Abstract
The chloroplast genome is an integral part of plant genomes in a species along with nuclear and mitochondrial genomes, contributing to adaptation, diversification, and evolution of plant lineages. In the family Caricaceae, only the Carica papaya chloroplast genome and its nuclear and mitochondrial genomes were sequenced, and no chloroplast genome-wide comparison across genera was conducted. Here, we sequenced and assembled the chloroplast genome of Vasconcellea pubescens A.DC. using Oxford Nanopore Technology. The size of the genome is 158,712 bp, smaller than 160,100 bp of the C. papaya chloroplast genome. And two structural haplotypes, LSC_IRa_SSCrc_IRb and LSC_IRa_SSC_IRb, were identified in both V. pubescens and C. papaya chloroplast genomes. The insertion-deletion mutations may play an important role in Ycf1 gene evolution in family Caricaceae. Ycf2 is the only one gene positively selected in the V. pubescens chloroplast genome. In the C. papaya chloroplast genome, there are 46 RNA editing loci with an average RNA editing efficiency of 63%. These findings will improve our understanding of the genomes of these two crops in the family Caricaceae and will contribute to crop improvement.
Similar content being viewed by others
Introduction
The family Caricaceae includes six genera and thirty-five species1,2,3. Among them, Vasconcellea pubescens A.DC, also named highland or mountain papaya, is a tropical crop native to the Ecuadorian Andes, distributed at high altitudes (above 1500 m). It was introduced to Chile about 60 years ago, and gradually became an important cash crop in central Chile4,5. V. pubescens has many economic values, and is commonly used to produce canned fruit, juice, jam and sweets5.
Chloroplasts are essential organelles in plants, which evolved from a cyanobacterium via endosymbiosis, specialized in photosynthesis. They are active metabolic centers, contributing to fatty acid and amino acid synthesis pathways6,7. The size of chloroplast genomes is small, ranging from 120 to 220 kb8. It consists of four regions, two inverted repeats (IRs), a large single-copy region (LSC) and a small single-copy region (SSC). LSC and SSC are separated by the IR regions. The structure of chloroplast genomes can be circular or linear, and the four regions can arrange differently7,9. There are 110–130 genes distributed in a chloroplast genome, which are involved in photosynthesis, transcription and translation processes10. The gene number and gene composition are conserved in chloroplast genomes of most plants, making them suitable for evolutionary analysis, Whereas variable regions of a genome provide sequence resources for developing molecular markers10,11,12,13.
RNA editing is a post-transcriptional modification of RNA, distinct from other modifications such as 5 prime capping and RNA splicing14. RNA editing events have been reported in many plant chloroplast genomes, such as maize, Spirodela, rice, and tobacco14,15,16,17. In plants, most RNA editing events detected in chloroplast mRNAs show cytidine to uridine conversion or sometimes an adenosine to inosine transition15. RNA editing in chloroplast mRNAs is more likely a rescue of mutation event than producing a new protein15,16. Members of PPR protein family appear to be involved, and a cis-element is required for editing site recognition19,20,21,22,23. RNA editing sites tend to favor the first and second bases of a codon, since most detected editing sites are the first two bases15,23.
Ycf1 is a gene with unknown function in chloroplast genomes. At structural level, it is usually predicted to contain six to eight trans-membrane domains at the N terminus and a variable hydrophilic C-terminal structural domain25. Moreover, due to high variability of Ycf1 gene sequences, it can be used as a promising plasmid DNA fragment for barcode development11,12. The function of YCF1 is still unclear, though there are some controversial reports that YCF1 included in the translocon at the inner envelope membrane of chloroplasts (TIC) system24,26,27,28,29,30,31.
Genomic researches on chloroplasts have been growing exponentially. Thanks to rapid development of sequencing technologies, the cost of sequencing is decreasing, and more chloroplast genomes will be deciphered and published. Long sequence reads facilitate assembly of chloroplast genomes. Here, we assembled and annotated the mountain papaya chloroplast genome using Oxford Nanopore Technology (ONT) reads, compared it with the C. papaya chloroplast genome, explored RNA editing profile of C. papaya chloroplast, and analyzed the evolution of the Ycf1 gene in the Caricaceae family.
Materials and methods
Sample collection and sequencing
Young leaves of V. pubescens were collected from a greenhouse in Fujian Agriculture and Forestry University, Fuzhou, China, and some of them were sent to Biomarker Technologies Corporation (Beijing) for DNA extraction, library construction, and Oxford Nanopore technology sequencing. The others were used to extract DNA in our lab using CTAB protocol and then sent to Biomarker Technologies Corporation (Beijing) Co., Ltd for library construction and next generation sequencing.
Chloroplast genome assembly and annotation
All sequence data from Nanopore sequencing was first corrected by Canu-1.732 and the reads were mapped to C. papaya chloroplast genome (GenBank: EU431223.1), which was downloaded from NCBI, using BLASR software33 with the parameters minMatch 15, minAlnLength 5000. All mapped reads were assembled via Smartdenovo (https://github.com/ruanjue/smartdenovo) to generate the first version of V. pubescens chloroplast genome. NGS data was then blasted to the first version genome and the reads mapped were used for chloroplast genome assembly using SPAdes3.13.134, to generate the second version of the V. pubescens chloroplast genome. The two versions were compared and mutually corrected. The unclear parts of the genome were corrected by Sanger sequencing to generate the final version of the V. pubescens chloroplast genome. Gene annotation was performed using Geseq35 and CPGAVAS236, and annotation results were manual corrected. The revised annotation gbf file was uploaded to OGDRAW37 to draw the chloroplast genome map.
Chloroplast genome analysis
The C. papaya and V. pubescens chloroplast genomes were aligned using the MAFFT7.0 software38 and the result was uploaded to DnaSP6.039 for DNA polymorphism analysis. GC content and codon usage bias analysis were performed in the GALAXY40 platform. We used mVISTA41 to compare chloroplast genomes of C. papaya and V. pubescens. MISA v1.042 (misa.ini parmeter: 1–10 2–5 3–4 4–3 5–3 6–3) and REPuter43 (minimal repeat size, 18 bp) were used to analyze simple sequence repeats (SSR) and repetitive sequences. PREP-cp44 was used to predict RNA editing sites of protein coding genes. Single nucleotide polymorphisms and indels were detected using DnaSP6.0 and a python script (https://www.biostars.org/p/119214/). Non-synonymous and synonymous rates of substitution analysis were done using ParaAT45 and KaKs_Calculator46. Haplotype detection was done using Cp-hap9 with both C. papaya and V. pubescens Nanopore reads (> 30 kb). The haplotype structure was confirmed using PCR and Sanger Sequencing. All related figures were drawn using R 3.4.0, IBS47, DNAMAN 6.0 software (Lynnon Biosoft) or OFFICE 2010.
Ycf1 gene analysis
Ycf1 gene sequences from V. pubescens and C. papaya were aligned, and identical sequences were selected to design primers to amplify Ycf1 genes using leaf DNA from Vasconcellea monoica, Jacaratia spinosa, Jarilla caudata, Jarilla heterophella, and Jarilla chocola. The phylogenetic tree of six Ycf1 genes was constructed by MEGA7.048 and MA model. Arabidopsis Ycf1 gene was selected as an out-group for analysis. DNA polymorphism and Ka/Ks ratio of Ycf1 genes were calculated, and YCF1 protein sequences were input to calculate dN/dS value using MEGA7.0 and easycodeML49. The trans-membrane region was predicted using TMHMM Server v. 2.050. The conserved motifs of Ycf1 genes were searched using MEME suit51.
Variant calling for RNA editing site
Four replications of RNA-seq data (unpublished) from C. papaya leaves were used for variant calling. The paired-end reads were mapped to the C. papaya chloroplast genome using HISAT252 and were sorted and converted into a bam file using samtools50,51. Variant calling with quality (> or = 20) and depth (> 10) were conducted using samtools (mpileup and bcftools). Variants in at least two replications were selected as final RNA editing candidate sites. Using DP4 value from final variant calling VCF file, we calculated RNA editing efficiency by edited reads divided by total mapped reads.
Result
Global comparison of V. pubescens and C. papaya chloroplast genomes
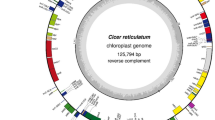
There were 124,170 ONT reads mapped to the C. papaya chloroplast genome, representing 10.3% of the total corrected ONT reads (1,195,269). These reads were assembled using Smartdenovo returned 2.4 M consensus unitigs, and the N50 is 118,474 bp. After self-blast, two unitigs were selected to assemble a closed circle chloroplast genome. We also did polishing and manually comparing with the chloroplast genome assembled with Illumina NGS data. Finally, we assembled the final version of the V. pubescens chloroplast genome. The size of V. pubescens chloroplast genome is 158,712 bp, 1388 bp smaller than the 160,100 bp in C. papaya chloroplast genome (deposited in GenBank: MT062856). The chloroplast genome of V. pubescens contains 131 genes, the same number as that of C. papaya, including 82 protein-coding genes, 2 pseudo genes (Ycf1* and infA*), 8 rRNA and 37 tRNA genes (Fig. 1, Table 1). In the V. pubescens chloroplast genome, the sizes of the LSC region and SSC region are 87,991 bp and 17,841 bp, respectively, which are separated by two IR regions. The IR region of V. pubescens is 26,440 bp, larger than that of C. papaya’s 26,325 bp. The rps19 genes in both V. pubescens and C. papaya crossed the LSC and IRb boundaries (Fig. 2). In the V. pubescens chloroplast genome, 15 protein-coding genes have introns, of which 11 genes have one intron and four genes have two introns, the same as those in C. papaya (S2.3). The GC content is the same at 37% in both chloroplast genomes.The untranslated region (UTR) had the highest GC content, whereas the lowest is in the intergenic region. The GC content of exons and introns was somewhere in between (Fig S1).
Gene map of the Vasconcellea pubescens. Thick lines indicate the extent of the inverted repeat regions (IRa and IRb), which separate the genome into small (SSC) and large (LSC) single copy regions. Genes drawn inside the circle are transcribed clockwise, and those outside are transcribed counter clockwise. Different colors represent different functional groups of Genes.
Structural variation and DNA polymorphism in V. pubescens and C. papaya
We compared V. pubescens and C. papaya chloroplast genome sequence similarity using mVISTA with C. papaya chloroplast as the reference. There is a high degree of synteny between the two chloroplast genomes, but sequences of genes, UTR, and CNS were highly variable (Fig S2). Three low consistency regions with less than 50% identity were found, two of which located in the LSC region (49,469–49,840 bp, 53,979–54,533 bp), while the third located in the SSC region (118,717–119,363 bp). In order to elucidate sequence nucleotide divergence between V. pubescens and C. papaya, we used DnaSP6.0 to analyze sequence nucleotide variability with a 600 bp sliding-window and a 200 bp step size. The highest value was observed in genes psbZ, trnG-GCC, between trnH-GUG and psbA, trnS-GCU and trnG-UCC, and gene rps16 (S2.11). We also found that average DNA sequence variation in SSC region is higher than that in LSC. Lower divergences are in IR regions (S2.4). We also investigated single nucleotide polymorphisms (SNPs) and indels between the two genomes with 2,321 SNPs and 387 indels found (S2.5).
SSR and repetitive sequences
Six types of SSRs were founded in the C. papaya chloroplast genome, including: single nucleotide, dinucleotide, trinucleotide, tetranucleotide, pentanucleotide, and hexanucleotide. Five types of SSRs were found in the V. pubescens chloroplast genome, including single nucleotides, dinucleotides, trinucleotides, tetranucleotides, and pentanucleotides. C. papaya had 103 SSRs compared to 84 in V. pubescens. C. papaya had a higher number of SSRs in each type except for the dinucleotide type. Repeated sequence information is important for phylogenetic analysis, which plays an important role in rearrangements of the genome54. 103 repetitive sequences were found in C. papaya, while 86 were found in V. pubescens. Similar to SSRs, all types of repetitive sequences in V. pubescens are fewer than those in C. papaya, which has 45 palindromic repeats, 34 direct repeats, 18 inverted repeats, and 6 complementary repeats, whereas V. pubescens has 36 palindromic repeats, 27 direct repeats, 14 inverted repeats, and 3 complementary repeats (Fig. 3).
RNA editing
In seed plants, RNA editing in the chloroplast converts cytidine to uridine (C to U RNA editing), some of which can affect protein function54,55. RNA editing in chloroplast was reported to contribute to chloroplast-to-nucleus signaling56,57. We predicted RNA editing sites in the V. pubescens and C. papaya chloroplast genomes. 52 RNA editing sites were detected in both V. pubescens and C. papaya, and coding genes of the subunits of the NAD(P)H dehydrogenase complex appear to undergo extensive RNA editing (S2.6). ndhB (12/12 loci) had the highest number of RNA edits, followed by ndhD (8/8 loci), ndhA (4/4 loci), rpoC2 (4/3 loci), ndhF (3/3 loci) and ndhG (3/3 loci). In V. pubescens, 56% of the editing sites occurred on the serine codon, and 54% of the editing events changed the target codon to the leucine codon. In C. papaya, 50% of the editing sites occurred on the serine codon and 50% of the editing sites changed to the leucine codon. For locations of editing sites, 80% of the RNA editing events in V. pubescens occurred at the second nucleotide of the codon, 20% at the first nucleotide, and no change was detected at the third nucleotide. The type and pattern of RNA editing in C. papaya were similar to those in V. pubescens. The start codon of ndhD was predicted to be edited from ACG to ATG, back into the normal start codon. Most RNA editing sites changed amino acids from polar to nonpolar, resulting in an increased protein hydrophobicity.
We used RNA-seq data to explore RNA editing events in C. papaya chloroplast genome and detected 46 RNA editing sites in 30 genes and 3 intergenic regions. All of them are C-to-U conversion. Of all editing loci, 15 (32%) located in the NDH components coding genes. The ribosomal protein-coding genes have 8 sites. Among them, the ndhD gene has the most frequent RNA editing sites (6 sites) and the rpoB has 3 sties. We detected only 2 loci in the ndhB gene, though this gene was predicted to be the one with the highest editing frequency by PREP. We calculated RNA editing efficiency based on the DP4 value in the VCF file, which is variable, ranging from 15 to 98%, with an average editing efficiency of 63%. RNA editing efficiency is also different between the same gene editing sites. For example, RNA editing efficiency of the ndhD ranges from 34 to 66% (Fig. 4). We selected three sites for verification based on either they showed low quality during the filter step in some replications or they are not detected in some replications, and all of them were verified by Sanger sequencing results, which showed a duplication peak or different sequencing results from different transformants in the editing sites (Fig. S4–S6).
RNA editing efficiency plot of Carica papaya chloroplast RNA. Different colors respect for different Region of chloroplast genome, red for LSC, green for IRa, yellow for SSC and blue for IRb region. X axis represent RNA efficiency, and the number after the gene name in the Y axis represent the editing site.
Codon usage bias
Codon usage bias (CUB) means different frequencies among synonymous codons. Analyzing CUB can help us to understand molecular evolution, environmental adaptation, and genomic features59. The chloroplast genome CUB is highly conserved and varies among different genes60. We explored the codon usage pattern of the chloroplast genome of V. pubescens and C. papaya using those protein-coding genes that have more than 300 nucleotides61. We first calculated relative synonymous codon usage (RSCU) value for each gene and marked codons with priority usage (RSCU > 1) with an asterisk (S2.7). We then investigated the base composition of each codon at each position in V. pubescens. The mean GC content was 47.4% in the first position, 40.2% in the second position, 28.7% in the third position, and 38.7% for all codons (Fig. 5A). The correlation between GC12 and GC3 was not significant (R2 = 0.0284), indicating that selective pressure does not affect the CUB in V. pubescens chloroplast genome (Fig. 5B). The codon adaptation index (CAI) is a measure of directional synonymous codon usage bias proposed by Paul et. al in 198760. We investigated the CAI of each gene in V. pubescens chloroplast genome by dividing the genes into three types: photosynthesis-related genes (photo-genes), genetic system-related genes (Genet-genes), and other genes. The CAI of the photo-genes was higher than that of other genes (Fig. 5C), and the CAI of genetic genes was the lowest58,61. The effective number of codons (ENC) values are used to quantify how far codon usage of a gene departs from equal usage of synonymous codons62. The ENCs of V. pubescens chloroplast genes ranged from 38.36 to 56.88, which implies that CUB of genes is different and relatively weak (Fig. 5D). The relationship between base composition and ENC was investigated using ENC plots. Most genes were off the standard curve, indicating that no base composition was affected CUB.
Vasconcellea pubescens chloroplast genome codon usage pattern related plot. (A) GC content of different codon sites. (B) Neutrality plot (GC12 against GC3). (C) The codon adaptation index (CAI) value of different function gene sets. (D) Relationship between GC3 and effective number of codons (ENC) (ENC-plot).
Structural haplotypes
We detected two haplotypes of the chloroplast genome from both C. papaya and V. pubescens, and the directions of the haplotypes are LSC_IRa_SSCrc_IRb and LSC_IRa_SSC_IRb (“rc” means reverse and complementary) (Fig. 6B). In V. pubescens, the ONT reads covering the LSC_IRa_SSCrc_IRb type were 6435 and the reads covering the LSC_IRa_SSC_IRb type were 6485. Meanwhile, in C. papaya, the numbers were 2047 and 2203, respectively (Fig. 6A). We further designed primers to verify the existence of the two types of chloroplast genome structures, PCR products of expected size were obtained. Sanger sequencing results were also confirmed that the PCR products either covering the end part of IR or the start part of SSC region or covering the end part of IR and start part of SSCrc region (S3). Using combination of the start part and the end part of the LSC region sequence (2371 bp) as a query to blast all the V. pubescens ONT reads, we detected one ONT read (we named it “ONT1”, 44,186 bp) that covered the end and start part of LSC sequence (S2.12). We then extracted and used the ONT1 sequence as a query to blast the V. pubescens chloroplast genome (S2.12), it does cover the start part of the LSC (from site 1 to site 16,440) and the end part of the LSC (from site 59,366 to site 87,980), and the LSC region start from 1 to 87,991. Based on these results, we conclude that there was a closed circle LSC form of chloroplast structure exist in the cell. However, the amount of this atypical type chloroplast DNA may be very small, since we only got one ONT read that covered the combined fragment. The other 505 blast hits represented ONT reads that covered only one of the two separate parts of the combined fragment or covered both but the ONT read represents the normal LSC fragment.
Stucture haplotypes plot. The number of Nanopore reads that supported the two structure haplotypes of Carica papaya and Vasconcellea pubescens chloroplast genome, respectively (A). Two different structure haplotypes of the chloroplast genome detected in this study, the dashed line with different colors illustrated Nanopore reads with different lengths which covered different regions of the two haplotypes chloroplast molecular (B).
Non-synonymous and synonymous rate of substitution
The evolutionary rates of all 78 protein coding genes were analyzed according to non-synonymous and synonymous rates of substitution (Ka and Ks). We generated 61 Ka/Ks ratio values (S2.8). The mean Ka/Ks ratio was 0.198, and most of the ratios were lower than 0.5, indicating purifying selection. The Ka/Ks of Ycf2 gene, however, was greater than 1, indicating positive selection.
Ycf1 gene
The function of YCF1 remains unknown. We compared Ycf1 gene sequences of V. pubescens and C. papaya, and designed primers to amplify Ycf1 orthologous gene of several other Caricaceae species. Finally, we got all seven Ycf1 gene sequences from four genera in family Caricaceae, and the four genera are Jarilla, Carica, Vasconcella, and Jacaratia (S2.9). Multiple alignment results showed that 5 prime (1–605 bp) and 3 prime regions (about 100 bp) of all Ycf1 genes were nearly identical, except variable 3 prime regions. Using the TMHMM website to predict trans-membrane helices, we identified six trans-membrane helices in all YCF1 proteins, as previously reported (Fig. S7)29. We also found a motif RLEDLACMNRYW through MEME in 3 prime regions (about 100 bp), which is consistent with previously reported conserved motif in the carboxy terminus of YCF1 in Ste-YCF125. We further investigated the indels (insertion-deletion mutations) and SNP distribution profile among all seven Ycf1 genes, and all four genera’s Ycf1 genes had their genus-specific indels and such situation was not found in the SNPs. Nine indels were detected and although the Carica genus has only one species, it contained five of the nine indels, including 4 insertions and 1 deletion (Table 2, S4). The genera Vasconcella and Jacaratia each had one genus-specific insertion, and the genus Jarilla had two genus-specific insertions. We then added Arabidopsis Ycf1 gene sequence for indel analysis, and all insertions and deletions were genus-specific, occured after the divergence of Brassicaceae and Caricaceae (S5). We also selected eight Ycf1 genes from three different genera in family Lauraceae, and alignment results did not show similar results (S6).
The evolution of Ycf1 gene was analyzed by calculating synonymous (Ks) and nonsynonymous (Ka) values of Ycf1 sequences from seven species in four genera. The results showed that Ka/Ks < 1, indicating that the Ycf1 was under purifying selection. We then examined site-specific evolution of the Ycf1 gene, and only one locus (311 K 0.961*) was positively selected according to the M8 model. The phylogenetic tree of Ycf1 genes shows that Jacaratia spinosa was close to the Vasconcellea genus (V. pubescens and V. monoica), while C. papaya was close to Jarilla genus (Jarilla heterophella, Jarilla chocola and Jarilla caudata) (Fig. S8).
The photosystem biogenesis regulator 1 (PBR1) strongly regulates the expression of the chloroplast gene Ycf1 by binding to the 5′UTR of the chloroplast gene Ycf1 mRNA in Arabidopsis. We also compared the 5′UTR of V. pubescens, C. papaya, and A. thaliana Ycf1 genes. All 5′UTR sequences have the same size (73 bp), with only one difference detected in the position 43 where V. pubescens and C. papaya have C base while A. thaliana has T base.
Discussion
The chloroplast genome of highland papaya V. pubescens was sequenced and assembled using Nanopore long reads combined with Illumina short reads. This long read sequences greatly facilitated the assembly of the chloroplast genome, and also revealed structural variations of the chloroplast genome. Our assembly results showed that two unitigs were sufficient to produce the whole circular chloroplast genome when assembled using the Smartdenovo soft. Meanwhile, we also tested the newly published NECAT2 software, which produced only one contig that covered almost the whole V. pubescens chloroplast genome (99% similar to the final version V. pubescens chloroplast genome, after polishing with the NGS data). All results showed advantage of long reads. Two haplotypes, LSC_IRa_SSCrc_IRb and LSC_IRa_SSC_IRb, were detected in both V. pubescens and C. papaya. Though different versions of the chloroplast genome structure of some plants have been reported recently9, there is no previous report in the Caricaceae family. The change of the chloroplast structures was assumed to be the result of flipped recombination9,64,65. We also found one over 40 kb ONT read that covered the beginning segment and ending segment of the LSC region. The amount of circular type LSC maybe very small, since there is only one ONT read among the 506 ONT reads matching the complete query sequence in the same direction. Chloroplast structure plasticity was reported in higher plants and chloroplast DNA without IRs was also found8. Our results further confirmed the structure plasticity of the chloroplast.
The chloroplast genome of V. pubescens was smaller than that of C. papaya, but its IR regions was longer than C. papaya. Unequal recombination and replication slippage can result in expansion and contraction of the IR regions, leading to variations in chloroplast genome size59,63,64. Five highly variable regions were identified in the V. pubescens and C. papaya chloroplast genomes, which are psbZ, trnG-GCC, trnH-GUG and psbA, trnS-GCU and trnG-UCC, and rps16. These variable sequences between the two chloroplast genomes can be used to develop molecular markers that have been widely used for new species identification in other plant lineages53,59,65,66,67.
RNA editing is an important tool for gene expression regulation, and most RNA editing events in chloroplast genomes are converting C to U15,54,55. Members of the pentatricopeptide repeat (PPR) protein family play an important role in RNA editing in chloroplast genomes, which has been reported in Arabidopsis, rice, and maize14,68,69,70,71,72. Prediction of RNA editing loci in chloroplasts helps us to understand chloroplast regulatory mechanisms and design strategies for chloroplast genome engineering. With PREP prediction, we detected 52 RNA editing loci in 18 protein-coding genes in V. pubescens and C. papaya. We used RNA-seq data from C. papaya leaves for variant calling, and identified 46 RNA editing loci located in 30 genes and 3 intergenic regions of the chloroplast genome. Some of these loci were verified by Sanger sequencing. RNA editing frequently occurs in gene-encoding components of the NDH complex, particularly ndhD gene. This is similar to the Ginkgo biloba chloroplast genome, supporting the hypothesis that the highest RNA editing events occur in the NDH complex genes in seed plants17,73,74. RNA editing efficiency were also calculated which showed differences across all genes with an average efficiency of 63%. RNA editing efficiency in Spirodela polyrhiza averaged 76%, higher than that in C. papaya17. The changes in RNA editing efficiency on genes or at different loci of the same gene imply complex RNA editing regulatory mechanisms, which need to be explored. RNA editing could cause protein structure change and regulate the photosynthesis process18,73. Therefore, the detailed analysis of RNA editing loci in the chloroplast genome is quite necessary.
Ycf1 genes were amplified from seven species in four genera of the Caricaceae family. The N terminal (605 bp) and C terminal (100 bp) of the Ycf1 genes are nearly identical, which enables us to amplify Ycf1 from different species, and also implies the evolutionary conservation of both regions of Ycf1 genes in Caricaceae family, whereas the other regions of the Ycf1 genes showed very high variation, making them good candidates for barcode development. All seven Ycf1 genes from different genera had genus-specific indels. These results implied that indels may play an important role in genus-specific evolution of Ycf1 genes in Caricaceae. Six trans-membrane helices were found as previously reported3. Analysis of full length Ycf1 genes showed that the Ycf1 genes were under purification selection, whereas site-specific evolution analysis results showed that position 311 K was under positive selection with a p < 0.05. This suggests that a specific locus of the gene might evolve independently under a specific driving force, or the 311 K locus might be an important boundary for YCF1.
Data availability
Raw data of high-throughput sequencing and Oxford Nanopore sequencing were deposited in NCBI SRA. (SRA accession: PRJNA605960, https://www.ncbi.nlm.nih.gov/sra/PRJNA605960).
References
Salvatierra-González, M. A. & Jana-Ayala, C. Floral expression and pollen germination ability in productive mountain papaya (Vasconcellea pubescens A.DC.) orchards. Chil. J. Agric. Res. 76, 132–142 (2016).
Ming, R., Yu, Q. & Moore, P. H. Sex determination in papaya. Semin. Cell Dev. Biol. 18, 401–408 (2007).
Ray, M. & Moore, P. Genetics and Genomics of Papaya (Springer, Berlin, 2014).
Gaete-Eastman, C. et al. Expression of an ethylene-related expansin gene during softening of mountain papaya fruit (Vasconcellea pubescens). Postharvest Biol. Technol. 53, 58–65 (2009).
Chong-Pérez, B. et al. Regeneration of highland papaya (Vasconcellea pubescens) from anther culture. Appl. Plant Sci. 6, 3–10 (2018).
Li, W., Zhang, C., Guo, X., Liu, Q. & Wang, K. Complete chloroplast genome of Camellia japonica genome structures, comparative and phylogenetic analysis. PLoS ONE 14, 1–18 (2019).
Daniell, H., Lin, C. S., Yu, M. & Chang, W. J. Chloroplast genomes: diversity, evolution, and applications in genetic engineering. Genome Biol. 17, 1–29 (2016).
Lilly, J. W., Havey, M. J., Jackson, S. A. & Jiang, J. Cytogenomic analyses reveal the structural plasticity of the chloroplast genome in higher plants. Plant Cell. 13, 245–254 (2001).
Wang, W. & Lanfear, R. Long-reads reveal that the chloroplast genom8e exists in two distinct versions in most plants. Genome Biol. Evol. 11, 1–31 (2019).
Amiryousefi, A., Hyvönen, J. & Poczai, P. The chloroplast genome sequence of bittersweet (Solanum dulcamara): Plastid genome structure evolution in Solanaceae. PLoS ONE 13, 1–23 (2018).
Dong, W. et al. ycf1, the most promising plastid DNA barcode of land plants. Sci. Rep. 5, 8348 (2015).
Neubig, K. M. et al. Phylogenetic utility of ycf1 in orchids: a plastid gene more variable than matK. Plant Syst. Evol. 277, 75–84 (2009).
Hernández-León, S., Gernandt, D. S., Pérez de la Rosa, J. A. & Jardón-Barbolla, L. Phylogenetic relationships and species delimitation in Pinus section Trifoliae inferrred from plastid DNA. PLoS ONE 8, 1–14 (2013).
Gommans, W. M., Mullen, S. P. & Maas, S. RNA editing: a driving force for adaptive evolution?. BioEssays 31, 1137–1145 (2010).
Xiao, H. et al. A rice dual-localized pentatricopeptide repeat protein is involved in organellar RNA editing together with OsMORFs. J. Exp. Bot. 69, 2923–2936 (2018).
Sasaki, T., Yukawa, Y., Miyamoto, T., Obokata, J. & Sugiura, M. Identification of RNA editing sites in chloroplast transcripts from the maternal and paternal progenitors of tobacco (Nicotiana tabacum): comparative analysis shows the involvement of distinct trans-factors for ndhB editing. Mol. Biol. Evol. 20, 1028–1035 (2003).
Wang, W., Zhang, W., Wu, Y., Maliga, P. & Messing, J. RNA editing in chloroplasts of Spirodela polyrhiza, an aquatic monocotelydonous species. PLoS ONE 10, 1–13 (2015).
Maier, R. M., Neckermann, K., Igloi, G. L. & Koössel, H. Complete sequence of the maize chloroplast genome: gene content, hotspots of divergence and fine tuning of genetic information by transcript editing. J. Mol. Biol. 251, 614–628 (1995).
Tang, W. Regulation of RNA editing in chloroplast. Open Biotechnol. J. 12, 16–24 (2018).
Mizuho Ichinose, M. S. The DYW domains of pentatricopeptide repeat RNA editing factors contribute to discriminate target and non-target editing sites. Plant Cell Physiol. 59, 1652–1659 (2018).
Härtel, B. et al. MEF10 is required for RNA editing at nad2-842 in mitochondria of Arabidopsis thaliana and interacts with MORF8. Plant Mol. Biol. 81, 337–346 (2013).
Guillaumot, D. et al. Two interacting PPR proteins are major Arabidopsis editing factors in plastid and mitochondria. Proc. Natl. Acad. Sci. 114, 8877–8882 (2017).
Shikanai, T. RNA editing in plant organelles: machinery, physiological function and evolution. Cell. Mol. Life Sci. 63, 698–708 (2006).
Yura, K. & Go, M. Correlation between amino acid residues converted by RNA editing and functional residues in protein three-dimensional structures in plant organelles. BMC Plant Biol. 8, 1–11 (2008).
De Vries, J., Archibald, J. M. & Gould, S. B. The carboxy terminus of YCF1 contains a motif conserved throughout > 500 Myr of streptophyte evolution. Genome Biol. Evol. 9, 473–479 (2017).
Nakai, M. Ycf1: a green TIC: response to the de Vries et al. commentary. Plant Cell 27, 1834–1838 (2015).
Nakai, M. New perspectives on chloroplast protein import. Plant Cell Physiol. 59, 1111–1119 (2018).
Bölter, B. & Soll, J. Ycf1/Tic214 is not essential for the accumulation of plastid proteins. Mol. Plant 10, 219–221 (2017).
de Vries, J., Sousa, F. L., Bölter, B., Soll, J. & Goulda, S. B. YCF1: a green TIC?. Plant Cell 27, 1827–1833 (2015).
Kikuchi, S. et al. Uncovering the protein translocon at the chloroplast inner envelope membrane. Science 339, 571–574 (2013).
Nakai, M. The TIC complex uncovered: the alternative view on the molecular mechanism of protein translocation across the inner envelope membrane of chloroplasts. Biochim. Biophys. Acta Bioenerg. 1847, 957–967 (2015).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 25, 1–11 (2014).
Chaisson, M. J. & Tesler, G. Mapping single molecule sequencing reads using basic local alignment with successive refinement (BLASR): application and theory. BMC Bioinform. 13, 238 (2012).
Nurk, S. et al. Assembling genomes and mini-metagenomes from highly chimeric reads. Lect. Notes Comput. Sci. 782, 158–170 (2013).
Tillich, M. et al. GeSeq-versatile and accurate annotation of organelle genomes. Nucleic Acids Res. 45, W6–W11 (2017).
Shi, L. et al. CPGAVAS2, an integrated plastome sequence annotator and analyzer. Nucleic Acids Res. 47, W65–W73 (2019).
Greiner, S., Lehwark, P. & Bock, R. OrganellarGenomeDRAW (OGDRAW) version 1.3.1: expanded toolkit for the graphical visualization of organellar genomes. Nucleic Acids Res. 47, W59–W64 (2019).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Rozas, J. et al. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol. Biol. Evol. 34, 3299–3302 (2017).
Afgan, E. et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2018 update. Nucleic Acids Res. 46, W537–W544 (2018).
Frazer, K. A., Pachter, L., Poliakov, A., Rubin, E. M. & Dubchak, I. VISTA: computational tools for comparative genomics. Nucleic Acids Res. 32, 273–279 (2004).
Beier, S., Thiel, T., Münch, T., Scholz, U. & Mascher, M. MISA-web: a web server for microsatellite prediction. Bioinformatics 33, 2583–2585 (2017).
Kurtz, S. REPuter: the manifold applications of repeat analysis on a genomic scale. Nucleic Acids Res. 29, 4633–4642 (2001).
Mower, J. P. The PREP suite: predictive RNA editors for plant mitochondrial genes, chloroplast genes and user-defined alignments. Nucleic Acids Res. 37, 12–14 (2009).
Zhang, Z. et al. ParaAT: a parallel tool for constructing multiple protein-coding DNA alignments. Biochem. Biophys. Res. Commun. 419, 779–781 (2012).
Zhang, Z. et al. KaKs_calculator: calculating ka and ks through model selection and model averaging. Genomics Proteomics Bioinform. 4, 259–263 (2006).
Liu, W. et al. IBS: an illustrator for the presentation and visualization of biological sequences. Bioinformatics 31, 3359–3361 (2015).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874 (2016).
Gao, F. et al. EasyCodeML: a visual tool for analysis of selection using CodeML. Ecol. Evol. 9, 3891–3898 (2019).
Krogh, A., Larsson, B., Von Heijne, G. & Sonnhammer, E. L. L. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J. Mol. Biol. 305, 567–580 (2001).
Bailey, T. L. et al. MEME Suite: tools for motif discovery and searching. Nucleic Acids Res. 37, 202–208 (2009).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11, 1650–1667 (2016).
Li, H. et al. The sequence alignment/map format and SAM tools. Bioinformatics 25, 2078–2079 (2009).
Khan, A. et al. First complete chloroplast genomics and comparative phylogenetic analysis of Commiphora gileadensis and C. foliacea: Myrrh producing trees. PLoS ONE 14, 1–21 (2019).
Sugiura, M. RNA editing in chloroplasts. In RNA Editing. Nucleic Acids and Molecular Biology (ed. Göringer, H. U.) 123–142 (Springer, Berlin, 2008).
Tillich, M., Lehwark, P., Morton, B. R. & Maier, U. G. The evolution of chloroplast RNA editing. Mol. Biol. Evol. 23, 1912–1921 (2006).
Larkin, R. M. RNA editing implicated in chloroplast-to-nucleus communication. Proc. Natl. Acad. Sci. U.S.A. 116, 9701–9703 (2019).
Zhao, X., Huang, J. & Chory, J. GUN1 interacts with MORF2 to regulate plastid RNA editing during retrograde signaling. Proc. Natl. Acad. Sci. U.S.A. 116, 10162–10167 (2019).
Zhang, R. et al. Differences in codon usage bias between photosynthesis-related genes and genetic system-related genes of chloroplast genomes in cultivated and wild Solanum species. Int. J. Mol. Sci. 19, 3142 (2018).
Sharp, P. M. & Li, W. H. The codon adaptation index-a measure of directional synonymous codon usage bias, and its potential applications. Nucleic Acids Res. 15, 1281–1295 (1987).
Yan, M. et al. The complete chloroplast genomes of Punica granatum and a comparison with other species in Lythraceae. Int. J. Mol. Sci. 20, 2886 (2019).
Wright, F. The ‘effective number of codons’ used in a gene. Gene 87, 23–29 (1990).
Kim, D. et al. Flip-flop organization in the chloroplast genome of Capsosiphon fulvescens (Ulvophyceae, Chlorophyta). J. Phycol. 55, 214–223 (2019).
Stein, D. B., Palmer, J. D. & Thompson, W. F. Structural evolution and flip-flop recombination of chloroplast DNA in the fern genus Osmunda. Curr. Genet. 10, 835–841 (1986).
Shahzadi, I. et al. Chloroplast genome sequences of Artemisia maritima and Artemisia absinthium: comparative analyses, mutational hotspots in genus Artemisia and phylogeny in family Asteraceae. Genomics https://doi.org/10.1016/j.yg-eno.2019.08.016 (2019).
Menezes, A. P. A. et al. Chloroplast genomes of Byrsonima species (Malpighiaceae): comparative analysis and screening of high divergence sequences. Sci. Rep. 8, 1–12 (2018).
Yue, F., Cui, L., dePamphilis, C. W., Moret, B. M. E. & Tang, J. Gene rearrangement analysis and ancestral order inference from chloroplast genomes with inverted repeat. BMC Genomics 9, 1–9 (2008).
Xue, S. et al. Comparative analysis of the complete chloroplast genome among Prunus mume, P. armeniaca, and P. salicina. Hortic. Res. 6, 1–13 (2019).
Wallinger, C. et al. Rapid plant identification using species- and group-specific primers targeting chloroplast DNA. PLoS ONE 7, e29473 (2012).
Santos, C. & Pereira, F. Identification of plant species using variable length chloroplast DNA sequences. Forensic Sci. Int. Genet. 36, 1–12 (2018).
Andrés-Colás, N. et al. Multiple PPR protein interactions are involved in the RNA editing system in Arabidopsis mitochondria and plastids. Proc. Natl. Acad. Sci. 114, 8883–8888 (2017).
Hammani, K., Takenaka, M., Miranda, R. & Barkan, A. A PPR protein in the PLS subfamily stabilizes the 5′-end of processed rpl16 mRNAs in maize chloroplasts. Nucleic Acids Res. 44, 4278–4288 (2016).
Shikanai, T. RNA editing in plants: Machinery and flexibility of site recognition. Biochim. Biophys. Acta Bioenerg. 1847, 779–785 (2015).
He, P., Huang, S., Xiao, G., Zhang, Y. & Yu, J. Abundant RNA editing sites of chloroplast protein-coding genes in Ginkgo biloba and an evolutionary pattern analysis. BMC Plant Biol. 16, 1–12 (2016).
Acknowledgements
This work was supported by startup fund from Fujian Agriculture and Forestry University. It was also supported by Natural Science Foundation of China (31701889) and Natural Science Foundation of Fujian Province of China (2018J01601).
Author information
Authors and Affiliations
Contributions
Z.Lin and R.M. designed the project. Z.Lin performed most of the data analysis and experiments. Z.Liao helped to prepare Fig. 6, R.L. helped to prepared Fig. 5, Y.D. helped in genome assembly strategy design. Z.Lin wrote the manuscript. Y.D., P.Z., X.M, and R.M. revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lin, Z., Zhou, P., Ma, X. et al. Comparative analysis of chloroplast genomes in Vasconcellea pubescens A.DC. and Carica papaya L.. Sci Rep 10, 15799 (2020). https://doi.org/10.1038/s41598-020-72769-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-72769-y
- Springer Nature Limited