Abstract
Plant growth-promoting bacteria (PGPB) may trigger tolerance against biotic/abiotic stresses and growth enhancement in plants. In this study, an endophytic bacterial strain from rapeseed was isolated to assess its role in enhancing plant growth and tolerance to abiotic stresses, as well as banded leaf and sheath blight disease in maize. Based on 16S rDNA and BIOLOG test analysis, the 330-2 strain was identified as Bacillus subtilis. The strain produced indole-3-acetic acid, siderophores, lytic enzymes and solubilized different sources of organic/inorganic phosphates and zinc. Furthermore, the strain strongly suppressed the in vitro growth of Rhizoctonia solani AG1-IA, Botrytis cinerea, Fusarium oxysporum, Alternaria alternata, Cochliobolus heterostrophus, and Nigrospora oryzae. The strain also significantly increased the seedling growth (ranging 14–37%) of rice and maize. Removing PCR analysis indicated that 114 genes were differentially expressed, among which 10%, 32% and 10% were involved in antibiotic production (e.g., srfAA, bae, fen, mln, and dfnI), metabolism (e.g., gltA, pabA, and ggt) and transportation of nutrients (e.g., fhu, glpT, and gltT), respectively. In summary, these results clearly indicate the effectiveness and mechanisms of B. subtilis strain 330-2 in enhancing plant growth, as well as tolerance to biotic/abiotic stresses, which suggests that the strain has great potential for commercialization as a vital biological control agent.
Similar content being viewed by others
Introduction
Plant growth-promoting bacteria (PGPB)1 colonize the plant rhizosphere, which enhances the plant growth and control of soil-borne diseases2. The PGPB associated with plant roots are often classified by the locations where they colonize, i.e., rhizosphere and endosphere3. The PGPB living in the rhizosphere and endosphere are known as rhizobacteria and endophytes, respectively. Bacillus species are important members of PGPB, which induce plants to tolerate abiotic and/or biotic stresses in a comprehensive manner4. Bacillus spp. have also been commercialized as biofertilizers and biocontrol agents5. The endophytic Bacillus spp. do not cause any visible damage or morphological alterations to the host. Therefore, these bacteria can be beneficial for the survival of the host species against environmental stresses and microbial competition6, 7. Furthermore, these bacteria may promote the growth of the plant through the formation of nodules with non-specific hosts8.
Bacillus spp. produce indole-3-acetic acid (IAA), which helps with nitrogen fixation from the atmosphere, siderophore production, solubilization of potassium (K), zinc (Zn), and phosphate (P) from the soil, and increasing the soil porosity7, 9 Along with the micro- and macro-nutrient supply, endophytic Bacillus spp. protect the plants from pathogens and play roles as antagonists7. They inhibit the activities of pathogens by producing diverse antimicrobial compounds, including siderophores10, hydrolytic enzymes11 and antibiotics12, volatile organic compounds (VOCs)13, and lipopeptides14 that are associated with the observed biocontrol activity against plant pathogens15, 16.
Rhizoctonia solani Kühn (teleomorph: Thanatephorus cucumeris (Frank) Donk) is one of the most prevalent soil-borne pathogen, which causes a significant economic losses in several economically important crops such as maize, rice and soybean17. Banded leaf and sheath blight (BLSB), caused by R. solani, is the most threatening disease, which may cause complete crop failure18. B. cinerea is an aggressive pathogen with a wide host range and uses multiple weapons to invade its host plants19, while F. oxysporum causes two major diseases of maize such as Fusarium ear rot and Gibberella ear rot, both of which can result in mycotoxic contamination of maize grains20. Under temperate and moist conditions, C. heterostropus causes southern leaf blight on corn and have devastating effect on other major cereal crops21. In addition to these devastating diseases, there are many other economically important diseases such as, rice grain spot and root rot of maize caused by N. oryzae under stress conditions22. On the other hand, A. alternata is one of the most common contaminating fungal pathogen, detected in cereal grains before harvest and may contribute to decrease in grain quality23.
The isolation and identification of differentially expressed genes was performed using several methods, including differential analysis of library expression (DAZLE)24, representational difference analysis (RDA)25, differential display and related techniques26, enzymatic degradation subtraction27, techniques involving physical removal of common sequences28, linker capture subtraction29, and suppression subtractive hybridization (SSH)30. These methods are vital but also possess some intrinsic drawbacks, such as the fact that the sequence must be known in advance. This leads to the generation of many false positives, sequencing that produces very short sequences, lack of reproducibility, and post-sequencing data analysis expenses31,32,33. Recently, a novel technique called removing polymerase chain reaction (R-PCR) has been used to efficiently remove the undesirable genes from a gene population33. The R-PCR reaction is the reverse process of a PCR. In PCR, the desired genes are amplified cycle by cycle, whereas in R-PCR, the undesired genes get removed cycle by cycle. The R-PCR permits rapid identification of differentially expressed genes, while excluding the false positive and false negative clones. Therefore, we performed this experiment to identify the antagonism-related genes from B. subtilis strain 330-2 using the most advanced R-PCR technique.
Therefore, one endophytic Bacillus was isolated from a rapeseed that had broad-spectrum antagonistic activity against several phytopathogenic fungi, including R. solani, F. oxysporum, B. cinerea, A. alternata, C. heterostrophus and N. oryzae. This study was aimed at (1); Isolation, identification, and characterization of the bacterial B. subtilis strain 330-2. (2); Evaluation of the antagonism of B. subtilis 330-2 on different plant pathogens. (3); Identification of the antagonism-related genes using R-PCR.
Results
Identification and properties of the strain
The isolated strain was catalase-positive, gram-positive, rod-shaped, aerobic, and motile and had distinct fermentation profiles for different carbon sources (Supplementary Table 1). Based on the observed phenotypic characteristics, the isolated strain was grouped into the genus Bacillus. Furthermore, it exhibited high levels of similarity (99%) to the closest known species in the database. Phylogenetic analysis of the 16S rDNA gene sequence revealed that the isolated 330-2 strain was Bacillus subtilis. In addition to B. subtilis, another species of the genus Bacillus, B. amyloliquefaciens, showed (99%) similarity with the strain during the BLAST search. The phylogenetic tree analysis using Mega5 version 5.2 showed that strain 330-2 was more closely related to B. subtilis and B. amyloliquefaciens (Fig. 1). To further confirm the strain, a BIOLOG test was performed.
An amplified 16S rDNA gene fragment from the isolated strain 330-2 was sequenced and blast searched through NCBI database. Closely related sequences were downloaded and aligned. These sequences were analyzed using the Maximum likelihood method. The GenBank accession number of each isolate is given in parentheses. Bootstrap values based on 1000 replicates are shown next to the branches.
BIOLOG identification of the bacterial strain
Phenotypic analyses was done using 96-well BIOLOG GENIII MicroPlates assay. The list of carbon substrates used by B. subtilis sp. is presented (Fig. 2 and Supplementary Table 1). The results of the 16S rDNA sequencing and BIOLOG identification indicated that the strain used in this study was B. subtilis.
The BIOLOG detection results showing the comparison of the B. subtilis strain 330-2 and B. subtilis strain 168. The negative control (A1); the positive control (A10). The well with a faint purple color shows the positive reaction, the half-filled light blue circle shows the moderate reaction, while the blank circle shows the negative reaction.
Production of hydrolytic enzymes and metabolites by bacterial antagonists
Progression of the pink color in the presence or absence of tryptophan in the culture broth showed the production of IAA. Maximum IAA production was recorded as 24.13 μg ml−1 in the B. subtilis strain 330-2, followed by 2.10 μg ml−1 in Bacillus subtilis strain 168 broth culture (B. subtilis strain 168 was taken as control). The B. subtilis strain 330-2 also produced the siderophore through the development of orange halo zones around their spot of inoculation on Chrome-azurol S (CAS) medium, while B. subtilis strain 168 did not produce a siderophore.
The B. subtilis strain 330-2 showed a positive result for phosphate solubilization by spot inoculation in the solid state of Pikovskaya’s media, but did not solubilize the Di-calcium phosphate (DCP) by spot inoculation on Pikovskaya’s medium. On the plates, B. subtilis strain 330-2 P-solubilizing zones around the bacterial colony were 4.2 and 6.3 mm using Tri-calcium phosphate (TCP) and zinc phosphate (ZP), respectively. However, B. subtilis strain 168 could not solubilize the phosphate in Pikovskaya’s solid medium. Both strains were tested for phytase production, whereas B. subtilis strain 330-2 solubilized calcium and sodium phytate by formation of halo zones around its spot of inoculation (12.7 and 9.6 mm for calcium and sodium phytate, respectively), suggesting the release of free P. The B. subtilis strain 168 did not solubilize the sodium phytate, except on calcium phytate-amended medium through the formation of a 6.6 mm halo zone around the bacterial colony.
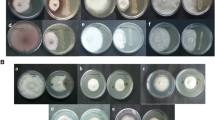
The B. subtilis strain 330-2 solubilized zinc oxide and zinc phosphate, which led to the formation of clear halo zones of 12.26 and 13.5 mm around the bacterial spot on the respective medium compared to B. subtilis strain 168, which did not solubilize zinc oxide and zinc phosphate. Both strains showed positive protease activity by producing clear halo zones of 29.5 mm and 17.8 mm around the spot inoculation. Both strains grew on laminrain azure and cellulose-amended minimal media, except strain B. subtilis strain 168, which did not grow on cellulose-amended medium (Table 1 and Fig. 3).
Production of hydrolytic enzymes and metabolites by B. subtilis strain 330-2 and B. subtilis strain 168. B. subtilis strain 330-2 (a); B. subtilis strain 168 (b); Siderophore production (1); Tri-calcium phosphate (2); Zinc phosphate (3); Zinc oxide (4); Zinc phosphate (5); Ca-phytate (6); Na-phytate (7); Laminarinase (8); Cellulase (9) and Protease production (10). All experiments were conducted three times through three independent trials.
In vitro antagonistic activity assays
The B. subtilis strain 330-2 illustrated the highest antifungal activity against all the phytopathogens listed (Fig. 4). The data on mycelium growth was recorded based on the respective control treatment (4–7 days) and revealed considerable reductions in the growth of fungal mycelia under the influence of B. subtilis strain 330-2. The maximum inhibition (%) in the radial growth by B. subtilis strain 330-2 was 49.53% for C. heterostrophus, followed by 47.96% for A. alternata, 44.57% for B. cinerea, 42.25% for F. oxysporum, 37.74% for N. oryzae, and 36.28% for R. solani. When compared with the controls, each of the treatments showed significant difference (P < 0.05).
Effects of B. subtilis strain 330-2. (a) Antagonistic effects of B. subtilis strain 330-2 against fungal pathogens; C. heterostrophus (1); A. alternate (2); B. cinerea (3); F. oxysporum (4); N. oryzae (5); R. solaniAG1 (6); and with their respective control (CK) on dual culture. (b) % inhibition of B. subtilis strain 330-2 against fungal pathogens. Data were statistically analyzed and the small alphabetical letters (a, b, c…) above the mean bars show the significant differences (P < 0.05) among treatments against different fungal pathogens. Each assay was performed in triplicate.
Plant growth enhanced by B. subtilis strain 330-2
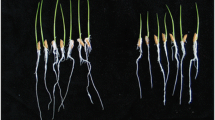
The B. subtilis strain 330-2 was chosen to determine its beneficial effects on rice and maize growth under greenhouse conditions. The performance of the rice and maize shoots were far better than the controls when bio-primed with the B. subtilis strain 330-2 (Fig. 5a–c and g). B. subtilis strain 330-2 was found to enhance the shoot and root length, shoot and root fresh weight and shoot and root dry weight of rice significantly (P < 0.05) by 37.41, 23.90, 39.81, 38.31, 36.79 and 12.21%, respectively, compared with the control. The respective traits for maize were 21.98, 14.19, 41.95, 42.09, 41.66 and 29.15% under the influence B. subtilis strain 330-2 (Fig. 5d–f and h).
Effect of B. subtilis strain 330-2 on the seedling growth performance of rice and maize. (a and d) Seedling elongation of rice and maize, (b and e) seedling fresh weight of rice and maize, (c and f) seedling dry weight of rice and maize, and (g and h) pictorial view of 3-weeks and 40 days old rice and maize seedling in the control and B. subtilis strain 330-2 treatment, respectively. Data were statistically analyzed and the vertical bars above indicate the standard error of three replicates. Small alphabetical letters (a, b…) above the mean bars show the significant differences (P < 0.05) among the treatments within specific parameters. FW: Fresh weight, DW: Dry weight, CK: Control, Tr: Treated with B. subtilis strain 330-2.
Biosafety and plant growth promoting effect of B. subtilis strain 330-2
The addition of B. subtilis strain 330-2 resulted in significantly (P < 0.05) increased plant height, as well as an increase in the fresh and dry weight (Supplementary Fig. 1a–c and d) compared to the non-inoculated plants. The massive inoculation of the B. subtilis strain 330-2 did not have a detrimental effect.
Generation of differentially expressed clones by R-PCR
The sequence identities between the tester and driver were found to be <87% as determined under diversifying differentiation with a specified selection33. Therefore, 87% identity was used as one of the evaluating criteria to determine R-PCR cloned genes. In the recent study, the R-PCR efficiency was estimated to be 92.10% (Supplementary Table 3). To underpin the removal mechanisms of R-PCR at the genomic DNA sequence level, both the tester and driver sequences were compared using MEGA 5 version 5.2 and it was found that, in the three main cases, the driver could not eliminate the tester (Fig. 6 and Supplementary Fig. 3). The sequence identities of the R-PCR cloned genes between tester and driver are presented (Supplementary Table 4).
The driver in R-PCR could not eliminate the tester in three cases: (a) the recognition cutting sites of the MseI and ApeKI (in red box) have mutations, (b) the tester contains enough bases that do not match the driver and (c) the sequence of the tester or driver is partially absent (or completely absent in the driver). *Identical.
Identification and functional categorization of differentially expressed genes in B. subtilis strain 330-2
Gene Ontology (GO) was performed to explore the functions of the differentially expressed genes. In total, 114 differentially expressed genes were assigned to three main functional categories (biological process, cellular component and molecular function) (Supplementary Table 2 and Supplementary Fig. 2a–c). We focused on the groups that were associated with stress tolerance, defense and plant growth enhancement (Fig. 7).
In the present study, five genes that were associated with cell motility (hag, mcpA), biofilm formation and root colonization (srfAA, wapA, spo0A) were differentially expressed under treatment with B. subtilis strain 330-2. B. subtilis strain 330-2, which was observed to regulate the expression of different NRPSs, including surfactin (srfAA), bacillaene (bae) and fengycin (fen). In addition to NRPS, B. subtilis strain 330-2 triggered the production of secondary metabolites, including antibiotics such as macrolactin (mln), difficidin (dfn), iturin A (ituA), bacillibactin (dhbF), penicillin binding protein 2B (pbpB) and beta-lactamase (penP), which are important manifestations of a plant’s defense mechanism and can cope with competing microorganisms and inhibit the growth of phytopathogenic fungi or bacteria.
Different genes associated with tolerance to heat (elongation factor Tu; aspartokinase II; and dihydroorotase, pyrC), salinity (cardiolipin synthase, ywiE; glutaminase-1, ybgJ; phosphoglyceratemutase, gpmI; 4-hydroxy-3-methylbut-2-enyl diphosphate reductase, ispH and glutamate synthase (NADPH/NADH), gltA), drought (cystathionine beta-lyase and cysteine desulfhydrase) and cold stresses (sporulation cortex protein, coxA) were differentially expressed in B. subtilis strain 330-2. Interestingly, NADH-dependent butanol dehydrogenase A (yugJ) and formate dehydrogenase (yjgC) are commonly involved in tolerance to heat, salinity and oxidative stresses. Likewise, glutamate symporter (gltT) and S-adenosylmethionine synthase (metK) are commonly involved in salinity and drought stress (Fig. 8). These results clearly suggest the beneficial role of B. subtilis strain 330-2 in enhancing the plant’s tolerance to different abiotic stresses.
Five genes involved in abiotic stress tolerance, e.g., pabA, the precursor of tryptophan (indole-3-acetic acid synthesis); bioA, responsible for biotin biosynthesis; narI, nitrate reductase production; pckA was shown to be involved in nitrogen fixation, ispH is responsible for isoprenoid biosynthesis, were found to be differentially expressed in B. subtilis strain 330-2 were found to be differentially expressed in B. subtilis strain 330-2. All these processes are critical for the normal growth and development of plants.
Four genes involved in cellulose degradation (epsB, ywqC, ydjE etc.), as well as xylan transport and utilization (xynA), were differentially expressed in the B. subtilis strain 330-2 (Supplementary Table 2).
Discussion
The present study was conducted to isolate an antagonistic B. subtilis strain 330-2 that can efficiently control a wide range of phytopathogens. The biochemical, physiological and phylogenetic analyses of the 16S rDNA gene sequences and BIOLOG test confirmed that strain 330-2 was B. subtilis. The BIOLOG micro plate assay has mainly been used for bacterial biocontrol agents in unraveling their specific carbon sources. The isolation of endophytic Bacillus species from various crops has been extensively studied7, 34. B. subtilis strain 330-2 produced significant amounts of IAA (24.13 µg ml−1), which might play an important role in seedling growth and development35. Eighty percent of the soil rhizosphere bacteria can produce IAA, whereas almost all (98%) of the PGPR strains isolated from the plant rhizosphere were able to produce IAA36. Significant amounts of in vitro IAA production by Bacillus spp. have also been documented by Singh et al.37. The production of siderophores by B. subtilis strain 330-2 demonstrated its antagonistic nature against various pathogens along with plant growth regulation. In the present study, B. subtilis strain 330-2 was found to produce phytase and was a good solubilizer of phosphate and zinc38. Phytase has been identified and characterized previously from several gram-positive and gram-negative soil bacteria, such as B. subtilis 39, B. laevolacticus, Pseudomonas sp.40, 41, and Enterobacter sp.42. Extracellular phytase from B. amyloliquefaciens FZB45 enhanced the growth of maize seedlings under in vitro conditions and, in another study, phytase producing bacterial isolates promoted the plant growth of Indian mustard43, 44.
Because of the restricted radial growth of the selected phytopathogens, formation of inhibition zones by B. subtilis strain 330-2 might be linked to the production of antimicrobial compounds such as antibiotics, cellulase, laminarinase, protease, siderophore production, as well as nutrient competition37. Notably, B. subtilis strain 330-2 secreted a complex of hydrolytic enzymes, including β-1,3-glucanase, β-1,4-glucanase and proteases, which possibly degrade the contents of the fungal cell wall, such as β-1,3-glucan and glucosidic bonds. However, rare literature is available on the production of hydrolytic enzymes by microorganisms45, 46. Therefore, it is assumed that the cell wall lysis of the pathogenic fungi is due to the coordinated action of hydrolytic enzymes, such as laminarase, cellulase and protease47. In the present study, B. subtilis strain 330-2 was quite effective against all six of the pathogens tested, indicating its potential efficacy as a biocontrol agent and suggesting its use as a bioinoculant for banded leaf sheath in both rice and maize.
The efficiency of the isolated strain in enhancing plant growth was further assessed in a screen-house experiment. B. subtilis strain 330-2 was found to produce a considerable amount of IAA and significantly enhance the growth of plants compared with the control. For instance, compared with the control, the B. subtilis strain 330-2-treated rice/maize seedlings showed 12–34% enhancements in seedling elongation and biomass accumulation (Fig. 5a–f). It has been stated that the endophytic Bacillus (B. megaterium strain HNSQJYH124, B. subtilis strain HNSQJYH135, and B. atrophaeus strain HNSQJYH170) significantly enhanced root length, shoot/root fresh and dry weight and chlorophyll content in wheat seedlings when inoculated with Bacillus 7. B. subtilis and P. aeruginosa increased the plant growth of African spinach, okra and tomato in terms of shoot length and dry biomass48. More recently, it was also found that the inoculation of plants with PGPB enhanced the root colonization and plant growth4, 49. The mechanisms underlying the promotion of plant growth are diverse, including synthesis of phytohormones and signaling molecules, improvement of the plant mineral nutrition and production of siderophores and volatile compounds50. It has been reported that Bacillus spp. can elicit induced systemic resistance (ISR), which may result in the promotion of plant growth51. It was clear from the results that B. subtilis strain 330-2 is safe for the health of plants without any adverse effects. It is suggested that the B. subtilis strain 330-2 used in this study could be used as a PGPB52.
To assess the differential expression of genes, the most advanced and authentic R-PCR method was used to compare the B. subtilis strain 330-2 with B. subtilis 168. In total, 114 single clones were differentially expressed by R-PCR, suggesting the involvement of many genes in biotic and abiotic stress tolerance, as well as growth enhancement. Approximately 10% of fragments, including polyketide synthases (PKS) and non-ribosomal peptide synthetases (NRPSs), were involved in the synthesis of secondary metabolites and defense mechanisms for the PGPB-induced suppression of plant pathogens (Supplementary Table 2). Biosynthesis of bioactive lipopeptides plays an important role in the interaction of Bacillus species and plants either by suppressing the soil-borne pathogens in the rhizosphere or/and enhancing the plant defense mechanisms. Biosynthesis of bioactive lipopeptides occurs due to various non-ribosomal peptide synthetases (NRPSs) that have antagonistic activities53. Activation of several PKS/NRPS genes related to the production of antibiotics (e.g., srfAA) plays a key role in root colonization by emulsifying, foaming and decreasing the surface tension of solids to assure proper root colonization and a suitable environment that facilitates proliferation54. In a previous study, it has been reported that srfAA, macrolactin polyketide synthase, difficidin synthase and fengycin synthetase were induced in response to the root exudates and were directly linked to the biological control activity of B. amyloliquefaciens FZB4255. Bacteria in the rhizosphere sense the root exudates that are released by the plants via methyl-accepting proteins, which activates motility-related genes (e.g., hag, mcpA) and then move to the root surface for attachment56. In the present study, several genes associated with cell motility and biofilm formation (e.g., epsB, wapA, spo0A) were differentially expressed in B. subtilis strain 330-2. They are suspected to enhance the ability of B. subtilis strain 330-2 to exploit various plant-derived polysaccharides in the rhizosphere. These differentially expressed genes might favor host plant sensing and defense against pathogens in plants treated with B. subtilis strain 330-2. Iron has been reported to play a key role in bacterial biofilm formation57. In this study, the ferrichrome transport system permease protein (fhuG) gene was differentially expressed by strain 330-2 (Supplementary Table 2), which is known to be involved in scavenging iron from environmental sources58.
In the natural environment, B. subtilis is frequently faced with different stress factors, including salinity, oxidative stress, heat stress, and deprivation of nutrients59. These stress factors negatively influence the growth and productivity of living organisms, including plants and bacteria, leading to significant economic losses globally. Thus, it is useful to develop stress-tolerant species and to understand the underlying mechanisms60. Nine genes associated with abiotic stress tolerance (e.g., yuaI, gpmI, glpT, coxA, glcD, yceD, yugJ, ggt, metK) were differentially expressed in B. subtilis strain 330-2. Specifically, approximately 7, 5, 3, 2, and 1 genes related to tolerance against salinity, oxidative, heat, drought cold and oxidative stress, respectively, were differentially expressed in B. subtilis strain 330-2 (Fig. 8). These differentially expressed genes belonged to diverse functional classes, such as photosynthesis, lipid metabolism (G3P), stress/defense carbohydrate metabolism, competition and membrane transport (Supplementary Table 2). In the past, it has been reported that inoculation with PGPR enhanced the drought tolerance via inducing transcription of drought response genes61, affecting the phytohormonal balance62 and sugar accumulation63. Glycerol-3-phosphate (glpT), an important component of carbohydrates as well as lipid metabolic processes, enhanced the basal defense against Colletotrichum higginsianum 64. Likewise, the involvement of yuaI, and gpmI against salinity stress is evident65, 66. In the present study we found that the sporulation cortex protein (coxA) was differentially expressed in the strain 330-2, which is involved in low temperature stress67.
The genes differentially expressed by B. subtilis strain330-2 are involved in oxidative stress (Supplementary Table 2) responses and the activation of different cellular defense mechanisms. These genes primarily include glycolate oxidase (glcD), tellurium resistant (yceD), which is important for tellurite resistance68, putative NADPH-dependent butanol dehydrogenase (yugJ), which acts in direct detoxification of hydrogen peroxide69, and gamma-glutamyltranspeptidase (ggt), for resistance to organic peroxides70. Furthermore, we observed that metK, the precursor of ethylene and polyamine biosynthesis71, was differentially expressed by B. subtilis strain 330-2. This gene has been known to trigger tolerance against salinity72 and drought stresses73. In conclusion, we have presented strong evidence that using the R-PCR approach allowed the identification of a substantial number of resistant genes in biotic/abiotic stresses.
Methods
Strains and growth conditions
The Bacillus subtilis strain 330-2 used in this study was isolated from the rapeseed in Wuhan, China and maintained at −80 °C. The Bacillus subtilis strain 168 and phytopathogenic fungi, Rhizoctonia solani AG1-IA, Fusarium oxysporum, Botrytis cinerea, Alternaria alternata, Cochliobolus heterostrophus, and Nigrospora oryzae, were obtained from the Department of Plant Pathology, Huazhong Agricultural University, Wuhan, 430070, P.R. China. The bacterial strain B. subtilis strain 168 was initially streaked from −80 °C glycerol stocks onto LB medium plates and then was cultured from a fresh single colony into LB broth. The fungal cultures were maintained on potato dextrose agar (PDA) by sub-culturing at regular interval and incubated at 28 °C.
Plant and soil sample collection
Samples of maize (Zea mays L.), wheat (Triticum astevam L.), rice (Oryza sativa L.), wild rapeseed (Brassica napus) and rhizosphere soils were collected from the field of Huazhong Agricultural University, Wuhan, 430070, P.R. China. Briefly, for endophytic isolation plant tissue samples were washed with running tap water to remove adherent soil particles and microbes. Samples were cut into 1–2 cm small pieces aseptically and surface sterilized with 70% ethanol for 1 min, 1.2% NaClO solution for 5 min, followed by 70% ethanol for 1 min. Then, samples were washed three times with sterilized distilled water and surface dried on sterilized filter paper. Surface sterilized samples were ground to a slurry with a sterilized mortar and pestle containing sterilized quartz sand. One gram each of rhizopheric soil, crushed roots, crushed leaves and crushed stem were homogenized separately in 9 ml saline (0.85% NaCl) in 20 ml test tube. The suspensions were homogenized, serial diluted up to 10−5. For each of these dilutions, 0.1 ml was spread on different Luria Bertani (LB) medium plates and then incubated at 28 °C for 4 days. The water from the last rinse was used as a control. Then, the colonies with different morphology were selected and purified by sub-culturing using maximum possible antiseptic condition74. Pure cultures were examined for growth studies.
Isolation and identification of the bacterial strain
The bacterial B. subtilis strain 330-2 used in this experiment was isolated from wild rapeseed. B. subtilis strain 330-2 indicated the highest inhibitory effect in vitro against R. solani and other fungi. Phenotypic profiling of the B. subtilis strain 330-2 was tested using BIOLOG GenIII MicroPlates (Model EL311, BioTek Instruments, USA), containing 71 carbon source utilization assays and 23 chemical sensitivity assays according to the manufacturer’s protocol. The 10 μl microbial suspension was inoculated into each well of the GENIII MicroPlates. The plates were incubated at 30 °C for 24 to 48 h and observed for color development at 12 h intervals. Data were recorded at a 590 nm wavelength and the color development in the plates was read and compared in the BIOLOG database (Microlog SystemTm, Release 4.0) to identify the strain to species level75. The 16S rDNA gene of the strain was amplified using universal primer, both forward 27F (5′-AGAGTTTGATCCTGGCTCAG-3′) and reverse 1492R (5′-GGTTACCTTGTTACGACTT-3′)76. These primers were synthesized by T-Singke, Wuhan, China. The total volume (50 μl) of the PCR reaction mixture contained 5 μl of 10 × Taq buffer, 1.5 μl of MgCl2 (25 mmol L−1), 4 μl of each dNTPs (10 mmol L−1), 2 μl of each primer (30 pmol), 1 μl of DNA template, 0.25 μl of Taq DNA polymerase, and 35.75 μl of ddH2O. The PCR reaction was performed using MyCycler Thermal Cycler (Bio-RAD, Foster, California, USA) with the following thermal program: 95 °C for 5 min, followed by 94 °C for 30 s, 50 °C for 30 s, and 72 °C for 1 min 30 s. These steps were repeated for 30 cycles, and a final 10 min extension at 72 °C. The molecular size of the resulting PCR product was analyzed on a 1.0% agarose gel to confirm that the fragment was 1.5 kb, and the amplified 16S rDNA fragment was sequenced by T-Singke, Wuhan, China. The 16S rDNA sequence results were analyzed and compared on the website, http://www.ncbi.nlm.nih.gov 77.
Indole-3-acetic acid (IAA) production detection
Detection of IAA production in the B. subtilis strain 330-2 and B. subtilis strain 168 were determined by the method reported by Brick et al.78. The strains were grown on LB broth and incubated at 28 °C for 48 h. Well-grown cultures were centrifuged at 10,000 rpm for 15 min at 4 °C to collect the supernatant. The supernatant (2 ml) was mixed with two drops of O-phosphoric acid and 4 ml of Salkowski reagent (50 ml, 35% of perchloric acid, 1 ml 0.5 M FeCl3 solution). The appearance of a pink color in the supernatant confirmed the production of IAA. After 30 min, the optical density (OD) of the mixture was read at an absorbance of 530 nm in a NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific Inc., Wilmington, USA). The amount of IAA was estimated using a standard curve.
Siderophore production detection
Siderophore production of the strains was determined on blue agar plates containing Chrome-azurol S (CAS) medium following Schwyn and Neilands’ method79. The bacterial strains (24 h-old culture) were spotted on CAS medium and incubated at 28 °C for 4 days. Formation of an orange halo around the colony was the indicator of siderophore production.
Phosphate solubilization
Phosphate solubilization of the isolated strain 330-2 and B. subtilis strain 168 was detected by spotting these strains separately on Pikovskaya’s (PVK) agar plates using zinc phosphate (ZP), di-calcium phosphate (DCP), and tri-calcium phosphate (TCP) as sources of insoluble inorganic phosphate. Plates were incubated at 28 °C for 7 days80. The halo zone and colony diameter were measured and data were recorded. The phosphate solubilization index (S.I.) was calculated using the equation: S.I. = (colony diameter + halo zone diameter)/colony diameter81.
Phytase production detection
Phytase production of the strains was evaluated by spot inoculation of log phase culture on phytase screening media using sodium and calcium phytate as sources of insoluble organic phosphate. The plates were incubated at 28 °C for 7 days. The halo zone around the colonies was observed on the plates.
Zinc solubilization
To evaluate zinc solubilization abilities, the log phase culture of the bacterial strains was spotted on Tris-minimal medium plates containing zinc phosphate and zinc oxide as sources of insoluble inorganic zinc. The inoculated plates were then incubated at 28 °C for 7 days. Clearing zones around the colonies were observed for zinc solubilization.
Cell wall degrading enzymes
β-1,3-glucanase and β-1,4-glucanase activities were studied using minimal medium (MM) containing laminarin azure (Sigma–Aldrich Co., USA) and cellulose powder as sole sources of carbon, respectively. Laminarin azure and cellulose amended minimal medium indicate the production of β-1,3-glucanase and β-1,4-glucanase, respectively. The medium plates were incubated at 28 °C for 7 days. Protease production was assessed by spotting each of the bacterial strains on Skim Milk Agar (SMA) medium plates. The bacterial strains that produced protease were identified by halo zones around the bacterial colonies82.
In vitro antagonistic activity assays
The strain was tested for its ability to inhibit the growth of R. solani AG1-IA, B. cinerea, F. oxysporum, A. alternata, C. heterostrophus, and N. oryzae in vitro. The B. subtilis strain 330-2 was dabbed at two corners 3 cm from the center by stabbing the plate with sterilized tooth pick. After 24 h, fresh mycelium of test fungi (0.8-cm diameter) was placed in the center of the PDA plates. All treatments were conducted in triplicate. Plates were incubated at 28 °C. The inhibition of fungal growth was monitored by recording the diameter of the inhibition zone (mm).The percentage of inhibition was calculated using the following formula83.
where “C” is the mycelium diameter (cm) of the fungus growing on the control plates (without bacteria) and “T” is the mycelium diameter (cm) of the fungus growing in the bacterial-treated plates. The experiment was repeated thrice with three biological replicates each time.
Plant growth assay
B. subtilis strain 330-2 was grown in 100 ml LB medium in a rotary shaker (170 rpm) at 37 °C, until they reached an optical density at 600 nm (OD600) of 0.8. The bacterial cells were harvested by centrifugation at 6000 × g for 5 min at 4 °C and resuspended in 100 ml of distilled water and adjusted to (1010 CFU ml−1). Seeds of rice (Oryza sativa, L.) (variety: Huanghuazhan) and corn (Zea mays L.) (variety: B73) were obtained from the College of Plant Science and Technology of Huazhong Agricultural University, Wuhan, China. The seeds were surface sterilized with 70% alcohol for 1 min, followed by 0.5% NaClO solution for 1.30 min and 70% alcohol for 1 min. The seeds were then washed three times with distilled water. The sterilized seeds were bio-primed with bacterial culture for 4 h (control seeds were treated with distilled water). Seeds were then placed in the sterile petri dishes and dried back to their original moisture contents at room temperature. After bio-priming, seed germination tests were carried out. Ten rice seeds and three maize seeds for each treatment were placed in plastic pots of 7 cm diameter and 15 cm diameter, respectively and were filled with sterilized soil. Each treatment was replicated three times, respectively. Then, pots were incubated in a growth chamber at 28 °C with a photoperiod of 16 h light and 8 h dark. Data were calculated for shoot length, root length, fresh weight and dry weight.
B. subtilis strain 330-2 effects in tobacco plants
The addition of the PGPB, which aimed to trigger plant growth, should not possess any detrimental effect on other plant species. B. subtilis strain 330-2 was tested in tobacco plants as a well-studied model and widely cultivated crop species84. Tobacco seeds were surface sterilized and grown in autoclaved soil in plastic pots (7 cm diameter). When the seedling shoots reached 1 cm, they were inoculated with a bacterial suspension (108–109 CFU ml−1). Four seedlings per condition were sampled on 7, 14, and 21 days, and data were recorded on plant height, fresh weight, and dry weight post inoculation. M9 buffer without bacterial inoculum was used as a negative control.
Generation of differentially expressed clones by R-PCR
The removing PCR (R-PCR) is a restriction enzyme-based method, which eliminates unwanted genes from the gene population very efficiently. Using this method, the undesirable genes can be removed by means of removing drivers. This recently developed R-PCR technique was used to isolate the differentially expressed antagonism-related genes by comparing genomic DNA populations of the B. subtilis strain 330-2 and B. subtilis strain 168. To identify specific genes in B. subtilis strain 330-2, the B. subtilis strain 330-2 and B. subtilis strain 168 were taken as a tester and driver, respectively. The B. subtilis strain 168 is a well-known model strain without roles in enhancing plant growth and tolerance to biotic/abiotic stresses. The R-PCR was performed between the ‘driver’ and ‘tester’ according to the previous method of Huan et al.33. All PCR steps were performed on MyCycler Thermal Cycler (Bio-RAD, Foster, California, USA). Bands of the second R-PCR were excised from the gel, and PCR products were purified using the QIAquick gel purification kit (QIAGEN, Germany). The purified PCR products were cloned in pmD18-T Easy vector following manufacturer’s protocol (Promega) and transformed into DH5α cells. Plates were incubated at 37 °C until small colonies were visible. Sequencing of the positive clones were done by T-Singke, Wuhan, China using primers both forward M13F (−47) (5′-CGCCAGGGTTTTCCCAGTCACGAC-3′) and reverse M13R (−48) (5′-AGCGGATAACAATTTCACACAGGA-3′).
Sequences analysis
A total of 180 white-clones were picked from the library. The inserted fragments were amplified, and fragments of the differentially expressed clones were sequenced using the primer pair of M13F and M13R. Sequences were trimmed of vector using DNAMAN software. Homology searches of all sequences were queried in the GenBank database with nucleotide blasts available at the NCBI (http://www.ncbi.nlm.nih.gov/BLAST).
Statistical analysis
Data were analyzed by one way analysis of variance (ANOVA) using STATISTIX software (8.1) (Analytical Software, Tallahassee, FL, USA). The mean variance was analyzed using a least significant difference (LSD) test at 0.05 probability level.
References
Bashan, Y. & Holguin, G. Proposal for the division of plant growth-promoting rhizobacteria into two classifications: biocontrol-PGPB (plant growth-promoting bacteria) and PGPB. Soil Biol. Biochem. 30, 1225–1228, doi:10.1016/S0038-0717(97)00187-9 (1998).
Compant, S., Clément, C. & Sessitsch, A. Plant growth-promoting bacteria in the rhizo- and endosphere of plants: Their role, colonization, mechanisms involved and prospects for utilization. Soil Biol. Biochem. 42, 669–678, doi:10.1016/j.soilbio.2009.11.024 (2010).
Timm, C. M. et al. Metabolic functions of Pseudomonas fluorescens strains from Populus deltoides depend on rhizosphere or endosphere isolation compartment. Front. Microbiol. 6, 1118, doi:10.3389/fmicb.2015.01118 (2015).
deSouza, R., Ambrosini, A. & Passaglia, L. M. P. Plant growth-promoting bacteria as inoculants in agricultural soils. Genet. Mol. Biol. 38, 401–419, doi:10.1590/S1415-475738420150053 (2015).
Yao, A. V. et al. Effect of FZB 24® Bacillus subtilis as a biofertilizer on cotton yields in field tests. Arch. Phytopathol. Plant Pro. 39, 323–328, doi:10.1080/03235400600655347 (2006).
Geetha, R., Falguni, S., Anjana, J. D. & Archana, G. Enhanced growth and nodulation of pigeon pea by co-inoculation of Bacillus strains with Rhizobium spp. Biores. Technol. 99, 4544–4550, doi:10.1016/j.biortech.2007.06.057 (2008).
Zhao, L. et al. Screening and characterization of endophytic Bacillus and Paenibacillus strains from medicinal plant Lonicera japonica for use as potential plant growth promoters. Braz. J. Microbiol. 46, 977–989, doi:10.1590/S1517-838246420140024 (2015).
Zhao, L. et al. Identification and characterization of the endophytic plant growth prompter Bacillus cereus strain mq23 isolated from Sophora alopecuroides root nodules. Braz. J. Microbiol. 42, 567–575, doi:10.1590/S1517-838220110002000022 (2011).
Zahid, M., Abbasi, M. K., Hameed, S. & Rahim, N. Isolation and identification of indigenous plant growth promoting rhizobacteria from Himalayan region of Kashmir and their effect on improving growth and nutrient contents of maize (Zea mays L.). Front. Microbiol. 6, 207, doi:10.3389/fmicb.2015.00207 (2015).
Saraf, M., Pandya, U. & Thakkar, A. Role of allelochemicals in plant growth promoting rhizobacteria for biocontrol of phytopathogens. Microbiol. Res. 169, 18–29, doi:10.1016/j.micres.2013.08.009 (2014).
Kumar, P., Dubey, R. C. & Maheshwari, D. K. Bacillus strains isolated from rhizosphere showed plant growth promoting and antagonistic activity against phytopathogens. Microbiogical. Res. 167, 493–499, doi:10.1016/j.micres.2012.05.002 (2012).
Shafi, J., Tian, H. & Ji, M. Bacillus species as versatile weapons for plant pathogens: a review. Biotech. Biotechnol. Equip. 31, 1–14, doi:10.1080/13102818.2017.1286950 (2017).
Chaurasia, B. et al. Diffusible and volatile compounds produced by an antagonistic Bacillus subtilis strain cause structural deformations in pathogenic fungi in vitro. Microbiol. Res. 160, 75–81, doi:10.1016/j.micres.2004.09.0,13 (2005).
Gond, S. K., Bergen, M. S., Torres, M. S. & White, J. F. Jr. Endophytic Bacillus spp. produce antifungal lipopeptides and induce host defence gene expression in maize. Microbiol. Res. 172, 79–87, doi:10.1016/j.micres.2014.11.004 (2015).
Yuan, J., Raza, W., Shen, Q. & Huang, Q. Antifungal Activity of Bacillus amyloliquefaciens NJN-6 Volatile Compounds against Fusarium oxysporum f. sp. cubense. Appl. Environ. Microbiol. 78, 5942–5944, doi:10.1128/AEM. 01357-12 (2012).
Chowdhury, P. S., Hartmann, A., Gao, X. & Borriss, R. Biocontrol mechanism by root-associated Bacillus amyloliquefaciens FZB42-a review. Front. Microbiol. 6, 780, doi:10.3389/fmicb.2015.00780 (2015).
García, G., Portal, V., Onco, M. A. & Rubio, V. S. Review. Biology and Systematics of the form genus Rhizoctonia. Spanish J. Agric. Res. 4, 55–79, doi:10.5424/sjar/2006041-178 (2006).
Tsror, L. Biology, epidemiology and management of Rhizoctonia solani on potato. J. Phytopathol. 158, 649–658, doi:10.1111/j.1439-0434.2010.01671.x (2010).
AbuQamar, S. F., Moustafa, K. & Tran, L. P. Omics’ and Plant Responses to Botrytis cinerea. Front. Plant Sci. 7, 1658, doi:10.3389/fpls.2016.01658 (2016).
Munkvold, G. P. Epidemiology of Fusarium diseases and their mycotoxins in maize ears. Epi. Mycotoxin Producing Fungi. 27, 705–713, doi:10.1007/978-94-017-1452-5_5 (2003).
Dai, Y. L. et al. First Report of Southern Leaf Blight Caused by Cochliobolus heterostrophus on Corn (Zea mays) in Fujian Province, China. Plant Disease 100, 1781, doi:10.1094/PDIS-12-15-1490-PDN (2016).
Mew, T.W. & Gonzales, P. A handbook of rice seedborne fungi. Ed. International Rice Research Institute (IRRI) Los Baños, Philipinas. 83 (2002).
Ismaiel, A. A. & Papenbrock, J. Mycotoxins: Producing Fungi and Mechanisms of Phytotoxicity. Agric. 5, 492–537, doi:10.3390/agriculture5030492 (2015).
Li, H., Gu, X., Dawson, V. L. & Dawson, T. M. Identification of calcium- and nitric oxide-regulated genes by differential analysis of library expression (DAzLE). PNAS 101, 647–652, doi:10.1073/pnas.0305145101 (2004).
Lisitsyn, N., Lisitsyn, N. & Wigler, M. Cloning the differences between two complex genomes. Science 259, 946–951, doi:10.1126/science.8438152 (1993).
Sokolov, B. P. & Prockop, D. J. A rapid and simple PCR-based method for isolation of cDNAs from differentially expressed genes. Nucl. Acids Res. 22, 4009–4015, doi:10.1093/nar/22.19.4009 (1994).
Zeng, J., Gorski, R. A. & Hamer, D. Differential cDNA cloning by enzymatic degrading subtraction (EDS). Nucl. Acids Res. 22, 4381–4385, doi:10.1093/nar/22.21.4381 (1994).
Deleersnijder, W. et al. Isolation of markers for chondro-osteogenic differentiation using cDNA library subtraction. Molecular cloning and characterization of a gene belonging to a novel multigene family of integral membrane proteins. J. Biol. Chem. 271, 19475–19482, doi:10.1074/jbc.271.32.19475 (1996).
Yang, M. & Sytkowski, A. J. Cloning differentially expressed genes by linker capture subtraction. Anal. Biochem. 237, 109–114, doi:10.1006/abio.1996.0207 (1996).
Diatchenko, L. et al. Suppression subtractive hybridization: a method for generating differentially regulated or tissue-specific cDNA probes and libraries. PNAS 93, 6025–6030, doi:10.1073/pnas.93.12.6025 (1996).
Goetz, F. W. The “ups” and “downs” in Using Subtractive Cloning Techniques to Isolate Regulated Genes in Fish. Integ. Comp. Biol. 43, 786–793, doi:10.1093/icb/43.6.786 (2003).
Baginsky, S., Hennig, L., Zimmermann, P. & Gruissem, W. Gene expression analysis, proteomics, and network discovery. Plant Physiol. 152, 402–410, doi:10.1104/pp.109.150433 (2010).
Huan, J., Wan, K., Liu, Y., Dong, W. & Wang, G. Removing PCR for the elimination of undesired DNA fragments cycle by cycle. Sci. Rep. 3, 2303, doi:10.1038/srep02303 (2013).
Castro, R. A. et al. Isolation and enzyme bioprospection of endophytic bacteria associated with plants of Brazilian mangrove ecosystem. Springer Plus 3, 382, doi:10.1186/2193-1801-3-382 (2014).
Seo, P. J. & Park, C. M. Auxin homeostasis during lateral root development under drought condition. Plant Signal Behav. 4, 1002–1004, doi:10.4161/psb.4.10.9716 (2009).
Arruda, L. et al. Screening of rhizobacteria from maize (Zea mays L.) in Rio Grande do Sul State (South Brazil) and analysis of their potential to improve plant growth. Appl. Soil Ecol. 63, 15–22, doi:10.1016/j.apsoil.2012.09.001 (2013).
Singh, N., Pandey, P., Dubey, R. C. & Maheshwari, D. K. Biological control of root rot fungus Macrophomina phaseolina and growth enhancement of Pinus roxburghii (Sarg.) by rhizosphere competent Bacillus subtilis BN1. World J. Microbiol. Biotechnol. 24, 1669–1679, doi:10.1007/s11274-008-9680-z (2008).
Iqbal, U., Jamil, N., Ali, I. & Hasnain, S. Effect of zinc-phosphate-solubilizing bacterial isolates on growth of Vigna radiata. Ann. Microbiol. 60, 243–248, doi:10.1007/s13213-010-0033-4 (2010).
Kerovuo, J., Lauraeus, M., Nurminen, P., Kalkkinen, N. & Apajalahti, J. Isolation, characterization, molecular gene cloning, and sequencing of a novel phytase from Bacillus subtilis. Appl. Environ. Microbiol. 64, 2079–2085 (1998).
Gulati, H. K., Chadha, B. S. & Saini, H. S. Production and characterization of thermostable alkaline phytase from Bacillus laevolacticus isolated from rhizosphere soil. J. Ind. Microbiol. Biot. 34, 91–98, doi:10.1007/s10295-006-0171-7 (2007).
Richardson, A. E. & Hadobas, P. A. Soil isolates of Pseudomonas spp. that utilize inositol phosphates. Can. J. Microbiol. 43, 509–516, doi:10.1139/m97-073 (1997).
Yoon, S. J. et al. Isolation and identification of phytase producing bacterium, Enterobacter sp. 4, and enzymatic properties of phytase enzyme. Enzyme Microb. Technol. 18, 449–454, doi:10.1016/0141-0229(95)00131-X (1996).
Idriss, E. E. et al. Extracellular phytase activity of Bacillus amyloliquefaciens FZB45 contributes to its plant-growth-promoting effect. Microbiol 148, 2097–2109, doi:10.1099/00221287-148-7-2.97 (2002).
Kumar, V. et al. Isolation of phytase-producing bacteria from Himalayan soils and their effect on growth and phosphorus uptake of Indian mustard (Brassica juncea). World J. Microbiol. Biotechnol. 29, 1361–9, doi:10.1007/s11274-013-1299-z (2013).
Gruber, S. & Seidl-Seiboth, V. Self versus non-self: fungal cell wall degradation in. Trichoderma. Microbiol. 158, 26–34, doi:10.1099/mic.0.052613-0 (2012).
Gupta, C. P., Kumar, B., Dubey, R. C. & Maheshwari, D. K. Chitinase mediated destructive antagonistic potential of Pseudomonas aeruginosa GRC against Sclerotinia sclerotiorum causing charcoal rot of peanut. Bio. Control 51, 821–835, doi:10.1007/s10526-006-9000-1 (2006).
Chaiharn, M., Chunhaleuchanon, S., Kozo, A. & Lumyong, S. Screening of rhizobacteria for their plant growth promoting activities. KMITL Sci. Tech. J. 8, 18–23 (2008).
Adesemoye, A. O., Obini, M. & Ugoji, E. O. Comparison of plant growth-promotion with Pseudomonas aeruginosa and Bacillus subtilis in three vegetables. Braz. J. Microbiol. 39, 423–426, doi:10.1590/S1517-83822008000300003 (2008).
Santoyo, G., Moreno-Hagelsieb, G., Orozco-Mosqueda, M. D. & Glick, B. R. Plant growth-promoting bacterial endophytes. Microbiol. Res. 183, 92–99, doi:10.1016/j.micres.2015.11.008 (2016).
Maksimov, I. V., Abizgildina, R. R. & Pusenkova, L. I. Plant growth promoting rhizobacteria as alternative to chemical crop protectors from pathogens (review). Appl. Biochem. Microbiol. 47, 333–345, doi:10.1134/S0003683811040090 (2011).
Kloepper, J. W., Ryu, C. M. & Zhang, S. Induced Systemic Resistance and Promotion of Plant Growth by Bacillus spp. Phytopathol. 94, 1259–1266, doi:10.1094/PHYTO.2004.94.11.1259 (2004).
Vílchez, J. I., Navas, A., González-López, J., Arcos, S. C. & Manzanera, M. Biosafety Test for Plant Growth-Promoting Bacteria: Proposed Environmental and Human Safety Index (EHSI) Protocol. Front. Microbiol. 6, 1514, doi:10.3389/fmicb.2015.01514 (2016).
Ongena, M. & Jacques, P. Bacillus lipopeptides: versatile weapons for plant disease biocontrol. Trends Microbiol. 16, 115–125, doi:10.1016/j.tim.2007.12.009 (2007).
Niazi, A. et al. Genome Analysis of Bacillus amyloliquefaciens subsp. plantarum UCMB5113: A Rhizobacterium That Improves Plant Growth and Stress Management. PLoS One 9, e104651, doi:10.1371/journal.pone.0104651 (2014).
Fan, B. et al. Transcriptomic profiling of Bacillus amyloliquefaciens FZB42 in response to maize root exudates. BMC Microbiol. 12, 116, doi:10.1186/1471-2180-12-116 (2012).
deWeert, S. et al. Flagella-driven chemotaxis towards exudate components is an important trait for tomato root colonization by Pseudomonas fluorescens. Mol. Plant Microbiol. Int. 15, 1173–1180, doi:10.1094/MPMI.2002.15.11.1173 (2002).
Avidan, O., Satanower, S. & Banin, E. Iron and bacterial biofilm development. Microbial. Mats 14, 359–383, doi:10.1128/JB.01601-09 (2010).
Hantke, K. Iron and metal regulation in bacteria. Curr. Opin. Microbiol. 4, 172–177, doi:10.1016/S1369-5274(00)00184-3 (2001).
Chaves, M. M. & Oliveira, M. M. Mechanisms underlying plant resilience to water deficits: prospects for water-saving agriculture. J. Exp. Bot. 55, 2365–2384, doi:10.1093/jxb/erh269 (2004).
Das, P. et al. Salt stress tolerant genes in halophilic and halotolerant bacteria: Paradigm for salt stress adaptation and osmoprotection. Int. J. Curr. Microbiol. App. Sci. 4, 642–658 (2015).
Sarma, R. K. & Saikia, R. Alleviation of drought stress in mung bean by strain Pseudomonas aeruginosa GGRJ21. Plant and Soil 377, 111–126, doi:10.1007/s11104-013-1981-9 (2014).
Figueiredo, M. V. B., Burity, H. A., Martínez, C. R. & Chanway, C. P. Alleviation of drought stress in the common bean (Phaseolus vulgaris L.) by co-inoculation with Paenibacillus polymyxa and Rhizobium tropici. Appl. Soil Ecol. 40, 182–188, doi:10.1016/j.apsoil.2008.04.005 (2008).
Sandhya, V., Ali, S. Z., Grover, M., Reddy, G. & Venkateswarlu, B. Effect of plant growth promoting Pseudomonas spp. on compatible solutes, antioxidant status and plant growth of maize under drought stress. Plant Growth Regul. 62, 21–30, doi:10.1007/s10725-010-9479-4 (2010).
Chanda, B. et al. Glycerol-3-Phosphate Levels Are Associated with Basal Resistance to the Hemibiotrophic Fungus Colletotrichum higginsianum in Arabidopsis. Plant Physiol. 147, 2017–2029, doi:10.1104/pp.108.121335 (2008).
Hahne, H. et al. Comprehensive Proteomics and Transcriptomics Analysis of Bacillus subtilis Salt Stress Adaptation. J. Bacteriol. 192, 870–882, doi:10.1128/JB.01106-09 (2010).
Wang, L. et al. Proteomic analysis of salt-responsive proteins in the leaves of mangrove Kandeliacandel during short-term stress. PLoS ONE 9, e83141, doi:10.1371/journal.pone.0083141 (2014).
Budde, I., Steil, L., Scharf, C., Völke, U. & Bremer, E. Adaptation of Bacillus subtilis to growth at low temperature: a combined transcriptomic and proteomic appraisal. Microbiol. 152, 831–853, doi:10.1099/mic.0.28530-0 (2006).
Rojas, C. M. et al. Glycolate oxidase modulates reactive oxygen species-mediated signal transduction during nonhost resistance in Nicotianabenthamiana and Arabidopsis. Plant Cell 24, 336–352, doi:10.1105/tpc.111.093245 (2012).
Chae, H. Z., Chung, S. J. & Rhee, S. G. Thioredoxin-dependent peroxide reductase from yeast. J. Biol. Chem. 269, 27670–27678 (1994).
Ghelardi, E. et al. Contribution of surfactin and SwrA to flagellin expression, swimming, and surface motility in Bacillus subtilis. Appl. Environ. Microbiol. 78, 6540–6544, doi:10.1128/AEM.01341-12 (2012).
Quan, J. et al. Molecular Cloning, Characterization and Expression Analysis of the SAMS Gene during Adventitious Root Development in IBA-Induced Tetraploid Black Locust. PLoS One 9, e108709, doi:10.1371/journal.pone.0108709 (2014).
Espartero, J., Pintor-Toro, J. A. & Pardo, J. M. Differential accumulation of S-adenosylmethionine synthetase transcripts in response to salt stress. Plant Mol. Biol. 25, 217–227, doi:10.1007/BF00023239 (1994).
Mayne, M. B., Coleman, J. R. & Blumwald, E. Differential expression during drought conditioning of a root-specific S-adenosylmethionine synthetase from jack pine (Pinusbanksiana Lamb.) seedlings. Plant Cell Environ. 19, 958–966, doi:10.1111/j.1365-3040.1996.tb00460.x (1996).
Paynter, Q. Evaluating the impact of a biological control agent Carmenta mimosa on the woody wetland weed Mimosa pigra in Australia. J. Appl. Ecol. 42, 1054–1062, doi:10.1111/j.1365-2664.2005.01105.x (2005).
Kumar, K., Natarajan, A., Someshwar, B., Kutthum, M. & Ramesh, C. S. Isolation and characterization of rhizobacteria associated with coastal agricultural ecosystem of rhizosphere soils of cultivated vegetable crops. World J. Microbiol. Biotechnol. 27, 1625–1632, doi:10.1007/s11274-010-0616-z (2011).
Reddy, G. S., Aggarwal, R. K. & Matsumoto, G. I. & Shivaji, S. Arthrobacter flavus sp. nov., a psychrophilic bacterium isolated from a pond in McMurdo Dry Valley, Antarctica. Int. J. Syst. Evol. Microbiol. 50, 1553–1561, doi:10.1099/00207713-50-4-1553 (2000).
Mignard, S. & Flandrois, J. P. 16S rRNA sequencing in routine bacterial identification: a 30-month experiment. J. Microbiol. Methods 67, 574–581, doi:10.1016/j.mimet.2006.05.009 (2006).
Brick, J. M., Bostock, R. M. & Silverstone, S. E. Rapid in situ assay for indole acetic acid production by bacteria immobilized on a nitrocellulose membrane. Appl. Environ. Microbiol. 57, 535–538 (1991).
Schwyn, B. & Neilands, J. Universal chemical assay for the detection and determination of siderophores. Anal. Biochem. 160, 47–56, doi:10.1016/0003-2697(87)90612-9 (1987).
Pikovskaya, R. I. Mobilization of phosphorus and soil in connection with the vital activity of some microbial species. Mikrobiologii 17, 362–370 (1948).
Premono, E. M., Moawad, M. A. & Vlek, P. L. G. Effect of phosphate solubilizing Pseudomonas putida on the growth of maize and its survival in the rhizosphere. Indo. J. Crop Sci. 11, 13–23 (1996).
Kumar, R. S. et al. Characterization of Antifungal Metabolite Produced by a New Strain Pseudomonas aeruginosa pupa3 that Exhibits Broad Spectrum Antifungal Activity and Biofertilizing Traits. J. Appl. Microbiol. 98, 145–154, doi:10.1111/j.1365-2672.2004.02435.x (2005).
Shakeel, M., Rais, A., Hassan, M. N. & Hafeez, F. Y. Root Associated Bacillus sp. Improves Growth, Yield and Zinc Translocation for Basmati Rice (Oryza sativa) Varieties. Front. Microbiol. 6, 1286, doi:10.3389/fmicb.2015.01286 (2015).
Mayak, S., Tirosh, T. & Glick, B. R. Stimulation of the growth of tomato, pepper and mung bean plants by the plant growth-promoting bacterium Enterobacter cloacae CAL3. Biol. Agric. Hort. 19, 261–274, doi:10.1080/01448765.2001.9754929 (2001).
Acknowledgements
We would like to thank Drs. Guoying Wang and Yunjun Liu for comments and suggestions on the experimental design and data analysis. This work was supported by the National Major Project for Transgenic Organism Breeding (2011ZX08003-001 and 2016ZX08003-001) and the Hubei Provincial Technology Innovation Program (2016ABA093).
Author information
Authors and Affiliations
Contributions
Z.A. carried out most of the experiments described in this paper. J.W. and L.C. isolated the strain. Z.A., J.W., L.C. and W.D. analyzed the data. Z.A. and W.D. designed the experiments and wrote the paper.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ahmad, Z., Wu, J., Chen, L. et al. Isolated Bacillus subtilis strain 330-2 and its antagonistic genes identified by the removing PCR. Sci Rep 7, 1777 (2017). https://doi.org/10.1038/s41598-017-01940-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-01940-9
- Springer Nature Limited
This article is cited by
-
Combined application of microbial inoculant and kelp-soaking wastewater promotes wheat seedlings growth and improves structural diversity of rhizosphere microbial community
Scientific Reports (2023)
-
Isolation and evaluation of Qatari soil rhizobacteria for antagonistic potential against phytopathogens and growth promotion in tomato plants
Scientific Reports (2023)
-
Control of Magnaporthe oryzae and Rice Growth Promotion by Bacillus subtilis JN005
Journal of Plant Growth Regulation (2022)