Abstract
The fundamental and assorted roles of β-1,3-glucans in nature are underpinned on diverse chemistry and molecular structures, demanding sophisticated and intricate enzymatic systems for their processing. In this work, the selectivity and modes of action of a glycoside hydrolase family active on β-1,3-glucans were systematically investigated combining sequence similarity network, phylogeny, X-ray crystallography, enzyme kinetics, mutagenesis and molecular dynamics. This family exhibits a minimalist and versatile (α/β)-barrel scaffold, which can harbor distinguishing exo or endo modes of action, including an ancillary-binding site for the anchoring of triple-helical β-1,3-glucans. The substrate binding occurs via a hydrophobic knuckle complementary to the canonical curved conformation of β-1,3-glucans or through a substrate conformational change imposed by the active-site topology of some fungal enzymes. Together, these findings expand our understanding of the enzymatic arsenal of bacteria and fungi for the breakdown and modification of β-1,3-glucans, which can be exploited for biotechnological applications.






Similar content being viewed by others
Data availability
Structural data have been deposited in the Protein Data Bank (https://www.rcsb.org/) under accession codes 6UAQ (AmGH128_I), 6UAR (AmGH128_I, L3), 6UAS (AmGH128_I/E199A, L5+GLC), 6UFL (AmGH128_I/E199Q, L6), 6UFZ (AmGH128_I/E199Q), 6UAT (AmGH128_I/E102A, L5), 6UAU (AmGH128_I/E102A, L3 + L2), 6UAV (PvGH128_II), 6UAW (PvGH128_II, L3), 6UAX (ScGH128_II), 6UAY (BgGH128_III), 6UAZ (BgGH128_III, GLC), 6UB0 (BgGH128_III, L2), 6UB1 (BgGH128_III, L3), 6UB2 (LeGH128_IV), 6UB3 (LeGH128_IV, L2), 6UB4 (LeGH128_IV, L3(C2)), 6UB5 (LeGH128_IV, L3 (P21)), 6UB6 (LeGH128_IV, L4), 6UB7 (CnGH128_V), 6UB8 (AnGH128_VI), 6UBA (AnGH128_VI, L3), 6UBB (AnGH128_VI, exo-site), 6UBC (CnGH128_VII) and 6UBD (TgGH128_VII). All other data generated or analyzed during this study are included in this published article (and its Supplementary information files) or are available from the corresponding author on reasonable request.
Change history
25 June 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41589-020-0590-1
References
Stone, B. A. in Chemistry, Biochemistry, and Biology of 1-3 Beta Glucans and Related Polysaccharides (eds Bacic, A., Fincher, G. B. & Stone, B. A.) 5–46 (Academic Press, 2009).
Gidley, M. J. & Nishinari, K. in Chemistry, Biochemistry, and Biology of 1-3 Beta Glucans and Related Polysaccharides 47–118 (Academic Press, 2009).
McIntosh, M., Stone, B. A. & Stanisich, V. A. Curdlan and other bacterial (1–>3)-beta-d-glucans. Appl. Microbiol. Biotechnol. 68, 163–173 (2005).
Kang, X. et al. Molecular architecture of fungal cell walls revealed by solid-state NMR. Nat. Commun. 9, 2747 (2018).
Helbert, W. et al. Discovery of novel carbohydrate-active enzymes through the rational exploration of the protein sequences space. Proc. Natl Acad. Sci. USA 116, 6063–6068 (2019).
Ishida, T. et al. Crystal structure of glycoside hydrolase family 55 β-1,3-glucanase from the basidiomycete Phanerochaete chrysosporium. J. Biol. Chem. 284, 10100–10109 (2009).
Bianchetti, C. M. et al. Active site and laminarin binding in glycoside hydrolase family 55. J. Biol. Chem. 290, 11819–11832 (2015).
Papageorgiou, A. C., Chen, J. & Li, D. Crystal structure and biological implications of a glycoside hydrolase family 55 beta-1,3-glucanase from Chaetomium thermophilum. Biochim. Biophys. Acta Proteins Proteom. 1865, 1030–1038 (2017).
Wu, H. M. et al. Structure, mechanistic action, and essential residues of a GH-64 enzyme, laminaripentaose-producing beta-1,3-glucanase. J. Biol. Chem. 284, 26708–26715 (2009).
Qin, Z. et al. The recognition mechanism of triple-helical β-1,3-glucan by a β-1,3-glucanase. Chem. Commun. 53, 9368–9371 (2017).
Zhou, P. et al. The structure of a glycoside hydrolase family 81 endo-beta-1,3-glucanase. Acta Crystallogr. D. 69, 2027–2038 (2013).
Pluvinage, B., Fillo, A., Massel, P. & Boraston, A. B. Structural analysis of a family 81 glycoside hydrolase implicates its recognition of beta-1,3-glucan quaternary. Structure 25, 1348–1359 e3 (2017).
Sakamoto, Y., Nakade, K. & Konno, N. Endo-beta-1,3-glucanase GLU1, from the fruiting body of Lentinula edodes, belongs to a new glycoside hydrolase family. Appl. Environ. Microbiol. 77, 8350–8354 (2011).
Masuda, T. et al. Subatomic structure of hyper-sweet thaumatin D21N mutant reveals the importance of flexible conformations for enhanced sweetness. Biochimie 157, 57–63 (2019).
Terrapon, N. et al. PULDB: the expanded database of polysaccharide utilization loci. Nucleic Acids Res. 46, D677–D683 (2018).
Boraston, A. B., Warren, R. A. & Kilburn, D. G. Beta-1,3-glucan binding by a thermostable carbohydrate-binding module from thermotoga maritima. Biochemistry 40, 14679–14685 (2001).
van Bueren, A. L., Morland, C., Gilbert, H. J. & Boraston, A. B. Family 6 carbohydrate binding modules recognize the non-reducing end of beta-1,3-linked glucans by presenting a unique ligand binding surface. J. Biol. Chem. 280, 530–537 (2005).
Jam, M. et al. Unraveling the multivalent binding of a marine family 6 carbohydrate-binding module with its native laminarin ligand. FEBS J. 283, 1863–1879 (2016).
Brunecky, R. et al. Revealing nature’s cellulase diversity: the digestion mechanism of Caldicellulosiruptor bescii CelA. Science 342, 1513–1516 (2013).
Henrissat, B. et al. Conserved catalytic machinery and the prediction of a common fold for several families of glycosyl hydrolases. Proc. Natl Acad. Sci. USA 92, 7090–7094 (1995).
Koshland, D. E. Jr Stereochemistry and the mechanism of enzymatic reactions. Bio. Rev. 28, 416–436 (1953).
Davies, G. & Henrissat, B. Structures and mechanisms of glycosyl hydrolases. Structure 3, 853–859 (1995).
Kim, H. W. & Ishikawa, K. Functional analysis of hyperthermophilic endocellulase from Pyrococcus horikoshii by crystallographic snapshots. Biochem. J. 437, 223–230 (2011).
Planas, A. Bacterial 1,3-1,4-beta-glucanases: structure, function and protein engineering. Biochim. Biophys. Acta 1543, 361–382 (2000).
Gloster, T. M. et al. Characterization and three-dimensional structures of two distinct bacterial xyloglucanases from families GH5 and GH12. J. Biol. Chem. 282, 19177–19189 (2007).
Gueguen, Y., Voorhorst, W. G., van der Oost, J. & de Vos, W. M. Molecular and biochemical characterization of an endo-beta-1,3- glucanase of the hyperthermophilic Archaeon pyrococcus furiosus. J. Biol. Chem. 272, 31258–31264 (1997).
Kumagai, Y. & Ojima, T. Isolation and characterization of two types of beta-1,3-glucanases from the common sea hare Aplysia kurodai. Comp. Biochem. Physiol. B. 155, 138–144 (2010).
Nakabayashi, M. et al. Structure of the gene encoding laminaripentaose-producing β-1,3-glucanase (LPHase) of Streptomyces matensis DIC-108. J. Ferment. Bioeng. 85, 459–464 (1998).
Jamois, F. et al. Glucan-like synthetic oligosaccharides: iterative synthesis of linear oligo-beta-(1,3)-glucans and immunostimulatory effects. Glycobiology 15, 393–407 (2005).
Miyanishi, N., Iwamoto, Y., Watanabe, E. & Odaz, T. Induction of TNF-alpha production from human peripheral blood monocytes with beta-1,3-glucan oligomer prepared from laminarin with beta-1,3-glucanase from Bacillus clausii NM-1. J. Biosci. Bioeng. 95, 192–195 (2003).
Cockburn, D. & Svensson, B. in Carbohydrate Chemistry Vol. 39, 204–221 (The Royal Society of Chemistry, 2013).
Viborg, A. H. et al. A subfamily roadmap for functional glycogenomics of the evolutionarily diverse glycoside hydrolase family 16 (GH16). J. Biol. Chem. 294, 15973–15986 (2019).
Juge, N., Payan, F. & Williamson, G. XIP-I, a xylanase inhibitor protein from wheat: a novel protein function. Biochim. Biophys. Acta 1696, 203–211 (2004).
Patil, D. N. et al. Structural investigation of a novel N-acetyl glucosamine binding chi-lectin which reveals evolutionary relationship with class III chitinases. PLoS ONE 8, e63779 (2013).
Gerlt, J. A. et al. Enzyme function initiative-enzyme similarity tool (EFI-EST): a web tool for generating protein sequence similarity networks. Biochim. Biophys. Acta 1854, 1019–1037 (2015).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Finn, R. D., Clements, J. & Eddy, S. R. HMMER web server: interactive sequence similarity searching. Nucleic Acids Res. 39, W29–W37 (2011).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152 (2012).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree: computing large minimum evolution trees with profiles instead of a distance matrix. Mol. Biol. Evol. 26, 1641–1650 (2009).
Nandi, L. G., Guerra, J. P., Bellettini, I. C., Machado, V. G. & Minatti, E. Properties of aqueous solutions of lentinan in the absence and presence of zwitterionic surfactants. Carbohydr. Polym. 98, 1–7 (2013).
Miller, G. L. Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal. Chem. 31, 426–428 (1959).
Dos Santos, C. R. et al. The mechanism by which a distinguishing arabinofuranosidase can cope with internal di-substitutions in arabinoxylans. Biotechnol. Biofuels 11, 223 (2018).
Dauter, Z., Dauter, M. & Rajashankar, K. R. Novel approach to phasing proteins: derivatization by short cryo-soaking with halides. Acta Crystallogr. D. 56, 232–237 (2000).
Kabsch, W. XDS. Acta Crystallogr. D. 66, 125–132 (2010).
Sheldrick, G. M. A short history of SHELX. Acta Crystallogr. A. 64, 112–122 (2008).
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D. 58, 1948–1954 (2002).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. 66, 486–501 (2010).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D. 53, 240–255 (1997).
Agirre, J. et al. Privateer: software for the conformational validation of carbohydrate structures. Nat. Struct. Mol. Biol. 22, 833–834 (2015).
Franke, D. et al. ATSAS 2.8: a comprehensive data analysis suite for small-angle scattering from macromolecular solutions. J. Appl. Crystallogr. 50, 1212–1225 (2017).
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
Lee, S. et al. CHARMM36 united atom chain model for lipids and surfactants. J. Phys. Chem. B. 118, 547–556 (2014).
Jorgensen, W., Chandrasekhar, J., Madura, J., Impey, R. & Klein, M. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Martinez, L., Andrade, R., Birgin, E. G. & Martinez, J. M. PACKMOL: a package for building initial configurations for molecular dynamics simulations. J. Comput. Chem. 30, 2157–2164 (2009).
Anandakrishnan, R., Aguilar, B. & Onufriev, A. V. H++ 3.0: automating pK prediction and the preparation of biomolecular structures for atomistic molecular modeling and simulations. Nucleic Acids Res. 40, W537–W541 (2012).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: an N⋅log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Feller, S. E., Zhang, Y., Pastor, R. W. & Brooks, B. R. Constant pressure molecular dynamics simulation: the Langevin piston method. J. Chem. Phys. 103, 4613–4621 (1995).
Hoover, W. G. Constant-pressure equations of motion. Phys. Rev. A. 34, 2499–2500 (1986).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Acknowledgements
We thank M.L. Sforça and J.A. Aricetti for the support in data collection of 1H-NMR spectra and lentinan purification, respectively. We thank C.S. Farah for the access and support in the operation of the rotating anode X-ray generator available at the Chemistry Institute, University of São Paulo. Use of the Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, is supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences under Contract No. DE-AC02-76SF00515. The Stanford Synchrotron Radiation Lights Structural Molecular Biology Program is supported by the Department of Energy Office of Biological and Environmental Research, and by the National Institutes of Health, National Institute of General Medical Sciences (P41GM103393). The contents of this publication are solely the responsibility of the authors and do not necessarily represent the official views of the NIGMS or NIH. We acknowledge the LNLS for the provision of time on the MX2, SAXS1 and SAXS2 beamlines, the Brazilian Biosciences National Laboratory (LNBio) for accessibility to crystallization (Robolab), NMR and spectroscopy facilities and the Brazilian Biorenewables National Laboratory (LNBR) for the use of the characterization of macromolecules facility. LNLS, LNBio and LNBR are operated by the Brazilian Center for Research in Energy and Materials for the Brazilian Ministry for Science, Technology, Innovations and Communications. This research was supported by grants from Fundação de Amparo à Pesquisa do Estado de São Paulo (grant nos. 2015/26982-0 to M.T.M. and 2013/08293-7 to M.S.S.) and Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) (grant no. 306135/2016-7 to M.T.M.).
Author information
Authors and Affiliations
Contributions
M.T.M. initiated the study and directed the project. S.E.T.G., E.T.P. and M.S.S. devised and performed the molecular dynamic simulations. P.A.C.R.C. and G.F.P. performed the bioinformatics analyses. R.A.S.P. and F.C.G. carried out the mass spectrometry analyses. C.R.S., P.A.C.R.C., E.A.L., F.M., P.S.V., T.L.R.C., L.C., R.L.C., M.P.M., M.N.D., B.P.S. and A.T.J. expressed and purified the enzymes and performed the structural and functional characterization. C.R.S., P.S.V., G.F.P., M.S.S. and M.T.M. wrote the paper with input from P.A.C.R.C. and T.L.R.C. All authors analyzed the results and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
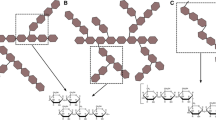
Extended Data Fig. 1 Domain organization in the GH128 family.
Domain architecture according to PFAM annotation. Subgroups in which the modular organization is present are indicated in roman numbers. Glyco_hydro_cc (PF11790), Sig-70 (PF04542), CBM6 (PF03422), F5_F8 (PF00754), WSC (PF01822), CBM4 (PF02018), Lectin (PF00652), DUFF (PF18099), Glyco_hydro_16 (PF00722) and PDK (PF00801).
Extended Data Fig. 2 Surface-binding sites in the subgroups IV and VI.
Comparison of the surface-binding sites in the subgroups IV (a and b) and VI (c and d). Relative position of the surface-binding sites to the nearest substrate-binding site (a and c). Protein-carbohydrate interactions (b and d).
Supplementary information
Supplementary Information
Supplementary Tables 1–11 and Figs. 1–30.
Rights and permissions
About this article
Cite this article
Santos, C.R., Costa, P.A.C.R., Vieira, P.S. et al. Structural insights into β-1,3-glucan cleavage by a glycoside hydrolase family. Nat Chem Biol 16, 920–929 (2020). https://doi.org/10.1038/s41589-020-0554-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-0554-5
- Springer Nature America, Inc.
This article is cited by
-
Investigating diversity and similarity between CBM13 modules and ricin-B lectin domains using sequence similarity networks
BMC Genomics (2024)
-
Structural insights into curdlan degradation via a glycoside hydrolase containing a disruptive carbohydrate-binding module
Biotechnology for Biofuels and Bioproducts (2024)
-
Gut microbiome of the largest living rodent harbors unprecedented enzymatic systems to degrade plant polysaccharides
Nature Communications (2022)
-
Enzymes knuckle down to the job
Nature Chemical Biology (2020)