Abstract
Erwinia tracheiphila is a bacterial plant pathogen that causes a fatal wilt infection in some cucurbit crop plants. Wilt symptoms are thought to be caused by systemic bacterial colonization through xylem that impedes sap flow. However, the genetic determinants of within-plant movement are unknown for this pathogen species. Here, we find that E. tracheiphila has horizontally acquired an operon with a microbial expansin (exlx) gene adjacent to a glycoside hydrolase family 5 (gh5) gene. Plant inoculation experiments with deletion mutants in the individual genes (Δexlx and Δgh5) and the full operon (Δexlx–gh5) resulted in decreased severity of wilt symptoms, decreased mortality rate, and impaired systemic colonization compared to the Wt strain. Co-inoculation experiments with Wt and Δexlx–gh5 rescued the movement defect of the mutant strain, suggesting that expansin and GH5 function extracellularly. Together, these results show that expansin–GH5 contributes to systemic movement through xylem, leading to rapid wilt symptom development and higher rates of plant death. The presence of expansin genes in diverse species of bacterial and fungal wilt-inducing pathogens suggests that microbial expansin proteins may be an under-appreciated virulence factor for many pathogen species.
Similar content being viewed by others
Introduction
The surfaces of all land plants are colonized by complex microbial communities. For a microbe, the ability to colonize a plant increases access to the nutritional resources produced by that plant1,2. This has driven the evolution of diverse molecular mechanisms for plant colonization throughout commensal, beneficial and pathogenic microbes3. Proteins called ‘expansins’ are particularly intriguing, and genes coding for expansins are being identified in the genomes of an increasing number of plant-associated bacterial and fungal species4,5,6,7. Expansins are non-enzymatic, two-domain proteins of ~ 250 amino acids. The expansin N-terminal domain is related to glycoside hydrolase family 45 functional domains and the C-terminal domain is related to grass pollen allergens8,9,10. In all species of land plants and green algae, expansin-coding genes are ubiquitous and fulfill the essential role of non-enzymatically loosening cell wall cellulose and enabling cell wall extension during normal growth8,11,12,13,14,15.
Expansin-encoding genes have also been identified in hundreds of taxonomically diverse bacterial and fungal species which do not have cellulosic cell walls, but interact with live or dead plant or algal matter4,6,7,16. In these microbial species, expansin proteins are hypothesized to promote colonization of plants through interactions with structural cellulose and/or hemicellulose in plant cell walls4,7,17. However, the functions and importance of microbial expansins for plant colonization have been empirically investigated in very few microbial species4. For several microbes that are non-pathogenic, expansins increase colonization efficiency of plant surfaces4,17,18,19. The microbial expansin from the plant commensal bacterium Bacillus subtilis (BsEXLX1) has only a fraction of the cellulose loosening activity against plant cell walls in vitro compared to plant expansins. Yet, BsEXLX1 deletion mutants are either severely impaired or unable to successfully colonize the surface of maize roots17,20. In some species of plant beneficial fungi, expansins (also referred to as ‘swollenins’) increase fungal mutualistic capabilities towards plant hosts18,19. Expansin function has also been investigated in several species of plant pathogens. In the bacterial plant pathogen Ralstonia solanacearum, an expansin deletion mutant has decreased virulence21. Expansin from Pectobacterium atrosepticum contributes to virulence in potato, and induces plant defense responses22. For Clavibacter michiganensis, studies have described contradictory expansin roles for virulence and ability to colonize xylem21,23,24,25,26. Outside of these few examples, fundamental questions surrounding how microbial expansins mediate plant colonization, and the molecular mechanism(s) by which these proteins interact with plant structural carbohydrates remain unknown for almost all species4,6,7,16,17,21,24,27,28.
The bacterial plant pathogen Erwinia tracheiphila Smith (Enterobacteriaceae), the causative agent of bacterial wilt of cucurbits, contains an operon with an expansin gene (exlx) and a glycoside hydrolase family 5 gene (gh5)7,29,30,31. E. tracheiphila colonizes host plant xylem and causes a fatal bacterial wilt infection in only two genera of cucurbit host plants: cultivars of Cucurbita spp. (summer and winter squash) and Cucumis spp. (cucumber and muskmelon)32. The E. tracheiphila genome has undergone dramatic structural changes consistent with an evolutionarily recent emergence into a novel host plant population, including the horizontal acquisition of multiple genes likely important for virulence32,33,34,35,36. Unlike most bacterial plant pathogens, E. tracheiphila cannot persist in environmental reservoirs34,37,38. Instead, E. tracheiphila is obligately transmitted by two species of highly specialized leaf beetle vectors39,40,41,42. Pathogen transmission can occur when E. tracheiphila cells in frass from infective beetles is deposited near recent foliar feeding wounds or on floral nectaries40,43. Bacteria can then move systemically through xylem and block sap flow to induce systemic wilting (Fig. 1), which is followed by plant death within 2–3 weeks after the first wilt symptoms appear33,44,45,46. E. tracheiphila costs cucurbit farmers millions of dollars annually through direct yield losses and the expense of indirect control measures32. Despite the economic burden caused by E. tracheiphila, no genetic determinants of bacterial pathogenesis or virulence have yet been empirically determined.
Xylem colonization by E. tracheiphila. (a) Visible strain of E. tracheiphila (Wt) oozing from xylem in all vascular bundles of a symptomatic plant after a horizontal stem cross section cut. (b) ×20 confocal microscopy image of a longitudinal section of a symptomatic, E. tracheiphila infected Cucurbita pepo stem. Image is falsely colored so that plant structures are shown in blue and live E. tracheiphila bacterial cells are red.
Here, we reconstruct the evolutionary histories of both the exlx and gh5 open reading frames (ORFs) in E. tracheiphila, and characterize the role of expansin–GH5 for colonization of squash (Cucurbita pepo) host plants. We find that the phylogenies of both exlx and gh5 are consistent with horizontal acquisition by E. tracheiphila. In planta inoculation experiments with deletion mutants for exlx and gh5 show these genes significantly increase the ability of E. tracheiphila to systemically colonize xylem (Fig. 1), induce wilt symptoms, and cause high rates of plant death. Co-inoculation experiments with Wt and deletion mutants suggest that these proteins likely function extracellularly, probably as an assembled EXLX–GH5 complex. Together, these results suggest that the Et–exlx–gh5 locus is a non-canonical yet potent virulence factor, and horizontal acquisition of this locus was a key event driving the recent emergence of E. tracheiphila32,35 as a virulent plant pathogen that causes high mortality by inducing wilt symptoms and plant death by blocking the flow of xylem sap (Fig. 1). These findings highlight the continued risk of horizontal gene transfer driving an increase in pathogen virulence, and the continuing vulnerability of agricultural populations to invasion by virulent new pathogens.
Results
Identification of a locus with an expansin gene in Erwinia tracheiphila
We identified a locus with two open reading frames (ORFs) flanked by mobile DNA elements during manual curation of ab initio gene predictions in the E. tracheiphila reference strains (Fig. 2A)29,30. The first ORF, Et–exlx (AXF78871.1), is predicted to encode a protein product with 243 amino acids containing both domains found in canonical expansin proteins14,47. The second ORF, Et–gh5 (AXF77819.1), has 315 codons and is predicted to encode a putatively pseudogenized endo-1,4-beta-xylanase A precursor (EC 3.2.1.8) with a glycoside hydrolase family 5 (GH5) functional domain (www.CAZy.org)48. Many Et–gh5 homologs in the NCBI nr database contain 415–450 amino acids, and RAST ab initio gene annotation predicts that the truncation to 315 amino acids eliminates cellulase activity and renders Et–gh5 non-enzymatic49. The sequences of both ORFs predict a signal peptide for secretion and a Signal Peptidase cleavage site50, suggesting that each protein product is individually secreted.
Genomic context and phylogenies of the expansin and glycoside hydrolase 5 genes in Erwinia tracheiphila. (A) Genomic context of the expansin (exlx) and glycoside hydrolase 5 (gh5) open reading frames (ORFs) in Erwinia tracheiphila. The ORFs and intergenic spaces are drawn to scale, with the black line representing position on the chromosome, and each ORF as an arrow color-coded according to ab initio annotated function. The scale bar is the length in nucleotides of the ORFs and intergenic spaces. (B) Distribution of expansin (exlx) homologs in a taxonomically representative set of bacterial species. Branches are colored according to taxonomic assignments. The tree was reconstructed using maximum likelihood and should be considered unrooted. Numbers at nodes are bootstrap pseudoreplicates, and the scale bar is the number of amino acid substitutions per site. (C) Distribution of glycoside hydrolase 5 (gh5) homologs in a taxonomically representative set of species. Branches are colored according to taxonomic assignments, using the same color assignments as Fig. 1b. The tree was reconstructed using maximum likelihood and should be considered unrooted. Numbers at nodes are bootstrap pseudoreplicates, and the scale bar is the number of amino acid substitutions per site.
The phylogenies of expansin and GH5 are consistent with horizontal gene transfer
In E. tracheiphila, the exlx and gh5 ORFs are flanked by mobile DNA elements (Fig. 2A). Because mobile DNA are common agents of horizontal gene transfer35,51,52, we reconstructed the phylogenies of both the exlx and gh5 genes and compared the phylogenies of these two genes to the bacterial species phylogeny7,35,51,52,53,54,55,56,57. We find that the phylogenies of both Et–exlx and Et–gh5 genes are in conflict with the species phylogeny, which is consistent with horizontal acquisition of these genes by E. tracheiphila (Fig. 2B,C). E. tracheiphila and Pantoea stewartii are the only species with microbial expansin homologs from the Erwinia and Pantoea genera, respectively (Fig. 2B). The E. tracheiphila and P. stewartii expansin homologs group with the other Enterobacterial plant pathogens (Pectobacterium spp. and Dickeya spp.), suggesting horizontal transfer has occurred between these species58.
The bacterial expansin phylogeny is consistent with additional horizontal gene transfer events, which is in agreement with previous studies6,7. There is a second group of γ-proteobacterial expansin homologs in Xanthomonadaceae, and these two distinct groups of γ-proteobacterial exlx homologs are separated by a group of Firmicutes exlx homologs7. The phylogenetic reconstruction is also consistent with an expansin acquisition by the β-proteobacterial plant pathogen Ralstonia solancearum from a Xanthomonadaceae donor (Fig. 2B)6,7.
The Et–gh5 phylogeny is also consistent with multiple horizontal gene transfer events (Fig. 2C). Overall, Et–gh5 homologs have a relatively sparse distribution in some Enterobacteriaceae, Firmicutes, Myxobacteria, Bacteroidetes and β-proteobacteria, and in some species of plant pathogenic nematodes57. In Enterobacteriaceae, the gh5 homologs separate into three distinct groups. One group is comprised of E. tracheiphila, Pantoea stewartii, Dickeya dianthicola, and Phaseolibacter sp. (recently reclassified to Enterobacteriaceae59). The gh5 homologs from the other plant-pathogenic Dickeya spp. comprise a second group of Enterobacterial gh5 homologs, and plant-pathogenic Pectobacterium and Brennaria spp. are a third group (Fig. 2C). These three groups of Enterobacterial gh5 homologs are dispersed among the gh5 homologs from Firmicutes, Bacteroidetes, Myxobacteria and other lineages of γ-proteobacteria, which is consistent with multiple HGT events between these bacterial lineages.
Co-occurrence patterns of expansin and glycoside hydrolase 5 genes in bacteria
In bacteria, the exlx gene can be found in species that (1) do not harbor a gh5 anywhere in their genomes, (2) harbor a gh5 ORF in the same genome but physically distant from exlx, or (3) encode a gh5 adjacent to an exlx in the same locus (Fig. 3). A gh5 homolog is present in the genomes of multiple plant-pathogenic Pectobacterium and Dickeya species that harbor an exlx homolog, but these ORFs do not co-occur in the same operon (Fig. 2B,C). Only three enterobacterial species (E. tracheiphila, Pantoea stewartii, and Dickeya dianthicola) harbor exlx and gh5 as two distinct ORFs that co-occur in the same operon. In these three species, the exlx and gh5 genes have distinct signal peptides and are separated by ~ 50 nucleotides. In P. stewartii, the exlx–gh5 locus is on a plasmid (pDSJ08), which may increase the probability of acting as a donor for horizontal gene transfer.
Co-occurrence of expansin genes with carbohydrate active domains. The gyrB species tree of selected bacteria with an expansin gene, and several species without expansins. The expansin and carbohydrate active domains are depicted as arrows. The rectangles within the arrows indicate whether that ORF has an expansin domain, a carbohydrate active domain, or both. Homologous carbohydrate active domains are color-coded. Both expansin genes are shown for Streptomyces scabiei, the only microbial species to harbor two expansin homologs with signal peptides for secretion in one genome. Accession numbers of the depicted protein sequences, and accession numbers of several expansin homologs that do not have predicted signal peptides for secretion and are not depicted in the figure, can be found in https://github.com/lshapiro31/gh5.expansin.phylogenetics. The domains are drawn to scale. The tree was reconstructed using maximum likelihood with 100 bootstrap pseudoreplicates and should be considered unrooted.
A gh5 is fused to an exlx as a single ORF in many plant pathogenic Xanthomonadaceae7,60. This gh5 domain in Xanthomonadaceae is non-homologous to the GH5 domain in Enterobacteriaceae, and the exlx and gh5 domain structure in E. tracheiphila, P. stewartii and D. dianthicola is in reverse orientation compared to the gh5–exlx domain order in Xanthomonadaceae. A distinct gh5 domain truncated to 289 amino acids is found in Clavibacter michiganensis (CelA), and this is the only known microbial expansin that is fused to both a GH5 and CBM2 domain in a single coding sequence25. Some isolates of C. michiganensis contain an additional exlx ORF that is not fused to either a GH5 or CBM2 domain6,16,21. The three non-homologous gh5 domains that are adjacent or fused to exlx genes in distinct bacterial lineages and with distinct domain architectures is consistent with at least three independent fusion events, and may be an example of functional convergence.
Expansin and GH5 genes both contribute to Erwinia tracheiphila virulence
To evaluate the individual and combined roles of expansin and GH5 for E. tracheiphila virulence, as well as the possible synergistic effects of both proteins, we generated a deletion mutant of the complete operon (strain Δexlx–gh5), as well as single-ORF mutants in only the expansin ORF (strain Δexlx) and only the gh5 ORF (strain Δgh5). Strains that complemented the three deletion mutations (Δexlx–gh5(cEXLX-GH5), Δexlx(cEXLX) and Δgh5(cEXLX-GH5), respectively) were also constructed (Supplemental Table S1). Variation in virulence was quantified via differences in days until (1) development of wilt symptoms localized to the inoculated leaf, (2) systemic spread of wilt symptoms to a second non-inoculated leaf, and (3) plant death.
Plants inoculated with Wt suffered high mortality (85%; 17 out of 20 plants), but significantly fewer plants inoculated with Δexlx–gh5 died (22%; 5 of 22) (Fig. 4a, Table 1). Plants inoculated with Δexlx–gh5 had delayed development of initial wilt symptoms in the inoculated leaf, and delayed appearance of systemic wilt symptoms in a non-inoculated leaf compared to plants inoculated with Wt (Fig. 4a, Tables 1 and 2). Further, wilt symptoms in plants inoculated with Δexlx–gh5 were more likely to be localized to the inoculated leaf (i.e., symptoms did not progress to systemic infection or plant death) compared to plants inoculated with Wt (Fig. 5). The complemented strain Δexlx–gh5(cEXLX-GH5) had restored ability to induce wilting symptoms at a second non-inoculated leaf, and partially restored the mortality rate (59%; 13 out of 22; Fig. 4a, Tables 1 and 2).
Contribution of the Erwinia tracheiphila exlx–gh5 locus to wilt symptom development and plant death. (a) In planta inoculation experiment comparing virulence of Wt, Δexlx–gh5, Δexlx–gh5(cEXLX-GH5), Δexlx and Δexlx(cEXLX) strains. (b) A second independent in planta inoculation experiment comparing virulence of Wt, Δgh5, and Δgh5(cEXLX-GH5) strains. In both (a) and (b), inoculated plants were monitored for first appearance of wilt symptoms in the inoculated leaf, first appearance of systemic wilt symptoms in a second non-inoculated leaf and plant death for 23 days post inoculation (DPI). Summary statistics are in Table 1 and statistical analyses are in Table 2.
Visual comparison of wilt symptoms in squash seedlings after inoculation with either the Δexlx–gh5 mutant or wild type Erwinia tracheiphila. (a) Non-inoculated squash seedling with no wilt symptoms (b) Squash seedling inoculated with wild type E. tracheiphila that has developed systemic wilt symptoms (c) Representative symptoms caused by inoculation with the Δexlx–gh5 mutant strain, where wilt often remains localized to the inoculated leaf without causing systemic wilt symptoms.
Individual deletions of the Et–exlx and Et–gh5 ORFs also caused a decrease in virulence and mortality compared to Wt (Fig. 4, Tables 1 and 2). Plants inoculated with either Δexlx or Δgh5 exhibited significant delays in the initial appearance of wilt symptoms in the inoculated leaf, delays in the appearance of systemic wilt symptoms in a second leaf and reduced mortality compared to Wt (Fig. 4, Tables 1 and 2). Genetic complementation of Δexlx (strain Δexlx(cEXLX)) caused partial recovery of virulence by restoring the ability to cause systemic wilt symptoms and plant death (Fig. 4a, Tables 1 and 2). Complementation of Δgh5 (strain Δexlx–gh5(EXLX–GH5)) restored the Wt ability to cause wilt symptoms in the inoculated leaf, systemic wilt symptoms in a second leaf, and plant death (Fig. 4b, Tables 1 and 2).
The Δexlx–gh5 mutant is impaired in systemic movement
The correlation between within-plant movement of E. tracheiphila to systemic wilt symptom development and plant death has been hypothesized, but not yet demonstrated32,61. It is assumed that systemic movement of bacteria through xylem—along with bacterial replication far from the initial inoculation point—is necessary to occlude xylem vessels to cause wilt symptoms and plant death (Fig. 1)32,61. Both the Δexlx and Δgh5 single mutants and the Δexlx–gh5 double mutant caused a delay in systemic symptom development (Fig. 4), suggesting both Et–exlx and Et–gh5 ORFs are needed for normal within-plant movement. Because both ORFs appear to be needed for Wt wilt symptom development, we compared the Δexlx–gh5 to the Wt strain to test how severity of wilt symptoms correlates with within-plant colonization.
To test whether Δexlx–gh5 has impaired within-host movement and colonization ability, squash seedlings were inoculated with either Wt or Δexlx–gh5. At 12 DPI, bacteria were quantified at two sites in each plant: the petiole of the inoculated leaf and the petiole of a second, non-inoculated leaf. At 12 DPI, all of the plants inoculated with the Wt strain were systemically wilting, while plants inoculated with Δexlx–gh5 were either asymptomatic or had wilt symptoms only in the inoculated leaf (Fig. 6a). At the inoculation site of all plants, the Wt and Δexlx–gh5 both reached similar cell counts (> 109 CFU/g for Wt, and 108–109 CFU/g for Δexlx–gh5) (Fig. 6a). However, in the petiole of a second, non-inoculated leaf Δexlx–gh5 only reached cell counts of 103 CFU/g, while the Wt reached 109 CFU/g (Fig. 6a). The Δexlx–gh5 strain does not have a growth deficiency in vitro compared to the Wt (Supplemental Figure S1), showing that the attenuation of wilt symptom development and decrease in plant death rates (Figs. 4 and 5) is due to impaired systemic movement of Δexlx–gh5 and not a difference in intrinsic growth rates. This experiment shows that attenuated virulence in Δexlx–gh5 mutant (Fig. 4) is related to a decreased ability in systemic xylem colonization62,63.
Colonization of Wt and Δexlx–gh5 strains in plant xylem. (a) Squash seedlings were inoculated with either Wt or Δexlx–gh5. At 12 days post inoculation (DPI), bacterial concentration was determined in the inoculation site and in a petiole of a second, non-inoculated leaf. (Sample sizes, n = 9 per treatment). (b) In trans complementation of Δexlx–gh5 and Wt. Plants were co-inoculated with a 1:1 mix of Wt & Δexlx–gh5. (Sample sizes, n = 11 per treatment). Y-axis is the log10 CFU/g fresh weight and is scaled to the lower limit of detection for the assay (log10 CFU/g fresh weight = 2.6). Bars show mean ± SE, and circles are individual biological replicates. **P < 0.005; ns non-significant.
The Et–EXLX–GH5 protein functions extracellularly
To test whether the protein products of the Et–exlx–gh5 locus functions extracellularly (as predicted by the presence of signal peptides for both Et–exlx and Et–gh5), plants were co-inoculated with a 1:1 mix of Wt and Δexlx–gh5. The aim of this experiment was to assess if the systemic colonization defect of Δexlx–gh5 could be rescued by the presence of the expansin and GH5 proteins produced by the Wt strain.
A total of 15 plants were co-inoculated with both Wt and Δexlx–gh5. CFU counts of both strains were quantified at 1 DPI at the inoculation site of four plants (which were then discarded) to ensure both strains were inoculated in equal concentrations, leaving 11 experimental plants (Supplemental Figure S2). At 12 DPI, two of the Wt:Δexlx–gh5 co-inoculated plants were asymptomatic, two had symptoms only in the inoculated leaf, and seven had systemic wilt symptoms. At 12 DPI, CFU counts of both the Wt and Δexlx–gh5 strains from all of the co-inoculated plants were determined in both the local inoculation site and a petiole of a second, non-inoculated leaf. The average cell count for Wt and Δexlx–gh5 in the petiole of a second, non-inoculated leaf both reached similar values of 108 CFU/g (Fig. 6b). This is a notably higher than Δexlx–gh5 reaches at the same 12-day time point when inoculated alone (103–104 CFU/g) (Supplemental Figure S3). Three independent experiments (shown in Figs. 4a, 6A and Supplemental Figure S3) resulted in the consistent trend of delayed symptom development in the deletion mutant compared to the Wt strain. The presence of a signal peptide for secretion in both proteins, and the ability of the Wt strain to rescue the systemic colonization defect of Δexlx–gh5 together suggests that EXLX and GH5 are secreted and function extracellularly.
The Δexlx and Δgh5 single deletion mutants were also co-inoculated in a 1:1 mix to test whether the EXLX and GH5 proteins function extracellularly together as a single complex, or as two independent proteins. The Δexlx deletion mutant is expected to still secrete an intact GH5 protein, and the Δgh5 deletion mutant is expected to still secrete an intact EXLX protein. If the EXLX and GH5 proteins function independently, the two strains would complement each other in trans. However, this reciprocal complementation was not achieved (Supplemental Figure S4; Supplemental Tables S2 and S3). This result suggests that both proteins need to be produced by the same cell, and that they could function as a single EXLX–GH5 protein complex that assembles before or during secretion. Further assays are needed to confirm that expansin and GH5 are secreted, and whether they assemble into—and function as—a single protein in planta.
Erwinia tracheiphila does not have cellulase or xylanase activity
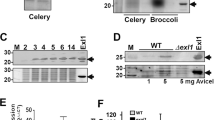
No expansin protein from any plant, bacteria, fungi, or other microbial eukaryote has detectable enzymatic activity4,10,16,64. However, glycoside hydrolases are enzymes that break the glycosidic bond between two or more carbohydrate subunits, and the predominant target of these enzymes is cellulose48. It is therefore possible that the gh5 ORF adjacent to or fused to bacterial expansin genes in some species may confer enzymatic activity. To test whether the gh5 ORF confers carbohydrate degrading ability to E. tracheiphila, the Wt strain (with the intact exlx–gh5 locus) was evaluated for enzymatic degradation of cellulose and xylan, the two main structural components of plant cell walls and the putative targets of active GH5 enzymes65. Neither E. tracheiphila culture supernatant nor colonies had detectable hydrolytic activity against cellulose or xylan (Supplemental Figure S5). These results indicate that the function of the GH5 protein is not the enzymatic degradation of plant structural carbohydrates.
Mutants in flagella and Type IV pili of Erwinia tracheiphila display wild type virulence
Type IV Pili and flagella are used by some bacterial plant pathogens during systemic movement through xylem66,67,68,69. To assess whether these cellular components may also contribute to E. tracheiphila xylem colonization, deletion mutants were generated for Type IV Pili (ΔT4P) and flagella (ΔfliC) (Supplemental Table S1). In squash inoculation experiments, there was no difference in the development of wilt symptoms or death rates in groups of plants inoculated with either ΔT4P, ΔfliC, or Wt (Fig. 7, Supplemental Tables S4 and S5). This indicates that neither Type IV Pili nor flagellar movement contribute to xylem colonization by E. tracheiphila, although it is still possible that these loci contribute in other, more subtle, ways to pathogenesis.
Comparison of virulence between wild type, flagellar deletion mutant (ΔfliC) and Type IV Pili deletion mutant (ΔT4P). Squash seedlings were inoculated with either Wt, ΔfliC or ΔT4P strain. Inoculated plants were monitored for first appearance of wilt symptoms in the inoculated leaf, first appearance of systemic wilt symptoms in a second non-inoculated leaf and plant death for 25 days post inoculation (DPI). Summary and statistical analyses are in Tables S4 and S5.
Discussion
Here, we find that the emerging plant pathogen E. tracheiphila has horizontally acquired an exlx–gh5 locus that functions as a virulence factor by conferring the ability to efficiently colonize xylem and cause high rates of plant death. The ability of a pathogen to move systemically through host vasculature—either plant xylem or animal cardiovascular systems—is a high-virulence phenotype, and is associated with development of more severe symptoms than localized infections70,71. The ability of a pathogen to reach a high titre and be distributed throughout the host's vasculature is also necessary for vector transmission by providing more opportunities for acquisition70,72. In E. tracheiphila, bacterial colonization of xylem blocks sap flow, inducing wilt symptoms and causing plant death. The development of systemic wilt symptoms by E. tracheiphila also induces a chemical volatile phenotype in symptomatic wild squash (Cucurbita pepo ssp. texana) that attracts significantly more foraging vectors to wilting leaves45,73, and a physical phenotype that facilitates insect vector feeding—and increased pathogen acquisition opportunities—from symptomatic foliage40,45. This increase in virulence conferred by the Et–exlx–gh5 locus induces more severe symptoms in infected plants that both attract obligate insect vectors to infected plants, and facilitates preferential feeding on wilting tissue once they arrive. Together, this suggests that the horizontal acquisition of the exlx–gh5 locus was a key step in the recent emergence of E. tracheiphila as a virulent wilt-inducing pathogen that is obligately insect vector transmitted32,35.
In ~ 10% of the bacterial species that harbor expansin genes, the expansin is fused to domains of carbohydrate active proteins. The formation of new genes via fusions of multiple modular domains is a key source of evolutionary innovation for organisms across the tree of life74,75. A notably high proportion of bacterial species where an expansin co-occurrs with a gh5 domain are pathogenic to plants, and can move through xylem7. This study suggests there may be further emergent properties of the expansin–GH5 protein complex that are uniquely adaptive for xylem-colonizing plant pathogenic bacteria. Healthy plants have effective physical barriers to allow the flow of xylem sap while excluding bacteria. Pit membranes between adjacent tracheids and perforation plates between xylem vessels are openings on a nanometric scale, while most bacteria are ~ 1 µm76. One hypothesis is that bacterial expansins may non-enzymatically ‘loosen’ the cellulose and pectin matrix at the perforation plates or at the pit membranes in order to increase their size enough to allow the passage of bacterial cells77,78,79,80. The ability of E. tracheiphila Wt strain to complement Δexlx–gh5 in trans supports the working hypothesis that the expansin–GH5 proteins function extracellularly by interacting with xylem structural carbohydrates that would normally prevent bacterial passage. This also suggests that, while the GH5 enzymatic activity has been lost due to truncation of the Et–gh5 gene, the remaining fragment may have been neofunctionalized and is providing an essential (though mechanistically undefined) role in virulence. One possibility is that the GH5 functional domain may physically (but non-enzymatically) interact with plant structural carbohydrates at perforation plates or pit membranes in a way that aids expansin function for loosening of cellulose microfibrils, or vice versa. In vitro assays for polysaccharide solubilizing activity (e.g. on isolated vascular bundles) are needed to support a non-enzymatic mechanism for expansin–GH5. It is also possible that microbial expansins and expansin–GH5 constructs may also affect virulence via swarming and attachment processes, as has been found for Pectobacterium spp.21,22. Further detailed work will be needed to understand the mechanism(s) of action for host interactions and virulence among the diverse microbial species with expansin genes6,7.
The phylogenies of the Et–exlx and Et–gh5 ORFs, and their genomic architecture as distinct genes in the same operon, may offer mechanistic insight into how bacterial expansins fuse to carbohydrate active domains. In many Firmicutes, Pectobacterium spp. and most Dickeya spp. plant pathogens, the exlx and gh5 ORFs are present in the same genome, but are not located directly adjacent to each other in the same operon. Only in E. tracheiphila, P. stewartii, and D. dadantii is the exlx homolog directly adjacent—but not fused to—the gh5 homolog. This suggests that during a horizontal gene transfer event between an Enterobacteriaceae donor and recipient, an expansin integrated by random chance adjacent to a GH5, and the two ORFs in this operon are now being horizontally transferred together. The assembled protein complex produced by the exlx–gh5 locus may provide a more efficient mode of action for movement through xylem, promoting the fitness of the host bacteria and providing opportunities for further horizontal transfer of this construct as a single virulence island. From a shared promoter and only ~ 50 nucleotide separation, a fusion of exlx and gh5 into a single ORF is possible from a small deletion mutation. We also note that all three of the bacterial plant pathogens with this construct are agricultural pathogens emerging into intensively cultivated, homogeneous crop plant populations. Erwinia tracheiphila has recently emerged into cucurbit agricultural populations32,35 and Pantoea stewartii infects sweet corn81. Both E. tracheiphila and P. stewartii only occur in temperate Eastern North America—one of the world’s most intensively cultivated regions—despite global distribution of susceptible host plants82. D. dianthicola causes a virulent wilt disease and is emerging into cultivated potato crops, and is also geographically restricted to Eastern North America and Europe83,84,85. The exlx–gh5 gene architecture in Enterobacteriaceae, the gh5–exlx gene architecture in Xanthomonadaceae and cbm2–gh5–exlx gene architecture in C. michaganensis (Actinobacteria) is consistent with multiple independent co-occurrences of bacterial expansins with evolutionarily distinct gh5 domains, and may be an example of functional convergence.
There is constant risk that agro-ecosystems will be invaded by virulent microorganisms, and the increasing homogeneity in crop plant populations may select for novel pathogens with non-canonical virulence mechanisms. The recent realization that microbial expansin genes are present in phylogenetically diverse xylem-colonizing bacterial and fungal species—including almost all of the most economically damaging bacterial and fungal wilt pathogens – and the function of expansins to increase E. tracheiphila virulence suggest these genes may be an under-appreciated virulence factor. The emergence of virulent plant pathogens that systemically colonize xylem is especially alarming because plants do not have inherent genetic resistance against xylem-dwelling vascular pathogens86. The increase in E. tracheiphila virulence conferred by expansin, the presence of this gene in many other bacterial and fungal wilt-inducing plant pathogen species, and the amenability of microbial expansins to horizontal gene transfer suggest this gene may be an under-appreciated virulence factor in taxonomically diverse agricultural pathogens.
Methods
Bacterial strains, culture media and plant cultivation
All bacterial strains used in this study are listed in Supplemental Table S1. Throughout this work, we used a rifampicin resistant variant of E. tracheiphila BHKYR (Wt)32. Escherichia coli TOP10 and PIR1 strains for used for routine cloning, and the E. coli strain S17-1λ was used as the donor for conjugation. E. tracheiphila was grown in KB liquid media or agar at room temperature (RT), and E. coli strains in LB media or agar at 37 °C, unless otherwise specified. Antibiotics were added to liquid or agar media at the following concentrations: rifampicin, 50 μg/ml; ampicillin or carbenicillin, 100 μg/ml; chloramphenicol 5 μg/ml; kanamycin 50 μg/ml. All in planta experiments were conducted with organic ‘Dixie’ variety crookneck squash bought from Johnny’s Seeds (https://www.johnnyseeds.com/). Plants were grown in potting mix in standard six cell seedling trays in a greenhouse environment set to 25 °C, 70% humidity, and a 12 h day:12 h night light cycle.
Visualization of fluorescent Erwinia tracheiphila in wilting squash seedlings
Erwinia tracheiphila BuffGH was transformed with a plasmid carrying the mCherry gene for visualization of fluorescent cells in symptomatic squash seedlings. Competent E. tracheiphila were prepared as described previously32,87. Briefly, cells were prepared by growing E. tracheiphila to an OD600 of 0.02. Cells were then washed with decreasing volumes, once with chilled sterile Milli-Q water and twice with 10% glycerol, and resuspended in 1/100 volume of chilled 10% glycerol. Plasmid pMP7605 was used for electroporation in a 0.2-cm cuvette, at 2.5 kV for 5.2–5.8 ms. Cells were incubated at room temperature without shaking for 1 h in 3 ml KB liquid and then plated in KB agar with ampicillin. Colonies of fluorescent E. tracheiphila Et (pMP605) were obtained after 5 days at room temperature. Ten µl of a Et (pMP7605) stationary culture were used for inoculating 2 week-old squash seedlings (at the two leaf stage), and confocal microscopic observations were performed once symptoms appear using fresh longitudinal cuts of the inoculated petiole.
Phylogenetic reconstruction of the exlx and gh5 genes and comparison of domain architecture
The amino acid sequences of the expansin (WP_046372116.1) and gh5 (WP_016193008.1) ORFs in the E. tracheiphila reference strain29 were used as queries to identify expansin and gh5 homologs using the BLASTP web interface88. A taxonomically representative sample of the top BLASTP hits for each gene were aligned using MAFFT v. 7.305b and default parameters89. The expansin alignment was trimmed visually such that the two canonical expansin domains were conserved in the alignment, and the gh5 alignment was trimmed with trimAI using the—automated 1 option90. For both alignments ProtTest v. 3.4.2 was used to identify the best-fitting substitution model by BIC score, which was WAG + G for the expansin gene alignment and LG + I + G for the GH5 alignment91. The GyrB species tree was constructed by using the E. tracheiphila GyrB sequence (KKF36621.1) as a query on the BLASTP web interface88. The GyrB amino acid sequences from species known to have an expansin gene or an expansin fusion to a domain from a carbohydrate active protein were downloaded and added to a multi-FASTA (https://github.com/lshapiro31/gh5.expansin.phylogenetics). The GyrB sequences were aligned with MAFFT v. 7.305b and default parameters89.
Phylogenetic trees were reconstructed using maximum likelihood with RAxML92 and the appropriate evolutionary model on the CIPRES server93. The expansin tree was reconstructed with 1000 bootstrap pseudoreplicates, and the GH5 and GyrB trees were reconstructed with 100 bootstrap pseudoreplicates. The bootstrapped pseudosamples were summarized with SumTrees v. 4.4.094. The resulting phylogeny was visualized in the R statistical environment using the ggtree library95,96. Amino acid sequences were analyzed with NCBI CBD tool to identify domain architecture47, and signal peptides were predicted with SignalP50. The genomic context of the Et–exlx–gh5 locus was visualized with genoPlotR97. Alignment files and phylogenetic scripts are available at https://github.com/lshapiro31/gh5.expansin.phylogenetics.
Construction of deletion mutants
Mutants with a deletion in the exlx–gh5 operon, exlx gene, gh5 gene, the Type IV pili operon and the fliC gene were generated from an E. tracheiphila isolate BHKYR parental strain by double homologous recombination, using the suicide plasmid pDS13298. This plasmid was improved by inserting in the XbaI site, a constitutive mCherry gene amplified from plasmid pMP760599 using primers JR72 and JR73 (Supplemental Table S6). The resulting plasmid (pJR74, Supplemental Table S1) allows rapid screening of conjugants colonies and colonies that have lost the plasmid. For the target genomic region to create each mutant, regions upstream of the target locus were amplified with primers pair F5 and R5, and downstream regions were amplified with primer pair F3 and R3 (See Supplemental Table S6 for specific primer names and sequences). An ampicillin resistance bla gene, coding for Beta-lactamase was amplified from pDK4687 using primers LS23 and LS24. Constructions consisting on each upstream and downstream region flanking the bla gene were used for exlx–gh5, gh5, fliC and Type 4 Pili mutants, while a construction with no flanked antibiotic cassette was prepared for the exlx deletion. All constructions were assembled using the Gibson Assembly Master Mix (New England Biolabs, Ipswich, MA), and then each was reamplified with nested primers containing SacI restriction site (primers SacI-F and SacI-R, Supplemental Table S6). Constructions for exlx–gh5, exlx, gh5, fliC and Type 4 Pili deletion were inserted into the SacI site of plasmid pJR74, obtaining plasmids pJR150, pJR323, pJR324, pJR74a and pJR149, respectively (Supplemental Table S1). These plasmids were transformed into Ec-PIR1 for preservation, and then into Ec-S17 for conjugation using E. tracheiphila as recipient. MCherry fluorescent E. tracheiphila conjugants were obtained in KB agar with rifampicin and chloramphenicol, then a few colonies were picked and grown in 3 ml liquid KB with chloramphenicol to stationary phase, and 100 µl were spread in KB agar with 5% sucrose and carbenicillin (or no antibiotic in the case of exlx deletion). Non-fluorescent, chloramphenicol-sensitive colonies were picked, PCR checked for the correct deletion and cryogenically stored in 15% glycerol at − 80 °C.
Genetic complementation
A new integration plasmid, specific for a neutral region in the chromosome of Et-BHKYR, was constructed from plasmid pJR74. To create this plasmid, two ≈ 0.8 Kb adjacent DNA fragments were PCR amplified form Et-BHKYR genomic DNA using primer pairs JR143–JR144, and JR145–JR146 (Supplemental Table S6). These fragments were ligated using the Gibson Assembly Master Mix (New England Biolabs, Ipswich MA), and reamplified using primers JR143 and JR146. The ≈ 1.6 Kb product was inserted in the SacI site of pJR74, generating plasmid pJR315 (Supplemental Table S1). Single cutting XhoI and BglII sites were engineered in the middle of the amplified neutral regions, which can be used for the insertion of complementation genes. For the complementation of the exlx–gh5 locus or the individual exlx gene, the genomic region together with its natural promoter were amplified from E. tracheiphila genomic DNA using primers JR152 and JR154, or JR152 and JR153 (Supplemental Table S6) respectively, and DNA products were inserted into the XhoI site of pJR315. Promoter regions for expression of the exlx–gh5 operon have not been characterized, but expression of individual ORFs in an operon is often directed from an upstream shared promoter. It is therefore reasonable to assume expression of the gh5 ORF is directed from a shared promoter region upstream of exlx100. For this reason, the single Δgh5 mutant was complemented with the full operon (exlx–eng) to include the promoter region of exlx. Each resulting plasmid was transformed into Ec-S17-1λ cells, which were then used as donors for conjugation with mutant strains Δexlx–gh5, Δexlx or Δgh5 as recipients. Conjugant colonies were used for negative selection with sucrose, as described above, and colonies carrying the exlx–gh5 operon or the exlx gene integrated in the expected site were confirmed by PCR.
In planta inoculation experiments
In planta virulence assays were performed by inoculating squash seedlings with E. tracheiphila Wt and derived strains, and monitoring wilt symptom development for approximately three weeks (between 21 and 25 days per experiment). To create inoculum, one bacterial colony of each strain was picked and added to 3 ml of liquid KB media with the appropriate antibiotic, and grown with shaking for 24 h. Then, 2–3-week-old squash seedlings (2–3 true leaves) were inoculated by manually inducing a wound where xylem was exposed in the petiole at the base of the first true leaf and adding 10 μl of culture containing ≈ 1 × 107 bacterial cells directly into the wound. In each experiment, an equal number of plants were inoculated in each group; however, plants that were inadvertently damaged during watering or symptom assessment were removed from experiments. Plants were kept at 25 °C, 70% humidity, and a 12 h day:12 h night light cycle 25 °C, and monitored daily for appearance of first symptoms in the inoculated leaf, appearance of wilt symptoms in a second non-inoculated leaf, and plant death.
In planta colony forming unit (CFU) counts
Bacterial colony forming units (CFU) counts were determined from plants inoculated with Et-BHKY Wt and derived strains. Bacterial cells can be obtained directly from petioles of infected plants (Fig. 1). Two cm samples of the petiole from the inoculated leaf, or from a second non-inoculated leaf were cut from the plants and washed briefly with 70% ethanol (EtOH). Excess EtOH was removed with a paper towel and petioles were surface-sterilized over a gas flame for 1 s and placed in a sterile plastic petri dish. From each petiole sample, 10–15 disks small disks (< 1 mm) were manually cut with a sterile blade and collected in a 2 ml microtube. The weight of each 2 ml tube with all leaf disks was recorded to be used for normalizing CFU per gram of plant tissue in each sample, and 500 μl of chilled PBS was added to each tube. After an incubation of 40 min on ice (vortexing every 10 min) 200 μl of PBS from each tube was pipetted into a new microtube and used for serial dilutions and plating onto KB agar with rifampicin. Bacterial CFU per gram of fresh tissue was calculated.
For obtaining CFUs of individual strains in plants co-inoculated with E. tracheiphila Wt and Δexlx–gh5 mutant, serial dilutions were plated in both KB with rifampicin and KB with rifampicin and carbenicillin agar plates. CFU of carbenicillin resistant colonies represent the Δexlx–gh5 strain. CFUs of Wt was determined as the count of total CFUs—carbenicillin resistant CFUs.
Statistics
Statistical analyses were performed using Prism version 7.0 (GraphPad Software, La Jolla California USA, www.graphpad.com). Curves following initial symptoms in first leaf, systemic wilt in second leaf, and plant death from each experiment were compared using the built-in Log-rank (Mantel–Cox) test for survival analysis. In the cases where significant differences were found (p < 0.05), pairwise comparisons were tested using the same analysis101. For comparisons of bacterial CFU in planta, CFU data and its log10-transformed values were checked for Gaussian distribution using the Shapiro–Wilk normality test. Since neither CFU data distribution nor the transformed Log10 values distribution passed the normality test, the Kruskal–Wallis non-parametric was used test to analyze if the medians vary significantly among experimental groups (p < 0.05). In the cases where differences were found, Dunn's multiple comparisons test was used to test for pairwise differences between groups.
Testing for cellulase and xylanase activity
Cellulase activity from cell-free supernatants of Wt and Δexlx–gh5 cultures were tested for extracellular enzymatic activity against cellulose. The Wt and Δexlx–gh5 strains were grown in 10 ml of liquid KB media for 48 H. Cultures were centrifuged at 7000 rpm for 10 min, and each supernatant was filter-sterilized. Supernatants and 1 mg/ml cellulase (Sigma), were spotted in 1% agar, 1% Carboxy Methyl Cellulose (CMC) plates. Plates were then incubated at 30 °C for 48 h, and flooded with Gram’s Iodine. Halos were imaged after 24 h, at RT.
A colony of E. tracheiphila grown on KB agar plates was used to test for extracellular xylanase activity. A xylanase producing strain of Streptomyces lividens was used as a positive control. Bacterial culture from each species was spotted on the surface of a KB agar plate, and grown at RT for 4 days. An overlay of 1% agar and 1% xylan was spread on top of the grown colonies, and plates were incubated at 30 °C for 48 h. Plates were flooded with 1% congo red and incubated for 10 min before discarding the Congo red solution. Plates were then flooded with 1 N NaOH, and incubated for 10 min. NaOH was discarded and plates were imaged after 24 h at room temperature.
References
Shi, S. et al. Effects of selected root exudate components on soil bacterial communities. FEMS Microbiol. Ecol. 77(3), 600–610 (2011).
Lindow, S. E. & Brandl, M. T. Microbiology of the Phyllosphere. Appl. Environ. Microbiol. 69(4), 1875–1883 (2003).
Miwa, H. & Okazaki, S. How effectors promote beneficial interactions. Curr. Opin. Plant Biol. 38, 148–154 (2017).
Cosgrove, D. J. Microbial expansins. Annu. Rev. Microbiol. 71(1), 479–497 (2017).
Georgelis, N., Nikolaidis, N. & Cosgrove, D. J. Biochemical analysis of expansin-like proteins from microbes. Carbohyd. Polym. 100, 17–23 (2014).
Nikolaidis, N., Doran, N. & Cosgrove, D. J. Plant expansins in bacteria and fungi: Evolution by horizontal gene transfer and independent domain fusion. Mol. Biol. Evol. 31(2), 376–386 (2013).
Chase, W. R. et al. Global cellulose biomass, horizontal gene transfers and domain fusions drive microbial expansin evolution. New Phytol. 226(3), 921–938 (2020).
Cosgrove, D. J. Loosening of plant cell walls by expansins. Nature 407(6802), 321–326 (2000).
Sampedro, J. & Cosgrove, D. J. The expansin superfamily. Genome Biol. 6(12), 242 (2005).
Cosgrove, D. J. Plant expansins: Diversity and interactions with plant cell walls. Curr. Opin. Plant Biol. 25, 162–172 (2015).
Chen, F. & Bradford, K. J. Expression of an expansin is associated with endosperm weakening during tomato seed germination. Plant Physiol. 124(3), 1265–1274 (2000).
Brummell, D. A. et al. Modification of expansin protein abundance in tomato fruit alters softening and cell wall polymer metabolism during ripening. Plant Cell 11(11), 2203–2216 (1999).
Cho, H.-T. & Cosgrove, D. J. Altered expression of expansin modulates leaf growth and pedicel abscission in Arabidopsis thaliana. Proc. Natl. Acad. Sci. 97(17), 9783–9788 (2000).
Li, Y. et al. Plant expansins are a complex multigene family with an ancient evolutionary origin. Plant Physiol. 128(3), 854–864 (2002).
Cosgrove, D. J. et al. The growing world of expansins. Plant Cell Physiol. 43(12), 1436–1444 (2002).
Georgelis, N., Nikolaidis, N. & Cosgrove, D. J. Bacterial expansins and related proteins from the world of microbes. Appl. Microbiol. Biotechnol. 99(9), 3807–3823 (2015).
Kerff, F. et al. Crystal structure and activity of Bacillus subtilis YoaJ (EXLX1), a bacterial expansin that promotes root colonization. Proc. Natl. Acad. Sci. 105(44), 16876–16881 (2008).
Brotman, Y. et al. Role of swollenin, an expansin-like protein from Trichoderma, in plant root colonization. Plant Physiol. 147(2), 779–789 (2008).
Saloheimo, M. et al. Swollenin, a Trichoderma reesei protein with sequence similarity to the plant expansins, exhibits disruption activity on cellulosic materials. Eur. J. Biochem. 269(17), 4202–4211 (2002).
Kende, H. et al. Nomenclature for members of the expansin superfamily of genes and proteins. Plant Mol. Biol. 55(3), 311–314 (2004).
Tancos, M. A. et al. Plant-like bacterial expansins play contrasting roles in two tomato vascular pathogens. Mol. Plant Pathol. 19(5), 1210–1221 (2018).
Narváez-Barragán, D. A. et al. Expansin-like Exl1 from Pectobacterium is a virulence factor required for host infection, and induces a defence plant response involving ROS, and jasmonate, ethylene and salicylic acid signalling pathways in Arabidopsis thaliana. Sci. Rep. 10(1), 1–14 (2020).
Chalupowicz, L. et al. Differential contribution of Clavibacter michiganensis ssp. michiganensis virulence factors to systemic and local infection in tomato. Mol. Plant Pathol. 18(3), 336–346 (2017).
Laine, M. J. et al. The cellulase encoded by the native plasmid of Clavibacter michiganensis ssp. sepedonicus plays a role in virulence and contains an expansin-like domain. Physiol. Mol. Plant Pathol. 57(5), 221–233 (2000).
Hwang, I. S. et al. Functional characterization of two cellulase genes in the Gram-positive pathogenic bacterium Clavibacter michiganensis for wilting in tomato. Mol. Plant Microbe Interact. 32(4), 491–501 (2019).
Hwang, I. S. et al. Multiple plasmid-borne virulence genes of Clavibacter michiganensis ssp. capsici critical for disease development in pepper. New Phytol. 217(3), 1177–1189 (2018).
Olarte-Lozano, M. et al. PcExl1 a novel acid expansin-like protein from the plant pathogen Pectobacterium carotovorum, binds cell walls differently to BsEXLX1. PLoS ONE 9(4), e95638 (2014).
Jahr, H. et al. The endo-β-1,4-glucanase CelA of Clavibacter michiganensis subsp. michiganensis is a pathogenicity determinant required for induction of bacterial wilt of tomato. Mol. Plant-Microbe Interact. 13(7), 703–714 (2000).
Shapiro, L. R. et al. Draft genome sequence of Erwinia tracheiphila, an economically important bacterial pathogen of cucurbits. genomeA 3(3), e00482-15 (2015).
Shapiro, L. R. et al. Draft genome sequence of an Erwinia tracheiphila isolate from an infected muskmelon (Cucumis melo). Microbiol. Res. Announcements 7(17), e01058-e1118 (2018).
Shapiro, L. A to ZYMV Guide to Erwinia tracheiphila Infection: An Ecological and Molecular Study Infection: An Ecological and Molecular Study (The Pennsylvania State University, State College, 2012).
Shapiro, L. R. et al. An introduced crop plant is driving diversification of the virulent bacterial pathogen Erwinia tracheiphila. mBio 9(5), e01307-18 (2018).
Shapiro, L.R. and K.E. Mauck, Chemically-mediated interactions among cucurbits, insects and microbes. in Chemical Ecology of Insects 55–90 (ed. J. Tabata). (CRC Press, 2018).
Smith, E. F. An Introduction to Bacterial Diseases of Plants (W.B. Saunders Company, Philadelphia, 1920).
Shapiro, L. R. et al. Horizontal gene acquisitions, mobile element proliferation, and genome decay in the host-restricted plant pathogen Erwinia tracheiphila. Genome Biol. Evol. 8(3), 649–664 (2016).
Moran, N. A. & Plague, G. R. Genomic changes following host restriction in bacteria. Curr. Opin. Genet. Dev. 14(6), 627–633 (2004).
Rand, F. V. & Enlows, E. M. A. Transmission and control of bacterial wilt of cucurbits. J. Agric. Res. 6(4), 7–434 (1916).
Rand, F. V. & Cash, L. C. Some insect relations of Bacillus tracheiphilus Erw. Sm. Phytopathology 10, 133–140 (1920).
de Mackiewicz, D. et al. Herbaceous weeds are not ecologically important reservoirs of Erwinia tracheiphila. Plant Dis. 82(5), 521–529 (1998).
Shapiro, L. et al. Dynamics of short-and long-term association between a bacterial plant pathogen and its arthropod vector. Sci. Rep. 4, 4155 (2014).
Garcia-Salazar, C. et al. ELISA versus immunolocalization to determine the association of Erwinia tracheiphila in Acalymma vittatum (Coleoptera: Chrysomelidae). Environ. Entomol. 29, 542–550 (2000).
Fleischer, S. J. et al. Serological estimates of the seasonal dynamics of Erwinia tracheiphila in Acalymma vittata (Coleoptera: Chrysomelidae). Environ. Entomol. 28, 470–476 (1999).
Sasu, M. et al. Floral transmission of Erwinia tracheiphila by cucumber beetles in a wild Cucurbita pepo. Environ. Entomol. 39(1), 140–148 (2010).
Shapiro, L. R. et al. Disease interactions in a shared host plant: Effects of pre-existing viral infection on cucurbit plant defense responses and resistance to bacterial wilt disease. PLoS ONE 8(10), e77393 (2013).
Shapiro, L. et al. Pathogen effects on vegetative and floral odours mediate vector attraction and host exposure in a complex pathosystem. Ecol. Lett. 15(12), 1430–1438 (2012).
Sasu, M. A. et al. Indirect costs of a nontarget pathogen mitigate the direct benefits of a virus-resistant transgene in wild Cucurbita. Proc. Natl. Acad. Sci. 106(45), 19067–19071 (2009).
Marchler-Bauer, A. et al. CDD/SPARCLE: Functional classification of proteins via subfamily domain architectures. Nucleic Acids Res. 45(D1), D200–D203 (2016).
Lombard, V. et al. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 42(D1), D490–D495 (2013).
Aziz, R. et al. The RAST server: Rapid annotations using subsystems technology. BMC Genom. 9(1), 75 (2008).
Armenteros, J. J. A. et al. SignalP 5.0 improves signal peptide predictions using deep neural networks. Nat. Biotechnol. 37, 420–423 (2019).
Thomas, C. M. & Nielsen, K. M. Mechanisms of, and barriers to, horizontal gene transfer between bacteria. Nat. Rev. Microbiol. 3(9), 711 (2005).
Zhaxybayeva, O. & Doolittle, W. F. Lateral gene transfer. Curr. Biol. 21(7), R242–R246 (2011).
Ochman, H., Lawrence, J. G. & Groisman, E. A. Lateral gene transfer and the nature of bacterial innovation. Nature 405(6784), 299–304 (2000).
Hug, L. A. et al. A new view of the tree of life. Nat. Microbiol. 1, 16048 (2016).
Polz, M. F., Alm, E. J. & Hanage, W. P. Horizontal gene transfer and the evolution of bacterial and archaeal population structure. Trends Genet. 29(3), 170–175 (2013).
Zhaxybayeva, O. et al. Phylogenetic analyses of cyanobacterial genomes: Quantification of horizontal gene transfer events. Genome Res. 16(9), 1099–1108 (2006).
Danchin, E. G. et al. Multiple lateral gene transfers and duplications have promoted plant parasitism ability in nematodes. Proc. Natl. Acad. Sci. 107(41), 17651–17656 (2010).
Smillie, C. S. et al. Ecology drives a global network of gene exchange connecting the human microbiome. Nature 480(7376), 241–244 (2011).
Halpern, M. et al. Transfer of Pseudomonas flectens Johnson 1956 to Phaseolibacter gen. nov., in the family Enterobacteriaceae, as Phaseolibacter flectens gen. nov., comb. nov. Int. J. Syst. Evolut. Microbiol. 63(1), 268–273 (2013).
Mensi, I. et al. Breaking dogmas: The plant vascular pathogen Xanthomonas albilineans is able to invade non-vascular tissues despite its reduced genome. R. Soc. Open Biol. 4(2), 130116 (2014).
Vrisman, C. M. et al. Differential colonization dynamics of cucurbit hosts by Erwinia tracheiphila. Phytopathology 106(7), 684–692 (2016).
Saile, E. et al. Role of extracellular polysaccharide and endoglucanase in root invasion and colonization of tomato plants by Ralstonia solanacearum. Phytopathology 87(12), 1264–1271 (1997).
Czajkowski, R. et al. Systemic colonization of potato plants by a soilborne, green fluorescent protein-tagged strain of Dickeya sp. biovar 3. Phytopathology. 100(2), 134–142 (2010).
Cosgrove, D. J. Catalysts of plant cell wall loosening. F1000Research (2016). https://doi.org/10.12688/f1000research.7180.1.
Malinovsky, F. G., Fangel, J. U. & Willats, W. G. The role of the cell wall in plant immunity. Front. Plant Sci. 5, 178 (2014).
Bahar, O., Goffer, T. & Burdman, S. Type IV pili are required for virulence, twitching motility, and biofilm formation of Acidovorax avenae subsp. citrulli. Mol. Plant Microbe Interact. 22(8), 909–920 (2009).
Burdman, S. et al. Involvement of type IV pili in pathogenicity of plant pathogenic bacteria. Genes 2(4), 706–735 (2011).
Cursino, L. et al. Identification of an operon, Pil–Chp, that controls twitching motility and virulence in Xylella fastidiosa. Mol. Plant Microbe Interact. 24(10), 1198–1206 (2011).
Bahar, O., Levi, N. & Burdman, S. The cucurbit pathogenic bacterium Acidovorax citrulli requires a polar flagellum for full virulence before and after host-tissue penetration. Mol. Plant Microbe Interact. 24(9), 1040–1050 (2011).
Ewald, P. W. Evolution of Infectious Disease (Oxford University Press, Oxford, 1993).
Mackinnon, M. & Read, A. F. Virulence in malaria: An evolutionary viewpoint. Philos. Trans. R. Soc. B Biol. Sci. 359(1446), 965–986 (2004).
Hinnebusch, B. J. et al. Role of the Yersinia pestis hemin storage (hms) locus in the transmission of plague by fleas. Science 273(5273), 367 (1996).
Yao, C., Zehnder, G., Bauske, E. & Kloepper, J. Relationship between cucumber beetle (Coleoptera: Chrysomelidae) density and incidence of bacterial wilt of cucurbits. J. Econ. Entomol. 89(2), 510–514 (1996).
Pasek, S., Risler, J.-L. & Brézellec, P. Gene fusion/fission is a major contributor to evolution of multi-domain bacterial proteins. Bioinformatics 22(12), 1418–1423 (2006).
Yang, S. & Bourne, P. E. The evolutionary history of protein domains viewed by species phylogeny. PLoS ONE 4(12), e8378 (2009).
Boutilier, M. S. et al. Water filtration using plant xylem. PLoS ONE 9(2), e89934 (2014).
Pérez-Donoso, A. G. et al. Cell wall-degrading enzymes enlarge the pore size of intervessel pit membranes in healthy and Xylella fastidiosa-infected grapevines. Plant Physiol. 152(3), 1748–1759 (2010).
Yadeta, K. & B. Thomma. The xylem as battleground for plant hosts and vascular wilt pathogens. Front. Plant Sci. 4, 97 (2013). https://doi.org/10.3389/fpls.2013.00097.
Secchi, F., Pagliarani, C. & Zwieniecki, M. A. The functional role of xylem parenchyma cells and aquaporins during recovery from severe water stress. Plant Cell Environ. 40(6), 858–871 (2017).
Christman, M. A. & Sperry, J. S. Single-vessel flow measurements indicate scalariform perforation plates confer higher flow resistance than previously estimated. Plant Cell Environ. 33(3), 431–443 (2010).
Roper, M. C. Pantoea stewartii subsp. stewartii: Lessons learned from a xylem-dwelling pathogen of sweet corn. Mol. Plant Pathol. 12(7), 628–637 (2011).
Foley, J. A. et al. Global consequences of land use. Science 309(5734), 570–574 (2005).
Nasaruddin, A. S. et al. First report of Dickeya dianthicola causing blackleg on potato in Texas. Plant Dis. 103(8), 2121 (2019).
Cai, W. et al. Draft genome sequences of two Dickeya dianthicola isolates from potato. Genome Announcements 6(11), e00115-e118 (2018).
Ma, X. et al. Pectobacterium and Dickeya responsible for potato blackleg disease in New York State in 2016. Plant Dis. 102(9), 1834–1840 (2018).
Bae, C. et al. Infection processes of xylem-colonizing pathogenic bacteria: Possible explanations for the scarcity of qualitative disease resistance genes against them in crops. Theor. Appl. Genet. 128(7), 1219–1229 (2015).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. 97(12), 6640–6645 (2000).
Altschul, S. F. et al. Basic local alignment search tool. J. Mol. Biol. 215(3), 403–410 (1990).
Katoh, K. et al. MAFFT: A novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30(14), 3059–3066 (2002).
Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. trimAl: A tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25(15), 1972–1973 (2009).
Abascal, F., Zardoya, R. & Posada, D. ProtTest: Selection of best-fit models of protein evolution. Bioinformatics 21(9), 2104–2105 (2005).
Stamatakis, A. RAxML-VI-HPC: Maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22(21), 2688–2690 (2006).
Miller, M.A., W. Pfeiffer, & T. Schwartz. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. in Proceedings of the Gateway Computing Environments Workshop (CGE). (New Orleans, LA, 2010).
Sukumaran, J. & Holder, M. T. DendroPy: A Python library for phylogenetic computing. Bioinformatics 26, 1569–1571 (2010).
R Core Team. R: A language and environment for statistical computing (R Foundation for Statistical Computing, Vienna, Austria, 2015). https://www.r-project.org/.
Yu, G. et al. ggtree: An R package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol. Evol. 8(1), 28–36 (2017).
Guy, L., Kultima, J. R. & Andersson, S. G. genoPlotR: Comparative gene and genome visualization in R. Bioinformatics 26(18), 2334–2335 (2010).
Philippe, N. et al. Improvement of pCVD442, a suicide plasmid for gene allele exchange in bacteria. Plasmid 51(3), 246–255 (2004).
Lagendijk, E. L. et al. Genetic tools for tagging Gram-negative bacteria with mCherry for visualization in vitro and in natural habitats, biofilm and pathogenicity studies. FEMS Microbiol. Lett. 305(1), 81–90 (2010).
Franklin, N. C. Genetic fusions for operon analysis. Annu. Rev. Genet. 12(1), 193–221 (1978).
Machin, D., Cheung, Y. B. & Parmar, M. Survival Analysis: A Practical Approach (Wiley, New York, 2006).
Acknowledgements
This work was supported by Fundación Mexico en Harvard, and Conacyt Grant 237414 to JR, NSF postdoctoral fellowship DBI-1202736 to LRS and NIH Grant GM58213 to RK. We thank all members of the Kolter lab, Einat Segev, William R. Chase and Olga Zhaxybayeva for valuable feedback and discussion. We thank the staff at the Harvard Arnold Arboretum for assistance in plant cultivation, use of growth facilities and use of the confocal microscope. Dominique Schneider kindly donated plasmid pDS132.
Author information
Authors and Affiliations
Contributions
J.R. and L.R.S. conceived the study. J.R., L.R.S. and R.K. designed experiments and interpreted experimental data. J.R. designed and conducted molecular protocols and lab experiments. L.R.S. conducted computational analyses and performed experiments. J.R. and L.R.S. wrote the first draft of the manuscript, and J.R., L.R.S. and R.K. added critical revisions.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rocha, J., Shapiro, L.R. & Kolter, R. A horizontally acquired expansin gene increases virulence of the emerging plant pathogen Erwinia tracheiphila. Sci Rep 10, 21743 (2020). https://doi.org/10.1038/s41598-020-78157-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-78157-w
- Springer Nature Limited
This article is cited by
-
Investigation of cold-active and mesophilic cellulases: opportunities awaited
Biomass Conversion and Biorefinery (2023)