Abstract
HIV-1 virion infectivity factor (Vif) promotes degradation of the antiviral APOBEC3 (A3) proteins through the host ubiquitin-proteasome pathway to enable viral immune evasion. Disrupting Vif-A3 interactions to reinstate the A3-catalyzed suppression of human immunodeficiency virus type 1 (HIV-1) replication is a potential approach for antiviral therapeutics. However, the molecular mechanisms by which Vif recognizes A3 proteins remain elusive. Here we report a cryo-EM structure of the Vif-targeted C-terminal domain of human A3F in complex with HIV-1 Vif and the cellular cofactor core-binding factor beta (CBFβ) at 3.9-Å resolution. The structure shows that Vif and CBFβ form a platform to recruit A3F, revealing a direct A3F-recruiting role of CBFβ beyond Vif stabilization, and captures multiple independent A3F-Vif interfaces. Together with our biochemical and cellular studies, our structural findings establish the molecular determinants that are critical for Vif-mediated neutralization of A3F and provide a comprehensive framework of how HIV-1 Vif hijacks the host protein degradation machinery to counteract viral restriction by A3F.




Similar content being viewed by others
Data Availability
The model of the truncated Vif–CBFβ–A3FCTDm complex has been deposited in the Worldwide Protein Data Bank with accession code PDB 6NIL. The cryo-EM maps of the truncated and full-length Vif–CBFβ–A3FCTDm complexes have been deposited in the EMDB with accession codes EMD-9380 and EMD-9381, respectively. Source data for Fig. 2c–e, Fig. 3c–g, Extended Data Fig. 1b,c, Extended Data Fig. 4c,d, Extended Data Fig. 5b,c, Extended Data Fig. 6a,b, and Extended Data Fig. 7b,c are available with the paper online. Other data are available from corresponding authors upon reasonable request.
References
Lecossier, D., Bouchonnet, F., Clavel, F. & Hance, A. J. Hypermutation of HIV-1 DNA in the absence of the Vif protein. Science 300, 1112 (2003).
Mangeat, B. et al. Broad antiretroviral defence by human APOBEC3G through lethal editing of nascent reverse transcripts. Nature 424, 99–103 (2003).
Zhang, H. et al. The cytidine deaminase CEM15 induces hypermutation in newly synthesized HIV-1 DNA. Nature 424, 94–98 (2003).
Jager, S. et al. Vif hijacks CBF-beta to degrade APOBEC3G and promote HIV-1 infection. Nature 481, 371–375 (2011).
Zhang, W., Du, J., Evans, S. L., Yu, Y. & Yu, X. F. T-cell differentiation factor CBF-β regulates HIV-1 Vif-mediated evasion of host restriction. Nature 481, 376–379 (2011).
Marin, M., Rose, K. M., Kozak, S. L. & Kabat, D. HIV-1 Vif protein binds the editing enzyme APOBEC3G and induces its degradation. Nat. Med. 9, 1398–1403 (2003).
Sheehy, A. M., Gaddis, N. C. & Malim, M. H. The antiretroviral enzyme APOBEC3G is degraded by the proteasome in response to HIV-1 Vif. Nat. Med. 9, 1404–1407 (2003).
Stopak, K., de Noronha, C., Yonemoto, W. & Greene, W. C. HIV-1 Vif blocks the antiviral activity of APOBEC3G by impairing both its translation and intracellular stability. Mol. Cell 12, 591–601 (2003).
Yu, X. et al. Induction of APOBEC3G ubiquitination and degradation by an HIV-1 Vif-Cul5-SCF complex. Science 302, 1056–1060 (2003).
Conticello, S. G., Harris, R. S. & Neuberger, M. S. The Vif protein of HIV triggers degradation of the human antiretroviral DNA deaminase APOBEC3G. Curr. Biol. 13, 2009–2013 (2003).
Conticello, S. G., Thomas, C. J., Petersen-Mahrt, S. K. & Neuberger, M. S. Evolution of the AID/APOBEC family of polynucleotide (deoxy)cytidine deaminases. Mol. Biol. Evol. 22, 367–377 (2005).
Feng, Y., Baig, T. T., Love, R. P. & Chelico, L. Suppression of APOBEC3-mediated restriction of HIV-1 by Vif. Front. Microbiol. 5, 450 (2014).
Chaipan, C., Smith, J. L., Hu, W. S. & Pathak, V. K. APOBEC3G restricts HIV-1 to a greater extent than APOBEC3F and APOBEC3DE in human primary CD4+ T cells and macrophages. J. Virol. 87, 444–453 (2013).
Navarro, F. et al. Complementary function of the two catalytic domains of APOBEC3G. Virology 333, 374–386 (2005).
Russell, R. A., Smith, J., Barr, R., Bhattacharyya, D. & Pathak, V. K. Distinct domains within APOBEC3G and APOBEC3F interact with separate regions of human immunodeficiency virus type 1 Vif. J. Virol. 83, 1992–2003 (2009).
Smith, J. L. & Pathak, V. K. Identification of specific determinants of human APOBEC3F, APOBEC3C, and APOBEC3DE and African green monkey APOBEC3F that interact with HIV-1 Vif. J. Virol. 84, 12599–12608 (2010).
Hache, G., Liddament, M. T. & Harris, R. S. The retroviral hypermutation specificity of APOBEC3F and APOBEC3G is governed by the C-terminal DNA cytosine deaminase domain. J. Biol. Chem. 280, 10920–10924 (2005).
Aydin, H., Taylor, M. W. & Lee, J. E. Structure-guided analysis of the human APOBEC3-HIV restrictome. Structure 22, 668–684 (2014).
Kitamura, S., Ode, H. & Iwatani, Y. Structural features of antiviral APOBEC3 proteins are linked to their functional activities. Front. Microbiol. 2, 258 (2011).
Russell, R. A. & Pathak, V. K. Identification of two distinct human immunodeficiency virus type 1 Vif determinants critical for interactions with human APOBEC3G and APOBEC3F. J. Virol. 81, 8201–8210 (2007).
Fribourgh, J. L. et al. Core binding factor beta plays a critical role by facilitating the assembly of the Vif-cullin 5 E3 ubiquitin ligase. J. Virol. 88, 3309–3319 (2014).
Kim, D. Y. et al. CBFβ stabilizes HIV Vif to counteract APOBEC3 at the expense of RUNX1 target gene expression. Mol. Cell 49, 632–644 (2013).
Guo, Y. et al. Structural basis for hijacking CBF-β and CUL5 E3 ligase complex by HIV-1 Vif. Nature 505, 229–233 (2014).
Bohn, M. F. et al. Crystal structure of the DNA cytosine deaminase APOBEC3F: the catalytically active and HIV-1 Vif-binding domain. Structure 21, 1042–1050 (2013).
He, Z., Zhang, W., Chen, G., Xu, R. & Yu, X. F. Characterization of conserved motifs in HIV-1 Vif required for APOBEC3G and APOBEC3F interaction. J. Mol. Biol. 381, 1000–1011 (2008).
Pery, E., Rajendran, K. S., Brazier, A. J. & Gabuzda, D. Regulation of APOBEC3 proteins by a novel YXXL motif in human immunodeficiency virus type 1 Vif and simian immunodeficiency virus SIVagm Vif. J. Virol. 83, 2374–2381 (2009).
Nakashima, M. et al. Structural insights into HIV-1 Vif-APOBEC3F interaction. J. Virol. 90, 1034–1047 (2016).
Yamashita, T., Kamada, K., Hatcho, K., Adachi, A. & Nomaguchi, M. Identification of amino acid residues in HIV-1 Vif critical for binding and exclusion of APOBEC3G/F. Microbes Infect. 10, 1142–1149 (2008).
Dang, Y., Davis, R. W., York, I. A. & Zheng, Y. H. Identification of 81LGxGxxIxW89 and 171EDRW174 domains from human immunodeficiency virus type 1 Vif that regulate APOBEC3G and APOBEC3F neutralizing activity. J. Virol. 84, 5741–5750 (2010).
Kitamura, S. et al. The APOBEC3C crystal structure and the interface for HIV-1 Vif binding. Nat. Struct. Mol. Biol. 19, 1005–1010 (2012).
Albin, J. S. et al. A single amino acid in human APOBEC3F alters susceptibility to HIV-1 Vif. J. Biol. Chem. 285, 40785–40792 (2010).
Siu, K. K., Sultana, A., Azimi, F. C. & Lee, J. E. Structural determinants of HIV-1 Vif susceptibility and DNA binding in APOBEC3F. Nat. Commun. 4, 2593 (2013).
Anderson, B. D. & Harris, R. S. Transcriptional regulation of APOBEC3 antiviral immunity through the CBF-β/RUNX axis. Sci. Adv. 1, e1500296 (2015).
Dang, Y., Wang, X., Zhou, T., York, I. A. & Zheng, Y. H. Identification of a novel WxSLVK motif in the N terminus of human immunodeficiency virus and simian immunodeficiency virus Vif that is critical for APOBEC3G and APOBEC3F neutralization. J. Virol. 83, 8544–8552 (2009).
Chen, G., He, Z., Wang, T., Xu, R. & Yu, X. F. A patch of positively charged amino acids surrounding the human immunodeficiency virus type 1 Vif SLVx4Yx9Y motif influences its interaction with APOBEC3G. J. Virol. 83, 8674–8682 (2009).
Kim, D. Y. The assembly of Vif ubiquitin E3 ligase for APOBEC3 degradation. Arch. Pharm. Res. 38, 435–445 (2015).
Richards, C. et al. The binding interface between human APOBEC3F and HIV-1 Vif elucidated by genetic and computational approaches. Cell Reports 13, 1781–1788 (2015).
LaRue, R. S. et al. Guidelines for naming nonprimate APOBEC3 genes and proteins. J. Virol. 83, 494–497 (2009).
Nakashima, M. et al. Mapping region of human restriction factor APOBEC3H critical for interaction with HIV-1 Vif. J. Mol. Biol. 429, 1262–1276 (2017).
Huthoff, H. & Malim, M. H. Identification of amino acid residues in APOBEC3G required for regulation by human immunodeficiency virus type 1 Vif and Virion encapsidation. J. Virol. 81, 3807–3815 (2007).
Bogerd, H. P., Doehle, B. P., Wiegand, H. L. & Cullen, B. R. A single amino acid difference in the host APOBEC3G protein controls the primate species specificity of HIV type 1 virion infectivity factor. Proc. Natl Acad. Sci. USA 101, 3770–3774 (2004).
Mangeat, B., Turelli, P., Liao, S. & Trono, D. A single amino acid determinant governs the species-specific sensitivity of APOBEC3G to Vif action. J. Biol. Chem. 279, 14481–14483 (2004).
Schrofelbauer, B., Chen, D. & Landau, N. R. A single amino acid of APOBEC3G controls its species-specific interaction with virion infectivity factor (Vif). Proc. Natl Acad. Sci. USA 101, 3927–3932 (2004).
Xu, H. et al. A single amino acid substitution in human APOBEC3G antiretroviral enzyme confers resistance to HIV-1 virion infectivity factor-induced depletion. Proc. Natl Acad. Sci. USA 101, 5652–5657 (2004).
Kouno, T. et al. Structure of the Vif-binding domain of the antiviral enzyme APOBEC3G. Nat. Struct. Mol. Biol. 22, 485–491 (2015).
Britan-Rosich, E., Nowarski, R. & Kotler, M. Multifaceted counter-APOBEC3G mechanisms employed by HIV-1 Vif. J. Mol. Biol. 410, 1065–1076 (2011).
Feng, Y., Love, R. P. & Chelico, L. HIV-1 viral infectivity factor (Vif) alters processive single-stranded DNA scanning of the retroviral restriction factor APOBEC3G. J. Biol. Chem. 288, 6083–6094 (2013).
Duda, D. M. et al. Structural insights into NEDD8 activation of cullin-RING ligases: conformational control of conjugation. Cell 134, 995–1006 (2008).
Mehle, A., Goncalves, J., Santa-Marta, M., McPike, M. & Gabuzda, D. Phosphorylation of a novel SOCS-box regulates assembly of the HIV-1 Vif-Cul5 complex that promotes APOBEC3G degradation. Genes Dev. 18, 2861–2866 (2004).
Harris, R. S. & Anderson, B. D. Evolutionary paradigms from ancient and ongoing conflicts between the lentiviral Vif protein and mammalian APOBEC3 enzymes. PLoS Pathog. 12, e1005958 (2016).
Hache, G., Shindo, K., Albin, J. S. & Harris, R. S. Evolution of HIV-1 isolates that use a novel Vif-independent mechanism to resist restriction by human APOBEC3G. Curr. Biol. 18, 819–824 (2008).
Albin, J. S., Hache, G., Hultquist, J. F., Brown, W. L. & Harris, R. S. Long-term restriction by APOBEC3F selects human immunodeficiency virus type 1 variants with restored Vif function. J. Virol. 84, 10209–10219 (2010).
Xiao, X., Li, S. X., Yang, H. & Chen, X. S. Crystal structures of APOBEC3G N-domain alone and its complex with DNA. Nat. Commun. 7, 12193 (2016).
Smith, J. L., Izumi, T., Borbet, T. C., Hagedorn, A. N. & Pathak, V. K. HIV-1 and HIV-2 Vif interact with human APOBEC3 proteins using completely different determinants. J Virol. 88, 9893–9908 (2014).
Nguyen, K. L. et al. Codon optimization of the HIV-1 vpu and vif genes stabilizes their mRNA and allows for highly efficient Rev-independent expression. Virology 319, 163–175 (2004).
Yee, J. K., Friedmann, T. & Burns, J. C. Generation of high-titer pseudotyped retroviral vectors with very broad host range. Methods Cell Biol. 43, 99–112 (1994).
Unutmaz, D., KewalRamani, V. N., Marmon, S. & Littman, D. R. Cytokine signals are sufficient for HIV-1 infection of resting human T lymphocytes. J. Exp. Med. 189, 1735–1746 (1999).
Desimmie, B. A., Smith, J. L., Matsuo, H., Hu, W. S. & Pathak, V. K. Identification of a tripartite interaction between the N-terminus of HIV-1 Vif and CBFβ that is critical for Vif function. Retrovirology 14, 19 (2017).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J .Struct. Biol. 180, 519–530 (2012).
Scheres, S. H. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nat. Methods 9, 853–854 (2012).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D Biol. Crystallogr. 67, 355–367 (2011).
Afonine, P. V. et al. New tools for the analysis and validation of cryo-EM maps and atomic models. Acta Crystallogr. D Struct. Biol. 74, 814–840 (2018).
Acknowledgements
We thank S. Wu, M. Llaguno and X. Liu at Yale Cryo-EM facilities and C. Wang and J. Liu for assistance with data collection. We thank K. Knecht, O. Buzovetsky, S. C. Devarkar and other Xiong lab members for discussions. This work was supported by National Institutes of Health grant AI116313 (Y.X.). This work was supported in part by the Intramural Research Program of the NIH, National Cancer Institute, Center for Cancer Research, and by an Intramural AIDS Targeted Antiviral Program grant and the Innovation Fund, Office of AIDS Research, NIH to V.K.P.
Author information
Authors and Affiliations
Contributions
Y.X., V.K.P., Y.H., and B.A.D. designed the experiments. Y.H. performed the biophysical and biochemical experiments, and B.A.D. performed the virological experiments. Data were analyzed by Y.X., Y.H., V.K.P., and B.A.D.; H.C.N., S.J.Z., T.C.C., J.C., J.W., H.W., and K.Z. contributed to experiments and discussions. Y.X., Y.H., V.K.P., and B.A.D. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Peer review information Inês Chen was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
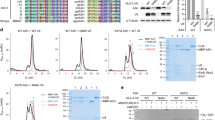
Extended Data Fig. 1 Biochemical and cellular characterizations of various Vif–CBFβ–A3FCTDm assemblies and the A3FCTDm-CBFβ interaction.
a, The Vif–CBFβ–A3FCTDm fusion complex with or without the Vif α-domain and corresponding interacting CBFβ C terminus stays as a tetramer in low-salt solution. The unfused Vif–CBFβ–A3FCTDm without these regions switches from monomer to tetramer at high protein concentration (146 μΜ loading concentration). b, No obvious shift for the elution peak was observed upon incubation of CBFβ and A3FCTDm compared to the CBFβ alone or A3FCTDm alone. The SDS-PAGE analysis of the peak fractions of CBFβ alone, A3FCTDm alone and CBFβ/A3FCTDm mixture is indicated. c, Co-immunoprecipitation (Co-IP) analysis of the interaction between A3F and CBFβ in the presence or absence of Vif in cells. Flag-A3F and CBFβ-myc were cotransfected with or without Vif-HA and co-immunoprecipitated using an anti-Flag antibody. In the absence of Vif, no binary A3F and CBFβ binding was observed. A representative blot from two independent experiments was shown.
Extended Data Fig. 2 Cryo-EM study of the Vif–CBFβ–A3FCTDm complex with or without the Vif α-domain and the corresponding interacting region of the CBFβ C-terminus.
a, The 5-Å cryo-EM reconstruction of Vif–CBFβ–A3FCTDm with (right) or without (left) the docked-in model (ribbon). The density corresponding to the flexible Vif α-domain and the corresponding interacting CBFβ C terminus (circled) is not visible. b, Left, the 3.9-Å cryo-EM reconstruction of the truncated Vif–CBFβ–A3FCTDm. Right, overlay of the cryo-EM models of Vif–CBFβ–A3FCTDm ternary complexes with (magenta) and without (yellow) the Vif α-domain and the corresponding interacting CBFβ C terminus shows that the removal of these regions does not affect the architecture of the ternary complex. c, Central slices of the top 3D classes of the Vif–CBFβ–A3FCTDm complex with (upper) or without (lower) the Vif α-domain and the corresponding interacting CBFβ C terminus indicate that removing these flexible regions reduces the tetramer flexibility. The location of the Vif α-domain and the corresponding interacting CBFβ C terminus is marked by yellow arrows in the first class average.
Extended Data Fig. 3 The relative conformational changes between Vif and CBFβ upon A3FCTDm binding, shown as rainbow putty representations of superpositions.
The color spectrum and the coil thickness represent the deviation of the aligned Cα atoms in the structures, which varies from 0 Å (blue) to ~10 Å (orange). The Vif C-terminal residues 173–176 missing in the Vif–E3 ligase structure are colored in red. The Vif–CBFβ structure without A3FCTDm binding used for superposition is extracted from the Vif–E3 ligase structure (PDB 4N9F).
Extended Data Fig. 4 Effects of Vif–CBFβ–A3FCTDm tetramer on A3F-Vif interaction, Vif-mediated A3F degradation, and viral infectivity.
a, Interternary complex interfaces between Vif and A3FCTDm detected in the tetrameric complex. Center, overview of the two interternary complex interfaces (indicated with arrows) involving one Vif molecule. Vif is shown in magenta, CBFβ in cyan, and A3FCTDm in green. One ternary complex containing the major Vif-A3F interface is marked by an oval. The detailed illustrations of the two interfaces are shown on the sides, with one involving the Vif α1 helix (right), and the other involves Vif residues (orange, marked by *) in the 55VxIPLx4-5L64 motif (left). b, SEC binding assays show that the tetramer formation depends on the presence of A3FCTDm (left) but not on the solubility-enhancing mutations of A3FCTDm (right). In contrast to the Vif–CBFβ–A3FCTDm complex, which switched from monomer to tetramer when reducing salt concentration, Vif-CBFβ alone stayed as monomer. Six A3FCTDm residues (196, 247, 248, 310, 314, 315) located near the observed tetramer interfaces were reverted back to wild-type amino acids to verify that this A3FCTDm variant (with four remaining point mutations away from the interface and without any disturbance to A3F structure) retained the ability to form tetramers. c, SEC binding (left) and MBP pulldown (right) assays show that the A3FCTDm D347R mutation disrupts the tetramer formation but not the individual Vif–CBFβ–A3FCTDm ternary complex. The loading controls are shown in Supplementary Fig. 1b,c. d, The effect of D347R mutation on A3F sensitivity to Vif-mediated degradation indicated by western blot (left), quantified A3F levels relative to no Vif (mean ± s.d.; n = 4 biologically independent experiments; middle), and relative infectivity (mean ± s.d.; n = 4 biologically independent experiments; right). A3FD347R retained wild-type A3F-like sensitivity to Vif-mediated degradation, which resulted in rescue of viral infectivity.
Extended Data Fig. 5 Vif R15 is located at the C terminus of A3FCTDm α2 helix interacting with the backbone carbonyls of the helix.
a, Upon A3F binding, Vif R15 flips away from the position (gray) pointing into the molecule core to electrostatically interact with the backbone carbonyls of the A3FCTDm α2 helix rather than the side chain carboxylates of D260/D261. b, Mutational analysis of the interactions by in vitro binding assay using MBP-tagged Vif–CBFβ–EloB–EloC variants to pull down A3FCTDm variants. The D260R/D261R double mutation did not affect the Vif interaction. The loading controls are shown in Supplementary Fig. 1b,c. c, The effect of D260A/D261A or D260R/D261R double mutants on A3F sensitivity to Vif-mediated degradation indicated by western blot (left), quantified A3F levels relative to no Vif (mean ± s.d.; n = 3 biologically independent experiments; middle), and relative infectivity (mean ± s.d.; n = 3 biologically independent experiments; right). Both alanine and arginine mutants did not confer resistance, indicating that the side chains of the residues are not involved in the Vif interaction.
Extended Data Fig. 6 Analysis of the effect of Vif K50E mutation on A3G degradation in cells.
a,b, Western blot (a) and quantified A3 levels relative to ‘no Vif’ (b) show that the Vif K50E mutant could not induce A3F degradation in the presence of either CBFβ wild-type or CBFβ E54K but could induce A3G degradation (mean ± s.d.; n = 3 biologically independent experiments).
Extended Data Fig. 7 HIV1-Vif R15 does not interact with A3F E289.
a, Vif R15 is located far away from A3F E289, which interacts with Vif K50 in the Vif–CBFβ–A3FCTDm structure. b, Mutational analysis of the interactions by in vitro binding assay using MBP-tagged Vif–CBFβ–EloB–EloC variants to pull down A3FCTDm variants. The loading controls are shown in Supplementary Fig. 1b,c. The charge-swapped Vif R15E/A3F E289K double mutation did not rescue the Vif-A3F interaction in vitro. c, The effect of Vif R15E or A3F E289K mutation on A3F sensitivity to Vif-mediated degradation indicated by western blot (left), quantified A3F levels relative to ‘no Vif’ (mean ± s.d.; n = 3 biologically independent experiments; middle), and relative viral infectivity (mean ± s.d.; n = 3 biologically independent experiments; right). In contrast to the prior report37, the charge-swapped Vif R15E/A3F E289K double mutation did not restore the Vif-mediated A3F degradation or viral infectivity in cells. The blot was cut as indicated (gray arrow), where one half was used to detect Flag-A3F and HSP90, and the other half was used to detect Vif-HA.
Extended Data Fig. 8 Effect of Vif–CBFβ–A3FCTDm complex on A3FCTDm deaminase activity.
a, UDG-based deamination assay of A3FCTDm (+: 5 μM; 15×: 75 μM) in the presence or absence of different molar excesses (2×: 10 μM; 30×: 150 μM; 40×: 200 μM) of MBP-tagged Vif–CBFβ–EloB–EloC variants. The inhibition of A3FCTDm deamination activity by a large excess of Vif–CBFβ–EloB–EloC variants was not caused by nonspecific binding interactions, as the same molar amount excess (40×) of BSA did not trigger the inhibition. b, One of the Vif–CBFβ–A3FCTDm tetramer interface involving the 55VxIPLx4-5L64 motif (Extended Data Fig. 4a, left) blocks the catalytic site of A3FCTDm (red and marked by an arrow). Vif is shown in magenta, CBFβ in cyan, and A3FCTDm in green. One Vif–CBFβ–A3FCTDm ternary complex with the major interface is circled with an oval.
Extended Data Fig. 9 Parameters of the cryo-EM reconstructions of Vif–CBFβ–A3FCTDm complexes and the final model.
a, FSC curves of the half-maps from gold standard refinements of the Vif–CBFβ–A3FCTDm complex with (cyan) or without (blue) the Vif α-domain and the corresponding interacting CBFβ C terminus. The FSC curve of the map and final model of the truncated Vif–CBFβ–A3FCTDm complex is in green. Resolution of the maps are determined by the cut-off values at FSC = 0.143. b, The Euler angle distribution of the classified particles of the truncated Vif–CBFβ–A3FCTDm complex used for the final 3D reconstruction. c, Color coded local resolution estimation of the D2 symmetrized map of the truncated Vif–CBFβ–A3FCTDm complex.
Extended Data Fig. 10 Detailed illustrations of the secondary structure elements.
a−c, Detailed illustrations of the secondary structure elements of Vif176 (a), CBFβ151 (b) and A3FCTD (c). The secondary structures are annotated on primary amino acid sequences (left) and tertiary structures (right). The tertiary structures for illustration are: Vif, extracted from PDB 4N9F; CBFβ151, extracted from our cryo-EM structure; A3FCTD, PDB 3WUS.
Supplementary information
Supplementary Information
Supplementary Figure 1
Source data
Source Data Fig. 2
Unprocessed western blots and gels
Source Data Fig. 2
Statistical source data
Source Data Fig. 3
Unprocessed western blots and gels
Source Data Fig. 3
Statistical source data
Source Data Extended Data Fig. 1
Unprocessed western blots and gels
Source Data Extended Data Fig. 4
Unprocessed western blots and gels
Source Data Extended Data Fig. 4
Statistical source data
Source Data Extended Data Fig. 5
Unprocessed western blots and gels
Source Data Extended Data Fig. 5
Statistical source data
Source Data Extended Data Fig. 6
Unprocessed western blots
Extended Data Fig. 6
Statistical source data
Source Data Extended Data Fig. 7
Unprocessed western blots and gels
Source Data Extended Data Fig. 7
Statistical source data
Rights and permissions
About this article
Cite this article
Hu, Y., Desimmie, B.A., Nguyen, H.C. et al. Structural basis of antagonism of human APOBEC3F by HIV-1 Vif. Nat Struct Mol Biol 26, 1176–1183 (2019). https://doi.org/10.1038/s41594-019-0343-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-019-0343-6
- Springer Nature America, Inc.
This article is cited by
-
Structural insights into PPP2R5A degradation by HIV-1 Vif
Nature Structural & Molecular Biology (2024)
-
The structural basis for HIV-1 Vif antagonism of human APOBEC3G
Nature (2023)
-
Structural basis of HIV-1 Vif-mediated E3 ligase targeting of host APOBEC3H
Nature Communications (2023)
-
Structural insights into RNA bridging between HIV-1 Vif and antiviral factor APOBEC3G
Nature Communications (2023)
-
CUL5-ARIH2 E3-E3 ubiquitin ligase structure reveals cullin-specific NEDD8 activation
Nature Chemical Biology (2021)