathologic, and other participant characteristics are summarized in Table 1. A total of 26 serum samples were analyzed, comprising two groups: 16 iCCAs and 10 matched CTRs for autoantibody profiling. In both groups, most samples were males (81.2% in iCCA and 80.0% in CTR). The mean age of the iCCA was 62.8 ± 6.9 years, while that of the CTR was 61.1 ± 4.8 years, indicating a similar age distribution between two groups (p = 0.5171, Independent t-test). Among clinical variables assessed, including risk factors and liver function indicators, only International Normalized Ratio (INR, p = 0.0217), Alkaline Phosphatase (ALP, p = 0.0001), and CA19-9 (p = 0.0031) showed statistically significant differences; other variables displayed no statistical significance (p > 0.05, Fisher’s exact test).
Identification of differentially expressed iCCA-associated autoantibodies using protein microarray
A heatmap in Supplementary Fig. S3a online summarizes the relative abundance of all 1622 autoantibodies. During the filtering process, 1427 proteins (87.98%) with mean expressions lower than 450 RFUs (optimized cutoff, see Supplementary Fig. S2 online) were excluded, leaving 195 proteins (12.02%) (Supplementary Fig. S3b online) for further analyses. These remaining 195 proteins were tested and compared between iCCA and CTR, showing 66 differentially expressed proteins at FDR ≤ 0.1 and exhibiting higher relative abundance in the iCCA group (log2 FC from 0.391 to 2.350, Supplementary Table S3 online). Among these, only 42 autoantibodies passed our criteria (FDR ≤ 0.1 and |log2 FC| ≥ 1, Fig. 2a) with an average log2 fold increase of 1.415 ± 0.292 in the iCCA group (log2 FC from 1.012 to 2.350, Supplementary Table S3 online). The relative abundance of these 42 autoantibodies was shown in Fig. 2b.
Comprehensive differential expression analysis of autoantibodies within the Thai cohort samples for potential biomarkers of iCCA diagnosis. (a) The volcano plot quantifies 195 filtered autoantibodies in iCCA vs CTR. Forty-two autoantibodies meeting the statistical cutoff criteria (FDR ≤ 0.1 and |log2 FC| ≥ 1) are considered as candidates. (b) The heatmap represents the relative expression levels of the 42 highly abundant autoantibodies that meet the criteria. (c) A bar chart displays individual protein AUCs, with ≥ 75% indicating strong diagnostic ability. ABNL abnormal, AveExpr average expression level, BMI body mass index, Expr expression level, NA not available, NL normal, OB obesity, OW overweight, UW underweight.
Selection of candidate biomarkers for iCCA detection
Given the limited sample size (n = 26) in comparison to the 42 candidate autoantibodies at this stage, overfitting is a concern. To prevent overfitting, a feature selection pipeline was implemented, ensuring that only iCCA-related features remained for the subsequent modeling stage. This approach aimed to strike a balance between the analysis's complexity and the available data, thereby optimizing the reliability and generalizability of the results. First, univariate ROC analysis for iCCA vs CTR classification was performed on the 42 candidates that had been identified (Fig. 2c). These autoantibodies were ranked based on their AUCs, which indicate their diagnostic performance. From this analysis, we identified 7 autoantibodies with AUCs ≥ 75%, namely NDE1, GFAP, KRT8, PYCR1, FADD, VIM, and KRT19. On average, they achieved a score of 78.1% (AUC range: 75.0–84.4%) and were chosen as the most relevant features (Fig. 2c).
To validate the 7 selected candidates, we examined tissue mRNA expression of their corresponding autoantibody-targeted antigens and compared it to autoantibody profiles (Fig. 3a). We hypothesized that the immune response leading to autoantibody production is triggered by elevated antigen expression, a consequence of abnormal gene expression in the tumor. Confirming increased mRNA expression of these antigens would substantiate the abundance of our candidate autoantibodies. While the patterns of some candidates display a negative correlation between the two profiles due to the complex relationship between autoantibodies and their targeted antigens, most candidates exhibited the same regulatory directions (Fig. 3a,b). However, GFAP was excluded from further analysis due to a different regulatory direction, as confirmed by the relative log2 FC between the autoantibody (1.235 log2 fold increase in iCCA) and the targeted antigen mRNA (− 0.125 log2 fold decrease in iCCA) expression profiles (Fig. 3b). Therefore, only 6 candidates remained, and each of them exhibited statistically significant expression in cancer cases with FDR < 0.05 (Fig. 3c), as well as the ability to distinguish between iCCA and CTR with AUCs > 70% (AUC range: 72.6–99.6%) in both AAb and mRNA profiles (details in Supplementary Fig. S4 online). Furthermore, we explored the level of serum CA19-9 in our cohort, as illustrated in Fig. 3d. The CA19-9 level exhibited a statistically significant increase in iCCA patients (median 5.61 log2 U/mL) compared to CTRs (median 3.05 log2 U/mL, p = 0.003, Mann–Whitney U test). The diagnostic performance of CA19-9 was further assessed, revealing an AUC of 83.8% (Supplementary Fig. S5 online). Overall, these results suggest that the selected autoantibodies, along with CA19-9, have a high likelihood of being true positive hits and could serve as potential biomarkers for diagnosing iCCA.
Candidate biomarker selection for iCCA diagnosis. (a) Heatmaps are used to compare the expression profiles of candidate autoantibodies with the mRNA expression of their corresponding targeted antigens. (b) Average log2 FC of candidate autoantibody and their targeted antigen gene expressions were quantified from the immunome and transcriptome cohorts, respectively. (c) Violin plots of selected candidates showing autoantibody and gene expression levels compared between iCCA/CTR and T/NT for immunome and transcriptome data, respectively. All candidates show levels of significance: FDR < 0.05; FDR < 0.001 (Mann–Whitney U test) for immunome and transcriptome data, respectively. (d) Violin plot of CA19-9 concentration retrieved from clinical data of immunome cohort. Level of significance: p = 0.003 (Mann–Whitney U test). Statistical significance is defined as a value of FDR and/or p ≤ 0.05. Expr expression level.
Distinct potential of NDE1 and CA19-9 as iCCA biomarkers
In our pursuit of an optimal biomarker panel to enhance iCCA diagnostic accuracy, we assessed relationships among candidate biomarkers using Spearman’s rank correlation (Fig. 4a). KRT8, PYCR1, FAD, VIM, and KRT19 exhibited very strong associations, with an average Spearman's rank correlation coefficient (ρ) of 0.817 ± 0.085 (p < 0.001). This high correlation suggests potential redundancy and multicollinearity among these candidates, indicating they convey similar diagnostic information. However, NDE1 and CA19-9 deviated from this trend, exhibiting moderate associations with other candidates, with lower average correlation coefficient ρ of 0.500 ± 0.103 (p < 0.05) and 0.467 ± 0.059 (p < 0.05), respectively. This correlation analysis highlights the unique potential of NDE1 and CA19-9 in the context of iCCA.
Enhanced diagnostic performance of autoantibody biomarker signature for iCCA detection. (a) Intercorrelation among selected hits (feature-feature matrix) using Spearman’s rank correlation method. (b) The PCA biplot of the selected candidate biomarkers. The arrows represent vector projections, indicating the strength of influence of each biomarker on the principal component. The eclipses denote a 95% confidence interval. (c) The bar chart illustrates the AUCs for both the top 15 diagnostic biomarker signatures and the last 7 individual biomarkers, serving as a baseline for comparison. The combination 'NDE1 + PYCR1 + VIM + CA19-9' was chosen as the most optimal signature for iCCA diagnosis. (d) ROC curves compare the selected optimal signatures with and without CA19-9, including the 'CA19-9 only' model. Statistical significance is defined as a value of p ≤ 0.05.
To delve further into predictive potential, we employed PCA analysis. This revealed distinct patterns among study groups, as illustrated in PCA biplot (Fig. 4b). The CTR group formed a tight cluster, indicating shared biomarker profiles. Conversely, the iCCA group displayed more pronounced heterogeneity, indicating diverse biomarker expression among patients. A noticeable overlap between iCCA and CTR groups underscored shared biomarkers with distinct expression levels. Furthermore, the vectors representing NDE1, and CA19-9 were notably longer compared to those of the other candidates. They had the most influence on the principal component (PC) 2 and showed relatively weak correlations. In contrast, vectors representing KRT8, KRT19, PYCR1, FADD, and VIM had similar lengths and were more influential in PC1, showing a positive correlation among them. These findings offer compelling evidence for the uniqueness of NDE1 and CA19-9 as biomarkers specific to iCCA.
Improved diagnostic accuracy of combined CA19-9 and autoantibody panels
Our final list of candidate biomarkers includes six autoantibodies and the CA19-9 antigen. Since a single biomarker is rarely sufficient for iCCA diagnosis, we generated and evaluated a total of 120 potential sets of biomarkers from all possible combinations of our candidates to assess their predictive performance, including AUCs for iCCA vs CTR classification. Among 120 signatures, 118 outperformed the CA19-9 alone, with 19 of them achieving superior diagnostic capabilities, exhibiting AUCs exceeding 95% (Supplementary Table S4 online). The top 15 highest AUCs for these biomarker combinations are depicted in Fig. 4c, with individual biomarker AUCs serving as the baseline reference. We demonstrated that the combination of three autoantibodies (NDE1, PYCR1, and VIM) along with the CA19-9 marker exhibited optimal performance in distinguishing iCCA from CTR, achieving the second-highest AUC of 96.9% (95% CI 93.9–100.0%, Fig. 4c). This signature, while not having the highest AUC, is preferable due to the inclusion of our aforementioned unique NDE1 and CA19-9 candidates. It utilizes two optimal thresholds at 0.206 and 0.225, yielding a sensitivity of 100.0% and a specificity of 87.5%, as well as a sensitivity of 93.8% and a specificity of 93.8%, respectively (see Supplementary Fig. S6 and Table S4 online). In Fig. 4d, the selected signature demonstrated superior performance compared to two other scenarios: the combination without CA19-9 (NDE1 + PYCR1 + VIM), which had an AUC of 92.6% (95% CI 81.9–100.0%), and CA19-9 alone, which achieved an AUC of 83.8% (95% CI 66.9–100.0%) (see Supplementary Figs. S7 and S5 online, respectively). We also conducted PCA analysis to confirm our selected signature's (NDE1 + PYCR1 + VIM + CA19-9) diagnostic ability (details in Supplementary Fig. S8 online). The PCA revealed noticeable separation between iCCA and CTR along the principal components, indicating strong discriminatory power. In summary, these results reinforce the value of autoantibody biomarkers as diagnostic tools alongside the existing CA19-9 marker for iCCA.
Functional enrichment analysis of iCCA associated autoantibody-targeted antigens
To uncover the potential roles of iCCA associated autoantibody-targeted antigens in common pathways, we conducted ORA analysis using the STRING network. We implemented the initial set of 66 candidates identified through t-test analysis (FDR ≤ 0.1) in this analysis, as the final six were deemed insufficient for meaningful results. The analysis revealed statistically significant enrichment terms related to the structure and composition of the cell’s cytoskeleton, specifically fibers and filaments (Supplementary Table S5 online). The protein–protein interaction (PPI) network showed more interactions than expected by chance, supported by a statistically significant PPI enrichment p-value of 0.0396. This indicates that the observed connections between submitted autoantibody-targeted antigens were important, providing insights into iCCA-related molecular interactions (Fig. 5a).
Functional enrichment results of proteins targeted by iCCA autoantibodies. (a) The PPI network of ORA analysis reveals twelve enriched clusters identified by the MCL clustering algorithm using STRING database. Among these clusters, eight (indicated by enclosed dotted lines and cluster numbers) exhibit statistically significant pathways (FDR ≤ 0.05). Nodes represent proteins, edges depict interactions, and line thickness indicates the degree of confidence in the prediction. Solid lines represent interactions supported by stronger or more direct evidence. Dashed lines indicate interactions supported by indirect or lower confidence evidence and represent inter-cluster edges. Clusters are color-coded for clarity. (b) Running score and pre-ranked list of GSEA results demonstrate five statistically significant GO terms with levels of significance: FDRs < 0.05. Statistical significance is defined as a value of FDR ≤ 0.05.
We further performed cluster analysis on the PPI network using the MCL technique, revealing 12 statistically significant clusters, each with a PPI enrichment p-value ≤ 0.05. Each cluster consisted of a distinctive set of at least two proteins, with a maximum of 7 proteins per cluster. Eight of these clusters showed enrichment in at least one statistically significant term (FDRs < 0.05) from the STRING database. Among these, Clusters 1, 5, and 7 were notable for their associations with metabolic pathways, including fructose and mannose metabolism, insulin regulation, and cholesterol biosynthesis. Conversely, clusters 2, 3 and 8 focused on proteins related to the cellular response mechanisms and the dynamic regulation of gene expression through transcriptional, epigenetic mechanisms, and protein synthesis. Finally, clusters 4 and 6 centered around proteins involved in cell structure and the cytoskeleton. These results highlight the intricate interconnections between metabolic pathways, cellular regulation, and cell structure (Fig. 5a and Supplementary Table S6 online). Additionally, GSEA analysis using the GO database identified five enriched functional annotations associated with iCCA (Fig. 5b). In short, these cellular processes are primarily involved in the organization of intermediate filaments and the formation of protein complexes. These processes play crucial roles in maintaining cellular structure, regulating signaling pathways, and controlling cell proliferation. This outcome is in coherence with the ORA analysis, reinforcing the significance of the identified processes in the context of iCCA development.
Discussion
Advanced iCCA poses a remarkable challenge as there are no curative treatments available, except for early surgical resection1. However, the standard diagnostic approaches for iCCA, which include CT, MRI, PET scans, and pathological examination, have limitations, including issues related to accuracy and potential complications10. In this context, CA19-9 has gained widespread attention as a biomarker for iCCA, but it exhibits only modest sensitivity and specificity7. A meta-analysis of 31 studies, each with varying control groups and cut-off values, revealed that CA19-9 has an overall pooled sensitivity of 72% (ranging from 38 to 100%) and a specificity of 84% (ranging from 31.35 to 100%), with an overall AUC of 83% for diagnosing CCA7. Our findings align well with these ranges, particularly when considering studies with healthy control groups similar to our study. Although CA19-9 has utility, its sensitivity could be improved, especially for early iCCA detection, where high sensitivity is critical for preventing false negatives and improving treatment outcomes. This highlights the continued necessity for the development of more reliable non-invasive diagnostic biomarkers for iCCA screening.
Our research has uncovered a promising array of autoantibody biomarker candidates, revealing 19 potential signatures with remarkable AUC values exceeding 95%. On average, these signatures demonstrated a sensitivity of 92.9%, a specificity of 87.3%, and an AUC of 95.9%, surpassing the reported values for CA19-9 alone7. These findings highlighted the potential of these signatures for iCCA diagnosis within our Thai cohort. However, it is worth noting the observed similarity in performance among these signatures, signifying their equal discriminatory power between iCCA and CTR. This likely stems from iCCA's inherent heterogeneity, making it challenging to pinpoint one diagnostic signature that can fully capture iCCA's diverse nature. iCCA's heterogeneity may result from factors like distinct molecular subtypes17 and/or varying disease progression among patients. Consequently, no specific set of biomarkers consistently outperforms others, as what works for one subgroup may not work as effectively for another due to their differences. Nonetheless, our focus is on effectively distinguishing iCCA from non-cancer individuals rather than classifying subtypes or states of the disease, which falls outside the scope of this research.
We also observed that out of the 66 differentially expressed autoantibodies we identified, the majority (89.4%) target intracellular antigens. Specifically, 59.1% target cytoplasmic proteins, 30.3% target nuclear proteins, and the remaining 10.6% target secreted and cell membrane proteins. These intracellular antigens, particularly nuclear proteins, are typically shielded from immune surveillance and are not involved in the negative selection process of B-cell maturation37. We speculate that their presence may result from the release of intracellular contents due to apoptotic or necrotic cells in cancer, potentially leading to an augmented immune response characterized by increased production of autoantibodies against them38. The analysis combining ORA with the MCL technique has revealed that our autoantibody-targeted antigens are linked to general cancer pathways shared across diverse cancer types. For example, metabolic reprogramming is a shared feature in many cancers, providing the energy and building blocks essential for cancer cell growth39. Cellular response mechanisms likely involve signaling cascades that contribute to cancer cell survival, proliferation, and evasion of the immune system40. However, the most prominent pathways, as confirmed by GSEA, are related to changes in cell structure (e.g., intermediate filament and cytoskeleton fiber), suggesting potential adaptations in cellular organization linked to cell motility, which is involved in cancer metastasis41. Thus, targeting these pathways for the development of biomarkers and therapies could be a promising approach to enhance diagnostic accuracy and potentially impede disease progression, thereby improving treatment effectiveness and clinical outcomes.
Two key structural proteins, KRT8 (CK8/K8) and KRT19 (CK19/K19), though not part of our final biomarker signature, show a significant increase in autoantibodies, hinting at potential clinical relevance for iCCA. As intermediate filament proteins, these two have already been extensively studied and proposed as potential therapeutic or biomarker candidates in various reports42,43,44,45. KRT8 has shown promise as a pan-cancer early biomarker in a comprehensive study involving over 17,000 samples, exhibiting significant overexpression in various cancers46. Meanwhile, KRT19, commonly utilized in immunohistochemical analysis, aids in distinguishing iCCA (K19 +) from hepatocellular carcinoma or HCC (K19-), reflecting its prevalence in iCCA47. Reports have also shown its association with poor prognosis in both iCCA and HCC45,48. Despite their roles in maintaining the structural integrity of epithelial cells49, the increased autoantibody levels against these keratins suggest possible abnormalities in their expression or modification in iCCA cells, which in turn may impact the structural stability of these cells. The continuous phosphorylation of keratin can prompt the reorganization of the keratin network in cells, resulting in the degradation of keratin structure, a characteristic of the Epithelial-Mesenchymal Transition (EMT) that has been associated with the aggressiveness, invasion, and metastasis of tumors49. Consequently, a plausible hypothesis emerges: in iCCA, EMT initiation might be triggered by the hyperphosphorylation of keratins. Thus, during apoptosis in cancer, these hyperphosphorylated keratins could stimulate an amplified immune response, evident in increased autoantibody production against them. With this knowledge, modulating keratin phosphorylation and reorganization represents a potential novel approach to control EMT and metastasis of iCCA. Notably, using kinase inhibitors tailored to keratins like KRT8, KRT19, and KRT15 may reduce their phosphorylation, thereby suppressing EMT and improving clinical outcomes49.
Two key candidates from our final autoantibody signature, VIM and NDE1, are also associated with EMT and cell migration processes50,51. Especially, VIM (vimentin), an intermediate filament protein, is known to induce EMT, thereby promoting metastasis50. It emerges as a potential therapeutic or biomarker candidate in various cancer types50. Abnormal VIM expression in iCCA is linked to poor prognosis and decreased overall survival rates, signifying a more aggressive tumor phenotype52. For NDE1, existing research predominantly focuses on its role in microtubule organization, mitosis, cell migration, and neuronal development51. Despite limited attention in the cancer field, NDE1 does hold potential relevance to cancer mechanisms. NDE1 regulates cell division and the cell cycle, interacting with proteins like LIS1 and dynein to control microtubule dynamics and ensure proper mitotic spindle formation51,53. Therefore, we hypothesize that aberrant NDE1 may contribute to mitotic defects, potentially resulting in chromosome aneuploidy54, and genomic instability55—characteristics frequently observed in cancer cells56. Furthermore, since both NDE1 and VIM are associated with cell migration and invasion50,51, their elevated autoantibodies might suggest a potential role in promoting EMT, contributing to the metastatic behavior of iCCA cells. Importantly, the novel status of NDE1 in iCCA, unexplored as a cancer biomarker or therapeutic target, holds transformative potential for future intervention and diagnosis.
The final candidate in our selected signature, PYCR1, exhibits increased expression across diverse malignancies and correlates with poor clinical outcomes57. Moreover, it is overexpressed in cancer-associated fibroblasts (CAFs), non-tumor cells implicated in regulating the tumor microenvironment by supporting collagen production for the extracellular matrix (ECM), which fuels cancer cell growth and metastatic dissemination58. It is known that iCCA tissue is recognized for its abundance of desmoplastic stroma, including CAFs, and the demonstrated crosstalk between them plays a pivotal role in tumor growth and development59. In our study, the elevated PYCR1 autoantibody level might suggest potential aberration in PYCR1 expression or modification in iCCA or CAF cells, contributing to tumor growth and metastatic spread. Despite unclear links between PYCR1 and iCCA progression, its role in remodeling the tumor microenvironment proposes a plausible association.
Finally, FADD, despite not being included in our final biomarker signature, is still regarded as one of our final candidates and warrants discussion. Prior research has shown that FADD, serving as a ubiquitous adaptor protein, actively participates in and modulates various signaling complexes such as necrosomes, endosomes, and inflammasomes60. Therefore, FADD holds a crucial role in apoptosis, inflammation, innate immunity, and carcinogenesis60. Reports have hinted at FADD's upregulation linked to tumor progression and poor prognoses in specific cancer cases61,62,63. A pan-cancer analysis indicated that FADD was highly expressed in CCA, showing exceptional diagnostic performance with an AUC of 94%64. The increased presence of FADD autoantibodies in our study suggests a potential abnormality in FADD expression, indicating its possible involvement in iCCA development. We speculate that this aberration could disrupt apoptotic signaling, allowing abnormal cells to evade cell death mechanisms. Additionally, FADD's role in inflammation might further contribute to iCCA progression by influencing the tumor microenvironment and promoting inflammation-related mechanisms.
These promising autoantibodies not only capture the diverse molecular landscape of iCCA but also pave the way for personalized therapeutic strategies. However, while elevated levels of autoantibodies might suggest their involvement, the precise mechanisms linking them to iCCA remain unclear. Autoantibody responses can be complex and may not always directly reflect the functions of the targeted proteins. Additionally, our study arrays exclusively focused on protein antigens, which cannot detect autoantibodies targeting non-protein antigens like CA19-9, a carbohydrate antigen. Although the literature does not provide significant evidence of autoantibodies specifically targeting CA19-9 in iCCA, the immune system could produce autoantibodies against such non-protein molecules. This suggests that our study may not capture the complete autoantibody landscape in iCCA. Future studies should incorporate platforms detecting autoantibodies against a broader range of antigens, including non-protein molecules, to provide a more comprehensive understanding of the autoantibody profile in iCCA.
Importantly, our study is also constrained by a small sample size with a specific focus on the Thai population. To address these limitations and validate the clinical applications of our findings, larger studies involving iCCA from diverse populations are essential. Moreover, while our study utilized mRNA expression data to validate candidate autoantigens, we acknowledge that protein expression analysis would provide a more direct assessment of antigen levels in tumor versus non-tumor tissues. Future research should incorporate protein expression studies to validate and extend our findings, thereby providing a more comprehensive understanding of the candidate autoantigens' roles in disease. These future investigations will elucidate the roles of autoantibodies in iCCA, ensuring their efficacy across a broader spectrum of cases and facilitating a comprehensive assessment of diagnostic sensitivity, specificity, and reliability.
In summary, our study emphasizes the significance of autoantibody biomarkers as valuable complements to the existing tools in the diagnosis of iCCA. The combination of an autoantibody signature panel, comprising NDE1, PYCR1, and VIM, along with the conventional biomarker CA19-9, holds the potential to improve diagnostic accuracy for iCCA.
Data availability
The immunome data supporting the findings of this study are not openly available due to the absence of a suitable repository but can be obtained from the corresponding author upon reasonable request. The transcriptome data supporting the findings can be found on Gene Expression Omnibus database under accession number GSE76297 and are available at the following URL: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE76297.
References
Chung, T. & Park, Y. N. Up-to-date pathologic classification and molecular characteristics of intrahepatic cholangiocarcinoma. Front. Med. 9, 857140. https://doi.org/10.3389/fmed.2022.857140 (2022).
Treeprasertsuk, S. et al. A significant cancer burden and high mortality of intrahepatic cholangiocarcinoma in Thailand: A nationwide database study. BMC Gastroenterol. 17, 3. https://doi.org/10.1186/s12876-016-0565-6 (2017).
Bridgewater, J. et al. Guidelines for the diagnosis and management of intrahepatic cholangiocarcinoma. J. Hepatol. 60, 1268–1289. https://doi.org/10.1016/j.jhep.2014.01.021 (2014).
Howlader, N. et al. SEER cancer statistics review, 1975–2017, based on November 2019 SEER data submission, Section: 14 Liver and bile duct, Table 14.18 (Surveillance Research Program, National Cancer Institute, 2020). https://seer.cancer.gov/csr/1975_2017/
Razumilava, N. & Gores, G. J. Classification, diagnosis, and management of cholangiocarcinoma. Clin. Gastroenterol. Hepatol. 11, 13-21.e11. https://doi.org/10.1016/j.cgh.2012.09.009 (2013).
Cerrito, L. et al. Contrast-enhanced imaging in the management of intrahepatic cholangiocarcinoma: State of art and future perspectives. Cancers (Basel) 15, 3393. https://doi.org/10.3390/cancers15133393 (2023).
Liang, B. et al. Diagnostic accuracy of serum CA19-9 in patients with cholangiocarcinoma: A systematic review and meta-analysis. Med. Sci. Monit. 21, 3555–3563. https://doi.org/10.12659/msm.895040 (2015).
Banales, J. M. et al. Expert consensus document: Cholangiocarcinoma: Current knowledge and future perspectives consensus statement from the European Network for the Study of Cholangiocarcinoma (ENS-CCA). Nat. Rev. Gastroenterol. Hepatol. 13, 261–280. https://doi.org/10.1038/nrgastro.2016.51 (2016).
Rizvi, S., Khan, S. A., Hallemeier, C. L., Kelley, R. K. & Gores, G. J. Cholangiocarcinoma—Evolving concepts and therapeutic strategies. Nat. Rev. Clin. Oncol. 15, 95–111. https://doi.org/10.1038/nrclinonc.2017.157 (2018).
Patel, T. Cholangiocarcinoma—Controversies and challenges. Nat. Rev. Gastroenterol. Hepatol. 8, 189–200. https://doi.org/10.1038/nrgastro.2011.20 (2011).
Fernández Madrid, F. Autoantibodies in breast cancer sera: Candidate biomarkers and reporters of tumorigenesis. Cancer Lett. 230, 187–198. https://doi.org/10.1016/j.canlet.2004.12.017 (2005).
Pedersen, J. W. & Wandall, H. H. Autoantibodies as biomarkers in cancer. Lab. Med. 42, 623–628. https://doi.org/10.1309/lm2t3ou3rzrthksn (2011).
Zaenker, P., Gray, E. S. & Ziman, M. R. Autoantibody production in cancer—The humoral Immune response toward autologous antigens in cancer patients. Autoimmun. Rev. 15, 477–483. https://doi.org/10.1016/j.autrev.2016.01.017 (2016).
Zaenker, P. & Ziman, M. R. Serologic autoantibodies as diagnostic cancer biomarkers—A review. Cancer Epidemiol. Biomark. Prev. 22, 2161–2181. https://doi.org/10.1158/1055-9965.EPI-13-0621 (2013).
Blackburn, J. M. & Shoko, A. Protein function microarrays for customised systems-oriented proteome analysis. Methods Mol. Biol. 785, 305–330. https://doi.org/10.1007/978-1-61779-286-1_21 (2011).
Blackburn, J. M. & Hart, D. J. Fabrication of protein function microarrays for systems-oriented proteomic analysis. Methods Mol. Biol. 310, 197–216. https://doi.org/10.1007/978-1-59259-948-6_14 (2005).
Chaisaingmongkol, J. et al. Common molecular subtypes among Asian hepatocellular carcinoma and cholangiocarcinoma. Cancer Cell 32, 57–7053. https://doi.org/10.1016/j.ccell.2017.05.009 (2017).
Adeola, H. A., Smith, M., Kaestner, L., Blackburn, J. M. & Zerbini, L. F. Novel potential serological prostate cancer biomarkers using CT100+ cancer antigen microarray platform in a multi-cultural South African cohort. Oncotarget 7, 13945–13964. https://doi.org/10.18632/oncotarget.7359 (2016).
Sumera, A. et al. A novel method to identify autoantibodies against putative target proteins in serum from beta-thalassemia major: A pilot study. Biomedicines https://doi.org/10.3390/biomedicines8050097 (2020).
Duarte, J., Serufuri, J. M., Mulder, N. & Blackburn, J. Protein function microarrays: Design, use and bioinformatic analysis in cancer biomarker discovery and quantitation. In Bioinformatics of Human Proteomics (ed. Wang, X.) 39–74 (Springer, Netherlands, 2013).
Welch, B. L. The generalization of ‘Student’s’ problem when several different population variances are involved. Biometrika 34, 28–35. https://doi.org/10.1093/biomet/34.1-2.28 (1947).
Meyer, B. et al. Identification of DNA methylation biomarkers with potential to predict response to neoadjuvant chemotherapy in triple-negative breast cancer. Clin. Epigenet. 13, 226. https://doi.org/10.1186/s13148-021-01210-6 (2021).
R: A language and environment for statistical computing v. 4.3.1 (R Foundation for Statistical Computing, 2023). https://www.R-project.org/.
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849. https://doi.org/10.1093/bioinformatics/btw313 (2016).
Kuhn, M. Building predictive models in R using the caret package. J. Stat. Softw. 28, 1–26. https://doi.org/10.18637/jss.v028.i05 (2008).
Robin, X. et al. pROC: An open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinform. 12, 77. https://doi.org/10.1186/1471-2105-12-77 (2011).
Breiman, L. Random forests. In Machine Learning Vol. 45 (ed. Schapire, R. E.) 5–32 (Kluwer Academic Publishers, 2001).
Webb, G. I. Naïve Bayes. In Encyclopedia of Machine Learning (eds Sammut, C. & Webb, G. I.) 713–714 (Springer US, 2010).
Friedman, J. H. Regularized discriminant analysis. J. Am. Stat. Assoc. 84, 165–175. https://doi.org/10.2307/2289860 (1989).
Chang, C.-C. & Lin, C.-J. LIBSVM: A library for support vector machines. ACM Trans. Intell. Syst. Technol. 2, 27. https://doi.org/10.1145/1961189.1961199 (2011).
Fix, E. & Hodges, J. L. Discriminatory analysis. Nonparametric discrimination: Consistency properties. Int. Stat. Rev. 57, 238–247. https://doi.org/10.2307/1403797 (1989).
Marron, J. S., Todd, M. J. & Ahn, J. Distance-weighted discrimination. J. Am. Stat. Assoc. 102, 1267–1271. https://doi.org/10.1198/016214507000001120 (2007).
Gelman, A., Jakulin, A., Pittau, M. G. & Su, Y.-S. A weakly informative default prior distribution for logistic and other regression models. Ann. Appl. Stat. 2, 1360–1383. https://doi.org/10.1214/08-AOAS191 (2008).
Szklarczyk, D. et al. The STRING database in 2023: Protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 51, D638-d646. https://doi.org/10.1093/nar/gkac1000 (2023).
Dongen, S. Graph clustering via a discrete uncoupling process. SIAM J. Matrix Anal. Appl. 30, 121–141. https://doi.org/10.1137/040608635 (2008).
Wu, T. et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation (Camb.) 2, 100141. https://doi.org/10.1016/j.xinn.2021.100141 (2021).
Giltiay, N. V., Chappell, C. P. & Clark, E. A. B-cell selection and the development of autoantibodies. Arthritis Res. Ther. 14, S1. https://doi.org/10.1186/ar3918 (2012).
Gamrekelashvili, J., Greten, T. F. & Korangy, F. Immunogenicity of necrotic cell death. Cell. Mol. Life Sci. 72, 273–283. https://doi.org/10.1007/s00018-014-1741-x (2015).
Phan, L. M., Yeung, S. C. & Lee, M. H. Cancer metabolic reprogramming: Importance, main features, and potentials for precise targeted anti-cancer therapies. Cancer Biol. Med. 11, 1–19. https://doi.org/10.7497/j.issn.2095-3941.2014.01.001 (2014).
Payne, K. K. Cellular stress responses and metabolic reprogramming in cancer progression and dormancy. Semin. Cancer Biol. 78, 45–48. https://doi.org/10.1016/j.semcancer.2021.06.004 (2022).
Ong, M. S. et al. Cytoskeletal proteins in cancer and intracellular stress: A therapeutic perspective. Cancers (Basel) 12, 238. https://doi.org/10.3390/cancers12010238 (2020).
Akiba, J. et al. The expression of arginase-1, keratin (K) 8 and K18 in combined hepatocellular-cholangiocarcinoma, subtypes with stem-cell features, intermediate-cell type. J. Clin. Pathol. 69, 846–851. https://doi.org/10.1136/jclinpath-2015-203491 (2016).
Komuta, M. et al. Clinicopathological study on cholangiolocellular carcinoma suggesting hepatic progenitor cell origin. Hepatology 47, 1544–1556. https://doi.org/10.1002/hep.22238 (2008).
Cogliati, B. et al. Identification of hepatic stem/progenitor cells in canine hepatocellular and cholangiocellular carcinoma. Vet. Comp. Oncol. 8, 112–121. https://doi.org/10.1111/j.1476-5829.2010.00210.x (2010).
Govaere, O. et al. Keratin 19: A key role player in the invasion of human hepatocellular carcinomas. Gut 63, 674–685. https://doi.org/10.1136/gutjnl-2012-304351 (2014).
Scott, M. K. D. et al. A multi-scale integrated analysis identifies KRT8 as a pan-cancer early biomarker. Pac. Symp. Biocomput. 26, 297–308. https://doi.org/10.1142/9789811232701_0028 (2021).
Komuta, M. Histological heterogeneity of primary liver cancers: Clinical relevance, diagnostic pitfalls and the pathologist’s role. Cancers (Basel) 13, 2871. https://doi.org/10.3390/cancers13122871 (2021).
Uenishi, T. et al. Serum cytokeratin 19 fragment (CYFRA21-1) as a prognostic factor in intrahepatic cholangiocarcinoma. Ann. Surg. Oncol. 15, 583–589. https://doi.org/10.1245/s10434-007-9650-y (2008).
Kim, H. J., Choi, W. J. & Lee, C. H. Phosphorylation and reorganization of keratin networks: Implications for carcinogenesis and epithelial mesenchymal transition. Biomol. Ther. (Seoul) 23, 301–312. https://doi.org/10.4062/biomolther.2015.032 (2015).
Satelli, A. & Li, S. Vimentin in cancer and its potential as a molecular target for cancer therapy. Cell. Mol. Life Sci. 68, 3033–3046. https://doi.org/10.1007/s00018-011-0735-1 (2011).
Wynshaw-Boris, A. Lissencephaly and LIS1: Insights into the molecular mechanisms of neuronal migration and development. Clin. Genet. 72, 296–304. https://doi.org/10.1111/j.1399-0004.2007.00888.x (2007).
Nakajima, T. et al. Intrahepatic cholangiocarcinoma with sarcomatous change: Clinicopathologic and immunohistochemical evaluation of seven cases. Cancer 72, 1872–1877. https://doi.org/10.1002/1097-0142(19930915)72:6%3c1872::aid-cncr2820720614%3e3.0.co;2-n (1993).
McKenney, R. J., Vershinin, M., Kunwar, A., Vallee, R. B. & Gross, S. P. LIS1 and NudE induce a persistent dynein force-producing state. Cell 141, 304–314. https://doi.org/10.1016/j.cell.2010.02.035 (2010).
Aumais, J. P. et al. Role for NudC, a dynein-associated nuclear movement protein, in mitosis and cytokinesis. J. Cell Sci. 116, 1991–2003. https://doi.org/10.1242/jcs.00412 (2003).
Houlihan, S. L. & Feng, Y. The scaffold protein Nde1 safeguards the brain genome during S phase of early neural progenitor differentiation. Elife 3, e03297. https://doi.org/10.7554/eLife.03297 (2014).
Schvartzman, J. M., Sotillo, R. & Benezra, R. Mitotic chromosomal instability and cancer: Mouse modelling of the human disease. Nat. Rev. Cancer 10, 102–115. https://doi.org/10.1038/nrc2781 (2010).
D’Aniello, C., Patriarca, E. J., Phang, J. M. & Minchiotti, G. Proline metabolism in tumor growth and metastatic progression. Front. Oncol. 10, 776. https://doi.org/10.3389/fonc.2020.00776 (2020).
Kay, E. J., Zanivan, S. & Rufini, A. Proline metabolism shapes the tumor microenvironment: From collagen deposition to immune evasion. Curr. Opin. Biotechnol. 84, 103011. https://doi.org/10.1016/j.copbio.2023.103011 (2023).
Montori, M. et al. Cancer-associated fibroblasts in cholangiocarcinoma: Current knowledge and possible implications for therapy. J. Clin. Med. https://doi.org/10.3390/jcm11216498 (2022).
Mouasni, S. & Tourneur, L. FADD at the crossroads between cancer and inflammation. Trends Immunol. 39, 1036–1053. https://doi.org/10.1016/j.it.2018.10.005 (2018).
Shimada, K. et al. Phosphorylation of FADD is critical for sensitivity to anticancer drug-induced apoptosis. Carcinogenesis 25, 1089–1097. https://doi.org/10.1093/carcin/bgh130 (2004).
Matsuyoshi, S., Shimada, K., Nakamura, M., Ishida, E. & Konishi, N. FADD phosphorylation is critical for cell cycle regulation in breast cancer cells. Br. J. Cancer 94, 532–539. https://doi.org/10.1038/sj.bjc.6602955 (2006).
Schrijvers, M. L. et al. FADD expression as a prognosticator in early-stage glottic squamous cell carcinoma of the larynx treated primarily with radiotherapy. Int. J. Radiat. Oncol. Biol. Phys. 83, 1220–1226. https://doi.org/10.1016/j.ijrobp.2011.09.060 (2012).
Wang, C., Jiang, X., Zhao, Q., Xie, Z. & Cai, H. The diagnostic or prognostic values of FADD in cancers based on pan-cancer analysis. Biomed. Rep. 19, 77. https://doi.org/10.3892/br.2023.1659 (2023).
Acknowledgements
We express our gratitude to the TIGER-LC Consortium members, as well as the patients and their families who generously contributed to this study.
Funding
This work was supported in part by the grant from Center of Excellence on Environmental Health and Toxicology (EHT), OPS, Ministry of Higher Education, Science, Research and Innovation, and Thailand Science Research and Innovation (TSRI), Chulabhorn Research Institute (Grant Nos. 48292/4691968 and 2536699/42113). X.W.W. was supported by the intramural program of the Center for Cancer Research, National Cancer Institute, U.S.A. W.K. was supported by Chulabhorn Graduate Scholarship (CGS2562/01).
Author information
Authors and Affiliations
Contributions
Conceptualization, M.R., M.F.; Methodology, W.K., J.C., Y.P., P.T., X.W.W., M.F.; Investigation, W.K.; Data curation: W.K.; Formal analysis: W.K.; Visualization, W.K.; Software, W.K., Y.P.; Writing—original draft, W.K.; Writing—review and editing, W.K., J.C., Y.P., P.T., X.W.W., M.R., M.F.; Supervision, M.R., M.F.; Funding acquisition, M.R., M.F.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kajornsrichon, W., Chaisaingmongkol, J., Pomyen, Y. et al. Identification of autoantibodies as potential non-invasive biomarkers for intrahepatic cholangiocarcinoma. Sci Rep 14, 20012 (2024). https://doi.org/10.1038/s41598-024-70595-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-70595-0
- Springer Nature Limited
Keywords
We’re sorry, something doesn't seem to be working properly.
Please try refreshing the page. If that doesn't work, please contact support so we can address the problem.