Abstract
Triticum militinae (2n = 4X = 28, AtAtGG), belonging to the secondary gene pool of wheat, is known to carry resistance to many diseases. Though some disease resistance genes were reported from T. timopheevii, the closest wild relative of T. militinae, there are no reports from T. militinae. Twenty-one T. militinae Derivatives (TMD lines) developed at the Division of Genetics, IARI, New Delhi, were evaluated for leaf and stripe rusts at seedling and adult plant stages. Eight TMD lines (6–4, 6–5, 11–6, 12–4, 12–8, 12–12, 13–7 and 13–9) showed seedling resistance to both leaf and stripe rusts while six TMD lines (7–5, 7–6, 11–5, 13–1, 13–3 and 13–4) showed seedling resistance to leaf rust but adult plant resistance to stripe rust and three TMD lines (9–1, 9–2 and 15) showed seedling resistance to leaf rust but susceptibility to stripe rust. Three TMD lines (2–7, 2–8 and 6–1) with adult plant resistance to leaf and stripe rusts were found to carry the known gene Lr34/Yr18. Ten TMD lines (7–5, 7–6, 9–1, 9–2, 11–5, 11–6, 12–12, 12–4, 12–8, and 15) with seedling resistance to leaf rust, showing absence of known genes Lr18 and Lr50 with linked markers requires further confirmation by the test of allelism studies. As not a single stripe rust resistance gene has been reported from T. militinae or its close relative T. timpopheevii, all the 8 TMD lines (6–4, 6–5, 11–6,12–4, 12–8, 12–12, 13–7 and 13–9) identified of carrying seedling resistance to stripe rust and 3 TMD lines (13–1, 13–3 and 13–4) identified of carrying adult plant resistance to stripe rust are expected to carry unknown genes. Also, all the TMD lines were found to be cytologically stable and thus can be used in inheritance and mapping studies.
Similar content being viewed by others
Introduction
Wheat (Triticum aestivum L.) is one of the most important cereal crops in the world and India. It serves as a major source of protein to lower- and middle-income nations and stands second to rice in meeting the calorie requirements. India stands second in terms of production after China. India produced 110.55 million tonnes of wheat during 2022–23, higher than the previous year1. Though India is seeing a trend of increase in wheat production every year, it is still having effort meeting the demands of an ever-growing population. The production and productivity of wheat is affected by several biotic and abiotic factors. Among biotic factors, wheat rusts caused by three Puccinia species (leaf rust or brown rust caused by Puccinia triticina, stem rust or black rust by Puccinia graminis f. sp. tritici and stripe rust or yellow rust by Puccinia striiformis Westend) are known to cause significant yield losses to wheat worldwide. The impact of rust on yield reduction in wheat ranges from 10% under moderate to 65% under intense epidemics2. However, 100% loss can occur due to stripe rust if it occurs on susceptible cultivars under favourable climatic conditions3,4. So, breeding for genetic resistance in wheat for these diseases is of utmost importance. Till now, a total of 83 leaf rust resistance (Lr) genes, 63 stem rust resistance (Sr) genes and 86 stripe rust resistance (Yr) genes have been documented5,6,7,8,9. Apart from these, several QTLs for leaf and stripe rust resistance have been identified and documented. However, many of these designated genes became ineffective over the years due to the evolution of new virulent pathotypes against them. Among the commonly used major leaf rust resistance genes like Lr1, Lr3, Lr9, Lr10, Lr13, Lr14a, Lr17, Lr19, Lr23, Lr24, Lr26 and Lr28, which exhibited all stage resistance, only Lr24 remained effective against prevalent pathotypes in India; however, races virulent on Lr24 have been reported from other parts of the world10. In the case of stripe rust, many previously deployed stripe rust resistance genes, such as Yr9 and Yr27, became ineffective due to the evolution of virulent pathotypes11,12,13.
Wild and related species of wheat carry enormous genetic variability that can be used in breeding programmes to broaden the genetic base of cultivated wheat. About half of the genes have been transferred to wheat through alien introgression14,15,16,17. Many wild species belonging to secondary and tertiary gene pools carry abundant genes resistant to biotic and abiotic stresses. However, they are still untapped because of the difficulty in making wide crosses and getting fertile F1 seeds16,18. One such wild species, Triticum militinae, belonging to the secondary gene pool of wheat, carries enormous genetic variability for disease resistance. T. militinae is considered a spontaneous mutant of T. timopheevii19. However, another view is that it originated from an introgressive hybridization between T. timopheevii and T. cathlicum Nevski (T. persicum Vav.)20. T. timopheevii has immunity to many diseases, such as leaf rust, stripe rust, stem rust, powdery mildew, loose smut, karnal bunt and dwarf smut21. However, till now, not a single disease resistance gene has been reported from T. militinae. As such, five-leaf rust resistance genes (Lr1822, Lr5023, LrTt124, LrTt225 and LrSelG1226), four stem rust resistance genes (Sr36, Sr40, SrTt3 and Sr37)27 and three powdery mildew resistance genes (Pm628, Pm2729 and Pm3730) were reported to be transferred from T. timopheevii to bread wheat. Breeders have always been interested in exploring this untapped wild species to identify useful resistance genes. At the Division of Genetics, IARI, New Delhi, the wheat group has developed 44 introgression lines (ILs), named TMD lines (T. militinae Derivatives). They were developed by crossing bread wheat lines, Chinese Spring and NI5439 with Triticum militinae acc. No. 117001 and backcrossing the F1s to Chinese Spring for three generations and further selfing of BC3F1 for seven generations. Earlier, Natraj et al. 2017 characterized three leaf rust resistance TMD lines (TMD6-4, TMD7-5 and TMD11-5) using SSR markers to identify the T. militinae introgression points in these three derivatives. In the current study, we have used 21 TMD lines (based on stability in their performance over the years) for screening at seedling and adult plant stages for leaf and stripe rusts. It gave us information on whether a particular derivative carries seedling or adult plant resistance or both. Also, the markers of known genes were used to screen these 21 TMD lines and to obtain initial knowledge of lines with unidentified sources of rust resistance.
Results
Cytological analysis
Cytological analysis of twenty-one TMD lines showed the presence of 42 chromosomes (21 bivalents) in all the derivatives during meiotic metaphase I. The presence of 21 bivalents indicates the cytological stability of these T. militinae derivatives and the absence of any numerical aberration in them. This study also indicates the suitability of these TMD lines for inheritance and mapping studies. Representative pictures showing 21 bivalents in some of the TMD lines are presented in Fig. 1.
Identification of leaf rust-resistant T. militinae derivatives
Out of 21 lines screened for leaf rust resistance, 17 lines (TMD6-4, TMD6-5, TMD7-5, TMD7-6, TMD9-1, TMD9-2, TMD11-5, TMD11-6, TMD12-4, TMD12-8, TMD12-12, TMD13-1, TMD13-3, TMD13-4, TMD13-7, TMD13-9 and TMD15) (Table 1) showed presence of SR (Seedling Resistant) genes as they were resistant at seedling and adult plant stages. Three lines (TMD2-7, TMD2-8 and TMD6-1) showed the presence of APR (Adult Plant Resistance) genes for leaf rust resistance as they were susceptible at the seedling stage but resistant at the adult plant stage (Table 1). One line, TMD17, was found to be susceptible to leaf rust at both stages. Seedling and adult plant reaction of TMD lines and their parental lines to leaf rust race 77–5 is presented in Fig. 2 and Table 1.
Screening of TMD lines and parental lines against leaf rust race 77–5 at the seedling stage. Here, 1: TMD2-7, 2: TMD2-8, 3: TMD6-1, 4: TMD6-4, 5: TMD6-5, 6: TMD7-4, 7: TMD7-5, 8: TMD9-1, 9: TMD9-2, 10: TMD11-5, 11: TMD11-6, 12: TMD12-4, 13: TMD12-8, 14: TMD12-12, 15: TMD13-1, 16: TMD13-3, 17: TMD13-4, 18: TMD13-7, 19: TMD13-9, 20: TMD15 and 21: TMD17.
Identification of stripe rust resistant T. militinae derivatives
Evaluation for stripe rust resistance identified eight lines (TMD6-4, TMD6-5, TMD11-6, TMD12-4, TMD12-8, TMD12-12, TMD13-7 and TMD13-9) with seedling resistance genes and nine lines (TMD2-7, TMD2-8, TMD6-1, TMD7-5, TMD7-6, TMD11-5, TMD13-1, TMD13-3 and TMD13-4) with APR genes. Four lines (TMD9-1, TMD9-2, TMD15 and TMD17) were susceptible to stripe rust at both the stages. Seedling and adult plant reaction of TMD lines, their parental lines and susceptible check Agra Local to stripe rust race 110S119 is presented in Fig. 3, Table 1 and Suppl. Fig. S1.
Screening of TMD lines and parental lines against stripe rust race 110S119 at the seedling stage. Here, 1:TMD2-7, 2:TMD2-8, 3:TMD6-1, 4:TMD6-4, 5:TMD6-5, 6:TMD7-4, 7: TMD7-5, 8: TMD9-1, 9: TMD9-2, 10: TMD11-5, 11: TMD11-6, 12: TMD12-4, 13: TMD12-8, 14: TMD12-12, 15: TMD13-1, 16: TMD13-3, 17: TMD13-4, 18: TMD13-7, 19: TMD13-9, 20: TMD15 and 21: TMD17.
Eight lines (TMD6-4, TMD6-5, TMD11-6, TMD12-4, TMD12-8, TMD12-12, TMD13-7 and TMD13-9) showed seedling resistance to both leaf and stripe rusts while three lines (TMD2-7, TMD2-8 and TMD6-1) showed adult plant resistance (APR) to both leaf and stripe rusts (Table 1). TMD17 showed susceptibility to both leaf and stripe rusts.
Screening with markers of known genes in T. militinae derivatives
Screening of TMD lines for Lr34/Yr18
As T. militinae derivatives were developed in the background of wheat cultivar Chinese Spring, which is known to carry linked genes Lr34/Yr18 (APR genes for leaf and stripe rust resistance), all the 21 TMD lines were screened with Lr34 specific marker, csLV3431. Out of 21 TMD lines, 10 lines showed the presence of Lr34/Yr18, while the remaining 11 lines showed its absence (Fig. 4; Table 2). Out of 10, three TMD lines (TMD2-7, TMD2-8 and TMD6-1) were identified as having adult plant resistance to both leaf and stripe rusts, three TMD lines (TMD7-5, TMD7-6 and TMD11-5) with seedling resistance to leaf rust and adult plant resistance to stripe rust and four TMD lines (TMD6-4, TMD6-5, TMD11-6 and TMD13-9) with seedling resistance to both leaf and stripe rusts (Table 1). Out of 10 marker positive TMD lines, six TMD lines (2–7, 2–8, 6–1, 7–5, 7–6 and 11–5) also showed leaf tip necrosis phenotype (phenotypic marker for Lr34/Yr18) at adult plant stage (Suppl. Fig. S1).
Screening of TMD lines for Lr18 and Lr50
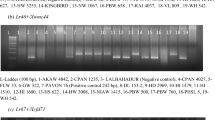
Screening of TMD lines with linked marker Xwmc7532 of Lr18 suggested the probable presence of Lr18 in 9 TMD lines (Table 2; Fig. 5). Apart from these, all other TMD lines showed absence of Lr18 in them. Out of these nine lines, two lines (TMD2-7 and TMD2-8) are identified as having adult plant resistance to both leaf and stripe rusts, four lines (TMD6-4, TMD6-5, TMD13-7 and TMD13-9) with seedling resistance to both leaf and stripe rusts and three lines (TMD13-1, TMD13-3 and TMD13-4) with seedling resistance to leaf rust but adult plant resistance to stripe rusts. T. militinae also showed a 180 bp band as of positive check T. timopheevii (Fig. 5), indicating the presence of Lr18 in T. militinae.
Screening with Xgwm38223 linked marker of Lr50 suggested the probable presence of Lr50 in eight T. militinae derivatives (Table 2; Fig. 6). Out of these, three lines (TMD2-7, TMD2-8, TMD6-1) are identified of having adult plant resistance to both leaf and stripe rusts, three lines (TMD6-5, TMD13-7 and TMD13-9) with seedling resistance to both leaf and stripe rusts and two lines (TMD13-1 and TMD13-3) with seedling resistance to leaf rusts but adult plant resistance to stripe rusts.
Discussion
Wild and related species of wheat carry enormous genetic variability that can be utilized in breeding programmes to broaden the genetic base of cultivated wheat. Triticum militinae, belonging to the secondary gene pool of wheat, carries enormous genetic variability for disease resistance. To date, no gene that confers rust resistance has been discovered in T. militinae. So, characterizing the resistance of T. militinae derivatives and knowing their cytological stability is important before using them in inheritance and mapping studies. T. militinae (AtAtGG; 2n = 4X = 28) is a tetraploid with 14 pairs of chromosomes, while T. aestivum (AABBDD; 2n = 6X = 42) is a hexaploid with 21 pairs of chromosomes. The transfer of genes from T. militinae to T. aestivum is possible through homologous and homoeologous pairing. 'G' genome of T. militinae has affinity to ‘B’ genome of wheat, which enable it to pair and exchange genomic regions33. Crossing of hexaploids with tetraploids produces sterility due to the production of univalents. With repeated backcrossing of F1 to hexaploid parent, there is a reduction in the number of univalents and an increase in the number of bivalents, which in turn increases the fertility of backcross derivatives. An increase in fertility levels from BC1 to BC2 generations is observed in a cross between T. aestivum and Ae. cylindrica34. T. militinae derivatives showed 21 bivalents as they have been developed by three generations of backcrossing and seven generations of selfing. The TMD lines with unidentified leaf and stripe rust resistance genes can now be used for inheritance and mapping studies as they are cytologically stable.
Evaluation of T. militinae derivatives for leaf and stripe rusts at seedling and adult plant stages and screening with gene based and linked markers of known genes provided insight into the distribution of known leaf rust resistance genes and their resistance pattern in T. militinae derivatives. Also, TMD lines with unknown rust-resistance genes were identified for further studies.
Out of seventeen TMD lines identified as carrying seedling resistance to leaf rust, ten TMD lines (7–5, 7–6, 9–1, 9–2, 11–5, 11–6, 12–12, 12–4, 12–8, and 15) found to carry the unknown gene(s) as they showed negative response to markers of known seedling resistance genes Lr18 and Lr50 from T. timopheevii. Out of ten, four lines (7–5, 7–6, 11–5 and 11–6) showed marker positivity to Lr34/Yr18, indicating a known adult plant resistance gene in addition to an unknown seedling resistance gene for leaf rust. The gene Lr34/Yr18 might have been introduced from Chinese Spring, which was used to develop TMD lines. The presence of Lr34/Yr18 in addition to Lr18 in three TMD lines (TMD6-4, TMD6-5 and TMD13-9) is expected to provide added seedling and adult plant resistances35.
Rest seven TMD lines with seedling resistance to leaf rust were found to be marker positive for Lr18, and some even showed the presence of Lr50 in them. Leaf rust resistance gene Lr18 provides seedling resistance to leaf rust but is temperature sensitive. It only provides effective resistance to its avirulent races at lower temperatures (18 °C) and becomes ineffective as the temperature goes beyond 25 °C36. So, to confirm the presence of Lr18, these lines need to be tested at different temperatures, or it can be confirmed through test of allelism studies. Even the lines that showed the absence of Lr18 can be tested similarly to confirm its absence. Although identified in some lines, the leaf rust resistance gene Lr50 is not effective against leaf rust race 77–5 used in the current study. Also, virulence towards Lr50 existed in races of P. triticina even before its deployment in wheat cultivars23. So, Lr50 can provide durable resistance only when it is pyramided along with other effective resistance genes23. So, the resistance pattern of TMD lines in the current study is mostly based on seedling resistance gene Lr18, adult plant resistance gene Lr34/Yr18 and/or some unknown genes in T. militinae.
Adult plant resistance to leaf and stripe rusts in three TMD lines (TMD2-7, TMD2-8 and TMD6-1) and identification of APR gene Lr34/Yr18 with marker csLV34 indicates resistance to leaf and stripe rusts are because of APR genes Lr34 and Yr18 respectively. Of the three, two lines, TMD2-7 and TMD2-8, showed the presence of Lr18 but did not show any seedling resistance to leaf rust. Apart from these two, all other TMD lines (6–4, 6–5, 13–1, 13–3, 13–4, 13–7 and 13–9) identified carrying Lr18 showed seedling resistance. The susceptibility of lines TMD2-7 and TMD2-8, even with the presence of Lr18, may be due to the occurrence of recombination events between the gene Lr18 and its marker XWmc75, present at a distance of 1.2 cM32. So, before using any of these 21 TMD lines in further studies, an allelism test with genetic stocks carrying Lr18 is required.
Screening for stripe rust resistance identified 8 TMD lines with seedling resistance and 9 TMD lines with APR to stripe rust. Out of 9 TMD lines with APR for stripe rust, 6 TMD lines (2–7, 2–8, 6–1, 7–5, 7–6 and 11–5) were found to carry the known gene Yr18/Lr34 from the wheat cultivar Chinese Spring, used in their development. Three TMD lines (13–1, 13–3 and 13–4) with negative responses to the Yr18/Lr34 marker indicate the presence of unknown adult plant resistance genes for stripe rust from T. militinae. Out of 8 TMD lines with seedling resistance to stripe rust, 4 TMD lines (6–4, 6–5, 11–6 and 13–9) also showed the presence of known APR gene, Yr18/Lr34 while 4 TMD lines (12–4, 12–8, 2–12 and 13–7) showed its absence. Therefore, all 8 TMD lines may carry unknown/unreported stripe rust resistance gene(s) from T. militinae, as till now, not a single stripe rust resistance gene has been reported from both T. militinae and T. timopheevii. So, the lines with unknown seedling and adult plant resistance genes for stripe rust can be explored soon for mapping and transfer in superior genetic backgrounds. The lines with unknown seedling resistance genes for stripe rust but with known APR gene, Yr18/Lr34, can be phenotyped in the seedling stage to tap the seedling stripe rust resistance gene while the other four (without Yr18/Lr34) can be tested at both the stages.
Except TMD 17, all other lines were found to carry resistance to either leaf or stripe rusts. This study also highlighted the presence of seedling and adult plant resistance genes in T. militinae derivatives, which can be used in future studies for mapping purposes and their utilization in breeding programmes to broaden the genetic base of wheat varieties.
Conclusion
The present study evaluated 21 T. militinae derivatives for leaf and stripe rusts at seedling and adult plant stages. This gave us information about lines carrying seedlings and adult plants' resistance to leaf rust, stripe rust, and leaf and stripe rust. To know whether the resistance in TMD lines is because of known genes or unknown genes, markers of two genes (Lr18 and Lr50) from T. timopheevii and one gene (Lr34/Yr18) from Chinese Spring were amplified. While ten TMD lines showed the probable presence of unknown genes for leaf rust resistance at the seedling stage, no lines with unknown genes for adult plant resistance to leaf rust could be identified. For stripe rust, while eight lines were identified as carrying unknown genes at the seedling stage, three were identified as carrying unknown genes at the adult plant stage. These lines were also found to be cytologically stable so that they can be utilized in inheritance and mapping studies in future.
Materials and methods
The plant materials used in the current study are twenty-one derivatives of T. militinae, parental lines, T. militinae (acc. No. 117001), Chinese Spring and NI5439 used in developing these derivatives and the susceptible check Agra Local. The T. militinae acc. no. 117001 was received in 1993 from the Institute of Plant Science Research, Norwich, Norfolk, England, Great Britain. Chinese Spring was used during the screening of TMD lines for leaf and stripe rust resistance and as a positive check for the gene Lr34/Yr18 during marker screening. The wheat genotypes NI5439, and Agra Local are two susceptible cultivars of India, susceptible to all three types of wheat rusts. While NI5439 and Agra Local were used as susceptible checks during screening for leaf and stripe rusts, only Agra Local was used when screening TMD lines for known markers. All the lines were screened with leaf rust pathotype 77–5 and stripe rust pathotype 110S119. In India, the leaf rust race 77–5 has remained most predominant for more than 20 years, but recently, the leaf rust race 77–9 became wider spread than 77–537. In the case of stripe rust, races 110S119, 238S119 and 46S119 are the three most prevalent and widespread races with a maximum frequency of 110S119 during 2021–2237. The race 77–5 is virulent on leaf rust resistance genes Lr1, Lr2a, Lr2c, Lr3a, Lr10, Lr13, Lr14a, Lr15, Lr17, Lr20, Lr23 and Lr26 and is avirulent on Lr9, Lr18, L19, Lr24 and Lr28. The stripe rust race 110S119 is virulent on stripe rust resistance genes Yr2, Yr2+, Yr3, Yr4, Yr6, Yr7, Yr8, Yr9, Yr6(un) Yr7(un) and avirulent on Yr1, Yr5, Yr9 + and Yr10. To identify new sources of leaf rust resistance, markers of designated leaf rust resistance genes, Lr18 and Lr50 from T. timopheevii, were screened in TMD lines, where T. timopheevii was used as a positive control. TMD lines were also screened with Lr34/Yr18 markers, as the Chinese Spring was used as a parental line during their development. TMD lines were screened for leaf and stripe rusts at seedling and adult plant stages. As most of the TMD lines showed resistance to either leaf, stripe or both diseases, their cytological stability was also studied at the meiotic metaphase stage so that they can be used in inheritance and mapping studies. The list of TMD lines used in the current study and their pedigree is given in Table 1. All methods were performed following the relevant guidelines/regulations/legislation.
Analysis of cytological stability of T. militinae derivatives
To study the cytological stability of T. militinae derivatives, immature spikes of appropriate size were collected during the booting stage. Anthers containing pollen mother cells (PMCs) at meiotic metaphase I were identified, fixed in Carnoy’s I fixative, and stored at 4 °C. For slide preparation, the anthers were hydrolyzed in a 1N HCl solution at 60 °C for 12 min. Following this, they were stained with Feulgen solution in the dark. Subsequently, slides were prepared using the squash method, utilizing 2% acetocarmine solution.
Screening for leaf and stripe rust resistance at the seedling stage
Screening for leaf and stripe rust resistance was carried out with pathotypes 77–5 and 110S119, respectively, in the glass house of the Division of Genetics. The screening experiments were repeated during 2018–19 and 2019–20. The screening materials included 21 TMD lines, their parental lines, T. militinae, Chinese Spring and NI5439 and the susceptible check Agra Local (AL). All the lines were sown in aluminium trays of 4 × 10 × 3 inches in size in the glass house at the Division of Genetics, IARI, New Delhi. Seedlings of ten days old with completely opened primary leaves were inoculated separately with leaf and stripe rust races. An inoculation mixture of concentration ̴6 × 105 spores/ml was prepared by adding urediospores in tap water. A drop of tween20 was added to the inoculation mixture to attain affinity and uniform spread of the urediospores on the leaf surface. After inoculation, the trays were kept in the incubation chamber under diffused light for 48 h and then shifted to glass house benches. Infection types on each introgression line were recorded 12–14 days after inoculation, following the standard scoring method38.
Screening for leaf and stripe rusts at the adult plant stage
For screening of TMD lines at the adult plant stage, all 21 TMD lines, along with their parental lines, were sown in leaf and stripe rust nurseries. In the rust nursery, each line was planted in 1 m rows, with infector rows planted after every 20 lines. Two rows of infectors were also planted in borders to have sufficient and uniform disease spread. Spores of leaf rust race 77–5 and stripe rust race 110S119 were sprayed in the respective rust nurseries as a suspension in water fortified with Tween20 (0.75 µl/ml) at an average concentration of 20 urediospores/microscopic field (10x × 10x). Inoculum suspension was also injected into the last internode of the plant with the help of a 2 ml hypodermic syringe at the boot leaf stage in the field before the emergence of the boot39. Plant response was recorded into five infection types40.
Screening with markers of known genes in T. militinae derivatives
To know whether these TMD lines carry known or unknown rust resistance genes, markers of leaf rust resistance genes Lr18 and Lr50 were used to screen TMD lines. Here, T. timopheevii was used as a positive check for Lr18 and Lr50 and Agra Local and NI5439 were used as negative checks. Though, no stripe rust resistance gene was reported from T. timopheevii or T. militinae, a marker of Yr18/Lr34 was also used in screening, as Chinese Spring was used as one of the parents in the development of TMD lines, which is known to carry linked stripe and leaf rust resistance gene Yr18/Lr34. Details of molecular markers used in the screening are presented in Table 3.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request. All methods were performed in accordance with the relevant guidelines/regulations/legislation by including a statement in the manuscript to this effect?
References
DES, MoA&FW, Government of India. https://desagri.gov.in/wp-content/uploads/2024/03/Time-Series-Production-2nd-AE-2023-24-English.pdf.
Saari, E. E. & Prescott, J. M. World distribution in relation to economic losses. In The Cereal Rusts, Diseases, Distribution, Epidemiology, and Control Vol. 2 (eds Roelfs, A. P. & Bushnell, W. R.) 259–298 (Academic Press, 1985).
Chen, X. M. Epidemiology and control of stripe rust (P. striiformis f. sp. tritici) on wheat. Can. J. Plant Pathol. 27, 314–337 (2005).
Bariana, H. et al. Adult plant stripe rust resistance gene Yr71 maps close to Lr24 in chromosome 3D of common wheat. Mol. Breed. 36, 98 (2016).
McIntosh, R. A., Dubcovsky, J., Rogers, W. J., Xia, X. C. & Raupp, W. J. Catalogue of gene symbols for wheat: 2020 supplement. Annu. Wheat Newsl. 66 (2020).
Kolmer, J. A., Bajgain, P., Rouse, M. N., Li, J. & Zhang, P. Mapping and characterization of the recessive leaf rust resistance gene Lr83 on wheat chromosome arm 1DS. Theor. Appl. Genet. 136(5), 115 (2023).
Mago, R. et al. Adult plant stem rust resistance in durum wheat Glossy Huguenot: Mapping, marker development and validation. Theor. Appl. Genet. 135, 1541–1550 (2022).
Klymiuk, V. et al. Discovery of stripe rust resistance with incomplete dominance in wild emmer wheat using bulked segregant analysis sequencing. Commun. Biol. 5, 826 (2022).
Zhu, Z. et al. Molecular characterization and validation of adult-plant stripe rust resistance gene Yr86 in Chinese wheat cultivar Zhongmai 895. Theor. Appl. Genet. 136, 142 (2023).
Huerta-Espino, J. et al. Global status of wheat leaf rust caused by Puccinia triticina. Euphytica 179(1), 143–160 (2011).
Line, R. F. & Qayoum, A. Virulence, Aggressiveness, Evolution, and Distribution of Races of Puccinia striiformis (the Cause of Stripe Rust of Wheat) in North America, 1968–87. Technical Bulletin Number 1788. United States Department of Agriculture, Agricultural Research Service, Washington DC (1992).
Chen, X. M., Penman, L., Wan, A. M. & Cheng, P. Virulence races of Puccinia striiformis f. sp. tritici in 2006 and 2007 and development of wheat stripe rust and distributions, dynamics, and evolutionary relationships of races from 2000 to 2007 in the United States. Can. J. Plant Pathol. 32(3), 315–333 (2010).
Tomar, S. M. S., Singh, S. K., Sivasamy, M. & Vinod,. Wheat rusts in India: Resistance breeding and gene deployment—a review. Indian J Genet. 74(2), 129–156 (2014).
Naik, B. K. et al. Molecular mapping and validation of the microsatellite markers linked to the Secale cereal derived leaf rust resistance gene Lr45 in wheat. Mol. Breed. 35, 1–10 (2015).
Niranjana, M. et al. Cytogenetic analysis and mapping of leaf rust resistance in Aegilops speltoides Tausch derived bread wheat line Selection2427 carrying putative gametocidal gene (s). Genome 60(12), 1076–1085 (2017).
Rani, K. et al. A novel leaf rust resistance gene introgressed from Aegilops markgrafii maps on chromosome arm 2AS of wheat. Theor. Appl. Genet. 133, 2685–2694 (2020).
Dinkar, V. et al. Molecular mapping of a new recessive wheat leaf rust resistance gene originating from Triticum spelta. Sci. Rep. 10, 22113 (2020).
Tomar, S. M. S., Vinod & Singh, B. Distant hybridization in wheat. Indian Agricultural Research Institute, New Delhi (2004).
Dorofeev, V. F. et al. AA World Wheat 75–87 (Agropromizdat, 1987).
Jarve, K., Jakobson, I. & Enno, T. Tetrapliod wheat species Triticum timopheevii and Triticum militinae in common wheat improvement. Acta Agron. Hung. 50, 463–477 (2002).
Nataraj, V. Cytogenetic characterization and molecular mapping of Triticum militinae derived leaf rust resistance in wheat. Ph. D. Thesis, Division of Genetics, IARI (2017).
Dyck, P. L. & Samborski, D. J. Genetics of resistance to leaf rust in the common wheat varieties Webster, Loros, Brevit, Carina, Malakof and Centenario. Can. J. Genet. Cytol. 10, 7–17 (1968).
Brown-Guedira, G. L., Singh, S. & Fritz, A. Performance and mapping of leaf rust resistance transferred to wheat from Triticum timopheevii subsp. armeniacum. Phytopathology 93(7), 784–789 (2003).
Leonova, I. et al. Identification of microsatellite markers for a leaf rust resistance gene introgressed into common wheat from Triticum timopheevii. Plant Breed. 123(1), 93–95 (2004).
Leonova, I. et al. Microsatellite mapping of a leaf rust resistance gene transferred to common wheat from Triticum timopheevii. Cereal Res. Commun. 38(2), 211–219 (2010).
Singh, A. K. et al. Genetics and mapping of a new leaf rust resistance gene in Triticum aestivum L. × Triticum timopheevii Zhuk. derivative ‘Selection G12’. J. Genet. 96(2), 291–297 (2017).
McIntosh, R. A. & Gyarfas, J. Triticum timopheevi as a source of resistance to wheat stem rust. Z. Pflanzenzucht. 66, 240–248 (1971).
Bro-Jorgensen, A. & Jensen, T. Gonococcal pharyngeal infections. Report of 110 cases. Br. J. Vener. Dis. 49(6), 491 (1973).
Jarve, K. et al. Chromosomal location of a Triticum timopheevii-derived powdery mildew resistance gene transferred to common wheat. Genome 43(2), 377–381 (2000).
Perugini, L. D., Murphy, J. P., Marshall, D. & Brown-Guedira, G. Pm37, a new broadly effective powdery mildew resistance gene from Triticum timopheevii. Theor. Appl. Genet. 116(3), 417–425 (2008).
Lagudah, E. S. et al. Molecular genetic characterization of the Lr34/Yr18 slow rusting resistance gene region in wheat. Theor. Appl. Genet. 114, 21–30 (2006).
Sadeghabad, A. A., Dadkhodaie, A., Heidari, B., Razi, H. & Mostowfizadeh-Ghalamfarsa, R. Microsatellite markers for the Triticum timopheevi-derived leaf rust resistance gene Lr18 on wheat 5BL chromosome. Breed. Sci. 67(2), 29–134 (2017).
Rodríguez, S., Maestra, B., Perera, E., Díez, M. & Naranjo, T. Pairing affinities of the B- and G-genome chromosomes of polyploid wheats with those of Aegilops speltoides. Genome 43(5), 814–819 (2000).
Zametra, R. S., Hansen, J. & Mallory-Smith, C. A. Potential for gene transfer between wheat (Triticum aestivum) and jointed goatgrass (Aegilops cyndrica). Weed Sci. 46, 313–317 (1998).
German, S. E. & Kolmer, J. A. Effect of gene Lr34 in the enhancement of resistance to leaf rust of wheat. Theor. Appl. Genet. 84(1–2), 97–105 (1992).
Dyck, P. L. & Johnson, R. Temperature sensitivity of genes for resistance in wheat to Puccinia recondita. Can. J. Plant Pathol. 5, 229–234 (1983).
ICAR-IIWBR 2022. Progress Report of All India Coordinated Wheat and Barley Improvement Project 2021–22. In Crop Protection (eds Kumar, S., Jasrotia, P., Kashyap, P. L., Kumar, R. & Singh, G. P.) ICAR- Indian Institute of Wheat and Barley Research, Karnal, Haryana, India.
Stakman, E. C., Stewart, D. M. & Loegering, W. Q. Identification of physiologic races of Puccinia graminis var. tritici. Agricultural Research Service E617. United States Department of Agriculture, Washington DC (1962).
Zadoks, J. C., Chang, T. T. & Konzak, C. F. A decimal code for the growth stages of cereals. Weed Res. 14(6), 415–421 (1974).
Johnston, C. O. & Browder, L. E. Seventh revision of the international register of physiologic races of Puccinia recondita f. sp. tritici. Plant Dis. Report. 50, 756–760 (1966).
Acknowledgements
The authors are grateful to the ICAR-Indian Institute of Wheat and Barley Research, Regional Station, Shimla, for providing pure inoculums of leaf and stripe rust pathogens and the ICAR-Indian Agricultural Research Institute, New Delhi, for facilitating the experiments.
Funding
This research work was funded by the Department of Biotechnology, Government of India. Project Title: Mapping and transfer of novel resistance genes for multiple biotic stresses in wheat (Triticum aestivum L.)” (Project code: 24-787).
Author information
Authors and Affiliations
Contributions
N.M. (Niharika) and V. conceived and designed the experiments; S.C., S.K.J., and N.M. (Niranjana) carried out the leaf rust phenotyping; S.C., R.K., and M.S.S carried out the stripe rust phenotyping; S.C. and S.B. performed the molecular work; S.C., M.K.C., and P.A analyzed the data; and N.M. wrote the paper. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chowdhury, S., Bansal, S., Jha, S.K. et al. Characterization and identification of sources of rust resistance in Triticum militinae derivatives. Sci Rep 14, 9408 (2024). https://doi.org/10.1038/s41598-024-59902-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-59902-x
- Springer Nature Limited