Abstract
Cyclic β-1,2-glucan synthase (CGS) is a key enzyme in production of cyclic β-1,2-glucans (CβGs) which are involved in bacterial infection or symbiosis to host organisms. Nevertheless, a mechanism of cyclization, the final step in the CGS reaction, has not been fully understood. Here we performed functional and structural analyses of the cyclization domain of CGS alone from Thermoanaerobacter italicus (TiCGSCy). We first found that β-glucosidase-resistant compounds are produced by TiCGSCy with linear β-1,2-glucans as substrates. The 1H-NMR analysis revealed that these products are CβGs. Next, action pattern analyses using β-1,2-glucooligosaccharides revealed a unique reaction pattern: exclusive transglycosylation without hydrolysis and a hexasaccharide being the minimum length of the substrate. These analyses also showed that longer substrate β-1,2-glucooligosaccharides are preferred, being consistent with the fact that CGSs generally produce CβGs with degrees of polymerization of around 20. Finally, the overall structure of the cyclization domain of TiCGSCy was found to be similar to those of β-1,2-glucanases in phylogenetically different groups. Meanwhile, the identified catalytic residues indicated clear differences in the reaction pathways between these enzymes. Overall, we propose a novel reaction mechanism of TiCGSCy. Thus, the present group of CGSs defines a new glycoside hydrolase family, GH189.
Key points
• It was clearly evidenced that cyclization domain alone produces cyclic β-1,2-glucans.
• The domain exclusively catalyzes transglycosylation without hydrolysis.
• The present catalytic domain defines as a new glycoside hydrolase family 189.
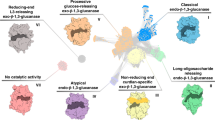
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
β-1,2-Glucan is a polysaccharide comprising glucose units linked by β-1,2-glucosidic bonds. In nature, cyclic forms are mainly found in bacteria such as Agrobacterium tumefaciens, Brucella abortus, and Ensifer meliloti (formerly Rhizobium meliloti and Sinorhizobium meliloti) (Dell et al. 1983; Bundle et al. 1988; Koizumi et al. 1984). Cyclic β-1,2-glucans (CβGs) play important roles in interactions between organisms such as infection of B. abortus and symbiosis of E. meliloti (Breedveld and Miller 1994; Haag et al. 2010; Dylan et al. 1990). Genes encoding enzymes responsible for CβG biosynthesis were identified from three microorganisms independently. These genes were found to encode cyclic β-1,2-glucan synthases (CGSs) homologous to each other (Zorreguieta and Ugalde 1986; Castro et al. 1996; Iannino et al. 1998). Among them, CGS from B. abortus (BaCGS) produces CβGs with degrees of polymerization (DPs) around 20 and is most extensively characterized (Bundle et al. 1988; Guidolin et al. 2009). BaCGS is composed of three regions responsible for the following steps: initiation (covalent bonding of glucose to CGS), elongation of linear β-1,2-glucan (LβGs) chains, adjustment of chain lengths of the glucans, and cyclization of the glucans by transglycosylation (Fig. S1) (Guidolin et al. 2009). The initiation and the elongation steps are carried out in the N-terminal domain classified into glycosyltransferase (GT) family 84 based on amino acid sequence homology by carbohydrate-active enzyme database (CAZy) (Coutinho et al. 2003; Drula et al. 2022; Guidolin et al. 2009). The C-terminal glycoside hydrolase (GH) family 94 domain adjusts the chain lengths of the elongated glucans (Ciocchini et al. 2006; Guidolin et al. 2009). Although the domain in the middle region is known to be involved in cyclization of the linear glucans of the optimum lengths (hereafter, this domain is called cyclization domain), a detailed reaction mechanism has not been unveiled.
Apart from the context of synthetic enzymes described above, β-1,2-glucan-degrading enzymes have been investigated (Abe et al. 2017; Tanaka et al. 2019; Nakajima 2023). Owing to establishment of a large-scale preparation of LβGs by using a 1,2-β-oligoglucan phosphorylase (SOGP) found in 2014 (Nakajima et al. 2014; Abe et al. 2015), prokaryotic and eukaryotic endo-β-1,2-glucanases (SGLs) that produce β-1,2-glucooligosaccharides (Sopns, n is DP) from β-1,2-glucans were explored. Consequently, a bacterial SGL from Chitinophaga pinensis (CpSGL) and a fungal SGL from Talaromyces funiculosus (TfSGL) were sequentially identified for the first time, leading to creation of new families (GH144 and GH162), respectively (Abe et al. 2017; Tanaka et al. 2019). Subsequently, functions and structures of several β-1,2-glucan-associated enzymes including the ones that are given new EC numbers were reported (Nakajima et al. 2016, 2017; Shimizu et al. 2018; Kobayashi et al. 2022). In addition, another SGL which belongs to neither GH144 nor GH162 has been identified from Escherichia coli very recently, leading to foundation of a new family, GH186 (Motouchi et al. 2023).
Interestingly, both CpSGL and TfSGL possess single (α/α)6-domains with similar overall structures although they belong to different families (Abe et al. 2017; Tanaka et al. 2019). Therefore, PSI-BLAST search was performed using CpSGL and TfSGL as queries to find further homologs with evolutional relationships. As a result, the cyclization domains of CGSs (CGSCys) came up although the domains do not belong to any GH families. In the previous study, we showed that the general acid, one of the two catalytic residues, of TfSGL (GH162) exhibits a unique catalytic mechanism that acts via a 3-hydroxy group of the glucose moiety (Tanaka et al. 2019). This mechanism rarely found in the anomer-inverting type is somehow highly conserved among cyclization domains of anomer-retaining CGSs. Therefore, CGSs may share a common mechanism with GH162 beyond inverting/retaining mechanisms. However, TfSGL and CGSCy are intrinsically different in terms of reaction mechanisms. TfSGL follows the anomer-inverting mechanism in which anomer of substrates changes when products are released, while CGS is considered to follow the anomer-retaining mechanism in that substrates and products share the same anomer (see https://www.cazypedia.org/index.php/Glycoside_hydrolases#Mechanistic_classification for detail) (The CAZypedia Consortium 2018; Davies and Henrissat 1995). Thus, we predicted that CGSCy follows a noncanonical reaction mechanism. In this study, we subcloned a region encoding the cyclization domain alone from CGS of Thermoanaerobacter italicus (TiCGS), a thermophilic bacterium, and explored biochemical functions and tertiary structure of the cyclization domain.
Materials and methods
Materials
The genomic DNA of T. italicus (DSM9252) was purchased from the National Institute of Technology and Evaluation (NITE, Tokyo, Japan). LβGs with the average DP of 77 (unless otherwise described, average DP of the β-1,2-glucans used in the present study is 77) and Sopns with DP of 2–10 were prepared using SOGP from Listeria innocua and CpSGL as described previously (Nakajima et al. 2014; Abe et al. 2015, 2017). CβGs with DPs of 17–24 were kindly donated by Dr. M. Hisamatsu of Mie University (Hisamatsu et al. 1984). Laminarin and carboxymethyl (CM)-cellulose were purchased from Sigma-Aldrich (MO, USA). CM-pachyman, CM-curdlan, lichenan, tamarind xyloglucan, arabinogalactan, arabinan, and polygalacturonic acid were purchased from Neogen (MI, USA).
Cloning, expression, and purification of TiCGSCy
A middle region (1005–1591 a.a.) of TiCGS (KEGG locus, Thit_1831) (TiCGSCy) was used for cloning (see the “Results” section for details). The gene region was inserted into the pET30a vector (Merck, NJ, USA) according to the manufacturer’s instructions so that histidine-tag derived from the vector is fused at C-terminus.
E. coli Rosetta2 (DE3) (Merck) was transformed using the constructed plasmid and cultured at 37 °C in LB medium containing 30 μg/ml kanamycin and 34 μg/ml chloramphenicol. After the optical density of the culture at 660 nm reached 0.8, protein expression was induced using 0.1 mM isopropyl-β-d-1-thiogalactopyranoside at 20 °C overnight. The harvested cells were lysed by sonication in 50 mM Tris–HCl buffer (pH 8.0) containing 150 mM NaCl. The supernatant was collected after centrifugation at 27,700 × g. Then the supernatant was filtrated with a 0.45-μm filter (Sartorius, Germany). The sample was loaded onto a HisTrap FF crude column (5 ml; Cytiva, MA, USA) equilibrated with 50 mM Tris–HCl buffer (pH 8.0) containing 150 mM NaCl (buffer A) using an AKTA explorer chromatography system (Cytiva). After unbound proteins were washed out using the same buffer containing 20 mM imidazole, TiCGSCy was eluted using a linear imidazole concentration gradient (20–300 mM) in buffer A. 2 M ammonium sulfate solution containing 100 mM sodium acetate buffer (pH 5.0) was added to the collected sample to obtain 1 M ammonium sulfate concentration. After unbound proteins were washed out using the 1 M ammonium sulfate containing 100 mM sodium acetate buffer (pH 5.0), TiCGSCy was eluted using a linear ammonium sulfate concentration gradient (1–0 M) in 100 mM sodium acetate buffer (pH 5.0). The enzyme solution was exchanged with 5 mM sodium acetate buffer (pH 5.0) using Amicon Ultra 10,000 molecular weight cut-off (Merck). The absorbance of the sample at 280 nm was measured using a spectrophotometer V-650 (Jasco, Tokyo, Japan), and the concentration of the enzyme was determined spectrophotometrically at 280 nm using the theoretical molecular mass of TiCGSCy (69,508 Da) and a molar extinction coefficient of 87,210 M−1·cm−1 calculated based on Pace et al. (Pace et al. 1995).
Size-exclusion chromatography
The enzyme solution concentrated with Amicon Ultra 10,000 molecular weight cut-off to 0.5 mg/ml (500 μl) was loaded onto a Superdex™ 200GL column (24 ml; Cytiva) equilibrated with 50 mM Tris–HCl buffer (pH 8.0) containing 150 mM NaCl, and then the target enzyme was eluted with the same buffer. This analysis by size-exclusion chromatography was carried out using an AKTA prime plus chromatography system (Cytiva). Ovalbumin (44 kDa), conalbumin (75 kDa), aldolase (158 kDa), ferritin (440 kDa), and thyroglobulin (669 kDa) (Cytiva) were used as molecular weight markers. Blue dextran 2000 (2,000 kDa) was used to determine the void volume of the column. The molecular weight of TiCGSCy was calculated using Eq. 1.
where Kav is the gel-phase distribution coefficient, Ve is the volume required to elute each protein, Vo is the volume required to elute blue dextran 2000, and Vt is the bed volume of the column.
Analysis of the cyclization activity of TiCGSCy
The enzymatic reaction of TiCGSCy on LβGs was performed in 20 mM sodium acetate buffer (pH 5.0) containing 1 mg/ml of TiCGSCy and 0.4% LβGs at 30 °C for one hour. After a heat treatment at 100 °C for 5 min, the sample (20 µl) was mixed with 20 µl of 0.1 mg/ml β-glucosidase from Bacteroides thetaiotaomicron (BGL) (Ishiguro et al. 2017) in 100 mM sodium acetate buffer (pH 5.0) and incubated at 40 °C for 30 min or 60 min. After a heat treatment at 100 °C for 5 min, the sample (20 µl) was mixed with 20 µl of 0.2 mg/ml CpSGL in 100 mM sodium acetate buffer (pH 5.0). The reaction mixture was incubated at 30 °C for an hour. Each reaction mixture was analyzed by thin layer chromatography (TLC).
TLC analysis
The reaction mixtures (0.5, 1, or 2 µl) were spotted onto TLC Silica Gel 60 F254 plates (Merck). As for analysis of cyclization activity, the plates were resolved with 70% acetonitrile. In the case of glucose and Sop2-5 produced by TiCGSCy, the plates were resolved with 75% acetonitrile. In the case of Sop6-10, the plates were resolved twice with the solution (1-butanol: acetic acid: deionized water = 2:1:1). The plates were then soaked in a 5% sulfuric acid/ethanol solution (w/v) and heated in an oven until the spots were clearly visualized.
The enzymatic reactions of TiCGSCy with each substrate (0.03% arabinogalactan, 0.2% CM-pachyman, 0.2% laminarin, 0.2% CM-cellulose, 0.2% CM-curdlan, 0.2% arabinan, 0.2% polygalacturonic acid, 0.2% LβGs or 0.2% tamarind xyloglucan, 5 mM glucose and Sop2–10) were carried out in 100 mM sodium acetate buffer (pH 5.0) containing 1 mg/ml of TiCGSCy at 30 °C. After a heat treatment at 100 °C for 5 min, the reaction products were detected by TLC.
NMR analysis
To collect the cyclic products of TiCGSCy derived from LβGs, the enzymatic reaction was performed at 30 °C for 42 h in 100 mM sodium acetate buffer (pH 5.0) containing 2 mg/ml of TiCGSCy and 5% LβGs with an average DP of 121 calculated from the number average molecular weight (Motouchi et al. 2023). After a heat treatment at 100 °C for 10 min, the supernatant (4 ml) was collected after centrifugation at 4,427 × g. Then the solution was mixed with 250 µl of 2.5 mg/ml BGL in 100 mM sodium acetate buffer (pH 5.0) and incubated at 30 °C for 27 h, which ensures completion of the reaction; the intensity of spot showing the polysaccharides no longer changes. After a heat treatment at 100 °C for 10 min, the sample was centrifugated at 4,427 × g. Then the supernatant was filtrated with a 0.45-μm filter (Sartorius). The sample was fractionated by size-exclusion chromatography using a Toyopearl HW-40F column (approximately 2 L gel), as described previously (Nakajima et al. 2014), and the fractions containing the target compound were freeze-dried using a FDU-2100 (EYELA, Tokyo, Japan). The resultant powder was dissolved in D2O, and acetone was added as a standard for calibration of chemical shifts. The chemical shifts were recorded relative to the signal of the methyl group of the internal standard acetone (2.22 ppm). As a reference, CβGs donated by Dr. Hisamatsu (Hisamatsu et al. 1984) were also dissolved in the same solvent. 1H-NMR spectra were recorded using a Bruker Avance 400 spectrometer (Bruker BioSpin).
Mass spectrometric analysis
The samples prepared for the NMR analysis were also analyzed by mass spectrometry. The positive electrospray-ionization mass spectra (ESI/MS) were recorded with samples dissolved in H2O containing 5 mM ammonium acetate on a X500R QTOF mass spectrometer (Sciex, Toronto, CA).
X-ray crystallography
The enzyme solution was concentrated to 17.6 mg/ml. The initial screening of TiCGSCy crystallization was performed using MembFac HT (Hampton research, CA, USA). The crystal for data collection was obtained by incubation of the mixture of 17.6 mg/ml TiCGSCy (2 μl) and a reservoir solution (2 μl) containing 0.1 M sodium cacodylate and 1.3 M sodium acetate (pH 6.5) at 20 °C for one month. The crystal was soaked in the reservoir solution supplemented with 25% (w/v) glycerol for cryoprotection and kept at 100 K in a nitrogen-gas stream during data collection. The X-ray diffraction data was collected on a beamline (BL-5A) at Photon Factory (Tsukuba, Japan). The TiCGSCy structure was determined by molecular replacement using a predicted TiCGSCy structure by AlphaFold2 (Jumper et al. 2021) as a model structure. The molecular replacement, auto model building, and refinement were performed using the MOLREP program (Vagin and Teplyakov 2010), REFMAC5 program (Murshudov et al. 1997), and Coot program (Emsley and Cowtan 2004), respectively. A structural homology search was performed with the DALI server (Holm 2020). The secondary structure was assigned with the DSSP program (Touw et al. 2015). The multiple amino acid alignment and the structure-based amino acid alignment with the secondary structures were visualized using the ESPript 3.0 server (http://espript.ibcp.fr/ESPript/ESPript/) (Robert and Gouet 2014). The overall structures of TiCGSCy, TfSGL and CpSGL were superimposed using the PDBeFold server (https://www.ebi.ac.uk/msd-srv/ssm/ssmcite.html) (Krissinel and Henrick 2004). All the structures in the figures were designed with the PyMOL program.
Mutational analysis
The plasmids of TiCGSCy mutants (E1442Q, E1442A, and E1356A) were constructed using a PrimeSTAR mutagenesis basal kit (Takara Bio) according to the manufacturer’s instructions. PCRs were performed using appropriate primer pairs (Table S2) and the template TiCGSCy plasmid. The transformation to E. coli, the expression, and purification of TiCGSCy mutants were performed in the same manner as that for the wild-type TiCGSCy. The enzymatic reactions were performed basically in the same manner as in the detection of cyclization activity of TiCGSCy.
Results
Purification of cyclization domain of CGS from T. italicus (TiCGSCy)
Domain configuration and biochemical function including cyclization activity of TiCGS were totally unknown. Therefore, multiple amino acid alignment was performed using CGS homologs including TiCGS and BaCGS (Fig. S2). The region of cyclization domain was expected to be residues 1005–1591 a.a. based on the minimum region that retains cyclization activity in BaCGS (Ciocchini et al. 2006; Guidolin et al. 2009). In addition, all transmembrane regions are within residues 1–1004 a.a. according to TMHMM-2.0 server (Krogh et al. 2001). Thus, the region (1005–1591 a.a.) was defined TiCGSCy, and TiCGSCy fused with histidine-tag at the C-terminus was produced as a recombinant protein. The recombinant TiCGSCy (hereafter simply called TiCGSCy) was purified by nickel affinity chromatography and hydrophobic chromatography, with which we obtained highly purified TiCGSCy that migrated as a single band at approximately 70 kDa in the SDS-PAGE analysis. It is consistent with a theoretical molecular mass of TiCGSCy (69.5 kDa).
Size-exclusion chromatography analysis of TiCGSCy
To investigate quaternary structure of TiCGSCy, size-exclusion chromatography was performed. TiCGSCy eluted at the retention time corresponding to 60.6 kDa, which is similar to the theoretical molecular mass shown above (Fig. S3). This result indicated that TiCGSCy exists as a monomer and TiCGS does not form multimer through interactions between the TiCGSCy domains.
Cyclization activity of TiCGSCy
To test the ability of purified TiCGSCy in cyclization of LβGs, the reaction products were analyzed by TLC. In the glycosylation (the former) step of the reaction by typical transglycosylases, linear glucans on the reducing end leave the catalytic site upon formation of the glycosyl-enzyme intermediate (Bissaro et al. 2015; Sinnott 1990; Van der Veen et al. 2000). In the deglycosylation (the latter) step, linear glucan products are expected to be produced in the inter-molecular transglycosylation called disproportionation. Note that the reaction with a water molecule in this step would result in hydrolysis but also releases a hydrolyzed linear product. On the other hand, cyclic products are produced when the non-reducing end of a sugar reacts with the covalent glycosyl-enzyme intermediate (reducing end) of the same molecule. If TiCGSCy possesses cyclization activity, both linear and cyclic glucan chains are expected to be produced. After incubation of LβGs with TiCGSCy, a broad smear band in DPs smaller than those of the substrate was detected (Fig. 1a). Next, the reaction products were subjected to the BGL that act exolytically on the non-reducing end of LβGs (Fig. 1b). Consequently, glucose was produced, but a bit smear band with relatively higher DPs remained on the TLC plate (Fig. 1a). The BGL-treated products were further treated with CpSGL, an endo-type enzyme that produces Sop2–4 (Fig. 1b), resulting in disappearance of the smear band and appearance of Sop2–4 instead (Fig. 1a). These results indicate that the products that formed the smear band after the BGL treatment were in cyclic forms, and thus, TiCGSCy possesses the cyclization activity. In addition, the smear band after the BGL treatment was detected at the position lower than the spot of the marker CβGs with DP17–24 (lane M2) in the TLC plate (Fig. 1a), suggesting that DPs of the cyclic products released by TiCGSCy are higher than 17–24. Furthermore, various polysaccharides were examined as candidate substrates by TLC analysis, but no reaction was detected (Fig. S4). This result suggested that the cyclization activity of TiCGSCy is highly specific to LβGs.
Products from catalysis of LβGs by TiCGSCy. a Detection of the reaction products by TLC analysis. Lane M1, 5 mM glucose and Sop2–5. Lane M2, 0.2% CβGs with DP17–24. Each sample (0.5–2 μl) was spotted on the plate. BGL and CpSGL represent treatment of products with BGL and/or CpSGL. The asterisk represents the origin of the TLC plate. b The present method to distinguish between cyclic and linear forms of β-1,2-glucans. The glucose molecules are illustrated with only the β-1,2-carbon skeleton and the hydroxy group at the reducing end. 'R' represents Sopns
NMR analysis of compounds produced from LβGs by TiCGSCy
In order to identify cyclic glucans produced by TiCGSCy, the reaction product from LβGs was treated with BGL, and the BGL-resistant products were then purified by size-exclusion chromatography. The chemical shifts of the resultants measured by 1H-NMR were almost the same as those of the reference (Figs. S5a and b) (Hisamatsu et al. 1984; Roset et al. 2006). In addition, chemical shifts derived from H-2 and H-4 at the non-reducing end glucose moiety and H-1 (α-anomer) at the reducing end glucose moiety as in the case of LβGs (Nakajima et al. 2014) (Fig. S5c) were not detected (Fig. S5a). These facts indicate that TiCGSCy produces CβGs, and this enzyme follows an anomer-retaining mechanism.
ESI/MS analysis of compounds produced from LβGs by TiCGSCy
To investigate the DP distribution of CβGs synthesized by TiCGSCy from LβGs, the NMR products (mentioned previously) were analyzed by the positive ESI/MS. As a result, multiple peaks corresponding to doubly and triply charged ions containing two or three ammonium ions were detected (Fig. S6a). Furthermore, these peaks matched the theoretical m/z of CβGs with DP17–26 (Fig. S6b). These results suggest that TiCGSCy synthesizes CβGs with DP17–26.
Action patterns of TiCGSCy
To clarify chain length specificity of substrates, various Sopns were adopted in the experiments. In the case of glucose and Sop2–5, no reaction product was detected by TLC analysis (Fig. 2a). On the contrary, Sopns with DPs 4 or higher were produced when Sopns with DPs 6 or higher were applied as substrates (Figs. 2b and 3). These results indicate that specific substrates of TiCGSCy in transglycosylation is Sopns with DPs 6 or higher.
TLC analyses of activity toward glucose and Sop2–6. a, b Lane M1, 5 mM glucose and Sop2–5. Lane M2, 5 mM Sop6–10. Each sample (0.2–1 μl) was spotted on the plate. c Possible reaction steps of TiCGSCy with a substrate Sop6. Subsite binding is based on TLC results. The hypothesized upper two patterns of the hydrolytic reaction were not actually observed because H2O molecules do not participate in attacking covalent intermediates. The glucose molecule at the reducing end is shown in gray
Generally, in the case of enzymes that produce cyclic glycan polymers, a nucleophilic amino acid sidechain initially binds covalently to a substrate to form a glycosyl-enzyme intermediate (Bissaro et al. 2015). This intermediate is then subjected to nucleophilic attack either by a water molecule, an intermolecular hydroxy group, or an intramolecular hydroxy group to cause hydrolysis, disproportionation or cyclization reaction, respectively (Bissaro et al. 2015; Sinnott 1990; Van der Veen et al. 2000). In the case of TiCGSCy with Sop6, Sop4 and Sop5 were detected as a result of reaction, which suggested that Sop6 binds to TiCGSCy at the catalytic site from subsite –2 to subsite + 4 or from –1 to + 5 (Figs. 2b and 2c). However, glucose and Sop2 (counterparts of Sop5 and Sop4, respectively, when Sop6 is hydrolyzed) were not detected. This result indicates that TiCGSCy catalyzes only transglycosylation without hydrolysis (Fig. 2c). Likewise, with Sop7–10 as substrates, glucose and Sop2–3 were not detected (Fig. 3). Therefore, TiCGSCy requires at least four subsites (from subsite + 1 to subsite + 4) occupied by glucose moieties for the reaction to proceed. The reaction mechanism of cyclization is fundamentally the same as that of transglycosylation. The only difference is whether the reaction is intra-molecular or inter-molecular. This is one of the reasons why linear products can be generated by CGSCy. The transglycosylation reaction results in linear products when intramolecular cyclization is not accomplished due to shortage in lengths of the substrate. As the minimum DP of the synthesized CβG is 17 according to the ESI/MS results, the initial reaction products from Sop6–10 are not cyclic.
In terms of the reaction velocities of substrates examined, the larger the DPs of the substrates were, the faster the amounts of the products reached to the similar level (Figs. 2 and 3). This result suggests that TiCGSCy prefers longer substrates, which is consistent with the biochemical property of CGS known to produce CβGs with DPs around 20. The amount of Sop4–6 produced from Sop7, Sop8 and Sop9 at the initial stage of the reactions were Sop4 > Sop5 > Sop6, while in the case of Sop10 as a substrate, it was Sop4–5 > Sop6 (Figs. 2 and 3). These results suggest that at least from subsite − 1 to subsite − 5 in subsite minus side are involved in substrate recognition.
Overall structure of TiCGSCy
A ligand-free structure of TiCGSCy was determined at 3.9 Å resolution (Table S1). An asymmetric unit contains almost identical four molecules (RMSD, 0.3 Å). The enzyme consists of a single (α/α)6-barrel domain with several inserted α-helices (Fig. 4a, c). According to DALI server (Holm 2020), CpSGL (RMSD, 2.4 Å; sequence identity, 17%; PDB ID, 5GZH), GH144 enzyme from Parabacteroides distasonis (RMSD, 2.4 Å; sequence identity, 18%; PDB ID, 5Z06), and TfSGL (RMSD, 2.7 Å; sequence identity, 12%; PDB ID, 6IMU) came up as top 3 structurally similar proteins in the case TiCGSCy is set as a query structure. TiCGSCy is structurally similar to these three enzymes although amino acid sequence identities are very low. Structure-based multiple amino acid alignment suggests that the additional α-helices in the middle region is unique to CGSs, and they are not found in SGLs (Fig. 4c). A large pocket observed in (α/α)6-barrel domain is expected to be a substrate binding site of TiCGSCy (Fig. 4b).
Comparison of substrate-binding site of TiCGSCy with CpSGL and TfSGL
To analyze a substrate binding mode of TiCGSCy, TiCGSCy was superimposed with two enzymes: TfSGL (GH162) complexed with a substrate (Sop7) (PDB ID: 6IMW) and CpSGL (GH144) complexed with a glucose and a Sop3 (PDB ID: 5GZK) (Tanaka et al. 2019; Abe et al. 2017). Consequently, the three overall structures are superimposed well (Fig. S7). The shape of the substrate pocket of TiCGSCy is similar to those of TfSGL and CpSGL in that the substrates observed in TfSGL and CpSGL complex structures can be potentially accommodated in the pocket, although the superimposed glucose moiety at subsite − 3 is a little too close to TiCGSCy (Fig. 5a). There is a sufficient space beyond 2-hydroxy group of the potential subsite − 4 (Fig. 5b), which is consistent with the fact that TiCGSCy prefers Sopns with larger DPs (Figs. 2 and 3). Contrarily, W1394 is likely to block binding of glucose moieties beyond subsite + 3 (Fig. 5c). Taking into account the result of action pattern analysis that subsite + 4 should be occupied (Figs. 2 and 3), side chain of W1394 may flip to make room for substrate binding. In addition, less information was retrieved by a sequence alignment among TiCGSCy homologs. Therefore, we further conducted structural alignment of TiCGSCy, TfSGL and CpSGL. As a result, several residues (G1104, W1109, L1406, Y1456, V1475, P1478 and G1509) are identified as conserved residues. Among these, W1109 and Y1456, which are located near subsite − 3, are assumed to be the key residues for substrate recognition (Figs. S2 and 7). Overall, it is suggested that the pocket in the (α/α)6-barrel domain is the substrate binding site.
Substrate pocket of TiCGSCy. The substrate pocket of TiCGSCy is shown semi-transparently in gray. Sop7 molecules shown as yellow sticks are placed by superimposition of TfSGL-Sop7 complex structure. Glucose and Sop3 molecules shown as light red sticks are placed by superimposition of CpSGL-glucose, Sop3 complex structure. Number labels represent subsite positions. b, c Close-up views around subsites − 4 and + 3. PDB IDs of TfSGL and CpSGL used throughout the manuscript are 6IMW and 5GZK, respectively
Catalytic residues of TiCGSCy
Canonical enzymes that synthesize cyclic carbohydrates take advantage of anomer-retaining mechanism to achieve transglycosylation reaction (see https://www.cazypedia.org/index.php/Transglycosylases for details) (The CAZypedia Consortium 2018; Sinnott 1990). First, an acidic residue (a nucleophile) attacks an anomeric carbon atom at subsite − 1 to form a glycosyl-enzyme covalently bonded intermediate, and an acid/base catalyst provides a proton to a scissile bond oxygen atom in a substrate to release a moiety at the reducing end from the scissile bond. This step is called glycosylation step. In the next step called deglycosylation, the intermediate is attacked by an intramolecular hydroxy group to complete a cyclization reaction mediated by an acid/base catalyst.
In the case of TiCGSCy, E1442 is found with a clear electron density at the position corresponding to that of the nucleophilic water in TfSGL, which attacks the anomeric carbon of the glucose moiety at subsite − 1 (Fig. 6). Meanwhile, no candidate acidic residue directly interacting with a scissile bond oxygen atom is found. However, E1356 of TiCGSCy is well-superimposed with E262 of TfSGL, a clearly evidenced catalytic residue acting on a scissile bond of a substrate through 3-OH of a glucose moiety at subsite + 2 (Fig. 6a). Electron density of E1356 was also observed clearly (Fig. 6b). Both E1442Q and E1442A mutants showed no cyclization activity toward LβGs. In addition, the BGL-resistant products remained as a weak spot at the origin on the TLC in the case of E1356A, indicating that E1356A mutant showed very low cyclization activity toward LβGs (Fig. S8). These results strongly suggest that E1442 is a catalytic residue that acts as a nucleophile, and E1356 is an acid/base catalyst.
Superimposition of catalytic residues and related residues in TiCGSCy, TfSGL, and CpSGL. a Sop7 in TfSGL-Sop7 complex and Sop3 in CpSGL in CpSGL-Glc, and Sop3 complex are partially visualized as yellow and light red sticks, respectively. Residues in TiCGSCy, TfSGL, and CpSGL are shown as thick cyan, purple and gray sticks, respectively. Residues in TiCGSCy, TfSGL, and CpSGL are labelled with bold letters, bold letters in parentheses and plane letters in parentheses, respectively. Water molecules observed in TfSGL-Sop7 complex are shown as red spheres. Gray dashed lines represent a route of the reaction pathway in TfSGL. Subsite positions are labelled − 1 and + 2. b A Fo-Fc omit map for E1356 and E1442 in TiCGSCy. The map is shown at the 4.0σ contour level and represented as cyan meshes
According to prediction of pKa by PROPKA3.5.0 (Olsson et al. 2011), pKa values of E1442 and E1356 in chain A were 6.72 and 8.98, respectively. E1400 highly conserved among CGSs is found in the vicinity of E1356 although E1400 is not conserved in TfSGL or CpSGL (Figs. S2 and 7). A negative charge of E1400 is considered to raise the pKa value of E1356. Contrarily, the basic residues H1536 and H1537 are found in close proximity to E1442, which probably contribute to the decrease in pKa value of E1442. In addition, these two histidine residues are also highly conserved among CGSs (Fig. S2). The difference of the pKa values between E1356 and E1442 suggests that these two residues are catalysts.
Discussion
In the present study, we explicitly showed that TiCGSCy domain alone produced CβGs by transglycosylation reaction without hydrolysis. Whether the final reaction product is cyclic or linear solely depends on the chain length of the substrate. ESI/MS analysis revealed that the minimum DP of BGL-resistant compounds (i.e., CβGs) synthesized from substrate LβGs by TiCGSCy was 17 (Fig. S6). On the other hand, action patterns in the TLC analysis of TiCGSCy (Figs. 2 and 3) suggested that a minimum DP of 4 is additionally required at the reducing end of the cleavage site. Taken together, the minimum DP of the substrate in the synthesis of cyclic sugars is 21 (= 17 + 4). Preference of this domain for Sopns with higher DPs is consistent with the chain lengths of reaction products by the intact CGSs (Hisamatsu 1992; Ciocchini et al. 2007; Guidolin et al. 2009).
Recently, a cryo-EM structure of an intact CGS from A. tumefaciens has been reported (Sedzicki et al. 2023). Nevertheless, a detailed reaction mechanism could not be determined due to the following reasons: Reaction products from LβGs have not been identified as CβGs; With the whole CGS, it was difficult to distinguish between activities of different domains, and the possibility that GH94 glycoside phosphorylase domain produced LβGs by transglycosylation in reversible reactions of phosphorolysis could not be excluded; A plausible reaction pathway to account for transglycosylation could not be drawn from the structure because the substrate chain appears to be placed in reverse orientation in comparison with that determined in TfSGL.
With clarified enzymatic functions and a solid reaction pathway of the sole TiCGSCy, the present study is the first demonstration of detailed reaction mechanism of the CGSCy domain. Based on the overall results of structural and functional analysis of TiCGSCy, the reaction pathway of the enzyme can be explained as follows (Figs. 6 and 8). First, in the glycosylation step, E1356 acts as a general acid to provide a proton to a scissile bond oxygen atom through 3-OH group of a glucose moiety at subsite + 2. Simultaneously, E1442 attacks an anomeric center at subsite − 1 as a nucleophile to form a glycosyl-enzyme intermediate. Next, E1356 acts as a general base to draw a proton of inter- or intra-molecular 2-OH group of a glucose moiety at subsite + 1 through 3-OH group of a glucose moiety at subsite + 2. The activated (deprotonated) 2-OH group at subsite + 1 attacks the anomeric carbon of covalently bonded glucose moiety at subsite − 1 to release transglycosylation products. If an intermolecular hydroxy group, which belongs to a different molecule, attacks an anomeric carbon atom, a product in a linear form is released. In the case of an intramolecular hydroxy group, a product in a cyclic form is released (Fig. 8).
Considering the unique reaction mechanism of TiCGSCy involving the 3-OH group of the glucosyl residue at the subsite + 2, it is evident that the well-ordered pre-association of the oligosaccharide acceptor is essential for TiCGSCy to perform effective transglycosylation. This process is unlikely to be replaced by individual water molecules, as they do not form such arrangements efficiently or frequently due to entropic factors. This might be the reason why TiCGSCy does not perform hydrolysis, and why the glycosyl-enzyme remains stable until oligosaccharide donors enter and bind to the positive subsites appropriately.
Furthermore, although such proton transfer called Grotthuss mechanism is non-canonical among GH families (de Grotthuss 1806; Cukierman 2006), GH162 and GH186 SGLs (TfSGL and OpgD from E. coli, respectively) (Tanaka et al. 2019; Motouchi et al. 2023), GH130 4-O-β-d-mannosyl-d-glucose phosphorylase (Nakae et al. 2013), and GH136 lacto-N-biosidase (Yamada et al. 2017) share this exceptional Grotthuss mechanism in GH families, supporting the proposed reaction mechanism of TiCGSCy.
Comparison of the reaction mechanisms between GH144, GH162, and CGS revealed that the general acid (E262 in GH162 TfSGL) is found also in GH144 CpSGL (E211) and TiCGSCy (E1356, in anomer-retaining mechanism, an acid/base) (Fig. 6a). These residues are also conserved according to multiple amino acid alignment (Fig. 7). Contrarily, D446 in TfSGL (a general base) is substituted with a hydrophobic residue in CpSGL that cannot act as a catalyst (Fig. 6a), indicating that GH162 and GH144 have distinct pathways although we should note that the reaction pathway of GH144 has not yet been fully determined (Tanaka et al. 2019). In TiCGSCy, D446 of TfSGL is substituted with H1536 (Fig. 6a). This histidine is also a proton dissociative residue highly conserved among CGSs (Fig. S2). Nevertheless, E1442 is the nucleophile and H1536 is rather considered to play an important role in supporting deprotonation of E1442. This observation clearly indicates the difference in the reaction pathways between GH162 and CGSs. Although GH162 TfSGL, GH144 CpSGL, and TiCGSCy share a similar overall fold, they belong to phylogenetically far different groups. Considering the clear differences in the reaction mechanism between the three groups, the group of CGSs including TiCGS define a new GH family, GH189.
As described above, superimposition of the present TiCGSCy structure with those of GH144 CpSGL and GH162 TfSGL revealed that arrangements of general acid or acid/base residues are common in the (α/α)6 motif (Figs. S9 and S10). Nevertheless, they are distinctively different from other GH clans (clan GH-G, L, M, O, P, and Q) with (α/α)6-folds. While the general acid or acid/base catalytic residues in CpSGL (GH144), TfSGL (GH162) and TiCGSCy (GH189) are located at almost the same position, they were never superimposed well with any of the counterparts in already-existing six GH clans (Fig. S10). This finding indicated that GH144, GH162, and GH189 are closely related to each other based on the arrangement of this key residue. Nevertheless, among them, the new enzyme TiCGSCy (GH189) has an anomer-retaining mechanism unlike the other two anomer-inverting enzymes. Because GH clans are defined basically according to both similarity in structures and reaction mechanisms, the members of groups that establish a new GH clan (clan GH-S) are GH144 and GH162. Meanwhile, GH189 is so far the only family related to clan GH-S. It would rather become a member of a potential GH clan when another new GH family of a retaining mechanism with a similar arrangement of a catalytic residue is found in the future.
The present study provides significant insights into biosynthesis of the physiologically important CβGs by further understanding of structures and functions of CGSs. Moreover, this finding is a large achievement to expand the field of carbohydrate-active enzymes by adding a new group of enzymes.
Data availability
The raw data for the TLC, size-exclusion chromatography of TiCGSCy, and the 1H-NMR and ESI/MS (referenced in Figs. 1–3, S3-S6 and S8) are available from the corresponding authors (email: n_tanaka@rs.tus.ac.jp; m-nakajima@rs.tus.ac.jp; tmasaike@rs.tus.ac.jp) upon reasonable request. The primer details provided in Table S2 can also be obtained through the same process. The collection and refinement statistics of the Xray crystallography presented in Table S1 are accessible from the Protein Data Bank (PDB, https://www.rcsb.org/). The data of structures in Fig. 4–7, including PDB ID 8WY1 for TiCGSCy, are also available from PDB.
References
Abe K, Nakajima M, Kitaoka M, Toyoizumi H, Takahashi Y, Sugimoto N, Nakai H, Taguchi H (2015) Large-scale preparation of 1,2-β-glucan using 1,2-β-oligoglucan phosphorylase. J Appl Glycosci 62:47–52. https://doi.org/10.5458/jag.jag.JAG-2014_011
Abe K, Nakajima M, Yamashita T, Matsunaga H, Kamisuki S, Nihira T, Takahashi Y, Sugimoto N, Miyanaga A, Nakai H, Arakawa T, Fushinobu S, Taguchi H (2017) Biochemical and structural analyses of a bacterial endo-β-1,2-glucanase reveal a new glycoside hydrolase family. J Biol Chem 292:7487–7506. https://doi.org/10.1074/jbc.M116.762724
Bissaro B, Monsan P, Fauré R, O’Donohue MJ (2015) Glycosynthesis in a waterworld: new insight into the molecular basis of transglycosylation in retaining glycoside hydrolases. Biochem J 467:17–35. https://doi.org/10.1042/BJ20141412
Breedveld MW, Miller KJ (1994) Cyclic β-glucans of members of the family Rhizobiaceae. Microbiol Rev 58:145–161
Bundle DR, Cherwonogrodzky JW, Perry MB (1988) Characterization of Brucella polysaccharide B. Infect Immun 56:1101–1106. https://doi.org/10.1128/iai.56.5.1101-1106.1988
Castro OA, Zorreguieta A, Ielmini V, Vega G, Ielpi L (1996) Cyclic β-(1,2)-glucan synthesis in Rhizobiaceae: roles of the 319-kilodalton protein intermediate. J Bacteriol 178:6043–6048. https://doi.org/10.1128/jb.178.20.6043-6048.1996
Ciocchini AE, Roset MS, Briones G, Iñón de Iannino N, Ugalde RA (2006) Identification of active site residues of the inverting glycosyltransferase Cgs required for the synthesis of cyclic β-1,2-glucan, a Brucella abortus virulence factor. Glycobiology 16:679–691. https://doi.org/10.1093/glycob/cwj113
Ciocchini AE, Guidolin LS, Casabuono AC, Couto AS, de Iannino NI, Ugalde RA (2007) A glycosyltransferase with a length-controlling activity as a mechanism to regulate the size of polysaccharides. Proc Natl Acad Sci U S A 104:16492–16497. https://doi.org/10.1073/pnas.0708025104
Coutinho PM, Deleury E, Davies GJ, Henrissat B (2003) An evolving hierarchical family classification for glycosyltransferases. J Mol Biol 328:307–317
Cukierman S (2006) Et tu, Grotthuss! and other unfinished stories. Biochim Biophys Acta 1757:876–885. https://doi.org/10.1016/j.bbabio.2005.12.001
Davies G, Henrissat B (1995) Structures and mechanisms of glycosyl hydrolases. Structure 3:853–859. https://doi.org/10.1016/S0969-2126(01)00220-9
de Grotthuss CJT (1806) Sur la décomposition de l’eau et des corps qu’elle tient en dissolution à l’aide de l’électricité galvanique. Ann Chim 58:54–73
Dell A, York WS, McNeil M, Darvill AG, Albersheim P (1983) The cyclic structure of β-d-(1→2)-linked d-glucans secreted by Rhizobia and Agrobacteria. Carbohydr Res 117:185–200. https://doi.org/10.1016/0008-6215(83)88086-0
Drula E, Garron ML, Dogan S, Lombard V, Henrissat B, Terrapon N (2022) The carbohydrate-active enzyme database: functions and literature. Nucleic Acids Res 50:D571–D577. https://doi.org/10.1093/nar/gkab1045
Dylan T, Nagpal P, Helinski DR, Ditta GS (1990) Symbiotic pseudorevertants of Rhizobium meliloti ndv mutants. J Bacterial 172:1409–1417. https://doi.org/10.1128/jb.172.3.1409-1417.1990
Emsley P, Cowtan K (2004) Coot : model-building tools for molecular graphics. Acta Crystallogr D Biol Crystallogr 60:2126–2132. https://doi.org/10.1107/S0907444904019158
Guidolin LS, Ciocchini AE, De Iannino NI, Ugalde RA (2009) Functional mapping of Brucella abortus cyclic β-1,2-glucan synthase: identification of the protein domain required for cyclization. J Bacteriol 191:1230–1238. https://doi.org/10.1128/JB.01108-08
Haag AF, Myka KK, Arnold MF, Caro-Hernández P, Ferguson GP (2010) Importance of lipopolysaccharide and cyclic β-1,2-glucans in Brucella-mammalian infections. Int J Microbiol 2010:124509. https://doi.org/10.1155/2010/124509
Hisamatsu M (1992) Cyclic (1→2)-β-d-glucans (cyclosophorans) produced by Agrobacterium and Rhizobium species. Carbohydr Res 231:137–146
Hisamatsu M, Amemura A, Harada T, Koizumi K, Utamura T, Okada Y (1984) Cyclic (1→2)-d-glucan produced by Agrobacterium and Rhizobium: the structure and the distribution of molecular weight. J Jpn Soc Starch Sci 31:117–123
Holm L (2020) DALI and the persistence of protein shape. Protein Sci 29:128–140. https://doi.org/10.1002/pro.3749
Iannino N, Briones G, Tolmasky M, Ugalde R (1998) Molecular cloning and characterization of cgs, the Brucella abortus cyclic β(1–2) glucan synthetase gene: genetic complementation of Rhizobium meliloti ndvB and Agrobacterium tumefaciens chvB mutants. J Bacteriol 180:4392–4400. https://doi.org/10.1128/JB.180.17.4392-4400.1998
Ishiguro R, Tanaka N, Abe K, Nakajima M, Maeda T, Miyanaga A, Takahashi Y, Sugimoto N, Nakai H, Taguchi H (2017) Function and structure relationships of a β-1,2-glucooligosaccharide-degrading β-glucosidase. FEBS Lett 591:3926–3936. https://doi.org/10.1002/1873-3468.12911
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, Tunyasuvunakool K, Bates R, Žídek A, Potapenko A, Bridgland A, Meyer C, Kohl SAA, Ballard AJ, Cowie A, Romera-Paredes B, Nikolov S, Jain R, Adler J, Back T, Petersen S, Reiman D, Clancy E, Zielinski M, Steinegger M, Pacholska M, Berghammer T, Bodenstein S, Silver D, Vinyals O, Senior AW, Kavukcuoglu K, Kohli P, Hassabis D (2021) Highly accurate protein structure prediction with AlphaFold. Nature 596:583–589. https://doi.org/10.1038/s41586-021-03819-2
Kobayashi K, Shimizu H, Tanaka N, Kuramochi K, Nakai H, Nakajima M, Taguchi H (2022) Characterization and structural analyses of a novel glycosyltransferase acting on the β-1,2-glucosidic linkages. J Biol Chem 298:101606. https://doi.org/10.1016/j.jbc.2022.101606
Koizumi K, Okada Y, Utamura T (1984) Further studies on the separation of cyclic (1→2)-β-d-glucans (cyclosophoraoses) produced by Rhizobium meliloti IFO 13336, and determination of their degrees of polymerization by high-performance liquid chromatography. J Chromatogr 299:215–224. https://doi.org/10.1016/S0021-9673(01)97833-1
Krissinel E, Henrick K (2004) Secondary-structure matching (SSM), a new tool for fast protein structure alignment in three dimensions. Acta Crystallogr D Biol Crystallogr 60:2256–2268. https://doi.org/10.1107/S0907444904026460
Krogh A, Larsson B, von Heijne G, Sonnhammer EL (2001) Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol 305:567–580. https://doi.org/10.1006/jmbi.2000.4315
Motouchi S, Kobayashi K, Nakai H, Nakajima M (2023) Identification of enzymatic functions of osmo-regulated periplasmic glucan biosynthesis proteins from Escherichia coli reveals a novel glycoside hydrolase family. Commun Biol 6:961. https://doi.org/10.1038/s42003-023-05336-6
Murshudov GN, Vagin AA, Dodson EJ (1997) Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr D Biol Crystallogr 53:240–255. https://doi.org/10.1107/S0907444996012255
Nakae S, Ito S, Higa M, Senoura T, Wasaki J, Hijikata A, Shionyu M, Ito S, Shirai T (2013) Structure of novel enzyme in mannan biodegradation process 4-O-β-d-mannosyl-d-glucose phosphorylase MGP. J Mol Biol 425. https://doi.org/10.1016/j.jmb.2013.08.002
Nakajima M (2023) β-1,2-Glucans and associated enzymes. Biologia (bratisl) 78:1741–1757. https://doi.org/10.1007/s11756-022-01205-5
Nakajima M, Toyoizumi H, Abe K, Nakai H, Taguchi H, Kitaoka M (2014) 1,2-β-oligoglucan phosphorylase from Listeria innocua. PLOS ONE 9:e92353. https://doi.org/10.1371/journal.pone.0092353
Nakajima M, Yoshida R, Miyanaga A, Abe K, Takahashi Y, Sugimoto N, Toyoizumi H, Nakai H, Kitaoka M, Taguchi H (2016) Functional and structural analysis of a β-glucosidase involved in β-1,2-glucan metabolism in Listeria innocua. PLOS ONE 11:e0148870. https://doi.org/10.1371/journal.pone.0148870
Nakajima M, Tanaka N, Furukawa N, Nihira T, Kodutsumi Y, Takahashi Y, Sugimoto N, Miyanaga A, Fushinobu S, Taguchi H, Nakai H (2017) Mechanistic insight into the substrate specificity of 1,2-β-oligoglucan phosphorylase from Lachnoclostridium phytofermentans. Sci Rep 7:42671. https://doi.org/10.1038/srep42671
Olsson MHM, Søndergaard CR, Rostkowski M, Jensen JH (2011) PROPKA3: Consistent treatment of internal and surface residues in empirical pKa predictions. J Chem Theory Comput 7. https://doi.org/10.1021/ct100578z
Pace CN, Vajdos F, Fee L, Grimsley G, Gray T (1995) How to measure and predict the molar absorption coefficient of a protein. Protein Sci 4:2411–2423. https://doi.org/10.1002/pro.5560041120
Robert X, Gouet P (2014) Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res 42:320–324. https://doi.org/10.1093/nar/gku316
Roset MS, Ciocchini AE, Ugalde RA, Iñón de Iannino N (2006) The Brucella abortus cyclic β-1,2-glucan virulence factor is substituted with O-ester-linked succinyl residues. J Bacteriol 188:5003–5013. https://doi.org/10.1128/JB.00086-06
Sedzicki J, Ni D, Lehmann F, Stahlberg H, Dehio C (2023) Structure-function analysis of the cyclic β-1,2-glucan synthase. bioRxiv. https://doi.org/10.1101/2023.05.05.539553
Shimizu H, Nakajima M, Miyanaga A, Takahashi Y, Tanaka N, Kobayashi K, Sugimoto N, Nakai H, Taguchi H (2018) Characterization and structural analysis of a novel exo-type enzyme acting on β-1,2-glucooligosaccharides from Parabacteroides distasonis. Biochemistry 57:3849–3860. https://doi.org/10.1021/acs.biochem.8b00385
Sinnott ML (1990) Catalytic mechanism of enzymic glycosyl transfer. Chem Rev 90:1171–1202. https://doi.org/10.1021/cr00105a006
Tanaka N, Nakajima M, Narukawa-Nara M, Matsunaga H, Kamisuki S, Aramasa H, Takahashi Y, Sugimoto N, Abe K, Terada T, Miyanaga A, Yamashita T, Sugawara F, Kamakura T, Komba S, Nakai H, Taguchi H (2019) Identification, characterization, and structural analyses of a fungal endo-β-1,2-glucanase reveal a new glycoside hydrolase family. J Biol Chem 294:7942–7965. https://doi.org/10.1074/jbc.RA118.007087
The CAZypedia Consortium (2018) Ten years of CAZypedia: a living encyclopedia of carbohydrate-active enzymes. Glycobiology 28:3–8. https://doi.org/10.1093/glycob/cwx089
Touw WG, Baakman C, Black J, te Beek TAH, Krieger E, Joosten RP, Vriend G (2015) A series of PDB-related databanks for everyday needs. Nucleic Acids Res 43:D364–D368. https://doi.org/10.1093/nar/gku1028
Vagin A, Teplyakov A (2010) Molecular replacement with MOLREP. Acta Crystallogr D Biol Crystallogr 66:22–25. https://doi.org/10.1107/S0907444909042589
Van der Veen BA, Uitdehaag JCM, Penninga D, Van Alebeek GJWM, Smith LM, Dijkstra BW, Dijkhuizen L (2000) Rational design of cyclodextrin glycosyltransferase from Bacillus circulans Strain 251 to increase α-cyclodextrin production. J Mol Biol 296:1027–1038. https://doi.org/10.1006/jmbi.2000.3528
Yamada C, Gotoh A, Sakanaka M, Hattie M, Stubbs KA, Katayama-Ikegami A, Hirose J, Kurihara S, Arakawa T, Kitaoka M, Okuda S, Katayama T, Fushinobu S (2017) Molecular insight into evolution of symbiosis between breast-fed infants and a member of the human gut microbiome Bifidobacterium longum. Cell Chem Biol 24. https://doi.org/10.1016/j.chembiol.2017.03.012
Zorreguieta A, Ugalde RA (1986) Formation in Rhizobium and Agrobacterium spp. of a 235-kilodalton protein intermediate in β-d(1–2) glucan synthesis. J Bacteriol 167:947–951. https://doi.org/10.1128/jb.167.3.947-951.1986
Acknowledgements
The CβGs with DP of 17–24 were kindly provided by Dr. Hisamatsu (Mie University).
Funding
Open Access funding provided by Tokyo University of Science. This work was supported by Photon Factory for X-ray data collection (Proposal No. 2020G527 and No. 2021G685).
Author information
Authors and Affiliations
Contributions
NT, MN, HN, HT, and TM conceived and designed research. NT and RS performed cyclization activity analysis, sequence analysis, and crystal structure analysis of TiCGSCy. NT performed action pattern analyses, NMR analyses, and ESI/MS analyses. KKu, SK, and NT acquired NMR data. KKo created the predicted structure of TiCGSCy for the initial structural analyses. NT, HN, and MN contributed to preparation of compounds for analysis. NT, RS, MN, and TM analyzed the data. NT, MN, and TM wrote the draft manuscript. All the authors read and approved the content of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors. All applicable international, national, and/or institutional guidelines for the use of Living modified organisms were followed in this article.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tanaka, N., Saito, R., Kobayashi, K. et al. Functional and structural analysis of a cyclization domain in a cyclic β-1,2-glucan synthase. Appl Microbiol Biotechnol 108, 187 (2024). https://doi.org/10.1007/s00253-024-13013-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00253-024-13013-9