Abstract
Structural maintenance of chromosome (SMC) protein complexes are the key organizers of the spatiotemporal structure of chromosomes. The condensin SMC complex has recently been shown to be a molecular motor that extrudes large loops of DNA, but the mechanism of this unique motor remains elusive. Using atomic force microscopy, we show that budding yeast condensin exhibits mainly open ‘O’ shapes and collapsed ‘B’ shapes, and it cycles dynamically between these two states over time, with ATP binding inducing the O to B transition. Condensin binds DNA via its globular domain and also via the hinge domain. We observe a single condensin complex at the stem of extruded DNA loops, where the neck size of the DNA loop correlates with the width of the condensin complex. The results are indicative of a type of scrunching model in which condensin extrudes DNA by a cyclic switching of its conformation between O and B shapes.






Similar content being viewed by others
Change history
08 October 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Nasmyth, K. & Haering, C. H. The structure and function of SMC and kleisin complexes. Annu. Rev. Biochem. 74, 595–648 (2005).
Haering, C. H. & Gruber, S. SnapShot: SMC protein complexes part I. Cell 164, 326.e1 (2016).
Hirano, T. Condensin-based chromosome organization from bacteria to vertebrates. Cell 164, 847–857 (2016).
Oliveira, R. A., Coelho, P. A. & Sunkel, C. E. The condensin I subunit Barren/CAP-H is essential for the structural integrity of centromeric heterochromatin during mitosis. Mol. Cell. Biol. 25, 8971–8984 (2005).
Houlard, M. et al. Condensin confers the longitudinal rigidity of chromosomes. Nat. Cell Biol. 17, 771–781 (2015).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320.e24 (2017).
Naumova, N. et al. Organization of the mitotic chromosome. Science 342, 948–953 (2013).
Sanborn, A. L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl Acad. Sci. USA 112, E6456–E6465 (2015).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
Dekker, J. & Mirny, L. The 3D genome as moderator of chromosomal communication. Cell 164, 1110–1121 (2016).
Alipour, E. & Marko, J. F. Self-organization of domain structures by DNA-loop-extruding enzymes. Nucleic Acids Res. 40, 11202–11212 (2012).
Goloborodko, A., Marko, J. F. & Mirny, L. A. Chromosome compaction by active loop extrusion. Biophys. J. 110, 2162–2168 (2016).
Terakawa, T. et al. The condensin complex is a mechanochemical motor that translocates along DNA. Science 358, 672–676 (2017).
Ganji, M. et al. Real-time imaging of DNA loop extrusion by condensin. Science 360, 102–105 (2018).
Eeftens, J. & Dekker, C. Catching DNA with hoops—biophysical approaches to clarify the mechanism of SMC proteins. Nat. Struct. Mol. Biol. 24, 1012–1020 (2017).
Diebold-Durand, M. L. et al. Structure of full-length SMC and rearrangements required for chromosome organization. Mol. Cell 67, 334–347.e5 (2017).
Kamada, K., Su’etsugu, M., Takada, H., Miyata, M. & Hirano, T. Overall shapes of the SMC–ScpAB complex are determined by balance between constraint and relaxation of its structural parts. Structure 25, 603–616.e4 (2017).
Soh, Y. M. et al. Molecular basis for SMC rod formation and its dissolution upon DNA binding. Mol. Cell 57, 290–303 (2015).
Bürmann, F. et al. A folded conformation of MukBEF and cohesin. Nat. Struct. Mol. Biol. 26, 227–236 (2019).
Lee, B. G. et al. Cryo-EM structures of holo condensin reveal a subunit flip-flop mechanism. Nat. Struct. Mol. Biol. https://doi.org/10.1038/s41594-020-0457-x (2020).
Yoshimura, S. H. et al. Condensin architecture and interaction with DNA. Curr. Biol. 12, 508–513 (2002).
Eeftens, J. M. et al. Condensin Smc2-Smc4 dimers are flexible and dynamic. Cell Rep. 14, 1813–1818 (2016).
Katan, A. J. & Dekker, C. High-speed AFM reveals the dynamics of single biomolecules at the nanometer scale. Cell 147, 979–982 (2011).
Kodera, N., Yamamoto, D., Ishikawa, R. & Ando, T. Video imaging of walking myosin V by high-speed atomic force microscopy. Nature 468, 72–76 (2010).
Uchihashi, T. et al. High-speed atomic force microscopy reveals rotary catalysis of rotorless F1-ATPase. Science 333, 755–758 (2011).
Shi, Z., Gao, H., Bai, X.-C. & Yu, H. Cryo-EM structure of the human cohesin-NIPBL-DNA complex. Science 368, 1454–1459 (2020).
Kschonsak, M. et al. Structural basis for a safety-belt mechanism that anchors condensin to chromosomes. Cell 171, 588–600 (2017).
Piazza, I. et al. Association of condensin with chromosomes depends on DNA binding by its HEAT-repeat subunits. Nat. Struct. Mol. Biol. 21, 560–568 (2014).
Kim, Y., Shi, Z., Zhang, H., Finkelstein, I. J. & Yu, H. Human cohesin compacts DNA by loop extrusion. Science 366, 1345–1349 (2019).
Japaridze, A. et al. Spatial confinement induces hairpins in nicked circular DNA. Nucleic Acids Res. 45, 4905–4914 (2017).
Anderson, D. E., Trujillo, K. M., Sung, P. & Erickson, H. P. Structure of the Rad50·Mre11 DNA repair complex from Saccharomyces cerevisiae by electron microscopy. J. Biol. Chem. 276, 37027–37033 (2001).
Moreno-Herrero, F. et al. Mesoscale conformational changes in the DNA-repair complex Rad50/Mre11/Nbs1 upon binding DNA. Nature 437, 440–443 (2005).
Park, Y. B. et al. Eukaryotic Rad50 functions as a rod-shaped dimer. Nat. Struct. Mol. Biol. 24, 248–257 (2017).
Lyumkis, D. Challenges and opportunities in cryo-EM single-particle analysis. J. Biol. Chem. 294, 5181–5197 (2019).
Noble, A. J. et al. Reducing effects of particle adsorption to the air–water interface in cryo-EM. Nat. Methods 15, 793–795 (2018).
D’Imprima, E. et al. Protein denaturation at the air-water interface and how to prevent it. Elife 8, e42747 (2019).
Griese, J. J., Witte, G. & Hopfner, K. P. Structure and DNA binding activity of the mouse condensin hinge domain highlight common and diverse features of SMC proteins. Nucleic Acids Res. 38, 3454–3465 (2010).
Chiu, A., Revenkova, E. & Jessberger, R. DNA interaction and dimerization of eukaryotic SMC hinge domains. J. Biol. Chem. 279, 26233–26242 (2004).
Kumar, R. et al. The bacterial condensin MukB compacts DNA by sequestering supercoils and stabilizing topologically isolated loops. J. Biol. Chem. 292, 16904–16920 (2017).
Haering, C. H., Löwe, J., Hochwagen, A. & Nasmyth, K. Molecular architecture of SMC proteins and the yeast cohesin complex. Mol. Cell 9, 773–788 (2002).
Li, Y., Schoeffler, A. J., Berger, J. M. & Oakley, M. G. The crystal structure of the hinge domain of the Escherichia coli structural maintenance of chromosomes protein MukB. J. Mol. Biol. 395, 11–19 (2010).
Niki, H. Open and closed questions about open and closed SMC. Structure 25, 569–570 (2017).
Ku, B., Lim, J. H., Shin, H. C., Shin, S. Y. & Oh, B. H. Crystal structure of the MukB hinge domain with coiled-coil stretches and its functional implications. Proteins Struct. Funct. Bioinforma. 78, 1483–1490 (2010).
Nichols, M. H. & Corces, V. G. A tethered-inchworm model of SMC DNA translocation. Nat. Struct. Mol. Biol. 25, 906–910 (2018).
Hassler, M., Shaltiel, I. A. & Haering, C. H. Towards a unified model of SMC complex function. Curr. Biol. 28, R1266–R1281 (2018).
Hwang, W. & Karplus, M. Structural basis for power stroke vs. Brownian ratchet mechanisms of motor proteins. Proc. Natl Acad. Sci. USA 116, 19777–19785 (2019).
Liu, Y. et al. ATP‐dependent DNA binding, unwinding, and resection by the Mre11/Rad50 complex. EMBO J. 35, 743–758 (2016).
Seifert, F. U., Lammens, K., Stoehr, G., Kessler, B. & Hopfner, K. Structural mechanism of ATP ‐dependent DNA binding and DNA end bridging by eukaryotic Rad50. EMBO J. 35, 759–772 (2016).
Woo, J. S. et al. Structural studies of a bacterial condensin complex reveal ATP-dependent disruption of intersubunit interactions. Cell 136, 85–96 (2009).
Erickson, H. P. Size and shape of protein molecules at the nanometer level determined by sedimentation, gel filtration, and electron microscopy. Biol. Proced. Online 11, 32–51 (2009).
Davidson, I. F. et al. DNA loop extrusion by human cohesin. Science 366, 1338–1345 (2019).
Schiener, J., Witt, S., Stark, M. & Guckenberger, R. Stabilized atomic force microscopy imaging in liquids using second harmonic of cantilever motion for setpoint control. Rev. Sci. Instrum. 75, 2564–2568 (2004).
Eeftens, J. M. et al. Real-time detection of condensin-driven DNA compaction reveals a multistep binding mechanism. EMBO J. 36, 3448–3457 (2017).
Rivetti, C., Guthold, M. & Bustamante, C. Scanning force microscopy of DNA deposited onto mica: equilibration versus kinetic trapping studied by statistical polymer chain analysis. J. Mol. Biol. 264, 919–932 (1996).
Uchihashi, T., Kodera, N. & Ando, T. Guide to video recording of structure dynamics and dynamic processes of proteins by high-speed atomic force microscopy. Nat. Protoc. 7, 1193–1206 (2012).
Nečas, D. & Klapetek, P. Gwyddion: an open-source software for SPM data analysis. Cent. Eur. J. Phys. 10, 181–188 (2012).
Gwynn, E. J. et al. Using DNA as a fiducial marker to study SMC complex interactions with the atomic force microscope. Biophys. J. 102, 839–848 (2012).
Japaridze, A. et al. Hyperplectonemes: a higher order compact and dynamic DNA self-organization. Nano Lett. 17, 1938–1948 (2017).
Acknowledgements
We thank S. Bisht, J. Eeftens, M. Ganji, E. Kim, B. Pradhan, S. H. Rah, I. Shaltiel, J. van der Torre, and W. W. W. Yang for discussions. This work was supported by ERC grants SynDiv 669598 (to C.D.), Marie Skłodowska-Curie grant agreement No. 753002 (to J.-K.R.), CondStruct 681365 (to C.H.H.), the Netherlands Organization for Scientific Research (NWO/OCW) as part of the Frontiers of Nanoscience and Basyc programs and the European Molecular Biology Laboratory.
Author information
Authors and Affiliations
Contributions
J.-K.R., C.D., A.J.K. and C.H.H. designed the experiments; J.-K.R., A.K. and T.W. performed the AFM experiments; E.V.D.S. and J.-K.R. purified condensin complex; J.-K.R., A.J.K. and T.W. contributed image analysis; J.-K.R. and R.D.G. performed the ATPase assay and single-molecule fluorescence assay; C.D. and C.H.H supervised the work; and J.-K.R., C.D. and A.J.K. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Anke Sparmann was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characterization of the functionality of yeast condensin.
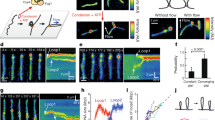
a, SDS page gel of the condensin proteins. b, ATPase activity of condensin with various controls (N = 3 independent experiments). Errors are s.d.. Uncropped image is available as source data. c, Loop extrusion assay with double tethered DNA showing that the condensin leads to the extrusion of a loop of DNA (N = 3 independent experiments and 200 molecules). d, Loop-extrusion probability with various negative controls (N > 200 molecules). In b and d, *** indicates P<0.001, assessed using two-tailed Student’s t-test.
Extended Data Fig. 2 Characterization of the condensin holocomplex.
a, Frequency distribution of the condensin volume (N = 3 independent experiments and 384 molecules) as measured from dry AFM images. Gaussian fitting yielded a mean value of 2420 ± 430 nm3. b, Mean volume of each shape conformation (N = 86, 200, 51, and 47 for each state, respectively). Errors are s.d.. There were no significant differences in molecular volume between the states, consistent with the fact that each contained the same subunits. c, Relative occurrence of different shapes of condensin in AFM images taken in the presence of ATP on polylysine-treated mica (N = 253) versus images on bare mica (N = 384). Before sample deposition, polylysine was treated for 3 min, and the unbound polylysine was rinsed using distilled water. Errors are s.d. obtained by counting errors. d, Representative images that classified as other. Image of a ‘kleisin-opened’ state with a disconnection between one of the SMC heads and the non-SMC subunits (top) (N = 21 molecules). The white arrow indicates the non-SMC domains while the orange arrow points to an SMC end. AFM image of a ‘hinge-opened’ state where the two SMC arms were disengaged at the hinge (‘hinge-opened shapes’) (N = 25 molecules). The orange arrows indicate the dissociated hinge domains.
Extended Data Fig. 3 Representative HS AFM image sequences of the condensin holocomplex in the presence of ATP.
a, b, Images were taking from movies acquired at 5 frames/s rate, from Supplementary Video 2 in a and Supplementary Video 3 in b (N = 6 molecules, 3 independent experiments, 2 independent protein purification, and 5,240 frames). Changes between O- and B-shapes were observed. O and B indicate O-shape and B-shape, respectively. Cartoons on the right of each image are for visual guidance. c, Number of distinguishable subunits in the globular domain (that is, excluding the hinge) (N = 5,240 frames from 6 movies). About 70% of the complexes featured 3 distinguishable subunits in the globular domain, and about 30% featured 4 subunits. The complex with 3 subunits likely is composed of a dimerized head domain and two heat-domains, while two head domains were resolved separately in the case of the complex with 4 subunits in the globular domain.
Extended Data Fig. 4 Temporal fluctuations in the hinge angle, and distance changes between the two SMC arms and between the two HEAT domains.
a, Snapshot image from a liquid AFM movie of a condensin holocomplex with the hinge angle indicated (N = 3,780 frames from 6 molecules). b, Representative trace of the hinge angle versus time. Right panel shows the distribution of the hinge angle from this trace (N = 622). c, Hinge angle distribution from 6 independent molecules (N = 3,780 frames). Fit is a lognormal distribution with a mean of 96° and a standard deviation of 48°, close to the hinge angle measured using dry AFM images (Fig. 1f). d, Snapshot image from a liquid AFM movie of a condensin holocomplex with the distance between two SMC arms indicated (N = 2,315 from 4 independent molecules). This mutual distance between the two SMC arms was measured along a line at the midpoint perpendicular (green) to a line (white) connecting the centers of mass at hinge and heads. e, Representative trace of the two SMC arms measured in a condensin holocomplex in the presence of ATP. Right panel shows the distribution of the distance between two SMC arms (N = 789). f, Distribution of the mutual distance between the two SMC arms obtained for 4 independent molecules (N = 2,315). Also plotted is the measured width for an individual SMC arm (FWHM of the cross-section of the arm) (N = 88). Fits are Gaussian distributions with 28.0 ± 8.1 nm and 6.6 ± 2.3 (mean ± s.d.) for the mutual distance and SMC arm thickness, respectively. g, Snapshot image from a liquid AFM movie with the Ycg1 and Ycs4 HEAT domains indicated for a condensin holocomplex where the two heads are dimerized (N = 2,577 from 4 independent molecules). h, Representative trace of the Ycg1-Ycs4 distance measured in a condensin holocomplex in the presence of ATP. Right panel shows the distribution of the Ycg1-Ycs4 distance from this trace (N = 784). i, The Ycg1-Ycs4 distance distributions obtained for 4 independent molecules (N = 2,577). Fit is Gaussian distribution with 12.0 ± 2.4 nm (mean ± s.d.).
Extended Data Fig. 5 Tip scanning direction does not bias the hinge motion.
In HS AFM imaging in liquid, tapping and scanning the sample with the tip can potentially affect experiments adversely, for example induce a movement due to the tip pushing the molecule. The exact magnitude of the tip-sample forces in fluid tapping mode AFM is difficult to assess during imaging, but nevertheless can be qualitatively estimated from the magnitude of the second harmonic of the tapping signal52. We precisely controlled this and kept it to the minimum that still allowed for sufficient imaging resolution. To assess whether this was sufficient to avoid influencing the molecule dynamics, we analyzed the distribution of frame-to-frame position changes of the hinge domain with respect to the head domains. Since the scanning motion is not isotropic, one might expect an uneven distribution of hinge positional changes in the fast and slow scanning directions. Instead, we find a highly isotropic distribution, indicating minimal tip-induced effects. a, The zigzag scanning direction of HS AFM imaging. The horizontal direction scanning motion (black lines) is much faster than the vertical scanning. After finishing the imaging of a frame, the tip moves back to the starting point (red line) and scanning restarts. b, Surface plot of the two-dimensional histograms of the hinge domain displacement at all time points (N > 4,000 frames from 6 movies on different molecules). A radially symmetric distribution is obtained. No bias is observed towards a certain direction, also not towards the horizontal scanning direction (X-axis). c, The distributions of X, Y displacements between adjacent images are nearly identical. d, Tables of standard deviation and skewness of the X, Y displacements. Almost no skewness was observed in both cases.
Extended Data Fig. 6 Relative occurrence of different states of DNA-bound wild type condensin without ATP and EQ mutant plus ATP.
a, Probability of the O-and B-shapes for condensin bound to DNA. b, Relative occurrence of each of the different binding modes. N = 83 and 347 molecules for wild type condensin without ATP and EQ mutant plus ATP, respectively. Errors are s.d. calculated by counting errors.
Extended Data Fig. 7 Binding probability and co-localization probability of condensin on DNA.
a, Schematic that illustrates the calculation of the probability for DNA and condensin to co-localize. Each condensin is approximated by a circumcircle with size Dp ~ 40 nm. The light blue shaded region indicates the area that should contain the center of the circumcircles for a randomly deposited condensin to co-localize with DNA. Condensin complexes inside the red circles overlap with DNA and would be counted as being bound in our measurements. Condensin complexes such as that in the blue circle are also included in the calculated co-localizing fraction, since their circumcircle overlaps with the DNA, even though the molecule itself does not touch the DNA. We deliberately err on the side of caution and do not correct for this, and the estimated value for the colocalization probability is therefore an upper bound. b, The DNA binding probability of condensin with ATP (red bar), the (inadvertent) colocalization probability of condensin (black bar), and DNA binding probability of condensin for negative controls (blue bars) The DNA binding probability of condensin is the observed binding probability, while the colocalization probability is the estimated probability that DNA and condensin accidently overlap on an AFM surface without biochemical interactions. N = 5, 5, 6, 8, and 5 independent experiments and errors are s.d., and *** indicates P<0.001, assessed using two-tailed Student’s t-test.
Extended Data Fig. 8 Loop size and occurrence of condensin at the stem of DNA loops.
a, DNA loop size distribution for data obtained with ATP, with ATPγS, without ATP, and without Ycg1 (N = 94, 22, 20, and 11 loops from 5, 6, 8, and 5 independent experiments, respectively). The importance of these data lies in the larger loop size observed with ATP as compared to the data in the other three controls. The actual value of the median loop size (0.54 μm in top panel) is in fact rather dependent on experimental conditions (for example incubation time) and limited by our finite imaging area (10 μm × 10 μm) and DNA overlap, which hampers clean identification of large loops in our experiments. Hence, we measured relative small loop sizes (< 1 μm) compared to earlier fluorescence microscopy experiments14. In the data presented here, we measured the size of loops where condensin localized at the stem. b, Loop size comparision in various conditions from a (median ± s.e.m.). c, Schematic to illustrate our estimate of the localization probability of condensin at the stem of the loop versus elsewhere on the DNA. Green circles denote the loop-stem area with a size Dp. Condensin occurring there (red circle) are counted ‘at the loop stem’, whereas all other DNA-bound condensin are counted ‘not at the loop stem’ (blue circle). d, Experimentally measured fractions of DNA-bound condensins that are observed at a loop stem (red), and not at a loop stem (blue) (N = 5 independent experiments). e, Fraction of DNA-bound condensins that are observed at a DNA loop stem (red), and the estimated accidental localization of condensin at a stem of DNA loops (green; N = 5 independent experiments) – see Material and Methods. Errors are s.e.m., and ** indicates P<0.01, assessed using two-tailed Student’s t-test.
Extended Data Fig. 9 Loops with an interwound DNA structure at the stem of loop.
a, Example of an interwound DNA loop. Two arrows indicate the starting and ending points of an interwound DNA region. Yellow arrow indicates a condensin complex located near the loop stem. b, Fraction of loops with an interwound region at the stem (N = 116 molecules from 3 independent experiments). c, Scatter diagram of the protein width vs the neck size of normal (non-interwound) DNA loops (N = 88 loops). d, Idem for DNA interwound loops at the stem (N = 27 loops). A neck size of ~9 nm is observed, which may be attributed to the two parallel DNA molecules (that feature a ~2 + 2 nm dsDNA width), convoluted by the AFM tip size.
Extended Data Fig. 10 Three different models of condensin-mediated DNA loop extrusion.
a, Tethered inchworm model. Here, the hinge domain anchors the DNA. The two heads are tethered by Brn1 (green), which also binds Ycg1 and Ycs4, the HEAT subunits that bind DNA. In a cyclic fashion, associated with ATP hydrolysis, these two DNA-binding domains bind and unbind from DNA (‘walk along the DNA’) to pull in target DNA. b, DNA pumping model. Ycg1 (orange) anchors DNA while the hinge domain binds to another spot along the DNA. After ATP hydrolysis, the dissociation of dimerized head domains induces a conformational change of O shape to I shape of the SMC arms. Zippering of the SMC arms pushes the hinge-bound DNA region towards the head region. After ATP binding to the head domains, the hinge domain binds to new target DNA region, and the cycle is repeated for further DNA loop extrusion. c, Scrunching model. Ycg1 (orange) anchors DNA, and the hinge domain binds to a different spot along the DNA. Through conformational changes between O shape and B shape, the hinge transfers its bound DNA to the head domain, whereupon the hinge finds a new DNA target site for further DNA loop extrusion.
Supplementary information
Supplementary Video 1
High-speed liquid AFM movies of condensin holocomplex, related to Fig. 2a–c. Movie speed: 4×.
Supplementary Video 2
High-speed liquid AFM movies of condensin holocomplex, related to Extended Data Fig. 4. Movie speed: 4×.
Supplementary Video 3
High-speed liquid AFM movies of condensin holocomplex, related to Extended Data Fig. 4. Movie speed: 4×.
Supplementary Video 4
High-speed liquid AFM movies of condensin holocomplex, related to Extended Data Fig. 4. Movie speed: 4×.
Supplementary Video 5
High-speed liquid AFM movies of EQ mutant with ATP, related to Fig. 2d,f. Movie speed: 4×.
Supplementary Video 6
High-speed liquid AFM movies of condensin holocomplex without ATP, related to Fig. 2g,h. Movie speed: 4×.
Supplementary Video 7
Liquid AFM movie of condensin at the stem of the DNA loop, related to Fig. 4a. Movie speed: 10×.
Source data
Source Data Fig. 1
Statistical source data
Source Data Fig. 2
Statistical source data
Source Data Fig. 3
Statistical source data
Source Data Fig. 4
Statistical source data
Source Data Fig. 5
Statistical source data
Source Data Extended Data Fig. 1
Uncropped SDS-PAGE gel
Rights and permissions
About this article
Cite this article
Ryu, JK., Katan, A.J., van der Sluis, E.O. et al. The condensin holocomplex cycles dynamically between open and collapsed states. Nat Struct Mol Biol 27, 1134–1141 (2020). https://doi.org/10.1038/s41594-020-0508-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-020-0508-3
- Springer Nature America, Inc.
This article is cited by
-
An extrinsic motor directs chromatin loop formation by cohesin
The EMBO Journal (2024)
-
Anisotropic scrunching of SMC with a baton-pass mechanism
Communications Biology (2024)
-
Motor domain of condensin and step formation in extruding loop of DNA
Journal of Biological Physics (2024)
-
Testing pseudotopological and nontopological models for SMC-driven DNA loop extrusion against roadblock-traversal experiments
Scientific Reports (2023)
-
Single cohesin molecules generate force by two distinct mechanisms
Nature Communications (2023)