Abstract
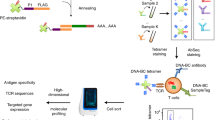
High-throughput 3′ single-cell RNA-sequencing (scRNA-seq) allows cost-effective, detailed characterization of individual immune cells from tissues. Current techniques, however, are limited in their ability to elucidate essential immune cell features, including variable sequences of T cell antigen receptors (TCRs) that confer antigen specificity. Here, we present a strategy that enables simultaneous analysis of TCR sequences and corresponding full transcriptomes from 3′-barcoded scRNA-seq samples. This approach is compatible with common 3′ scRNA-seq methods, and adaptable to processed samples post hoc. We applied the technique to identify transcriptional signatures associated with T cells sharing common TCRs from immunized mice and from patients with food allergy. We observed preferential phenotypes among subsets of expanded clonotypes, including type 2 helper CD4+ T cell (TH2) states associated with food allergy. These results demonstrate the utility of our method when studying diseases in which clonotype-driven responses are critical to understanding the underlying biology.




Similar content being viewed by others
Data availability
FASTQ file format data related to murine samples are available through GEO and BioProject under accession numbers GSE136028 and PRJNA560970. FASTQ file format data related to human samples is available through dbGaP under accession number phs001897.v1.p1. Source data files and associated metadata tables for Figs. 2–4 are available on http://shaleklab.com/resources/, https://github.com/mitlovelab/ or upon request. MSigDB results for Extended Data Fig. 1c, Extended Data Fig. 5a and Supplementary Fig. 3h are available as Supplementary Table 12. Full results from the differential gene expression comparison shown in Extended Data Fig. 2c are available as Supplementary Table 13. All recovered CDR3 sequences, and their frequencies, of TCR α and β chains from E7-immunized mice and patients with peanut allergy are available as Supplementary Tables 6 and 10. Gene expression matrices for E7-immunized mice and patients with peanut allergy are available as Supplementary Data 1 and 2.
Code availability
R scripts for generating all analysis, Matlab scripts for processing TCR sequencing data, as well as all updates, are available at http://shaleklab.com/resources/, https://github.com/mitlovelab/ or upon reasonable request.
References
Schrama, D., Ritter, C. & Becker, J. C. T cell receptor repertoire usage in cancer as a surrogate marker for immune responses. Semin. Immunopathol. 39, 255–268 (2017).
Lossius, A. et al. High-throughput sequencing of TCR repertoires in multiple sclerosis reveals intrathecal enrichment of EBV-reactive CD8+ T cells. Eur. J. Immunol. 44, 1–41 (2014).
Kirsch, I. R. et al. TCR sequencing facilitates diagnosis and identifies mature T cells as the cell of origin in CTCL. Sci. Transl. Med. 7, 1–13 (2015).
Carlson, C. S. et al. Using synthetic templates to design an unbiased multiplex PCR assay. Nat. Commun. 4, 2680 (2013).
Crosby, E. J. et al. Complimentary mechanisms of dual checkpoint blockade expand unique T-cell repertoires and activate adaptive anti-tumor immunity in triple-negative breast tumors. Oncoimmunology 7, e1421891 (2018).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Khodadoust, M. S. et al. Antigen presentation profiling reveals recognition of lymphoma immunoglobulin neoantigens. Nature 543, 723–727 (2017).
Avraham, R. et al. Pathogen cell-to-cell variability drives heterogeneity in host immune responses. Cell 162, 1309–1321 (2015).
Papalexi, E. & Satija, R. Single-cell RNA sequencing to explore immune cell heterogeneity. Nat. Rev. Immunol. 18, 35–45 (2017).
Shalek, A. K. et al. Single-cell transcriptomics reveals bimodality in expression and splicing in immune cells. Nature 498, 236–240 (2013).
Azizi, E. et al. Single-cell map of diverse immune phenotypes in the breast tumor microenvironment. Cell 174, 1293–1308.e36 (2018).
Zhang, L. et al. Lineage tracking reveals dynamic relationships of T cells in colorectal cancer. Nature 564, 268–272 (2018).
Han, A., Glanville, J., Hansmann, L. & Davis, M. M. Linking T-cell receptor sequence to functional phenotype at the single-cell level. Nat. Biotechnol. 32, 684–692 (2014).
Stubbington, M. J. T. et al. T cell fate and clonality inference from single-cell transcriptomes. Nat. Methods 13, 329–332 (2016).
Dash, P. et al. Paired analysis of TCRα and TCRβ chains at the single-cell level in mice. J. Clin. Invest. 121, 288–295 (2011).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Klein, A. M. et al. Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Cell 161, 1187–1201 (2015).
Gierahn, T. M. et al. Seq-well: portable, low-cost RNA sequencing of single cells at high throughput. Nat. Methods 14, 395–398 (2017).
Saikia, M. et al. Simultaneous multiplexed amplicon sequencing and transcriptome profiling in single cells. Nat. Methods 16, 59–62 (2019).
Singh, M. et al. High-throughput targeted long-read single cell sequencing reveals the clonal and transcriptional landscape of lymphocytes. Nat. Commun. 10, 3120 (2019).
Zemmour, D. et al. Single-cell gene expression reveals a landscape of regulatory T cell phenotypes shaped by the TCR. Nat. Immunol. 19, 291–301 (2018).
Jain, M. et al. Nanopore sequencing and assembly of a human genome with ultra-long reads. Nat. Biotechnol. 36, 338–345 (2018).
Gupta, I. et al. Single-cell isoform RNA sequencing characterizes isoforms in thousands of cerebellar cells. Nat. Biotechnol. 36, 1197–1202 (2018).
Hughes, T. K. et al. Highly efficient, massively-parallel single-cell RNA-seq reveals cellular states and molecular features of human skin pathology. Preprint at bioRxiv https://doi.org/10.1101/689273 (2019).
Blüthmann, H. et al. T-cell-specific deletion of T-cell receptor transgenes allows functional rearrangement of endogenous α- and β-genes. Nature 334, 156–159 (1988).
Dash, P. et al. Quantifiable predictive features define epitope-specific T cell receptor repertoires. Nature 547, 89–93 (2017).
Mousset, C. M. et al. Comprehensive phenotyping of T cells using flow cytometry. Cytometry A 95, 647–654 (2019).
Farber, D. L., Yudanin, N. A. & Restifo, N. P. Human memory T cells: generation, compartmentalization and homeostasis. Nat. Rev. Immunol. 14, 24–35 (2014).
Singer, M. et al. A distinct gene module for dysfunction uncoupled from activation in tumor-infiltrating T cells. Cell 166, 1500–1511.e9 (2016).
Huang, W. & August, A. The signaling symphony: T cell receptor tunes cytokine-mediated T cell differentiation. J. Leukoc. Biol. 97, 477–485 (2015).
Padovan, E. et al. Expression of two T cell receptor alpha chains: dual receptor T cells. Science 262, 422–424 (1993).
Bacher, P. & Scheffold, A. Flow-cytometric analysis of rare antigen-specific T cells. Cytometry A 83A, 692–701 (2013).
Chattopadhyay, P. K., Yu, J. & Roederer, M. Live-cell assay to detect antigen-specific CD4+ T-cell responses by CD154 expression. Nat. Protoc. 1, 1–6 (2006).
Syed, A., Kohli, A. & Nadeau, K. C. Food allergy diagnosis and therapy: where are we now? Immunotherapy 5, 931–944 (2013).
Seumois, G. et al. Transcriptional profiling of Th2 cells identifies pathogenic features associated with asthma. J. Immunol. 197, 655–664 (2016).
Mueller, S. N., Gebhardt, T., Carbone, F. R. & Heath, W. R. Memory T cell subsets, migration patterns, and tissue residence. Annu. Rev. Immunol. 31, 137–161 (2013).
Nish, S. A. et al. CD4+ T cell effector commitment coupled to self-renewal by asymmetric cell divisions. J. Exp. Med. 214, 39–47 (2017).
Wei, G. et al. Global mapping of H3K4me3 and H3K27me3 reveals specificity and plasticity in lineage fate determination of differentiating CD4+ T cells. Immunity 30, 155–167 (2009).
Foletta, V. C., Segal, D. H. & Cohen, D. R. Transcriptional regulation in the immune system: all roads lead to AP-1. J. Leukoc. Biol. 63, 139–152 (1998).
Müller, U. et al. Lack of IL-4 receptor expression on T helper cells reduces T helper 2 cell polyfunctionality and confers resistance in allergic bronchopulmonary mycosis. Mucosal Immunol. 5, 299–310 (2012).
Upadhyaya, B., Yin, Y., Hill, B. J., Douek, D. C. & Prussin, C. Hierarchical IL-5 expression defines a subpopulation of highly differentiated human Th2 cells. J. Immunol. 187, 3111–3120 (2011).
Ritvo, P.-G. et al. High-resolution repertoire analysis reveals a major bystander activation of Tfh and Tfr cells. Proc. Natl Acad. Sci. USA 115, 9604–9609 (2018).
Raj, A. & van Oudenaarden, A. Nature, nurture, or chance: stochastic gene expression and its consequences. Cell 135, 216–226 (2008).
Han, Q. et al. Polyfunctional responses by human T cells result from sequential release of cytokines. Proc. Natl Acad. Sci. USA 109, 1607–1612 (2012).
Stoeckius, M. et al. Cell hashing with barcoded antibodies enables multiplexing and doublet detection for single cell genomics. Genome Biol. 19, 224 (2018).
Schumacher, T. N. M., Gerlach, C. & van Heijst, J. W. J. Mapping the life histories of T cells. Nat. Rev. Immunol. 10, 621–631 (2010).
Rosati, E. et al. Overview of methodologies for T-cell receptor repertoire analysis. BMC Biotechnol. 17, 61 (2017).
Hermiston, M. L., Xu, Z. & Weiss, A. CD45: a critical regulator of signaling thresholds in immune cells. Annu. Rev. Immunol. 21, 107–137 (2003).
Rohland, N. & Reich, D. Cost-effective, high-throughput DNA sequencing libraries for multiplexed target capture. Genome Res. 22, 939–946 (2012).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Liberzon, A. et al. The Molecular Signatures Database hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Gu, Z., Gu, L., Eils, R., Schlesner, M. & Brors, B. circlize implements and enhances circular visualization in R. Bioinformatics 30, 2811–2812 (2014).
Acknowledgements
We thank P. Blainey and M. Birnbaum for fruitful discussions. This work was supported in part by the Koch Institute Support (core) NIH Grant P30-CA14051 from the National Cancer Institute, as well as the Koch Institute - Dana-Farber/Harvard Cancer Center Bridge Project. This work was also supported by the Food Allergy Science Initiative at the Broad Institute and the NIH (5P01AI039671, 5U19AI089992, U19AI095261). A.K.S. was supported by the Searle Scholars Program, the Beckman Young Investigator Program, the Pew-Stewart Scholars Program for Cancer Research, a Sloan Fellowship in Chemistry, the NIH (1DP2GM119419, 2U19AI089992, 2R01HL095791, 1U54CA217377, 2P01AI039671, 5U24AI118672, 2RM1HG006193, 1R33CA202820, 1R01AI138546, 1R01HL134539, 1R01DA046277, 1U2CCA23319501) and Agilent Technologies.
Author information
Authors and Affiliations
Contributions
A.A.T., T.M.G, A.K.S. and J.C.L. developed the concepts and designed the study. A.A.T. and T.M.G. performed the experiments. A.A.T. prepared the manuscript with input from all authors. A.A.T., T.M.G., B.M. and D.M.M. performed bioinformatics analyses. N.K.M. performed E7 immunization of mice. W.G.S. designed the clinical study and provided samples from peanut-allergic patients. B.R. performed stimulation and sorting of PBMCs from allergic patients.
Corresponding authors
Ethics declarations
Competing interests
A.A.T., T.M.G., J.C.L. and the Massachusetts Institute of Technology have filed a patent application (patent no. PCT/US2018/013443) that relates to T cell receptor recovery, compositions of matter and the outlined experimental and computational methods and uses thereof. J.C.L. and A.K.S. are co-founders and shareholders of Honeycomb Biotechnologies, Inc. T.M.G. is currently an employee of Honeycomb Biotechnologies, Inc.
Additional information
Peer review information Laurie A. Dempsey was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Stimulated and ex vivo cells are transcriptionally distinct.
a, tSNE visualization of all cells colored based on stimulation condition (n = 6,912 stimulated cells, dark grey; 7,512 ex vivo cells, light grey). b, Proportions of stimulated and ex vivo cells in each of the computationally determined clusters shown in Fig. 3a. Dash line indicates expected proportions assuming even distributions of cells from both conditions. c, Enriched MsigDb signatures of the four modules of genes identified in Fig. 3d. FDR q-values represent Benjamini and Hochberg-corrected, one-tailed hypergeometric P values. 50, 49, 35, and 48 genes are included in Module 1, 2, 3, and 4, respectively for enrichment calculation. See Supplementary Table 12 for more details. Data represent combined data from four independent experiments of four mice total (a-c).
Extended Data Fig. 2 Group 1 and 2 clonotypes differ in expansion and gene expression upon stimulation.
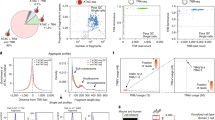
a, Clonal sizes of Group 1 and 2 clonotypes in the stimulated and ex vivo conditions shown in Fig. 3d. P value calculated by two-sample Mann-Whitney U test (Stimulated: n = 74 clonotype groups in Group 1; 37 Group 2 clonotypes. ex vivo: n = 82 Group 1 clonotypes; 40 Group 2 clonotypes). Box and whisker plots indicate the (box) 25th and 75th percentile along with (whisker) + /− 1.5*interquartile range. Violin plots represent estimated density of clonotypes. b, Gene expression fold changes between stimulated and ex vivo cells in (x-axis) Group 1 clonotypes and (y-axis) Group 2 clonotypes. Each point represents a shared gene across Group 1 and 2 clonotypes. Red line indicates fitted linear model. P value calculated by one-tailed F statistics (F(1,5908)) of the linear regression. n = 5908 genes. c, Volcano plots of differentially expressed genes between Group 1 and 2 clonotypes in the (left) stimulated and the (right) ex vivo conditions. P values were determined using a two-tailed likelihood ratio test, and adjusted by Bonferroni correction. Top 10 genes with positive or negative fold changes are labeled. Cells in Group 1 and 2 have been downsampled to 300 each (n = 300 cells for each of the groups). See Supplementary Table 13 for more details.
Extended Data Fig. 3 Analysis of shared clonotypes across four E7-HPV immunized mice.
a, Venn diagram of shared unique Tcrb clones across the four mice. b, Amino acid logo plot of TCRβ sequences that show high similarity among public clones shared by at least three of four animals. Individual sequences are shown in c. c, TCRα and β matching of highly similar clonotypes found in public clones shared by at least three of four animals (Supplementary Table 9). Bold outline indicates dual TCRα chains found in the same cells (Supplementary Table 3). Structural amino acids shown in grey. Number of cells shown in parenthesis. TCRα and β sequences that were detected in less than two cells were excluded for visualization.
Extended Data Fig. 4 Distinct patterns of gene expression correlate with pseudotime.
a, Expression of top 100 most significant genes visualized across pseudotime. Genes were clustered via Ward.D2 based on their patterns of expression. Data represent an individual experiment with 1847 single-cells from one patient (patient 77).
Extended Data Fig. 5 Psuedotime correlates with effector T cell signatures.
a, MsigDB analysis of genes enriched early (cluster 3 in Extended Data Fig 4a; n = 38 genes) or late (cluster 4 in Extended Data Fig 4a; n = 123 genes) on the pseudotemporal trajectory. Description indicates cell state enriched with the corresponding gene set in comparison to another cell state. FDR q-values represent Benjamini and Hochberg-corrected, one-tailed hypergeometric P values. See Supplementary Table 12 for more details. b, Pseudotime distribution of expanded clones shown in Fig. 4d. Number of cells for each clonotype group indicated in parenthesis. A total of 16 clonotype groups are shown. All box and whisker plots indicate the (box) 25th and 75th percentile along with (whisker) + /− 1.5*interquartile range (b). Data represent an individual experiment with 1847 single-cells from one patient (patient 77; a,b).
Supplementary information
Supplementary Information
Supplementary Figs. 1–5 and Tables 1,2,4,5,8 and 9.
Supplementary Tables
Supplementary Tables 3,6,7 and 10–13.
Supplementary Data 1
Digital gene expression matrices for data shown in Fig. 3.
Supplementary Data 2
Digital gene expression matrices for data shown in Fig. 4.
Rights and permissions
About this article
Cite this article
Tu, A.A., Gierahn, T.M., Monian, B. et al. TCR sequencing paired with massively parallel 3′ RNA-seq reveals clonotypic T cell signatures. Nat Immunol 20, 1692–1699 (2019). https://doi.org/10.1038/s41590-019-0544-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-019-0544-5
- Springer Nature America, Inc.
This article is cited by
-
Nonlinear dimensionality reduction based visualization of single-cell RNA sequencing data
Journal of Analytical Science and Technology (2024)
-
T-cell immunity induced and reshaped by an anti-HPV immuno-oncotherapeutic lentiviral vector
npj Vaccines (2024)
-
Evolution of T cells in the cancer-resistant naked mole-rat
Nature Communications (2024)
-
Single-cell analysis in rheumatic and allergic diseases: insights for clinical practice
Nature Reviews Immunology (2024)
-
Single cell RNA-seq: a novel tool to unravel virus-host interplay
VirusDisease (2024)