Abstract
Cutaneous leishmaniasis (CL) is a very common parasitic infection in subtropical areas worldwide. Throughout decades, there have been challenges in vaccine design and vaccination against CL. The present study introduced novel T-cell-based vaccine candidates containing IFN-γ Inducing epitopic fragments from Leishmania major (L. major) glycoprotein 46 (gp46), cathepsin L-like and B-like proteases, histone H2A, glucose-regulated protein 78 (grp78) and stress-inducible protein 1 (STI-1). For this aim, top-ranked human leukocyte antigen (HLA)-specific, IFN-γ Inducing, antigenic, CD4+ and CD8+ binders were highlighted. Four vaccine candidates were generated using different spacers (AAY, GPGPG, GDGDG) and adjuvants (RS-09 peptide, human IFN-γ, a combination of both, Mycobacterium tuberculosis Resuscitation promoting factor E (RpfE)). Based on the immune simulation profile, those with RS-09 peptide (Leish-App) and RpfE (Leish-Rpf) elicited robust immune responses and their tertiary structure were further refined. Also, molecular docking of the selected vaccine models with the human toll-like receptor 4 showed proper interactions, particularly for Leish-App, for which molecular dynamics simulations showed a stable connection with TLR-4. Upon codon optimization, both models were finally ligated into the pET28a( +) vector. In conclusion, two potent multi-epitope vaccine candidates were designed against CL and evaluated using comprehensive in silico methods, while further wet experiments are, also, recommended.
Similar content being viewed by others
Introduction
Leishmaniases are a heterogeneous group of parasitic diseases with considerable mortality, just second to the malaria, caused by the arthropod-borne flagellated protozoan, Leishmania spp., being vectored by the bites of infected phlebotomine sand flies1,2. Leishmania species are obligatory intracellular organisms, with macrophages as the main target cell3. Leishmaniases are represented with diverse clinical manifestations ranging from cutaneous lesions to the mucocutaneous involvement and fatal systemic infection. In total, 350 million individuals in 98 countries are threatened by different types of leishmaniasis. Cutaneous leishmaniasis (CL) is the most common form of the Disease, affecting 0.7–1.2 million people annually; Several species are known as the causative agents of CL, including Leishmania (L.) major, L. tropica, L. aethiopica, L. infantum and L. donovani with specific distribution in the Middle East (Iran, Iraq, Syria and Afghanistan), North Africa (Algeria) and South America (Brazil and Colombia)4. Rodents and humans are the principal reservoir hosts for L. major and L. tropica infections (Fig. 1). Actually, CL is a public health concern in some of the endemic areas and neither reservoir (humans, rodents) nor vector control is effective, Moreover, the available treatments are not safe and the efficacy is not high5. This issue justifies an urgent need for the discovery of novel vaccine targets and subsequent development of unprecedented vaccine candidates.
A variety of antigenic proteins are expressed in each life stages of Leishmania, i.e. amastigotes (within macrophages and dendritic cells), promastigotes and some intermediate forms (within sand fly gut), which might be directed towards vaccination strategies against leishmaniasis6. In previous studies, multiple L. major antigens have been discovered and used in immunization studies against leishmaniasis, most of which are conserved among other Leishmania species. Among these, some have been recognized with promising preventive effects, comprising glycoprotein 46 (gp46), cathepsin L-like and B-like proteases, histone H2A, glucose-regulated protein 78 (grp78) and stress-inducible protein 1 (STI-1)7,8. Cell-mediated immune responses may act as a double-edged sword during the course of CL lesion development and subsequent healing process; the IFN-γ cytokine, induced by type-I helper T-cells (Th1), is a potent killer of Leishmania parasites within the macrophages, through activation of macrophages and subsequent upsurge in nitric oxide (NO) and reactive oxygen species (ROS), while on the other hand Th2-mediated cytokines (Interleukin-4 (IL-4), IL-5 and IL-10) favor parasite survival, devastating the course of the infection9,10,11. Thus, in-depth exploration of antigenic vaccine candidates is advantageous to find novel immunodominant peptide targets, so-called as immunogenic epitopes.
Conventional vaccinology requires a vast amount of specialized biomolecular equipment, trained laboratory personnel, appropriate animal models, followed by long-lasting experimental intervals12,13,14. Advances in the biomedical informatics associated with novel genome sequencing technologies have led to the progression in biological databases and computational methods. This ongoing flow of information will increase our knowledge of host–pathogen interactions and aid in vaccinology-related research against infectious and non-infectious diseases. Such modalities give us the chance to recognize and organize novel antigenic proteins and specific B- and T-cell epitopes towards the development of safer and more effective next-generation vaccine candidates in a cost- and time-effective manner15,16,17,18. Therefore, in silico exploration of vaccine candidate antigens and their immunodominant epitope regions can provide important insights for future vaccinology studies. The present computer-based investigation was aimed at identification of immunogenic, IFN-γ-inducing epitopes with specific binding to the human leukocyte antigen (HLA) alleles in six antigenic proteins of L. major, i.e. gp46, CatL, CatB, H2A, grp78 and STI-1, in order to assemble novel T-cell-based multi-epitope vaccine candidates (MEVCs) against leishmaniasis.
Results
Selected human MHC-binders and engineering of MEVCs
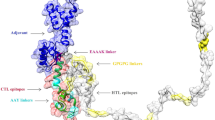
Following strict prediction and screening procedures applied to the six L. major vaccine candidate antigens, four CTL epitopes (CatB2-10, gp463-11, gp4627-35, gp4613-21,) and seven HTL epitopes (CatB12-26, grp7838-52, grp7837-51, gp46136-150, gp46137-151, gp46133-147, STI-110-24) were finally highlighted for designing MEVCs. In the present in silico study, four potent MEVCs were designed, based on different adjuvants and linkers, as follow: Seq1 or Leish-App) a TLR-4 agonist as RS-09 synthetic peptide (APPHALS) (N-terminal), His-Tag sequence (C-terminal), in addition to AAY and GPGPG linkers; Seq2 or Leish-Ifn) human IFN-γ as a genetic adjuvant (N-terminal), His-Tag sequence (C-terminal) with AAY and GPGPG linkers; Seq3 or Leish-Apfn) APPHALS (N-terminal) and human IFN-γ as adjuvants, with AAY and GPGPG linkers, and Seq4 or Leish-Rpf) M. tuberculosis resuscitation-promoting factor E (RpfE) as adjuvant (N-terminal), His-Tag sequence (C-terminal), and GDGDG as linker. In Seq1-3 constructs, “AAY” and “GPGPG” linkers were used to adjoin CTL and HTL epitopes, respectively, whereas in Seq4, “GDGDG” linker was used for both kind of epitopes. Also, “EAAAK” linker was used to connect sequences belonging to adjuvants to the rest of the MEVCs. The full details of the designed MEVC constructs are illustrated in Fig. 2.
Prediction of antigenicity, allergenicity, solubility and physico-chemical properties
Among four different MEVCs designed in the present study, the highest antigenicity predicted by VaxiJen server belonged to “Leish-App” and “Leish-Rpf” vaccine sequences, with 0.8376 and 0.7866, respectively. All MEVCs were approved to be non-allergenic in nature, as substantiated by AllergenFP v1.0 and AllerTOP v2.0 servers. The above-mentioned vaccine candidates possessed the highest solubility scores, with 0.545 (Leish-App) and 0.626 (Leish-Rpf), whilst the solubility of Leish-Apfn and Leish-Ifn vaccine candidates was 0.313 and 0.282, respectively, which was lower than the threshold (0.45). The lengths of Seq1-4 candidates, predicted using ProtParam server, were 218, 377, 366 and 391 residues, respectively. Also, Leish-Ifn and Leish-App had the highest and lowest MW, with 40.25 and 21.59 kDa, respectively. All of the designed candidates had extreme thermotolerance, as demonstrated by high aliphatic index. Moreover, all candidates, except Leish-App, had negative GRAVY scores, hence were recognized as hydrophilic in nature. Since the instability index of all vaccine candidates in the present study was below 40, they were assigned as “stable” polypeptides. Details of the physico-chemical properties of four designed MEVCs are provided in Table 1.
Structural analyses of different MEVCs
Regarding secondary structure analysis outputs using GOR IV online tool, random coils were the most prominent structures found in all MEVCs, with the exception of Leish-Apfn, where alpha helices were abundant (47.54%). The highest percentage of random coils were predicted in Leish-Rpf (59.34%), Leish-App (50.92%) and Leish-Ifn (44.83%), respectively (Supplementary Figures). In the following, a homology modelling prediction by I-TASSER server was used to illustrate the 3D model of the engineered MEVCs. A highest C-score usually supports a more reliable predicted model. On this basis, Leish-App model 1 (C-score: − 2.62), Leish-Ifn model 4 (C-score: − 2.28), Leish-Apfn model 2 (C-score: − 1.09) and Leish-Rpf model 3 (C-score: − 2.62) were selected among top five predicted models by I-TASSER web server (Supplementary Figures).
Simulation of the immune responses using MEVCs
Using C-ImmSim web server, three major immune-related parameters, including cytokine induction, Th cell population (cells per mm3) and Th cell population per state (cells per mm3) were evaluated for each vaccine candidate. Based on cytokines, IFN-γ in all candidates ranged between 4,00,000 ng/ml to 4,50,000 ng/ml, while IL-2, another Th1-specific cytokine, was higher (over 1,80,000 ng/ml) in Leish-App and Leish-Rpf vaccine candidates and lower rates (between 1,40,000 and 1,60,000 ng/ml) were elicited by the two remaining constructs. With respect to the Th cell population type, the highest number of specific memory T cells belonged to Leish-App (300 per mm3) and Leish-Rpf (between 250 and 300 per mm3) vaccine candidates. Additionally, Th cell immune responses by Leish-App and Leish-Rpf vaccine candidates showed the highest numbers of duplicating (near 500) and active (over 1500) cells per mm3. As compared with gp46 and LeIF as potent Leishmania vaccine candidate antigens, similar cytokine profile was elicited by the selected MEVCs, showing their potency (Supplementary Figures). Thus, based on the immune simulation results, Leish-App and Leish-Rpf vaccine models were chosen as the best immunogenic MEVCs for further analyses (Fig. 3).
Tertiary model refinement and validation
Pertinent to the GalaxyRefine server outputs, Leish-App model 4 was selected as the best model with acceptable rehashing and structural relaxation parameters, among other four refined models. The evaluated parameters for this model were as follows: GDT-HA of 0.9472, RMSD of 0.442, MolProbity of 2.513, Clash score of 24.8, Poor rotamers of 0.0 and Rama favored of 86.6. Regarding Leish-Rpf vaccine candidate, the refined model number 3 was selected, having GDT-HA of 0.9418, RMSD of 0.440, MolProbity of 2.380, Clash score of 17.3, Poor rotamers of 0.0 and Rama favored of 85.9. The quality improvement of the refined models for both candidates, in comparison with the crude models, was confirmed by the PROCHECK online tool. In the case of Leish-Rpf, the percentage of residue allocation in the crude and refined models, were as follows: 49.8% vs 72.7% (in the most favored regions), 42.1% vs 23.6% (in additional allowed regions), 5.7% vs 1.3% (in generously allowed regions) and 2.4% vs 2.4% (in disallowed regions). The crude model of Leish-App showed 72 residues (44.2%) in the most favored regions, followed by 76 residues (46.6%) in additional allowed regions, 7 residues (4.3%) in generously allowed regions as well as 8 residues (4.9%) in disallowed regions, whereas they were improved as 109 (66.9%), 45 (27.6%), 2 (1.2%) and 7 (4.3%) in the refined Leish-App model (Fig. 4).
Molecular docking analysis using human TLR-4 as receptor
Based on the ClusPro 2.0 web server for the protein–protein docking, the most populated docking clusters for Leish-App/TLR4 and Leish-Rpf/TLR4 had 217 and 40 members, respectively, with the highest binding scores of − 1121.4 and − 863.1. The non-bonded contacts (e.g., Van der Waals forces, hydrophobic forces, etc.) constituted the most part of interactions in Leish-App/TLR4 and Leish-Rpf/TLR4, with 154 and 189 contacts, respectively. The vaccines-receptor interactions have been illustrated in details in Fig. 5.
The representation of the whole docked complexes (vaccine-TLR4) and their respective residue-by-residue interactions. Regarding Leish-App/TLR-4 complex, there were 217 members with the lowest energy of − 1121.4, while in Leish-Rpf/TLR-4 complex, there observed 40 members with the lowest energy of − 863.1.
Stability of vaccine construct complexed with TLR-4 receptor
The simulation of the Leish-App-TLR4 complex was successfully done, based on the calculated average temperature of 300 K (range: 298–302 K), average pressure of 0.953073 bar (range: − 200 to + 200 bar), average potential energy of − 2695122.552 kJ/mol (range: − 270247 to − 2687123 kJ/mol) and average density of 1009 kg/m3 (range: 1006–1012) during 100 ns. Based on the total energy analysis, the energy values for the vaccine-ligand complex were converged, averaged − 2243714.486 kJ/mol. To understand the mobile nature of complex, the radius of gyration (Rg) was calculated, showing a constant rate during simulation for the receptor protein (3.195) and designed peptide (2.194), which means molecular stability during the process (Supplementary Figures). The RMSD plot of the human TLR-4 indicated values between 0.11 to 0.40 nm (Fig. 6A). The vaccine’s RMSD plot demonstrated an upward trend initially, with a peak at 0.84 nm after 46 ns of simulation, then reaching a relative plateau at 0.7 nm after 60 ns, due to the flexible nature of the receptor (Fig. 6B). In the following, the root mean square fluctuation (RMSF) values were plotted for both the receptor and Leish-App peptide. In the receptor protein, two peaks at the terminal regions indicated the highly-flexible parts (Fig. 6C). Also, due to the helical peptide structure, terminally-located residues were more flexible, while 5–25 residues showed more robust, hydrogenic bonds, hence lower fluctuation (Fig. 6D). The mean RMSF values for both peptide and protein were estimated to be 0.15 and 0.11, respectively. Hydrogen bond analysis, important for structural dynamics and complex stability, was done using gmx hbond order. The results showed that the peptide establishes stable hydrogen bonds (average: 9.4 bonds, up to 16 bonds) with the receptor during stimulation, leading to strong protein-peptide complex (Supplementary Figures).
Vaccine safety and in silico cloning
Based on the BLASTp tool output of the UniProtKB server, none of the selected vaccine sequences (Leish-App and Leish-Rpf) had homology with the human proteome, hence considered to be safe for administration. Since the cutting sites of Eco53KI and EcoRV were absent in both vaccine sequences, hence they were embedded at the N- and C-terminal of sequences, along with the Shine-Dalgarno sequence and start/stop codons. The reverse translated “Leish-App” and “Leish-Rpf” vaccine models were further codon optimized for enhanced expression in E. coli K12 strain using JCat web tool. A similar GC content was estimated in Leish-App and Leish-Rpf vaccine candidates, with 59.32% and 58.90, respectively, and the CAI values for both models were calculated to be 1. Ultimately, the final vaccine sequences were successfully cloned into the pET28a( +) plasmid using SnapGene v6.2.2. software. The total length of the formed clones was 4662 bp (Leish-App) and 5181 bp (Leish-Rpf) (Fig. 7).
Discussion
Leishmaniasis is highly prevalent in subtropical countries, with considerable burden every year19. Since vector and reservoir control strategies are not fully effective, development of efficacious vaccines seems to be an appropriate and sole preventive to reduce disease expansion20. Historically, leishmanization was shown to be the most effective strategy to control leishmaniasis21,22. Second and third generation vaccines such as the Leishmune (fucose-mannose ligand)23, Leish-Tec (Adenoviral-expressing L. donovani A2 protein)24, gp63 DNA vaccine25 as well as Leish-111f. recombinant vaccine (LeIF, LmSTI-1 and TSA) in Brazil, Peru and USA26 are promising examples of efficient immunization against leishmaniasis. Peptide-based immunogenic constructs, discovered by a number of in silico (B- and T-cell epitope prediction, protein localization, conservation analysis, etc.) and in vitro methods (Bio-panning assay, peptide-binding assay), constitute the next-generation vaccines, capable to efficiently target the appropriate classes of immune cells, in order to yield more effective immunity against a given pathogen27. This approach seems to be highly stable and reproducible, with decreased antigen complexity and lower costs for mass production. In this sense, the KSAC polyprotein has shown protection against L. major lesion formation in BALB/c mice28.
As mentioned earlier, Leishmania possesses a diverse array of antigenic compounds, which might be employed in rational vaccine design studies29. After preliminary in silico screening of 18 L. major vaccine candidate antigens, six proteins (gp46, CatL, CatB, H2A, grp78 and STI-1) were shown as the best targets for vaccination, since they had higher antigenicity scores and/or signal peptide/transmembrane domain. The surface glycoprotein gp46 is expressed in most parasite species and cells with reactivity against this protein were shown to produce a higher level of IFN-γ, conferring natural immunity30. It has, also, been demonstrated to elicit significant immune responses against L. amazonensis31 and L. major32 experimentally. Also, STI-1 is a novel antigen with copies of tetratricopeptide consensus motif with constitutive expression in both amastigote and promastigote stages33. Previously several studies have demonstrated its role as a potent vaccine candidate in immunization studies against CL34,35,36. The chaperonic protein, grp78, is functionally associated with the members of the highly-conserved 70 kDa stress protein family8. It has been shown that recombinant grp78 or its DNA induce protection in BALB/c, C57BL/6 and C3H/He mice against L. major infection37. Histones are evolutionarily-conserved, DNA-associated proteins, among which core histones can efficiently kill some Leishmania species, including L. major as antimicrobial peptides29. Cysteine proteases are key factors in extracellular matrix degradation and parasite propagation within host tissues38; among these, CatL and CatB molecules, found in all Leishmania species, play a pivotal role in parasite virulence39. Two expression plasmid containing cysteine proteinase genes (cpa and cpb) provided durable protection against CL through IFN-γ induction40. Based on the immunogenic capacity of these selected proteins, immunodominant HLA-binding epitopes were spotted in gp46, CatL, CatB, H2A, grp78 and STI-1 proteins of L. major and the potency of different designed MEVCs were evaluated using comprehensive in silico approaches.
At the first step, extensive epitope prediction and screening was performed using HLA reference set alleles, provided by the IEDB server. Herein, four T-cell-based MEVCs were designed and engineered using HLA-specific, IFN-γ Inducing epitopic fragments of six L. major antigens (gp46, CatL, CatB, H2A, grp78 and STI-1). Notably, B-cell epitopes were excluded from the prediction step, since humoral immune responses and higher levels of specific antibodies fail to provide protection against Leishmania infections, and may be strong predictors of parasite persistence41. Additionally, those CD8+, IFN-γ inducing T cell epitopes were predicted which are involved in parasite killing and lesion recovery41. A crucial part in the multi-epitope vaccine design is the utilization of appropriate linkers. Linkers or spacers are responsible for the flexibility of the polyprotein, its proper folding and segregation of functional domains, rendering a more stable protein structure42. The “AAY” linker, used to connect CTL epitopes in the current study, increases epitope partitioning and presentation through making the C-termini of CTL epitopes more accessible for binding43. Also, a glycine-rich linker, “GPGPG”, not only enhances the construct solubility, but also provides flexibility, high accessibility and free activity for adjacent domains. Of note, these epitopes are capable to induce HTL responses44–46. Consistent with our study, Khatoon et al.47 and Shams et al.48, also, employed AAY and GPGPG spacers to adjoin the epitope fragments of multi-epitope to be used against visceral leishmaniasis. In addition, “GDGDG” is known as a flexible linker, previously being used to link both CTL and HTL epitopes and design a multi-epitope candidate against L. major49. In the following, we used a rigid, non-flexible linker, “EAAAK”, after the adjuvant sequence, in order to prevent possible interaction with the rest of the vaccine sequence and prevent the formation of neo-epitopes50. Adjuvants are known as innate immune boosters or catalysts, and their utilization is implicated by the type of immune response required to be elicited. There are a wide range of genetic adjuvants, which can be embedded in the MEVCs51; in the present study, three different adjuvants were employed, such as a TLR-4 agonist (APPHALS), human IFN-γ and M. tuberculosis RpfE. The RS-09 synthetic peptide is an excellent adjuvant, previously used in many studies52,53,54. It allows the co-stimulation of CTL epitopes, providing a more robust immune response55. Moreover, another TLR-4 agonist, RpfE adjuvant, promotes dendritic cell maturation and enhances differentiation of naïve CD4+ T-cells toward Th1 population56. Also, the adjuvanticity of IFN-γ has been studied previously57,58,59.
The final stretch of four designed MEVCs was found to be 218 (Leish-App), 377 (Leish-Ifn), 366 (Leish-Apfn) and 391 (Leish-Rpf) residues. A good vaccine candidate should possess proper physico-chemical properties during production phase, along with strong immune stimulation. On this basis, the ProtParam web server was used for physico-chemical evaluation. The output showed that the largest MW belonged to Leish-Ifn (40.25 kDa), followed by Leish-Apfn (39.37 kDa) and Leish-Rpf (38.95 kDa), while the lowest MW was associated with the Leish-App candidate, with 21.59 kDa, rendering a more convenient extraction. With the exception of Leish-App, other designed vaccine candidates were shown to be highly hydrophilic in nature, based on a negative GRAVY score. Moreover, all candidates had stable molecular structure (instability index < 40) and were highly thermotolerant (high aliphatic index value). All candidates possessed no allergenic traits, through investigation by AllergenFP v1.0 and AllerTOP v2.0 servers, while two constructs (Leish-App and Leish-Rpf) demonstrated the highest antigenicity and solubility scores. Random coils were the most plentiful secondary structures in three candidates, except of Leish-Apfn, consistent with previous multi-epitope vaccine studies against L. major by Rabienia et al.49and against L. donovani by Khatoon et al.47.
In the present study, we selected two potent MEVCs, based on their immune profile shown by C-ImmSim web server. Actually, Leish-App and Leish-Rpf demonstrated extensive cell-mediate immune induction, in comparison with two other constructs, as evidenced by highest numbers of duplicating and active Th cells, specific memory T cells and elicited Th1-type cytokines (IFN-γ and IL-2). In comparison with Leishmania gp46 and LeIF protein sequences, as positive controls in immune simulation step, both vaccine candidates had similar expression of IFN-γ, as a representative of Th1-biased, protective response. Such immune excitement by these two vaccine candidates may partly arise from the immunogenic nature of the used adjuvants as potent TLR-4 agonists, i.e., RS-09 synthetic peptide and RpfE of M. tuberculosis. Next, the 3D structure of two selected MEVCs were further refined using GalaxyRefine server for global and regional structural relaxations. In the following, the Ramachandran plot analysis by the PROCHECK tool showed satisfactory results, with most residues tightly clustered in the most favored and additional allowed regions. Next, a protein–protein docking analysis was performed using ClusPro 2.0 server. Based on the member density and the estimated binding scores, Leish-App/TLR-4 docked complex possessed better interactions than Leish-Rpf/TLR-4. Accordingly, the MD simulation study was performed on Leish-App/TLR-4 complex. Deviations in the polypeptide chain structure, compared to the reference one (the structure after molecular docking) and during simulation were shown as RMSD plots. Our results revealed the stability of the vaccine-TLR4 complex during the MD simulation study. Also, the RMSF analysis was done to determine the most flexible residues; generally, most parts were relatively rigid and stable, particularly those residues located internally. Moreover, hydrogen bond calculation between the vaccine peptide and receptor protein demonstrated highly stable complex, with an average of 9.4 hydrogen bonds. It was noteworthy that both candidates showed no homology to the human proteome, hence were considered to be safe for human use. Ultimately, improvements in transcriptional and translational efficiency was done using codon optimization, directed towards high-level expression yield of the recombinant proteins. For this purpose, total GC content and CAI value of Leish-App and Leish-Rpf DNA sequences were analyzed against E. coli (strain K12) using JCat server. In the final step, two proposed MEVCs against L. major were successfully ligated into the pET28a( +) vector, being prepared for subsequent vaccine production.
In the literature, some studies have performed immunoinformatics-based predictions to design, engineer and evaluate different MEVCs against leishmaniases. In a study by Hashemzadeh et al. (2019), three L. infantum proteins, gp63, KMP-11 and HSP-70 were targeted for B- and T-cell epitope prediction and a 45.9 kDa polyprotein was designed using GGGGS and GSGSGS linkers, connected to two adjuvants (M. tuberculosis RpfE and RpfBG G5 domain)60. However, this study lacked the secondary structure analysis, molecular docking and vaccine immune profile evaluation. Rabienia et al.49 designed a 27.17 kDa MEVC using B-cell, T-cell and IFN-γ Inducing epitopes of L. major HASPB and KMP-11, in association with GDGDG linker and profilin as adjuvant; they showed proper and stable interaction between the vaccine model and TLR-11 receptor, without the prediction of immune stimulation. Another study by Yadav et al.61 designed a 71 kDa, stable and hydrophilic multi-component vaccine candidate using B- and T-cell epitopes derived from three L. donovani HyP proteins, prevailed by random coils, using AAY and KK spacers. Ropon-Palacios et al.62 investigated the in-silico binding of a novel multi-component vaccine (32.5 kDa), designed by 4 conserved epitopes from the Latin American species such as L. braziliensis, L. mexicana, L. panamensis and L. guyanensis, with TLR4/MD2 receptor complex, showing a stable interaction. In our study, four designed MEVCs were initially screened in terms of basic physico-chemical, antigenicity, allergenicity, solubility and immune profile characteristics, then two potent candidates were further evaluated regarding 3D structure refinement, molecular docking, vaccine safety and in silico cloning.
Conclusion
The safety and rational design nature of the multi-epitope vaccines have led researchers towards engineering more robust, stable and efficient vaccine candidates against many pathogens and cancers. Hence this novel field of immunoinformatics saves time and experimental resources and deserves further exploration. The aim of the present study was to design four novel MEVCs against CL through initial screening of 18 L. major vaccine candidate antigens (H1, H2A, H2B and H4, HSP60, HSP70, HSP83 (HSP90), HSP100, rP0, KMP11, TSA, LeIF, LACK, STI-1, gp46, CatL, CatB and grp78), and by arranging different linkers (GDGDG, GPGPG, AAY, EAAAK) and/or adjuvants (RS-09 synthetic peptide, RpfE and human IFN-γ) among IFN-γ Inducing T-cell epitopes. As a final word, two selected vaccine models in the present study (Leish-App and Leish-Rpf) demonstrated antigenic, allergenic, solubility, safety and physico-chemical properties. Moreover, adequate binding score and members were predicted between the vaccine candidates and TLR-4 receptor, along with robust cell-mediated immune stimulation. It is finally noteworthy that in vitro and in vivo experiments are demanded to validate the efficacy of the proposed MEVCs.
Methods
A schematic representation of the antigen selection and MEVC design is provided in Fig. 8.
Antigen selection and amino acid sequence retrieval
A preliminary study was done on 18 L. major vaccine candidate antigens explored in leishmaniasis vaccination studies, including histone proteins H1, H2A, H2B and H4, heat shock proteins HSP60, HSP70, HSP83 (HSP90), HSP100, ribosomal protein P0 (rP0), kinetoplast membrane protein 11 (KMP11), thiol-specific antioxidant (TSA), Leishmania elongation initiation factor (LeIF), Leishmania activated C-kinase antigen (LACK), STI-1, gp46, CatL, CatB and grp78 (data not shown); among these six antigens were shown to be highly antigenic (STI-1 and H2A) or possessed signal peptide and transmembrane domains (gp46, CatL, CatB, grp78), and they were included in the present study to design different antigenic MEVCs against CL. The amino acid sequence of six selected L. major proteins was retrieved through a leading high quality, comprehensive and freely-accessible resource of protein sequences and functional information, UniProt Knowledge Base63, available at https://www.uniprot.org/, with the following accession numbers: Q4Q6B6 (gp46), P90627 (CatB), Q4QI62 (CatL), Q4Q8E6 (grp78), Q4QEG6 (H2A) and Q4Q271 (STI-1).
Prediction of HLA-restricted IFN-γ Inducing Epitopes in six selected L. major antigens
Major histocompatibility complex class-II (MHC-II) binders, the so-called HTL epitopes, were predicted using MHC-II epitope prediction of the IEDB web server using recommended method (http://tools.iedb.org/mhcii/), “Human” as the target host and “HLA reference set alleles” option (Population coverage over 97%)64. The top-ten high-ranked epitopes possessing lower percentile rank were then screened regarding antigenicity and IFN-γ inducing epitopes, by using VaxiJen v2.0 (http://www.ddg-pharmfac.net/vaxijen/VaxiJen/VaxiJen.html) and IFNepitope (http://crdd.osdd.net/raghava/ifnepitope/) online tools, respectively. The latter employs a dataset of MHC-II-binding IFN-γ-inducers and non-inducers and the most accuracy can be reached by selecting hybrid model (> 81.39%), as we did in the current prediction (Supplementary Table 1). Those 9–10-mer CTL epitopes (MHC-I binders) specific to humans were predicted using IEDB MHC-I epitope prediction tool, available at http://tools.iedb.org/mhci/, using IEDB recommended method 2020.09 (NetMHCpan EL 4.1)64, with the selection of reference HLA allele set, including 16 class A alleles (01:01, 02:01, 02:03, 02:06, 03:01, 11:01, 23:01, 24:02, 26:01, 30:01, 30:02, 31:01, 32:01, 33:01, 68:01 and 68:02) and 11 class B alleles (07:02, 08:01, 15:01, 35:01, 40:01, 44:02, 44:03, 51:01, 53:01, 57 l:01 and 58:01)65. The top-ten high-affinity epitopes having a percentile rank < 1 were screened in terms of immunogenicity and IFN-γ induction using the immunogenicity tool in the IEDB server (http://tools.iedb.org/immunogenicity/) and IFNepitope server (Supplementary Table 2). All potent human CTL and HTL epitopes were, also, screened in terms of toxicity and allergenicity, using ToxinPred (http://crdd.osdd.net/raghava/toxinpred/) and AllergenFP v1.0 (https://www.ddg-pharmfac.net/AllergenFP/) web servers, respectively.
Design and assemblage of potent MEVCs
Accurately predicted and strictly screened human CTL (N = 4) and HTL (N = 7) epitopes were involved in the MEVC design process. In this study, three different adjuvants were used in designing four potent vaccine candidates, including a Toll-Like Receptor 4 (TLR-4) agonist (RS-09 peptide), human IFN-γ and Mycobacterium tuberculosis resuscitation-promoting factor E (RpfE). Moreover, different linkers such as “EAAAK”, “GDGDG”, “AAY’ and “GPGPG” were employed to connect adjuvants and/or immunodominant epitopes and design different MEVCs. Of note, a novel double-adjuvant vaccine candidate was introduced in the present study, incorporating RS-09 peptide and human IFN-γ at the N- and C-termini of the vaccine sequence, respectively.
Evaluation of allergenicity, antigenicity, solubility and physico-chemical properties of MEVCs
Two web servers were used to evaluate allergenicity, including AllergenFP v1.0 and AllerTOP v2.0 (https://www.ddg-pharmfac.net/AllerTOP/). The latter performs an 85.3% prediction through transforming the amino acid sequences into the integral vectors with equivalent lengths, based on auto cross covariance (ACC)66. Also, a novel alignment-independent, descriptor-based fingerprint method is used by AllergenFP v1.0 using physico-chemical and structural properties, making a prediction with 88% accuracy67. In the following, VaxiJen server was used to evaluate antigenicity of different MEVCs; which performs on the basis of ACC transformation of the sequences into uniform vectors, being truly depended on the proteins chemical nature and make predictions with 70–89% accuracy68. The Protein-Sol server, available at https://protein-sol.manchester.ac.uk/, was utilized for the prediction of protein solubility, so that protein scores over 0.45 are considered as soluble69. In the next step, major physico-chemical characteristics of the engineered MEVCs were predicted using ExPASy ProtParam web tool, available at https://web.expasy.org/protparam/. The server predicts several parameters, comprising molecular weight (MW), theoretical isoelectric point (pI), amino acid composition, in vitro and in vivo protein half-life, aliphatic index, instability index and the grand average of hydropathicity (GRAVY) score70.
Secondary and tertiary structure prediction for MEVCs
Secondary structure of MEVCs was predicted using Garnier–Osguthorpe–Robson IV (GOR IV) server (https://npsa-prabi.ibcp.fr/NPSA/npsa_gor4.html). This server predicts the number and percentage of residues in different secondary structures, including alpha helix, extended strand and random coil71. In the following, the Iterative Threading ASSEmbly Refinement (I-TASSER) server, available at https://zhanggroup.org/I-TASSER/, was used for three-dimensional (3D) modelling for the submitted MEVCs. It employs different 3D templates for the homology modelling of the input sequence, and finally provides tope five 3D models, having different C-scores. A C-score is an index for confidence of prediction, ranging between − 5 and 2, so that those models having higher C-scores can be considered as more reliable72.
Immune simulation profile of different MEVCs
The virtual immune simulation process elicited by four designed MEVs was predicted using C-ImmSim online server, available at http://150.146.2.1/C-IMMSIM/index.php. “These predictions are premised on PSSM for machine learning methods. The output implies to three stimulated regions including bone marrow, thymus and lymph node73. This computer-aided simulation was accomplished using default parameters with random seed 12,345, simulation volume 10 and simulation steps 100″. Based on the results, two highly immunogenic MEVCs were chosen for further analysis. For cytokine expression comparison, L. major gp46 (ID: Q4Q6B6) and LeIF (ID: W5XL77) protein sequences were selected from UniProt, as positive controls.
Tertiary model refinement and validations
The GalaxyRefine web server was employed to refine the 3D model of the best immunogenic MEVCs through mild/aggressive quality improvement. Several parameters are provided as output, encompassing global distance test-high accuracy (GDT-HA), root mean square deviation (RMSD), MolProbity, Clash score, Poor rotamers and Rama favored74. In the following, the quality of the rehashing process was validated, in comparison with crude models, using Ramachandran plot analysis; for this aim, the PROCHECK tool of the SAVES v6.0 server was used. “The server evaluates the stereochemical quality of a protein structure, producing a number of PostScript plots analyzing its overall and regional qualities”75.
Molecular interaction between selected MEVCs and TLR-4
For this aim, the 3D structure of human TLR-4/MD2 molecule (Accession No.: 3FXI) was retrieved via the PDB database of Research Collaboratory for Structural Bioinformatics (RCSB) (https://www.rcsb.org). Next, ClusPro 2.0 protein–protein docking web server, available at https://cluspro.bu.edu, was used to predict the affinity between the amino acid sequences of the best immunogenic MEVCs designed and engineered in the present study (as ligand) and human TLR-4 (as receptor) using default settings76. “The output of the server is provided as a top-rank cluster, among which the best docking pose is selected for visualization”77.
Molecular dynamics (MD) simulations
The MD study was carried out for Leish-App and human TLR-4 complex, using GROMCAS 2020 software. The biomolecular properties of Leishmania-associated multi-epitope vaccine construct complexed with TLR-4 were given to the software using pdb2gmx module. Next, the OPLS-AA/L all atom force field was selected, the complex was confined in a rectangular box, then dissolved in water. Using the solvate and editconf modules, the size of the system was determined based on the biomolecule extent, and the thickness of the water layer above and below the protein was considered to be 10 Å. The PME (Particle-mesh-Ewald) method was used to neutralize the system for long-range electrostatic interactions. Also, at this stage, KCl salt ions at a near physiological concentration (0.15 M) were added to the system for electrical neutralization. The number of Cl− and K+ ions was automatically determined by the ion accessible volume (V) and total system charge (Qsys). In the following, energy minimization (EM) was done in order to allow the system to reach the lowest possible energy and the most stable state possible prior to starting dynamics. This aim was done by PEM decreasing slope method and continued until the maximum force was smaller than 1000 kJ/mol/nm. Next, to prevent the system to crash, equilibration was applied between the solvent and the ions around the protein. Equilibration is often done in two steps. First, it takes place under a NVT (constant number of particles, volume and temperature) set, in which the temperature of the system must reach a plateau at the desired value. After stabilizing the temperature, pressure equilibration was performed under an NPT set using Parrinello-Rahman barostat, where the number of particles, pressure, and temperature are all constant. Finally, the simulation step was performed for 100 ns for the system containing the docked complex and the simulation quality was evaluated using the gmx check tool. The analysis was done with the gmx Energy tool.
Safety prediction, codon optimization and in silico cloning
For this aim, the BLAST online tool of the UniProtKB server (https://www.uniprot.org/blast) was utilized, in order to discriminate likely similar regions between selected vaccine candidate sequences and the respective organism. In this study, the human proteome was defined as target and identity rates over 35% mean homologous proteins with the human proteome78.
Efficient protein expression in the host bacterium, Escherichia coli, is a crucial step for subunit vaccine production. Accordingly, reverse translation of the selected MEVCs was done using the reverse translate tool of the Sequence Manipulation Suite (https://www.bioinformatics.org/sms2/rev_trans.html), followed by codon optimization using JCat server (http://www.jcat.de/). The JCat server assesses several important properties of the DNA sequence, such as GC content and codon adaptation index (CAI), which are significantly implicated in the expression of chimeric proteins in their respective hosts. Hence, in the present study, codon optimization was performed based on E. coli K12 strain79. Next, NEBcutter 2 server (https://nc2.neb.com/NEBcutter2/) was employed to evaluate the presence of cutting sites of different commercially available restriction enzymes within codon-adapted vaccine sequences. Ultimately, the cutting sites of two restriction enzymes, Eco53KI and EcoRV, were added to the 5′- and 3′-OH of the codon optimized vaccine sequence. Moreover, the Shine-Dalgarno sequence (AGGAGG) was added before the start codon to improve the yield of expression. The in-silico cloning process of the selected final multimeric vaccine models was accomplished using SnapGene v6.2.2. standalone software (from Insightful Science; available at https://www.snapgene.com).
Data availability
All data generated or analysed during this study are included in this published article (and its Supplementary Information files).
References
Khanjani, N. et al. Vaccines for preventing cutaneous leishmaniasis. Cochrane Database Syst. Rev. 2018, CD007634 (2018).
WHO. Leishmaniasis Fact Sheet (Accessed 7/13/2023). (World Health Organization, 2023).
Rostami, M. N. & Khamesipour, A. Potential biomarkers of immune protection in human leishmaniasis. Med. Microbiol. Immunol. 210, 81–100 (2021).
Postigo, J. A. R. Leishmaniasis in the world health organization eastern mediterranean region. Int. J. Antimicrob. Agents 36, S62–S65 (2010).
Health, M. O. Statistics of cutaneous leishmaniasis in Iran: Hearing before National Leishmaniasis Committee, office of zoonoses, center of disease control, Ministry of Health and Medical Education; (Sep. 30, 2004). (2004).
Kaye, P. M. et al. Overcoming roadblocks in the development of vaccines for leishmaniasis. Expert Rev. Vaccines 20, 1419–1430 (2021).
Dunning, N. Leishmania vaccines: From leishmanization to the era of DNA technology. Bioscience horizons 2, 73–82 (2009).
Nagill, R. & Kaur, S. Vaccine candidates for leishmaniasis: A review. Int. Immunopharmacol. 11, 1464–1488 (2011).
Gabriel, Á. et al. Cutaneous leishmaniasis: The complexity of host’s effective immune response against a polymorphic parasitic disease. J. Immunol. Res. 2019, 2603730 (2019).
Bogdan, C. & Röllinghoff, M. The immune response to Leishmania: Mechanisms of parasite control and evasion. Int. J. Parasitol. 28, 121–134 (1998).
Zhang, Y. et al. Metabotropic glutamate receptor 5-related autoimmune encephalitis with reversible splenial lesion syndrome following SARS-CoV-2 vaccination. Medicine 102(7), e32971. https://doi.org/10.1097/MD.0000000000032971 (2023).
Goodswen, S. J., Kennedy, P. J. & Ellis, J. T. A guide to current methodology and usage of reverse vaccinology towards in silico vaccine discovery. FEMS Microbiol. Rev. 47, fuad004 (2023).
Hu, F. et al. Spatiotemporal evolution of online attention to vaccines since 2011: an empirical study in China. Front. Public Health 10, 949482. https://doi.org/10.3389/fpubh.2022.949482 (2022).
Cui, Y. et al. MiR29a-3p improves acute lung injury by reducing alveolar epithelial cel PANoptosis. Aging dis. (2022).
Moxon, R., Reche, P. A. & Rappuoli, R. Reverse vaccinology. Front. Immunol. 10, 2776 (2019).
Xie, X. et al. New theoretical ISM-K2 Bayesian network model for evaluating vaccination effectiveness. J. Ambient Intelligence and Humanized Comput. (2023).
Zhang, Q. et al. X-linked Charcot-Marie-Tooth disease after SARS-CoV-2 vaccination mimicked stroke-like episodes: A case report. World J. Clin. Cases 11(2), 464–471. https://doi.org/10.12998/wjcc.v11.i2.464 (2023).
Yi, Z. et al. Propofol attenuates mast cell degranulation via inhibiting the miR‑221/PI3K/Akt/Ca2+ pathway. Exp. Ther. Med. https://doi.org/10.3892/etm.2018.6317 (2018).
Knight, C. A. et al. Leishmaniasis: Recent epidemiological studies in the Middle East. Front. Microbiol. 13, 1052478 (2023).
Malvolti, S., Malhame, M., Mantel, C. F., Le Rutte, E. A. & Kaye, P. M. Human leishmaniasis vaccines: Use cases, target population and potential global demand. PLoS Negl. Trop. Dis. 15, e0009742. https://doi.org/10.1371/journal.pntd.0009742 (2021).
Khamesipour, A. et al. Leishmanization: Use of an old method for evaluation of candidate vaccines against leishmaniasis. Vaccine 23, 3642–3648 (2005).
Mohebali, M., Nadim, A. & Khamesipour, A. An overview of leishmanization experience: A successful control measure and a tool to evaluate candidate vaccines. Acta Tropica 200, 105173 (2019).
Parra, L. et al. Safety trial using the Leishmune® vaccine against canine visceral leishmaniasis in Brazil. Vaccine 25, 2180–2186 (2007).
Evans, K. J. & Kedzierski, L. Development of vaccines against visceral leishmaniasis. J. Trop. Med. 2012, 892817 (2012).
Yang, D. et al. Oral Salmonella typhimurium (AroA-) vaccine expressing a major leishmanial surface protein (gp63) preferentially induces T helper 1 cells and protective immunity against leishmaniasis. J. Immunol. (Baltimore, Md.: 1950) 145, 2281–2285 (1990).
Ghalib, H. & Modabber, F. Consultation meeting on the development of therapeutic vaccines for post kala azar dermal leishmaniasis. Kinetoplastid Biol. Dis. 6, 1–14 (2007).
De Brito, R. C. et al. Peptide vaccines for leishmaniasis. Front. Immunol. 9, 1043 (2018).
Gomes, R. et al. KSAC, a defined Leishmania antigen, plus adjuvant protects against the virulence of L. major transmitted by its natural vector Phlebotomus duboscqi. PLoS Negl. Trop. Dis. 6, e1610 (2012).
Singh, B. & Sundar, S. Leishmaniasis: Vaccine candidates and perspectives. Vaccine 30, 3834–3842 (2012).
Kemp, M. et al. The Leishmania promastigote surface antigen-2 (PSA-2) is specifically recognised by Th1 cells in humans with naturally acquired immunity to L. major. FEMS Immunol. Med. Microbiol. 20, 209–218 (1998).
Champsi, J. & McMahon-Pratt, D. Membrane glycoprotein M-2 protects against Leishmania amazonensis infection. Infect. Immun. 56, 3272–3279 (1988).
Handman, E., Symons, F. M., Baldwin, T. M., Curtis, J. M. & Scheerlinck, J. Protective vaccination with promastigote surface antigen 2 from Leishmania major is mediated by a TH1 type of immune response. Infect. Immun. 63, 4261–4267 (1995).
Webb, J. R., Kaufmann, D., Campos-Neto, A. & Reed, S. G. Molecular cloning of a novel protein antigen of Leishmania major that elicits a potent immune response in experimental murine leishmaniasis. J. Immunol (Baltimore, Md.: 1950) 157, 5034–5041 (1996).
Badiee, A., Jaafari, M. R. & Khamesipour, A. Leishmania major: Immune response in BALB/c mice immunized with stress-inducible protein 1 encapsulated in liposomes. Exp. Parasitol. 115, 127–134 (2007).
Campos-Neto, A. et al. Protection against cutaneous leishmaniasis induced by recombinant antigens in murine and nonhuman primate models of the human disease. Infect. Immun. 69, 4103–4108 (2001).
Campos-Neto, A. et al. Vaccination with plasmid DNA encoding TSA/LmSTI1 leishmanial fusion proteins confers protection against Leishmania major infection in susceptible BALB/c mice. Infect. Immun. 70, 2828–2836 (2002).
Jensen, A. T., Curtis, J., Montgomery, J., Handman, E. & Theander, T. G. Molecular and immunological characterisation of the glucose regulated protein 78 of Leishmania donovani. Biochim. Biophys. Acta (BBA)-Protein Struct. Mol. Enzymol. 1549, 73–87 (2001).
Yao, C., Donelson, J. E. & Wilson, M. E. The major surface protease (MSP or GP63) of Leishmania sp. Biosynthesis, regulation of expression, and function. Mol. Biochem. Parasitol. 132, 1–16 (2003).
Selzer, P. M. et al. Cysteine protease inhibitors as chemotherapy: Lessons from a parasite target. Proc. Natl. Acad. Sci. 96, 11015–11022 (1999).
Rafati, S., Salmanian, A.-H., Taheri, T., Vafa, M. & Fasel, N. A protective cocktail vaccine against murine cutaneous leishmaniasis with DNA encoding cysteine proteinases of Leishmania major. Vaccine 19, 3369–3375 (2001).
Scott, P. & Novais, F. O. Cutaneous leishmaniasis: Immune responses in protection and pathogenesis. Nat. Rev. Immunol. 16, 581–592 (2016).
Parvizpour, S., Pourseif, M. M., Razmara, J., Rafi, M. A. & Omidi, Y. Epitope-based vaccine design: A comprehensive overview of bioinformatics approaches. Drug Discov. Today 25, 1034–1042 (2020).
Hasan, M. & Mia, M. Exploratory algorithm of a multi-epitope-based subunit vaccine candidate against cryptosporidium hominis: Reverse vaccinology-based immunoinformatic approach. Int. J. Peptide Res. Ther. 28, 134 (2022).
Tarrahimofrad, H., Rahimnahal, S., Zamani, J., Jahangirian, E. & Aminzadeh, S. Designing a multi-epitope vaccine to provoke the robust immune response against influenza A H7N9. Sci. Rep. 11, 24485 (2021).
Feng, S., Song, G., Liu, L., Liu, W., Liang, G., Song, Z. Allergen-specific immunotherapy induces monocyte-derived dendritic cells but attenuates their maturation and cytokine production in the lesional skin of an atopic dermatitis mouse model. J. Dermatology https://doi.org/10.1111/1346-8138.16582 (2022).
Liu, L., Song, G., Song, Z. Intrinsic Atopic Dermatitis and Extrinsic Atopic Dermatitis: Similarities and Differences. Clinical, Cosmetic and Investigational Dermatology, (2022).
Khatoon, N., Pandey, R. K. & Prajapati, V. K. Exploring Leishmania secretory proteins to design B and T cell multi-epitope subunit vaccine using immunoinformatics approach. Sci. Rep. 7, 1–12 (2017).
Shams, M. et al. Engineering a multi-epitope vaccine candidate against Leishmania infantum using comprehensive immunoinformatics methods. Biologia 77, 277–289 (2022).
Rabienia, M. et al. Exploring membrane proteins of Leishmania major to design a new multi-epitope vaccine using immunoinformatics approach. Eur. J. Pharm. Sci. 152, 105423 (2020).
Arai, R., Ueda, H., Kitayama, A., Kamiya, N. & Nagamune, T. Design of the linkers which effectively separate domains of a bifunctional fusion protein. Protein Eng. 14, 529–532 (2001).
Li, Q. et al. Built-in adjuvants for use in vaccines. Eur. J. Med. Chem. 227, 113917 (2022).
Khan, M. A. A. et al. An immunoinformatic approach driven by experimental proteomics: In silico design of a subunit candidate vaccine targeting secretory proteins of Leishmania donovani amastigotes. Parasites Vectors 13, 1–21 (2020).
Onile, O. S., Fadahunsi, A. I., Adekunle, A. A., Oyeyemi, B. F. & Anumudu, C. I. An immunoinformatics approach for the design of a multi-epitope subunit vaccine for urogenital schistosomiasis. PeerJ 8, e8795 (2020).
Tarang, S. et al. In silico design of a multivalent vaccine against Candida albicans. Sci. Rep. 10, 1–7 (2020).
Coler, R. N. et al. A synthetic adjuvant to enhance and expand immune responses to influenza vaccines. PloS one 5, e13677 (2010).
Choi, H. G. et al. Mycobacterium tuberculosis RpfE promotes simultaneous Th1-and Th17-type T-cell immunity via TLR4-dependent maturation of dendritic cells. Eur. J. Immunol. 45, 1957–1971 (2015).
Kim, J. et al. Coimmunization with IFN-gamma or IL-2, but not IL-13 or IL-4 cDNA can enhance Th1-type DNA vaccine-induced immune responses in vivo. J. Interf. Cytokine Res.: Off. J. Int. Soc. Interf. Cytokine Res. 20, 311–319 (2000).
Suárez-Méndez, R. et al. Adjuvant interferon gamma in patients with drug–resistant pulmonary tuberculosis: A pilot study. BMC Infect. Dis. 4, 1–8 (2004).
Zanetti, B. F., Ferreira, C. P., Vasconcelos, J. R. C. & Han, S. W. Adjuvant properties of IFN-γ and GM-CSF in the scFv6. C4 DNA vaccine against CEA-expressing tumors. Gene Ther. 30, 41–50 (2023).
Hashemzadeh, P., Ghorbanzadeh, V., Lashgarian, H. E., Kheirandish, F. & Dariushnejad, H. Harnessing bioinformatic approaches to design novel multi-epitope subunit vaccine against Leishmania infantum. Int. J. Pept. Res. Ther. 26, 1417–1428 (2020).
Yadav, S. et al. Design of a multi-epitope subunit vaccine for immune-protection against Leishmania parasite. Pathog. Glob. Health 114, 471–481 (2020).
Ropón-Palacios, G., Chenet-Zuta, M. E., Otazu, K., Olivos-Ramirez, G. E. & Camps, I. Novel multi-epitope protein containing conserved epitopes from different Leishmania species as potential vaccine candidate: Integrated immunoinformatics and molecular dynamics approach. Comput. Biol. Chem. 83, 107157 (2019).
UniProt. UniProt: A worldwide hub of protein knowledge. Nucleic Acids Res. 47, D506–D515 (2019).
Vita, R. et al. The immune epitope database (IEDB): 2018 update. Nucleic Acids Res. 47, D339–D343 (2019).
Pourseif, M. M. et al. A domain-based vaccine construct against SARS-CoV-2, the causative agent of COVID-19 pandemic: Development of self-amplifying mRNA and peptide vaccines. BioImpacts: BI 11, 65 (2021).
Dimitrov, I., Bangov, I., Flower, D. R. & Doytchinova, I. AllerTOP v. 2—a server for in silico prediction of allergens. J. Mol. Model. 20, 1–6 (2014).
Dimitrov, I., Naneva, L., Doytchinova, I. & Bangov, I. AllergenFP: Allergenicity prediction by descriptor fingerprints. Bioinformatics 30, 846–851 (2014).
Doytchinova, I. A. & Flower, D. R. VaxiJen: A server for prediction of protective antigens, tumour antigens and subunit vaccines. BMC Bioinform. 8, 1–7 (2007).
Hebditch, M., Carballo-Amador, M. A., Charonis, S., Curtis, R. & Warwicker, J. Protein–Sol: A web tool for predicting protein solubility from sequence. Bioinformatics 33, 3098–3100 (2017).
Gasteiger, E. et al. Protein identification and analysis tools on the ExPASy server (Springer, 2005).
Garnier, J., Gibrat, J.-F. & Robson, B. in Methods Enzymol. Vol. 266, 540–553 (Elsevier, 1996).
Roy, A., Kucukural, A. & Zhang, Y. I-TASSER: A unified platform for automated protein structure and function prediction. Nat. Protoc. 5, 725–738 (2010).
Rapin, N., Lund, O., Bernaschi, M. & Castiglione, F. Computational immunology meets bioinformatics: The use of prediction tools for molecular binding in the simulation of the immune system. PLoS One 5, e9862 (2010).
Heo, L., Park, H. & Seok, C. GalaxyRefine: Protein structure refinement driven by side-chain repacking. Nucleic Acids Res. 41, W384–W388 (2013).
Laskowski, R., MacArthur, M. & Thornton, J. in International Tables for Crystallography Vol. F (eds M.G. Rossmann & E. Arnold) 722–725 (2006).
Kozakov, D. et al. The ClusPro web server for protein–protein docking. Nat. Protoc. 12, 255 (2017).
Shams, M. et al. Towards the first multiepitope vaccine candidate against Neospora caninum in mouse model: Immunoinformatic standpoint. BioMed Res. Int. 2022, 2644667 (2022).
Pundir, S., Martin, M. J., O’Donovan, C. & Consortium, U. UniProt tools. Curr. Protoc. Bioinform. 53, 1–29 (2016).
Grote, A. et al. JCat: A novel tool to adapt codon usage of a target gene to its potential expression host. Nucleic Acids Res. 33, W526–W531 (2005).
Acknowledgements
This work would not have been possible without the support of the "BioinfCamp.com" site. We want to thank "BioinfCamp" for sharing information about the vaccine designing process with us during this research.
Author information
Authors and Affiliations
Contributions
H.M., M.S. and E.R.B. conceived and designed the study protocol; E.R.B., M.A., M.M. and M.A. performed epitope selection, vaccine design and evaluation; M.K., S.Gh. and H.I. conducted immune simulation, molecular docking, molecular dynamic simulations, codon optimization, and in silico cloning. E.R.B., M.A. and H.M. drafted the manuscript. H.M., A. Kh. And M.S. reviewed and revised the manuscript draft. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Basmenj, E.R., Arastonejad, M., Mamizadeh, M. et al. Engineering and design of promising T-cell-based multi-epitope vaccine candidates against leishmaniasis. Sci Rep 13, 19421 (2023). https://doi.org/10.1038/s41598-023-46408-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-46408-1
- Springer Nature Limited