Abstract
Guava (Psidium guajava L.) is a semi-domesticated fruit tree of moderate importance in the Neotropics, utilized for millennia due to its nutritional and medicinal benefits, but its origin of domestication remains unknown. In this study, we examine genetic diversity and population structure in 215 plants from 11 countries in Mesoamerica, the Andes, and Amazonia using 25 nuclear microsatellite loci to propose an origin of domestication. Genetic analyses reveal one gene pool in Mesoamerica (Mexico) and four in South America (Brazilian Amazonia, Peruvian Amazonia and Andes, and Colombia), indicating greater differentiation among localities, possibly due to isolation between guava populations, particularly in the Amazonian and Andean regions. Moreover, Mesoamerican populations show high genetic diversity, with moderate genetic structure due to gene flow from northern South American populations. Dispersal scenarios suggest that Brazilian Amazonia is the probable origin of guava domestication, spreading from there to the Peruvian Andes, northern South America, Central America, and Mexico. These findings present the first evidence of guava domestication in the Americas, contributing to a deeper understanding of its evolutionary history.
Similar content being viewed by others
Introduction
In the Neotropics, there are approx. 8200 species either managed or domesticated to various degrees in Mesoamerica, the Andes, and the lowlands of South America, the majority of which are perennial1. This frequent use of long-lived species is due to their many valuable products such as roots, fleshy or starchy fruits, nuts, fibers, and oils1,2,3,4, which provide significant quantities of macro and micronutrients2. However, only a few of these perennial crops are known in international markets, and the majority are still cultivated mainly as subsistence crops for local consumption and sale2,5. Particularly in the Neotropics, fleshy fruits have been an essential dietary component of numerous human groups in pre- and post-Columbian times6,7,8. However, few studies on their genetic diversity and population structure have been performed, with some work on Annona cherimola Mill.9, Bactris gasipaes Kunth10,11, Chrysophyllum cainito L.12,13, Carica papaya L.14, Spondias purpurea L.15, and Theobroma cacao L.16,17,18. To bridge this knowledge gap and increase the number of Neotropical fruit species studied we conducted an analysis of the genetic diversity and population structure of guava.
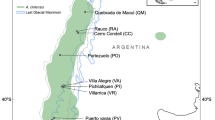
Guava (Psidium guajava L.) is a Neotropical semi-domesticated fruit tree species of some importance in the Americas and elsewhere19,20. It is distributed from Mexico and the Antilles to northwestern Argentina21. The fruit is the most used part of the plant, consumed fresh or used to make candies, dried fruits, jams, jellies, juices, pastes, soup bases, and syrup21,22. It is a good source of calcium, iron, niacin, pantothenic acid, phosphorus, riboflavin, and thiamine23. In folk medicine, guava is used to treat respiratory discomfort, gastrointestinal problems and help to expel the placenta after childbirth23,24. Guava grows in tropical dry forests and savannah-like vegetation, as well as in disturbed areas (roadsides and grasslands), small agroecological environments (homegardens and orchards), and larger-scale production systems21,22. It adapts easily to different rainfall conditions and soil types but does not tolerate flooded soils and is sensitive to low temperatures25.
During post-Columbian times, guava was the fruit tree most widely recorded by European chroniclers of the sixteenth century, who documented its presence in Mesoamerica and South America in both reputably wild and cultivated populations26. The European conquerors learned to use guava fruits and leaves as medicine and food8, which prevail among indigenous peoples until now. The oldest archaeological record of macro remains place guava in pre-Columbian contexts in Southwestern Amazonia (dates between 9490 and 6505 calibrated years before present [cal. BP])27 and the human settlements of the Peruvian coast (7000 cal. BP)28. The earliest macro remains (fruit fragments) found in Mexico are much more recent, dating to ca. 670 cal. BP29.
Despite its cultural, economic, and historical importance, guava has received little attention from geneticists30. Perennial trees are often propagated asexually3,4,31, which results in a reduction of sexual reproduction4,32 and, therefore, slower rates of evolution and less pronounced changes in domestication syndrome traits33,34. Guava wild populations are very hard to distinguish from tolerated or feral individuals that may form small populations (personal observations). Without a clearly defined wild ancestor, it is difficult to identify centers of origin of domestication, quantify changes due to human selection and trace routes of human-mediated dispersal. Our study aims to characterize the genetic diversity and population structure of guava across parts of its Neotropical distribution using SSR markers, looking for answers to the following questions: (a) What is the level of genetic variation among the sampled genotypes of guava? (b) How is this diversity structured? (c) Is there isolation by distance between locations? (d) Where is the most likely origin of domestication? (e) What is the history of dispersal of this species?
Results
Null alleles, Hardy–Weinberg equilibrium, and linkage disequilibrium
We removed 18 samples and discarded the locus mPgCIR08 which had more than 40% of missing data, leaving 197 samples and 24 loci for further analysis. We found no evidence of null alleles in our data set. All loci showed significant deviations from Hardy–Weinberg equilibrium (HWE) and linkage disequilibrium (LD) in more than one population (Supplementary Tables 1, 2). Because the loci in HWE and LD were not the same for all localities, all markers were retained for further analyses. The genotype accumulation curve (Supplementary Fig. 1) shows that the set of loci tested had sufficient power to discriminate between individuals. The curve revealed that 100% of the genotypes could be detected with 12 markers, hence the loci accurately estimated the diversity of our sample.
Genetic diversity and genetic differentiation
The PCA provided evidence of genetic structure of guava across its geographical range. The first two principal components explained 18.5% of the total variation (Fig. 1a). Amazonian guavas from Brazil and Peru (BRA-AM and PER-AM, respectively) formed well-defined clusters. In contrast, guava samples from Colombia (COL) and Venezuela (VEN) overlap with those from the Antilles (ANT), documenting the close relationship between these regions. Given that the centroids of the Colombian and Venezuelan clusters do not co-occur within their respective standard deviation ellipses, we decided to define the Colombian (COL) and Venezuela-Antilles (VEN-ANT) clusters separately. Similarly, we defined a Peruvian Andes (PER-AND) cluster, Central American (CenAme) cluster and a Mexican (MEX) cluster. We decided to discard the samples from southern Brazil (BRA-SP), because the origin of these samples is uncertain (Fig. 1a). Therefore, for subsequent analyses, we used 192 samples.
(a) Principal component analysis (PCA) of microsatellite genotype data from P. guajava individuals showing the clustering along principal component axis 1–2. (b) Discriminant analysis of principal components (DAPC) for eight guava localities. Localities: MEX (Mexico), CenAme (Central America), ANT (The Antilles), VEN (Venezuela), COL (Colombia), BRA-SP (São Paulo, Brazil), BRA-AM (Brazilian Amazonia), PER-AM (Peruvian Amazonia), PER-AND (Peruvian Andes).
The DAPC analysis with 7 defined groups from the results of the PCA revealed a clear differentiation of guavas from Brazilian and Peruvian Amazonia and the Peruvian Andes (Fig. 1b). These results were consistent with the patterns obtained using PCA but with far clearer differentiations among clusters. Likewise, the Mesoamerican (MEX + CenAme) and the northern South American clusters (COL + VEN-ANT) appeared as admixed groups (Fig. 1b). We performed a second DAPC analysis excluding the Peruvian and Brazilian well-differentiated groups to depict the relationships among Mesoamerican and northern South American groups. This analysis showed that VEN-ANT cluster was well differentiated from COL, CenAme and MEX, which are more closely related (Supplementary Fig. 2).
The Mexican and Central American localities showed the highest HE values, while the lowest HE values were found in the Peruvian Andes and Peruvian Amazonia (Table 1). When considering all the samples, guava showed high values of total and unbiased expected heterozygosity, and low values of average observed heterozygosity (HO) (Table 1). The results for each locality maintain this pattern. The inbreeding coefficient showed high values for most of the localities (Table 1).
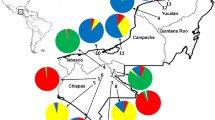
The results of the STRUCTURE analysis were consistent with the results from PCA and DAPC. Evanno and Jane's methods indicated an optimal value of K = 3 and K = 5 as the most likely numbers of genetic clusters (Supplementary Fig. 3). In K = 3, three genetic clusters are probable. Cluster 1 is predominant in Mexico, strongly represented in Central America, moderately represented in Colombia, and a minor component of Venezuela and the Antilles (Fig. 2). Cluster is 2 predominant in Brazilian Amazonia and a relatively minor component of all other localities. The Peruvian Andes, the only locality with 100% of the plants fully attributed to cluster 3 (Fig. 2), was strongly represented in Peruvian Amazonia, Colombia, Venezuela, and the Antilles, and present in Brazilian Amazonia, Central America, and Mexico. At K = 4, although it is not an optimal value according to Evanno and Jane's methods, a new genetic cluster appears in Colombia, foreshadowing K = 5 (Fig. 2). In K = 5, each of the five clusters is dominant in a separate locality: (1) Peruvian Amazonia; (2) Brazilian Amazonia; (3) Peruvian Andes; (4) Colombia; (5) Mexico. Venezuela and the Antilles are mixtures of Peruvian Amazonia and Colombia, while Central America is a mixture of Peruvian Amazonia, Colombia, and Mexico (Fig. 2).
Assignment probabilities of each of the 192 guava samples to each cluster inferred by STRUCTURE for K = 3, 4, and 5. Each sample is represented by a vertical bar, and color indicates the probability of belonging to each cluster. Samples are ordered according to the geographic region from southern to northern parts of the Americas.
This sequence of clusters from K = 3 to K = 5 suggests that western South America (BRA-AM → PER-AM → PER-AND) contains the origin of domesticated guava, given the number of clusters dominated by distinct genetic groups. From western South America, guava was then dispersed northward through Colombia to Central America and Mexico. Colombia is also the crossroads for the guava that arrived in Venezuela and later the Antilles. The fact that Mexico is always a clear cluster suggests that dispersal happened long enough ago for the thorough mixing of origins that became a distinct genetic group.
Estimates of Wright's F among the sampling localities indicated that guava diversity is higher within localities than among localities; nevertheless, we found intermediate levels of genetic differentiation with FST = 0.21 (Supplementary Table 3). FIT (0.64) and FIS (0.54) estimates were higher compared to FST (Supplementary Table 3). All fixation indexes were statistically significant. These results suggest that the frequency of heterozygotes is lower than expected under HWE.
The pairwise FST values suggested moderate to high genetic differentiation between localities (Fig. 3). The largest difference was observed for the Peruvian Andes, Peruvian Amazonia, and Brazilian Amazonia. Mesoamerica (MEX and CenAme) and northern South America (COL and VEN-ANT) showed less pairwise genetic differentiation (Fig. 3).
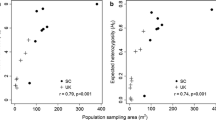
The Mantel test revealed no significant correlation between genetic and geographic distance matrices (R2 = − 0.287, p = 0.779), indicating a lack of isolation by distance. We found that PER-AND, PER-AM, and BRA-AM localities, which are geographically closer to each other, are genetically less similar (Supplementary Fig. 4; Supplementary Table 4). The observed low pairwise FST values indicates possible long-distance gene flow between MEX, CenAme, VEN, and COL (Supplementary Tables 4–5). According to AMOVA, the variation among samples within localities (41%, Φ = 0.52) is higher than between localities (21.04%, Φ = 0.62; Table 2).
Approximate Bayesian Computation (ABC) analyses indicated that the best supported dispersal hypothesis was scenario 2 (Fig. 4) with posterior probability of 0.999 and non-overlapping confidence intervals. This scenario showed low type I and type II error rates (0.00013; Supplementary Table 6), suggesting that domestication started in South America, specifically in Brazilian Amazonia (Brazil-AM), with dissemination to Mexico via Peruvian Amazonia (PER-AMA) and northern South America (COL).
Highest-probability scenario tested for dispersal of Psidium guajava in the Neotropics. Eight localities with effective population sizes N1 to N8 correspond to MEX (Mexico), CenAme (Central America), ANT (The Antilles), VEN (Venezuela), COL (Colombia), BRA-AM (Brazilian Amazonia), PER-AM (Peruvian Amazonia), PER-AND (Peruvian Andes), respectively. Time since divergence corresponds to t1 to t5.
Discussion
Perennial trees, characterized by their extended lifespan and delayed sexual reproduction, tend to exhibit a weak population structure35,36. Domestication profoundly influences the population dynamics and genetic structure of species, shaped by a complex set of evolutionary events involving both natural factors and ancestral and contemporary human activities. The present study provides the first evidence of the domestication history of guava in the Neotropics and assesses scenarios of the species' dispersal in the region.
Our analyses found that values of guava genetic diversity expressed as HE ranged from 0.44 to 0.70. These levels are comparable to those of other perennials with populations domesticated to some degree, such as A. cherimola, Olea europaea L., and Prunus armeniaca L.9,37,38. However, compared to other Psidium species, P. guajava has high genetic diversity, for example, in a single population in Southeast Brazil the maximum HE values (HE = 0.71) are comparable to those of P. guineense Sw. (HE = 0.74) and P. macahense O. Berg (HE = 0.63)39. In contrast, an insular Psidium species (P. galapageium Hook. f.) has moderate to low (HE = 0.275–0.570) genetic diversity40. P. cattleianum Afzel. ex Sabine also showed lower diversity values (HE = 0.117–0.326)41.
Observed heterozygosity (Ho) systematically showed lower values than expected heterozygosity, suggesting heterozygote deficiency among guava due to inbreeding. In different islands of the Galapagos42 and guava samples from a germplasm bank43 registered similar results. These findings may be explained, in part, by self-fertilization and vegetative propagation, which can occur in P. guajava44. Likewise, Robertson’s45 hypotheses could be useful to explain the heterozygosity decline in guava. He proposes that subdividing a population into several isolated groups would allow maximum genetic diversity (minimum global co-ancestry) to be achieved in the long term since different allelic variants will develop and become fixed in each group, becoming a genetic reservoir of variation. However, complete isolation leads to higher rates of local inbreeding with the possible consequence of inbreeding depression. Therefore, Robertson45 also suggests that occasional mixing of these subpopulations would minimize the overall rate of inbreeding. In support of this hypothesis, we found lower rates of global inbreeding for guava. Likewise, Mantel's analysis suggests long-distance gene flow, especially between the populations of northern South America (COL and VEN) and Mesoamerica (MEX and CenAme; Supplementary Tables 4–5), which would allow the reduction of inbreeding and its effects. Overall, the absence of isolation by distance, the broad range of FST values, the high FIS value, and the separate gene pools indicated by PCA, DAPC, and STRUCTURE suggest a metapopulation dynamic. Local cultivated guava populations may originate from the surrounding genetic variation and occasionally receive long-distance gene flow. Finally, further studies are needed to examine the cause of heterozygote deficiency in guava.
Contrary to our hypothesis of south-to-north decrease in diversity, the genetic diversity pattern, expressed as HE is just the opposite. A decreasing trend in HE was observed from Mexico and Central America (HE = 0.70) to the Peruvian Andes (PER-AND = 0.44). This pattern can be explained by the mixture of guavas from different regions occurring in Central America and Mexico. In these areas, most of the individuals show signatures of admixture of well-defined South American genetic groups. Hence, anthropic dispersal may have enhanced guava genetic diversity in Central America and Mexico. In addition, the diversity of environmental conditions, new biotic interactions, and selection pressures in Mesoamerica could have contributed to the maintenance of genetic variants that were present in the gene pool due to mutations. These events would explain an increased guava genetic diversity in response to new environmental conditions and challenges, a hypothesis that is testable by using ecological niche models46,47,48. In addition, whether this pattern points towards a center of genetic diversity or is the result of admixture among clusters is a matter to be evaluated by rating explicit demographic scenarios.
In our study, the genetic differentiation of P. guajava populations yielded an FST value of 0.207, indicating moderate differentiation, considering the wide geographical range across which the species is distributed. Likewise, the molecular variance is higher among individuals/within localities than among localities. Similar findings have been reported for other perennial fruit trees like A. cherimola, Diospyros kaki L.F., Juglans regia L., Mangifera indica L., O. europaea, P. persica L., and P. armeniaca9,37,38,49,50,51,52. In the case of guava, the observed FST value may be attributed to limited gene flow among the sampled localities, which span different regions of the Neotropics. Similarly, the pattern of variance identified here can be due to outcrossing and guava’s invasive (successional) character (Hamrick et al.53 and therein). In cultivated and invasive populations of guava, the genetic variation pattern is also similar, with higher genetic variance among individuals/between populations and clearly defined genetic groups42,54,55.
Regarding the genetic clustering found in our study, each of the most geographically isolated populations from South America (Peruvian Andes [PER-AND], and Brazilian and Peruvian Amazonia [BRA-AM; PER-AMA]) belongs to a distinct genetic group and shows greater differentiation in relation to other groups (FST; Fig. 3). Localities in northern South America, Central America, and Mesoamerica show lower values of genetic differentiation with some individuals being admixed. This scenario suggests a pattern of greater differentiation among localities in South America, probably due to the isolation between guava populations, especially between the Amazonian and Andean regions. In Amazonia, the vast expanse of tropical rainforest and relatively homogeneous climatic conditions have favored the domestication of a variety of crops such as Manihot esculenta Crantz, T. cacao, and various fruits and nuts. This region is been an independent center of plant domestication, where indigenous peoples have managed and cultivated numerous crops over millennia, resulting in notable genetic diversity within these crops1,6,27,56,57. In contrast, the Andean ecosystems' altitudinal and climatic variability has led to the genetic differentiation of plants adapted to specific microenvironments. This environmental diversity has promoted the evolution of plants with unique genetic traits necessary for surviving extreme conditions, resulting in a mosaic of locally adapted crops, each with distinct genetic variations1,6,27,56,57. Moreover, human interaction with the environment in both regions has played a crucial role. In Amazonia, landscape management practices, such as the creation of "terra preta" (Amazonian Dark Earths), have enriched the soil and fostered crop diversification. In the Andes, agricultural techniques such as terracing and irrigation, have enabled the adaptation and cultivation of plants on steep slopes and less fertile soils1,57,58.
Therefore, the localities evaluated here would have likely been exposed to specific evolutionary processes, considering the climatic and ecological characteristics of their geographical origin’s setting, thus promoting differentiation between them. Variable admixture levels among populations may also be the outcome of diverse trade routes and human migrations over time30, as is the case of J. regia50 and M. indica51.
According to the best-supported ABC scenario, Amazonia is the most probable area of domestication of guava, and the first dispersal route likely was from there towards the Peruvian Andes. This result agrees with the oldest archaeological guava macro remains found in Southwestern Amazonia in the Teotonio archeological site, in a layer between 9490 and 6505 cal. BP27. In South America, the lowlands of southwestern Amazonia are recognized as a relevant center of domestication1,27,56,57 and the place from which important species, such as manioc and peanut (Arachis hypogaea L.), dispersed towards the Peruvian dry coast58. Indeed, a significant number of archaeological guava remains dating from 6975 to 450 BP have been reported from the Peruvian dry coast (see Fig. 4 in Arévalo-Marín et al.30). This evidence also supports the hypothesis that guava could have spread through the Andes, from Amazonia to the Peruvian coast, as the best supported scenario in this study suggests. Therefore, more detailed archaeological and genetic studies that include samples from both the southwestern region and other areas of Amazonia would allow for a confirmation of the domestication area of guava.
In summary, our study provides an overview of the genetic diversity and population structure of guava in the Neotropics. The microsatellite markers and Bayesian clustering approaches identified the presence of one gene pool in Mesoamerican (Mexico) and four in South America (Brazilian Amazonia, Peruvian Amazonia and Andes, and Colombia). The high genetic differentiation between the Brazilian and Peruvian Amazonia and Peruvian Andes guava samples could be due to environmental differences, since guava subpopulations in distinct geographic settings may reflect divergent local adaptation. Niche analyses are needed to understand whether climatic events could explain these hypotheses, and genomic analyses would allow testing of hypotheses of local adaptation. On the other hand, the ABC approach identified Brazilian Amazonia as the potential area of guava domestication, with subsequent dispersal into western and northern South America and Mesoamerica where local diversification processes occurring in these last two regions could also underlie the observed diversity patterns. Follow-up studies that include defined populations of feral and cultivated guavas, and focused sampling in southwestern Amazonia and the Andes could help to unravel the guava domestication process.
Materials and methods
Material
We studied 215 guava plants from 11 countries. We collected 86 samples from Brazil, Colombia, Honduras, Mexico, and Venezuela in the guava germplasm bank of the Instituto Nacional de Investigaciones Forestales, Agrícolas y Pecuarias (INIFAP) in Aguascalientes, Mexico; 17 samples from the guava collection of the Tropical Agricultural Research and Higher Education Center (CATIE) in Turrialba, Costa Rica, from Costa Rica, El Salvador, Guatemala, and Honduras; and 26 samples from Brazil, Colombia, and Puerto Rico in the guava collection of the Corporación Colombiana de Investigación Agropecuaria (Agrosavia) in Palmira, Colombia. We also included samples collected outside germplasm banks from Brazilian Amazonia (38 plants), Peruvian Amazonia (15 plants), the Peruvian Andes (19 plants), and 14 samples from different localities in Venezuela. We considered samples collected in the Peruvian Andes and Brazilian Amazonia as tolerated or planted because these were collected in empty lots, roadsides, orchards, and gardens. Samples from Venezuela and Peruvian Amazonia were collected in areas far from plantations or crops; however, since it is difficult to distinguish between wild and feral guavas, each of these samples were considered feral. Because of this collection strategy, which is commonly used with cultivated plants, we are not dealing with biological populations, so we will call groups of plants from different areas “localities”. All methods were performed in accordance with the relevant guidelines and regulations, and appropriate permissions for the collection of plant material were obtained from all relevant parties.
Molecular methods
DNA was extracted from young leaves using a CTAB-based protocol59. Initially, all 215 individuals were genotyped using 25 nuclear microsatellite loci developed for P. guajava60,61. We combined five primers in each of the five multiplex reactions (see Supplementary Table 7 for primers and multiplex reaction details). PCRs were performed using the Platinum Multiplex PCR Master Mix (Thermo-Fisher, USA) following the manufacturer’s instructions for reaction assembly and program. Every reaction was driven to a 5.5 μL final volume containing 2.0 μL Platinum Multiplex PCR Master Mix, 2.0 μL PCR grade H2O, 0.5 μL G/C enhancer volume, 1.0 μL DNA template (50–200 ng/μL), and primer concentrations between 50 to 70 nM according to each product’s relative fluorescent units (RFU). Multiplex reactions required an annealing temperature of 55 °C for all primers; 40 cycles were used in every PCR reaction. When amplification was not successful, we repeated the PCR reactions using 0.04 μL Kapa polymerase (Kapa Taq HotStart), 2.0 μL Buffer Kapa, 2.0 μL PCR grade H2O, and 1.0 μL DNA template (50–200 ng/μL). The annealing temperature and the number of cycles were maintained. To prevent possible contamination, we used negative controls for each multiplex assembly. All products were verified in 2% agarose gel electrophoresis. PCRs were carried out in a MultiGene OptiMax (Labnet International, Inc., Edison, NJ, USA) or in a 2700 thermal cycler (Applied Biosystems, Foster City, CA, USA). Genotyping was achieved using the Microsatellite plugin (v. 1.4.7) of Geneious Prime 2022 (Dotmatics, NZ). Allele scoring was performed manually following Selkoe and Toonen62.
Null alleles, Hardy–Weinberg equilibrium, and linkage disequilibrium tests
We tested the presence and frequency of null alleles following Brookfield63 using the PopGenReport v.3.0.7 package64 in R. We calculated deviations from Hardy–Weinberg equilibrium (HWE) for each locus and separately for each locality. Also, we calculated HWE across all samples using the ‘hw.test’ function of the R package pegas v.1.165, with 1000 Monte Carlo permutations. Alpha levels to determine statistically significant deviations from Hardy–Weinberg proportions and independent sorting of genotypes were adjusted using the false discovery rate (FDR) approach developed by Benjamini and Hochberg66, using 0.05 alpha level. P-values were corrected for multiple comparisons using the Benjamini–Hochberg method66. We calculated a measure of correlation (r̄d67 using the function ‘ia’68 in the R package poppr v.2.9.3, for testing overall linkage disequilibrium. Using the function ‘genotype curve’ of the same package, we described the genotypic diversity in relation to different combinations of loci by a genotype accumulation curve to determine whether our sample provided a reasonable estimate of genetic diversity. The curve was generated by randomly sampling x loci and counting the number of multilocus genotypes (MLG) observed. This sampling was repeated r times from 1 to n − 1 loci, creating n − 1 distributions of observed MLGs69.
Genetic diversity and genetic differentiation
Genetic differentiation was examined using several complementary approaches. First, as an exploratory method, we performed a Principal Components Analysis (PCA) to summarize the genetic variation based on the microsatellite data set. Subsequently, we performed a Discriminant Analysis of Principal Components (DAPC)70. DAPC is an approach that optimizes the separation of individuals into predefined groups using a discriminant function of the principal component70. Based on DAPC, the membership probability was calculated for the overall genetic background of an individual. We used the components identified in the PCA analysis as predefined groups for the DAPC implementation. For implementing the PCA, we used the ‘dudi.pca’ function from ade4 v.1.7–22 R71 and visualized it with factoextra v.1.0.7 R72. For DAPC, we used adegenet v.2.2.1073 implemented in R.
We assessed standard measures of genetic diversity for the entire dataset and genetic groups according to DAPC results. The number of individuals (N), number of alleles (A), and the expected (HE) and observed (HO) heterozygosities were calculated using poppr v.2.9.368 in R. We estimated rarefied allelic richness using the ‘allel.rich’ function of PopGenReport v.3.0.764 in R. Private allele richness (AP) were calculated using a rarefaction approach74,75 implemented in ADZE 1.076.
As an additional test to calculate the group assignment probability for each sample, we performed genetic population structure analysis using the Bayesian approach implemented in STRUCTURE 2.3.477,78, based on the admixture model with correlated allele frequencies and information on the origin of localities (popinfo = 1). The admixture model was tested for K-values ranging from 1 to 8, since 8 is the number of sampled regions, with 10 independent runs per K value for the entire dataset. We used 1,000,000 Markov Chain Monte Carlo iterations with a burn-in length of 100,000. To determine the most probable value of K, we used Evanno’s ΔK method79 and mean LnP(K)80 implemented in Structure Harvester v0.6.9481. We used CLUMPP 1.1.282 with the Greedy algorithm to infer the optimal K-cluster affiliations of samples and StructuRly v.0.1.083 in R to generate bar graphs of the STRUCTURE software results.
Wright's F statistics84 (FIS, FIT, and FST) were estimated using the methods of Weir and Cockerham85. We also calculated the genetic differentiation among localities through a pairwise FST matrix. Both the F statistics and the paired FST matrix were calculated with 95% confidence intervals from 10,000 bootstrapping, using the ‘diffCalc’ function of diveRsity86. A Mantel test87 was used to assess isolation by distance (IBD) between pairs of guava localities. We used the geographic distance matrix transform from coordinates in Euclidean distance and calculated using the function ‘dist’ in the stats v.4.3.1 in R and a linearized pairwise FST matrix (FST/1 − FST) as genetic distance. The function ‘mantel.rtest’ from ade4 v.1.7-2288 was used to calculate the Mantel test, and scatter-plots were then generated with adegenet v.2.2.1073.
We also tested the degree of genetic differentiation between DAPC groups (determined here) and locations, performing the analysis of molecular variance (AMOVA) followed by an estimation of the extent of genetic differentiation with phi-statistics, both using the ‘poppr.amova’ function in poppr v.2.9.368. The significance of variance components was assessed using a permutation test implemented through the ‘randtest’ function in ape4 v.5.771,89,90 with 999 permutations.
Identification of the origin of domestication
We used nuclear microsatellite data to run the Approximate Bayesian Computation (ABC) framework91,92 implemented in DIYABC-RF GUI93 to model a possible branching order among guava localities that would represent the history of domestication of the lineage. We considered five scenarios (1) Mexico as a probable center of origin of domestication with dissemination to South America; (2) South America, specifically Brazilian Amazonia (Brazil-AM) as a probable center and later dissemination to Mexico via Peru; (3) Two independent centers of origin of domestication, one in the Peruvian Andes (Peru-An) and another in Mexico; (4) Peruvian Amazonia (Peru-AM) and Brazil-AM as independents centers of origin of domestication and dissemination towards northern South America with Central America and Mexico being of admixed destination; and (5) domestication in northern South America and dissemination to three areas (Mexico and Central America, Venezuela and Antilles, and Peru and Brazil) (Supplementary Fig. 5). The priors and conditions for each parameter can be found in the Supplementary Material 2; we considered a generation time of 10 years (probable fruiting time in natural conditions)44. We conducted previous runs to adjust the tested scenarios and the parameters94. For the final run, we obtained 500,000 simulated datasets, 500 trees, and 424 summary statistics. To identify the best supported scenario, we performed model check based on 500 pseudo-observed data sets (PODs) under each scenario to assess confidence in scenario choice, and to estimate the class specific error rates, which is the mean classification error rate93,95,96.
Data availability
The genotyping data generated in the present study will be released upon acceptance and is (privately) available at: https://figshare.com/s/47366c57067686695d91.
References
Clement, C. R. et al. Disentangling domestication from food production systems in the Neotropics. Quaternary 4, 4 (2021).
Kreitzman, M., Toensmeier, E., Chan, K. M. A., Smukler, S. & Ramankutty, N. Perennial staple crops: Yields, distribution, and nutrition in the global food system. Front. Sustain. Food Syst. 4, 1–21 (2020).
McClure, K. A., Sawler, J., Gardner, K. M., Money, D. & Myles, S. Genomics: A potential panacea for the perennial problem. Am. J. Bot. 101, 1780–1790 (2014).
Miller, A. J. & Gross, B. L. From forest to field: Perennial fruit crop domestication. Am. J. Bot. 98, 1389–1414 (2011).
Galluzzi, G. & Noriega, I. L. Conservation and use of genetic resources of underutilized crops in the Americas—A continental analysis. Sustainability 6, 980–1017 (2014).
Clement, C. R. 1492 and the loss of Amazonian crop genetic resources. II. Crop biogeography at contact. Econ. Bot. 53, 203–216 (1999).
Casas, A., Caballero, J., Mapes, C. & Zárate, S. Manejo de la vegetación, domesticación de plantas y origen de la agricultura en Mesoamérica. Bot. Sci. 61, 31–47 (1997).
Patiño, V. M. Plantas cultivadas y animales domésticos en América Equinoccial I: Frutales. (Cali: Impre. Departamental, 1963).
Larranaga, N. et al. A Mesoamerican origin of cherimoya (Annona cherimola Mill.): Implications for the conservation of plant genetic resources. Mol. Ecol. 26, 4116–4130 (2017).
Clement, C. R., Rival, L. & Cole, D. M. Domestication of peach palm (Bactris gasipaes): The roles of human mobility and migration. Mobil. Migr. Indig. Amazon. Contemp. Ethnoecol. Perspect. 11, 117–140 (2009).
Clement, C. R. et al. Origin and dispersal of domesticated peach palm. Front. Ecol. Evol. 5, 1–19 (2017).
Parker, I. M. et al. Domestication syndrome in caimito (Chrysophyllum cainito L.): Fruit and seed characteristics. Econ. Bot. 64, 161–175 (2010).
Petersen, J. J., Parker, I. M. & Potter, D. Origins and close relatives of a semi-domesticated neotropical fruit tree: Chrysophyllum cainito (Sapotaceae). Am. J. Bot. 99, 585–604 (2012).
Chávez-Pesqueira, M. & Núñez-Farfán, J. Genetic diversity and structure of wild populations of Carica papaya in northern Mesoamerica inferred by nuclear microsatellites and chloroplast markers. Ann. Bot. 118, 1293–1306 (2016).
Miller, A. J. & Schaal, B. A. Domestication and the distribution of genetic variation in wild and cultivated populations of the Mesoamerican fruit tree Spondias purpurea L. (Anacardiaceae). Mol. Ecol. 15, 1467–1480 (2006).
Motamayor, J. C. et al. Geographic and genetic population differentiation of the Amazonian chocolate tree (Theobroma cacao L.). PLoS One 3, e3311 (2008).
Cornejo, O. E. et al. Population genomic analyses of the chocolate tree, Theobroma cacao L., provide insights into its domestication process. Commun. Biol. 1, 167 (2018).
Zarrillo, S. et al. The use and domestication of Theobroma cacao during the mid-Holocene in the upper Amazon. Nat. Ecol. Evol. 2, 1879–1888 (2018).
Altendorf, S. Minor tropical fruits: Mainstreaming a niche market In Food Outlook Biannual Report on Global Food Markets 67–74 (2018).
Altendorf, S. Major tropical fruits market review 2018. http://www.fao.org/3/ca5692en/CA5692EN.pdf (2019).
Landrum, L. R. The genus Psidium (Myrtaceae) in the state of Bahia, Brazil. Canotia 13, 1–101 (2017).
Landrum, L. R. Psidium guajava L.: Taxonomy, relatives and possible origin. In Guava: Botany, Production and Uses (ed. Mitra, S.) 1–21 (Cab International, 2021). https://doi.org/10.1079/9781789247022.0001.
Hiwale, S. Guava (Psidium guajava). In Sustainable Horticulture in Semiarid Dry Lands 213–224 (Springer India, 2015). https://doi.org/10.1007/978-81-322-2244-6.
Gutiérrez, R. M., Mitchell, S. & Solis, R. V. Psidium guajava: A review of its traditional uses, phytochemistry and pharmacology. J. Ethnopharmacol. 117, 1–27 (2008).
Menzel, C. M. Guava: An exotic fruit with potential in Queensland. Qld. Agric. J. 111, 93–98 (1985).
Patiño, V. M. Historia y dispersión de los frutales nativos del neotrópico 191–198 (Centro Internacional de Agricultura Tropical, 2002).
Watling, J. et al. Direct archaeological evidence for southwestern Amazonia as an early plant domestication and food production centre. PLoS One 13, e0199868 (2018).
Cárdenas, M. El Periodo Precerámico en el valle de Chao. Boletín de Arqueología PUCP 141–169 (1999).
Smith, C. E. Plant remains. In The prehistory of Tehuacan Valley. Volume One: Environment and Subsistence (ed. Byers, D. S.) 220–255 (University of Texas Press, 1967).
Arévalo-Marín, E. et al. The Taming of Psidium guajava: Natural and cultural history of a Neotropical fruit. Front. Plant Sci. 12, 1–15 (2021).
Diez, C. M. et al. Olive domestication and diversification in the Mediterranean Basin. New Phytologist 206, 436–447 (2015).
Gaut, B. S., Díez, C. M. & Morrell, P. L. Genomics and the contrasting dynamics of annual and perennial domestication. Trends Genet. 31, 709–719 (2015).
Meyer, R. S., DuVal, A. E. & Jensen, H. R. Patterns and processes in crop domestication: An historical review and quantitative analysis of 203 global food crops. New Phytol. 196, 29–48 (2012).
Meyer, R. S. & Purugganan, M. D. Evolution of crop species: Genetics of domestication and diversification. Nat. Rev. Genet. 14, 840–852 (2013).
Hamrick, J. L. & Godt, M. J. W. Allozyme diversity in plant species. In Plant Population Genetics, Breeding, and Genetic Resources. 43–63 (1990).
Loveless, M. D. & Hamrick, J. L. Ecological determinants of genetic structure in plant populations. Annu. Rev. Ecol. Syst. 15, 65–95 (1984).
Bourguiba, H. et al. Genetic structure of a worldwide germplasm collection of Prunus armeniaca L. reveals three major diffusion routes for varieties coming from the species’ center of origin. Front. Plant Sci. 11, 638 (2020).
Breton, C., Tersac, M. & Bervillé, A. Genetic diversity and gene flow between the wild olive (oleaster, Olea europaea L.) and the olive: several Plio-Pleistocene refuge zones in the Mediterranean basin suggested by simple sequence repeats analysis. J. Biogeogr. 33, 1916–1928 (2006).
de Oliveira Bernardes, C. et al. Genetic Diversity and population structure of Psidium species from Restinga: A coastal and disturbed ecosystem of the Brazilian Atlantic Forest. Biochem. Genet. 60, 2503–2514 (2022).
Urquia, D., Pozo, G., Gutierrez, B., Rowntree, J. K. & de Lourdes Torres, M. Understanding the genetic diversity of the guayabillo (Psidium galapageium), an endemic plant of the Galapagos islands. Glob. Ecol. Conserv. 24, e01350 (2020).
Machado, R. M., de Oliveira, F. A., de Matos Alves, F., de Souza, A. P. & Forni-Martins, E. R. Population genetics of polyploid complex Psidium cattleyanum Sabine (Myrtaceae): Preliminary analyses based on new species-specific microsatellite loci and extension to other species of the genus. Biochem. Genet. 59, 219–234 (2021).
Urquía, D. et al. Psidium guajava in the Galapagos islands: population genetics and history of an invasive species. PLoS One 14, e0203737 (2019).
Sitther, V. et al. Genetic characterization of guava (Psidium guajava L.) germplasm in the United States using microsatellite markers. Genet. Resour. Crop Evol. 61, 829–839 (2014).
Crane, J. H. & Balerdi, C. F. Guava growing in the Florida home landscape. In Horticultural Sciences Department Document HS4, Florida Cooperative Extension Service Institute of Food and Agricultural Sciences, University of Florida 1–8 (2005).
Robertson, A. The effect of non-random mating within inbred lines on the rate of inbreeding. Genet. Res. (Camb.) 5, 164–167 (1964).
Ruiz-Gil, P. J. et al. Wild papaya shows evidence of gene flow from domesticated Maradol papaya in Mexico. Genet. Resour. Crop Evol. 70, 1–20 (2023).
Pérez-Valladares, C. X., Moreno-Calles, A. I., Mas, J. F. & Velazquez, A. Species distribution modeling as an approach to studying the processes of landscape domestication in central southern Mexico. Landsc. Ecol. 1–16 (2022).
Martínez-Ainsworth, N. E. et al. Fluctuation of ecological niches and geographic range shifts along chile pepper’s domestication gradient. Ecol. Evol. 13, e10731 (2023).
Cao, K. et al. Comparative population genomics reveals the domestication history of the peach, Prunus persica, and human influences on perennial fruit crops. Genome Biol. 15, 1–15 (2014).
Pollegioni, P. et al. Ancient humans influenced the current spatial genetic structure of common walnut populations in Asia. PLoS One 10, e0135980 (2015).
Warschefsky, E. J. & von Wettberg, E. J. B. Population genomic analysis of mango (Mangifera indica) suggests a complex history of domestication. New Phytologist 222, 2023–2037 (2019).
Xu, Y. et al. Genetic diversity and association analysis among germplasms of Diospyros kaki in Zhejiang Province based on SSR markers. Forests 12, 422 (2021).
Hamrick, J. L., Godt, M. J. W. & Sherman-Broyles, S. L. Factors influencing levels of genetic diversity in woody plant species. In Population Genetics of Forest Trees: Proceedings of the International Symposium on Population Genetics of Forest Trees Corvallis, Oregon, USA, July 31–August 2, 1990 95–124 (Springer, 1992).
Fagundes, B. S. et al. Transferability of microsatellite markers among Myrtaceae species and their use to obtain population genetics data to help the conservation of the Brazilian Atlantic Forest. Trop. Conserv. Sci. 9, 408–422 (2016).
Kumar, C. et al. Development of novel g-SSR markers in guava (Psidium guajava L.) cv. Allahabad Safeda and their application in genetic diversity, population structure and cross species transferability studies. PLoS One 15, e0237538 (2020).
Clement, C. R. 1492 and the loss of Amazonian crop genetic resources. I. The relation between domestication and human population decline. Econ. Bot. 53, 188–202 (1999).
Clement, C. R., de Cristo-Araújo, M., D’Eeckenbrugge, G. C., Alves Pereira, A. & Picanço-Rodrigues, D. Origin and domestication of native Amazonian crops. Diversity (Basel) 2, 72–106 (2010).
Piperno, D. R. The origins of plant cultivation and domestication in the New World tropics. Curr. Anthropol. 52, S453–S470 (2011).
Doyle, J. J. & Doyle, J. L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 19, 11–15 (1987).
Guavamap. Improvement of guava: linkage mapping and QTL analysis as a basis for marker-assisted selection. (2008).
Risterucci, A. M., Duval, M. F., Rohde, W. & Billotte, N. Isolation and characterization of microsatellite loci from Psidium guajava L. A. Mol. Ecol. Notes 5, 745–748 (2005).
Selkoe, K. A. & Toonen, R. J. Microsatellites for ecologists: A practical guide to using and evaluating microsatellite markers. Ecol. Lett. 9, 615–629 (2006).
Brookfield, J. F. Y. A simple new method for estimating null allele frequency from heterozygote deficiency. Mol. Ecol. 5, 453–455 (1996).
Adamack, A. T. & Gruber, B. PopGenReport: Simplifying basic population genetic analyses in R. Methods Ecol. Evol. 5, 384–387 (2014).
Paradis, E. pegas: An R package for population genetics with an integrated—Modular approach. Bioinformatics 26, 419–420 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B (Methodol.) 57, 289–300 (1995).
Agapow, P. & Burt, A. Indices of multilocus linkage disequilibrium. Mol. Ecol. Notes 1, 101–102 (2001).
Kamvar, Z. N., Tabima, J. F. & Grünwald, N. J. Poppr: An R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2, e281 (2014).
Kamvar, Z. N., Brooks, J. C. & Grünwald, N. J. Novel R tools for analysis of genome-wide population genetic data with emphasis on clonality. Front. Genet. 6, 208 (2015).
Jombart, T., Devillard, S. & Balloux, F. Discriminant analysis of principal components: A new method for the analysis of genetically structured populations. BMC Genet. 11, 1–15 (2010).
Dray, S. & Dufour, A.-B. The ade4 package: Implementing the duality diagram for ecologists. J. Stat. Softw. 22, 1–20 (2007).
Wickham, H. Data analysis. ggplot2: Elegant Graphics for Data Analysis 189–201 (2016).
Jombart, T. adegenet: A R package for the multivariate analysis of genetic markers. Bioinformatics 24, 1403–1405 (2008).
Hurlbert, S. H. The nonconcept of species diversity: A critique and alternative parameters. Ecology 52, 577–586 (1971).
Kalinowski, S. T. Counting alleles with rarefaction: Private alleles and hierarchical sampling designs. Conserv. Genet. 5, 539–543 (2004).
Szpiech, Z. A., Jakobsson, M. & Rosenberg, N. A. ADZE: A rarefaction approach for counting alleles private to combinations of populations. Bioinformatics 24, 2498–2504 (2008).
Falush, D., Stephens, M. & Pritchard, J. K. Inference of population structure using multilocus genotype data: Linked loci and correlated allele frequencies. Genetics 164, 1567–1587 (2003).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Evanno, G., Regnaut, S. & Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 14, 2611–2620 (2005).
Janes, J. K. et al. The K= 2 conundrum. Mol. Ecol. 26, 3594–3602 (2017).
Earl, D. A. & VonHoldt, B. M. STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 4, 359–361 (2012).
Jakobsson, M. & Rosenberg, N. A. CLUMPP: A cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23, 1801–1806 (2007).
Criscuolo, N. G. & Angelini, C. StructuRly: A novel shiny app to produce comprehensive, detailed and interactive plots for population genetic analysis. PLoS One 15, e0229330 (2020).
Wright, S. The genetical structure of populations. Ann. Eugen 15, 323–354 (1949).
Weir, B. S. & Cockerham, C. C. Estimating F-statistics for the analysis of population structure. Evolution (N Y) 38, 1358–1370 (1984).
Keenan, K., McGinnity, P., Cross, T. F., Crozier, W. W. & Prodöhl, P. A. diveRsity: An R package for the estimation and exploration of population genetics parameters and their associated errors. Methods Ecol. Evol. 4, 782–788 (2013).
Mantel, N. The detection of disease clustering and a generalized regression approach. Cancer Res. 27, 209–220 (1967).
Thioulouse, J. et al. Multivariate analysis of ecological data with ade4. (2018).
Bougeard, S. & Dray, S. Supervised multiblock analysis in R with the ade4 package. J. Stat. Softw. 86, 1–17 (2018).
Chessel, D., Dufour, A. B. & Thioulouse, J. The ade4 package-I-One-table methods. R News 4, 5–10 (2004).
Beaumont, M. A. Approximate Bayesian computation in evolution and ecology. Annu. Rev. Ecol. Evol. Syst. 41, 379–406 (2010).
Beaumont, M. A., Zhang, W. & Balding, D. J. Approximate Bayesian computation in population genetics. Genetics 162, 2025–2035 (2002).
Collin, F. et al. Extending approximate Bayesian computation with supervised machine learning to infer demographic history from genetic polymorphisms using DIYABC Random Forest. Mol. Ecol. Resour. 21, 2598–2613 (2021).
Bertorelle, G., Benazzo, A. & Mona, S. ABC as a flexible framework to estimate demography over space and time: Some cons, many pros. Mol. Ecol. 19, 2609–2625 (2010).
Cornuet, J.-M., Ravigné, V. & Estoup, A. Inference on population history and model checking using DNA sequence and microsatellite data with the software DIYABC (v1. 0). BMC Bioinform. 11, 1–11 (2010).
Robert, C. P., Cornuet, J.-M., Marin, J.-M. & Pillai, N. S. Lack of confidence in approximate Bayesian computation model choice. Proc. Natl. Acad. Sci. 108, 15112–15117 (2011).
Acknowledgements
This manuscript constitutes part of the doctoral project of the EA-M, who thanks the Posgrado en Ciencias Biológicas of the Universidad Nacional Autónoma de México (UNAM) and acknowledges the doctoral scholarship supported by the Consejo Nacional de Humanidades, Ciencia y Tecnología (CONAHCyT). We thank André Braga Junqueira, and Sara Deambrozi Coelho, Carlos Cordero Vargas, Yani Aranguren, and the staff at the Campo Experimental Pabellón (Aguascalientes) of INIFAP for field support during guava collections in Brazil, Costa Rica, Venezuela, and Mexico, respectively. We are grateful to the staff of Corporación Colombiana de Investigación Agropecuaria, especially Alvaro Caicedo Arana, Carolina González Almario, and Luisa Alejandra Rugeles Barandica for their support in setting up the agreement that allowed access to the guava samples in Colombia. EA-M thanks Shirley Camacho and Allison Muñoz for the field work made in Agrosavia. EA-M is grateful to Isolda-Luna Vega and Othón Alcántara-Ayala for access to computing and software, and Esteban Rincón Suárez for his help in organizing the graphics. Finally, we thank the administrative staff of IIES-UNAM, campus Morelia, Mexico especially, Benjamín Mora, Juan Carlos Mata, and Claudia Lenina Sánchez for their support in the importation of plant material.
Funding
This work was supported by the CONAHCyT [Grant number A1-S-14306], and the DGAPA-UNAM research project [Grant number IN206520 and IN224023]. CRC thanks the Brazilian Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) for a research fellowship [Grant number 303477/2018-0].
Author information
Authors and Affiliations
Contributions
Conceptualization: E.A.-M., A.C., and C.R.C.; Laboratory work: E.A.-M. and H.A.-S.; Field work: E.A.-M.; A.C.; C.R.C.; H.A.-S., G.F., J.S.P.-R.; Methodology: E.A.-M., H.A.-S., E.R.-S., G.C.-M. and L.J.-B.; Writing—original draft preparation: E.A.-M. and A.C.; Writing—review and editing: all the authors; Supervision: A.C. and C.R.C.; Funding acquisition: A.C. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Arévalo-Marín, E., Casas, A., Alvarado-Sizzo, H. et al. Genetic analyses and dispersal patterns unveil the Amazonian origin of guava domestication. Sci Rep 14, 15755 (2024). https://doi.org/10.1038/s41598-024-66495-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-66495-y
- Springer Nature Limited