Abstract
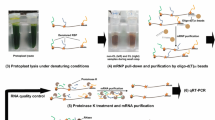
Despite crucial roles of RNA-binding proteins (RBPs) in plant physiology and development, methods for determining their transcriptome-wide binding landscape are less developed than those used in other model organisms. Cross-linking and immunoprecipitation (CLIP) methods (based on UV-mediated generation of covalent bonds between RNAs and cognate RBPs in vivo, purification of the cross-linked complexes and identification of the co-purified RNAs by high-throughput sequencing) have been applied mainly in mammalian cells growing in monolayers or in translucent tissue. We have developed plant iCLIP2, an efficient protocol for performing individual-nucleotide-resolution CLIP (iCLIP) in plants, tailored to overcome the experimental hurdles posed by plant tissue. We optimized the UV dosage to efficiently cross-link RNA and proteins in plants and expressed epitope-tagged RBPs under the control of their native promoters in loss-of-function mutants. We select epitopes for which nanobodies are available, allowing stringent conditions for immunopurification of the RNA–protein complexes to be established. To overcome the inherently high RNase content of plant cells, RNase inhibitors are added and the limited RNA fragmentation step is modified. We combine the optimized isolation of RBP-bound RNAs with iCLIP2, a streamlined protocol that greatly enhances the efficiency of library preparation for high-throughput sequencing. Plant researchers with experience in molecular biology and handling of RNA can complete this iCLIP2 protocol in ~5 d. Finally, we describe a bioinformatics workflow to determine targets of Arabidopsis RBPs from iCLIP data, covering all steps from downloading sequencing reads to identifying cross-linking events (https://github.com/malewins/Plant-iCLIPseq), and present the R/Bioconductor package BindingSiteFinder to extract reproducible binding sites (https://bioconductor.org/packages/release/bioc/html/BindingSiteFinder.html).
Key points
-
As in mammals, RBPs in plants are key regulators of the RNA life cycle. This protocol describes an optimized plant iCLIP method to define the transcriptome-wide RBP binding landscape at single-nucleotide resolution.
-
This plant iCLIP protocol, entailing UV cross-linking and nanobody-mediated precipitation of tagged RBP–RNA complexes, is optimized for efficient library preparation. A streamlined bioinformatics pipeline is also provided for the identification of RBP binding sites.














Similar content being viewed by others
Data availability
The AtGRP7 iCLIP dataset processed in the bioinformatics workflow is available through the Gene Expression Omnibus series GSE99427 or directly via SRA by the accession number SRR24391474. Source data are provided with this paper.
Code availability
All necessary code is publicly available under the Massachusetts Institute of Technology license. The R package BindingSiteFinder is accessible via the Bioconductor repository at https://bioconductor.org/packages/release/bioc/html/BindingSiteFinder.html. The scripts described in this publication are accessible via GitHub at https://github.com/malewins/Plant-iCLIPseq.
References
Singh, G., Pratt, G., Yeo, G. W. & Moore, M. J. The clothes make the mRNA: past and present trends in mRNP fashion. Annu. Rev. Biochem. 84, 325–354 (2015).
Hornyik, C., Terzi, L. C. & Simpson, G. G. The spen family protein FPA controls alternative cleavage and polyadenylation of RNA. Dev. Cell 18, 203–213 (2010).
Macknight, R. et al. FCA, a gene controlling flowering time in Arabidopsis, encodes a protein containing RNA-binding domains. Cell 89, 737–745 (1997).
Sugliani, M., Brambilla, V., Clerkx, E. J., Koornneef, M. & Soppe, W. J. The conserved splicing factor SUA controls alternative splicing of the developmental regulator ABI3 in Arabidopsis. Plant Cell 22, 1936–1946 (2010).
Carvalho, R. F., Carvalho, S. D. & Duque, P. The plant-specific SR45 protein negatively regulates glucose and ABA signaling during early seedling development in Arabidopsis. Plant Physiol. 154, 772–783 (2010).
Zhang, X.-N. & Mount, S. M. Two alternatively spliced isoforms of the Arabidopsis thaliana SR45 protein have distinct roles during normal plant development. Plant Physiol. 150, 1450–1458 (2009).
König, J. et al. iCLIP reveals the function of hnRNP particles in splicing at individual nucleotide resolution. Nat. Struct. Mol. Biol. 17, 909–915 (2010).
König, J., Zarnack, K., Luscombe, N. M. & Ule, J. Protein–RNA interactions: new genomic technologies and perspectives. Nat. Rev. Genet. 13, 77–83 (2011).
Wang, Z. et al. iCLIP predicts the dual splicing effects of TIA–RNA interactions. PLoS Biol. 8, e1000530 (2010).
Huppertz, I. et al. iCLIP: Protein–RNA interactions at nucleotide resolution. Methods 65, 274–287 (2014).
Heintzen, C., Nater, M., Apel, K. & Staiger, D. AtGRP7, a nuclear RNA-binding protein as a component of a circadian-regulated negative feedback loop in Arabidopsis thaliana. Proc. Natl Acad. Sci. USA 94, 8515–8520 (1997).
Hackmann, C. et al. Salicylic acid-dependent and -independent impact of an RNA-binding protein on plant immunity. Plant Cell Envir. 37, 696–706 (2014).
Löhr, B., Streitner, C., Steffen, A., Lange, T. & Staiger, D. A glycine-rich RNA-binding protein affects gibberellin biosynthesis in Arabidopsis. Mol. Biol. Rep. 41, 439–445 (2014).
Staiger, D. RNA-binding proteins and circadian rhythms in Arabidopsis thaliana. Philos. Trans. R. Soc. Lond. B Biol. Sci. 356, 1755–1759 (2001).
Schmidt, F. et al. A proteomic analysis of oligo(dT)-bound mRNP containing oxidative stress-induced Arabidopsis thaliana RNA-binding proteins ATGRP7 and ATGRP8. Mol. Biol. Rep. 37, 839–845 (2010).
Meyer, K. et al. Adaptation of iCLIP to plants determines the binding landscape of the clock-regulated RNA-binding protein AtGRP7. Genome Biol. 18, 204 (2017).
Mateos, J. L. & Staiger, D. Toward a systems view on RNA-binding proteins and associated RNAs in plants: guilt by association. Plant Cell 35, 1708–1726 (2023).
Arabidopsis Genome Initiative. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408, 796–815 (2000).
Streitner, C. et al. The small glycine-rich RNA-binding protein AtGRP7 promotes floral transition in Arabidopsis thaliana. Plant J. 56, 239–250 (2008).
Rothbauer, U. et al. A versatile nanotrap for biochemical and functional studies with fluorescent fusion proteins. Mol. Cell. Proteom. 7, 282–289 (2008).
Van Ende, R., Balzarini, S. & Geuten, K. Single and combined methods to specifically or bulk-purify RNA–protein complexes. Biomolecules 10, 1160 (2020).
Köster, T., Reichel, M. & Staiger, D. CLIP and RNA interactome studies to unravel genome-wide RNA-protein interactions in vivo in Arabidopsis thaliana. Methods 178, 63–71 (2020).
Widjaja, I. et al. Combining subproteome enrichment and Rubisco depletion enables identification of low abundance proteins differentially regulated during plant defense. Proteomics 9, 138–147 (2009).
Marondedze, C., Thomas, L., Serrano, N. L., Lilley, K. S. & Gehring, C. The RNA-binding protein repertoire of Arabidopsis thaliana. Sci. Rep. 6, 29766 (2016).
Zhang, Z. et al. UV crosslinked mRNA-binding proteins captured from leaf mesophyll protoplasts. Plant Methods 12, 42 (2016).
Bach-Pages, M. et al. Discovering the RNA-binding proteome of plant leaves with an improved RNA interactome capture method. Biomolecules 10, 661 (2020).
Köster, T., Marondedze, C., Meyer, K. & Staiger, D. RNA-binding proteins revisited—the emerging Arabidopsis mRNA interactome. Trends Plant Sci. 22, 512–526 (2017).
Briese, M. et al. A systems view of spliceosomal assembly and branchpoints with iCLIP. Nat. Struc. Mol. Biol. 26, 930–940 (2019).
Sutandy, F. X., Hildebrandt, A. & König, J. Profiling the binding sites of RNA-binding proteins with nucleotide resolution using iCLIP. Methods Mol. Biol. 1358, 175–195 (2016).
Van Nostrand, E. L. et al. Robust, cost-effective profiling of RNA binding protein targets with single-end enhanced crosslinking and immunoprecipitation (seCLIP). Methods Mol. Biol. 1648, 177–200 (2017).
Zarnegar, B. J. et al. irCLIP platform for efficient characterization of protein–RNA interactions. Nat. Methods 13, 489–492 (2016).
Buchbender, A. et al. Improved library preparation with the new iCLIP2 protocol. Methods 178, 33–48 (2020).
Van Nostrand, E. L. et al. Robust transcriptome-wide discovery of RNA-binding protein binding sites with enhanced CLIP (eCLIP). Nat. Methods 13, 508–514 (2016).
Köster, T. & Staiger, D. Plant individual nucleotide resolution crosslinking and immunoprecipitation to characterize RNA–protein complexes. Methods Mol. Biol. 2166, 255–268 (2020).
Busch, A., Brüggemann, M., Ebersberger, S. & Zarnack, K. iCLIP data analysis: a complete pipeline from sequencing reads to RBP binding sites. Methods 178, 49–62 (2020).
Arribas-Hernandez, L. et al. Principles of mRNA targeting via the Arabidopsis m(6)A-binding protein ECT2. eLife 10, e72375 (2021).
Lewinski, M., Bramkamp, Y., Köster, T. & Staiger, D. SEQing: web-based visualization of iCLIP and RNA-seq data in an interactive python framework. BMC Bioinforma. 21, 113 (2020).
Scutenaire, J. et al. The YTH domain protein ECT2 is an m6A reader required for normal trichome branching in Arabidopsis. Plant Cell 30, 986–1005 (2018).
Linder, B. et al. Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome. Nat. Methods 12, 767 (2015).
Parker, M. T. et al. Nanopore direct RNA sequencing maps the complexity of Arabidopsis mRNA processing and m6A modification. eLife 9, e49658 (2020).
Körtel, N. et al. Deep and accurate detection of m6A RNA modifications using miCLIP2 and m6Aboost machine learning. Nucleic Acids Res. 49, e92 (2021).
Wheeler, E. C., Van Nostrand, E. L. & Yeo, G. W. Advances and challenges in the detection of transcriptome-wide protein–RNA interactions. Wiley Interdiscip. Rev. RNA 1, e1436 (2017).
Hannigan, M. M., Zagore, L. L. & Licatalosi, D. D. Mapping transcriptome-wide protein–RNA interactions to elucidate RNA regulatory programs. Quant. Biol. 6, 228–238 (2018).
Terzi, L. C. & Simpson, G. G. Arabidopsis RNA immunoprecipitation. Plant J. 59, 163–168 (2009).
Xing, D., Wang, Y., Hamilton, M., Ben-Hur, A. & Reddy, A. S. N. Transcriptome-wide identification of RNA targets of Arabidopsis SERINE/ARGININE-RICH45 uncovers the unexpected roles of this RNA-binding protein in RNA processing. Plant Cell 27, 3294–3308 (2015).
Köster, T. & Staiger, D. in Arabidopsis Protocols (eds. Sanchez-Serrano, J. J. & Salinas, J.) 453–461 (Springer, 2021).
Zhang, Y. et al. Integrative genome-wide analysis reveals HLP1, a novel RNA-binding protein, regulates plant flowering by targeting alternative polyadenylation. Cell Res. 25, 864–876 (2015).
Licatalosi, D. D. et al. HITS-CLIP yields genome-wide insights into brain alternative RNA processing. Nature 456, 464–469 (2008).
Zhang, C. & Darnell, R. B. Mapping in vivo protein-RNA interactions at single-nucleotide resolution from HITS-CLIP data. Nat. Biotechnol. 29, 607–614 (2011).
Sugimoto, Y. et al. Analysis of CLIP and iCLIP methods for nucleotide-resolution studies of protein–RNA interactions. Genome Biol. 13, R67 (2012).
Wu, Z. et al. RNA-binding proteins At RZ-1B and At RZ-1C play a critical role in regulation of pre-mRNA splicing and gene expression during Arabidopsis development. Plant Cell 28, 55–73 (2016).
Van Nostrand, E. L. et al. Principles of RNA processing from analysis of enhanced CLIP maps for 150 RNA binding proteins. Genome Biol. 21, 90 (2020).
Van Nostrand, E. L. et al. A large-scale binding and functional map of human RNA-binding proteins. Nature 583, 711–719 (2020).
Burjoski, V. & Reddy, A. S. N. The landscape of RNA–protein interactions in plants: approaches and current status. Int. J. Mol. Sci. 22, 2845 (2021).
Brannan, K. W. et al. Robust single-cell discovery of RNA targets of RNA-binding proteins and ribosomes. Nat. Methods 18, 507–519 (2021).
McMahon, A. C. et al. TRIBE: hijacking an RNA-editing enzyme to identify cell-specific targets of RNA-binding proteins. Cell 165, 742–753 (2016).
Xu, W., Rahman, R. & Rosbash, M. Mechanistic implications of enhanced editing by a hyperTRIBE RNA-binding protein. RNA 24, 173–182 (2018).
Endo, M., Shimizu, H., Nohales, M. A., Araki, T. & Kay, S. A. Tissue-specific clocks in Arabidopsis show asymmetric coupling. Nature 514, 419–422 (2014).
Balzarini, S., Van Ende, R., Voet, A. & Geuten, K. A widely applicable and cost-effective method for specific RNA–protein complex isolation. Sci. Rep. 13, 6898 (2023).
Frohnmeyer, H. & Staiger, D. Ultraviolet-B radiation-mediated responses in plants. Balancing damage and protection. Plant Physiol. 133, 1420–1428 (2003).
Heijde, M. & Ulm, R. UV-B photoreceptor-mediated signalling in plants. Trends Plant Sci. 17, 230–237 (2012).
Rahman, R., Xu, W., Jin, H. & Rosbash, M. Identification of RNA-binding protein targets with HyperTRIBE. Nat. Protoc. 13, 1829–1849 (2018).
Padrón, A., Iwasaki, S. & Ingolia, N. T. Proximity RNA labeling by APEX-Seq reveals the organization of translation initiation complexes and repressive RNA granules. Mol. Cell 75, 875–887.e875 (2019).
Sutandy, F. X. R. et al. In vitro iCLIP-based modeling uncovers how the splicing factor U2AF2 relies on regulation by cofactors. Genome Res. 28, 699–713 (2018).
Sharma, D. et al. The kinetic landscape of an RNA-binding protein in cells. Nature 591, 152–156 (2021).
Zarnack, K. et al. Direct competition between hnRNP C and U2AF65 protects the transcriptome from the exonization of Alu elements. Cell 152, 453–466 (2013).
Chakrabarti, A., Haberman, N., Praznik, A., Luscombe, N. & Ule, J. Data science issues in studying protein–RNA interactions with CLIP technologies. Annu. Rev. Biomed. Data Sci. 1, 235–261 (2018).
Licatalosi, D. D. & Darnell, R. B. RNA processing and its regulation: global insights into biological networks. Nat. Rev. Genet 11, 75–87 (2010).
Castello, A. et al. Insights into RNA biology from an atlas of mammalian mRNA-binding proteins. Cell 149, 1393–1406 (2012).
Zhang, R. et al. A high quality Arabidopsis transcriptome for accurate transcript-level analysis of alternative splicing. Nucleic Acids Res. 45, 5061–5073 (2017).
Lamesch, P. et al. The Arabidopsis Information Resource (TAIR): improved gene annotation and new tools. Nucleic Acids Res. 40, D1202–D1210 (2012).
Gregory, T. R. in The Evolution of the Genome (ed. Gregory, T. R.) 3–87 (Academic Press, 2005).
Blue, S. M. et al. Transcriptome-wide identification of RNA-binding protein binding sites using seCLIP-seq. Nat. Protoc. 17, 1223–1265 (2022).
Thorvaldsdóttir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinforma. 14, 178–192 (2013).
Streitner, C. et al. An hnRNP-like RNA-binding protein affects alternative splicing by in vivo interaction with target transcripts in Arabidopsis thaliana. Nucleic Acids Res. 40, 11240–11255 d (2012).
Murashige, T. & Skoog, F. A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol. Plant. 15, 473–497 (1962).
Kent, W. J., Zweig, A. S., Barber, G., Hinrichs, A. S. & Karolchik, D. BigWig and BigBed: enabling browsing of large distributed datasets. Bioinformatics 26, 2204–2207 (2010).
Dodt, M., Roehr, J. T., Ahmed, R. & Dieterich, C. FLEXBAR—flexible barcode and adapter processing for next-generation sequencing platforms. Biology 1, 895–905 (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. GigaScience 10, giab008 (2021).
Krakau, S., Richard, H. & Marsico, A. PureCLIP: capturing target-specific protein–RNA interaction footprints from single-nucleotide CLIP-seq data. Genome Biol. 18, 240 (2017).
Acknowledgements
We thank K. Neudorf and E. Klemme for expert technical assistance throughout the development of the procedure. Work in our laboratories is supported by the German Research Foundation through grants STA653/14-1, 15-1 and 19-1 to D.S., SPP 1935 ZA881/5-2 and FOR 2333 ZA881/3-1 to K.Z., SPP 1935 KO4566/3-2 and FOR 2333 KO4566/5-1 to J.K., and by a postdoctoral fellowship of the Alexander von Humboldt Foundation to M.R.
Author information
Authors and Affiliations
Contributions
T.K., M.R. and K.M. developed the experimental methods. M.L. developed the processing part of the bioinformatics workflow. M.B. developed the R/Bioconductor package BindingSiteFinder. D.S. and J.K. directed the development of the experimental methods. K.Z. directed the development of the bioinformatics workflow. M.R. assembled part 1 of the procedure pertaining to the experimental protocol. M.L. and M.B. assembled part 2 of the procedure pertaining to the bioinformatics workflow. T.B. assembled the Materials section. K.M. produced the data in Fig. 2. D.S. wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Oliver Rossbach and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Meyer, K. et al. Genome Biol. 18, 204 (2017): https://doi.org/10.1186/s13059-017-1332-x
Arribas-Hernandez, L. et al. eLife 10, e72375 (2021): https://doi.org/10.7554/eLife.72375
Buchbender, A. et al. Methods 178, 33–48 (2020): https://doi.org/10.1016/j.ymeth.2019.10.003
Busch, A. et al. Methods 178, 49–62 (2020): https://doi.org/10.1016/j.ymeth.2019.11.008
Source data
Source Data Fig. 3
Unprocessed radiograms and blot.
Source Data Fig. 5
Unprocessed gel and blots.
Source Data Fig. 7
Unprocessed gel.
Source Data Fig. 8
Unprocessed gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lewinski, M., Brüggemann, M., Köster, T. et al. Mapping protein–RNA binding in plants with individual-nucleotide-resolution UV cross-linking and immunoprecipitation (plant iCLIP2). Nat Protoc 19, 1183–1234 (2024). https://doi.org/10.1038/s41596-023-00935-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-023-00935-3
- Springer Nature Limited
This article is cited by
-
Revealing the Arabidopsis AtGRP7 mRNA binding proteome by specific enhanced RNA interactome capture
BMC Plant Biology (2024)