Abstract
Using genome-wide clustered regularly interspaced short palindromic repeats (CRISPR) screens to understand endocrine drug resistance, we discovered ARID1A and other SWI/SNF complex components as the factors most critically required for response to two classes of estrogen receptor-alpha (ER) antagonists. In this context, SWI/SNF-specific gene deletion resulted in drug resistance. Unexpectedly, ARID1A was also the top candidate in regard to response to the bromodomain and extraterminal domain inhibitor JQ1, but in the opposite direction, with loss of ARID1A sensitizing breast cancer cells to bromodomain and extraterminal domain inhibition. We show that ARID1A is a repressor that binds chromatin at ER cis-regulatory elements. However, ARID1A elicits repressive activity in an enhancer-specific, but forkhead box A1-dependent and active, ER-independent manner. Deletion of ARID1A resulted in loss of histone deacetylase 1 binding, increased histone 4 lysine acetylation and subsequent BRD4-driven transcription and growth. ARID1A mutations are more frequent in treatment-resistant disease, and our findings provide mechanistic insight into this process while revealing rational treatment strategies for these patients.






Similar content being viewed by others
Data availability
Change history
21 February 2020
The Supplementary Information file initially published online was missing the Supplementary Figures and Note. The error has been corrected in the HTML version of the article.
31 January 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41588-020-0582-9
References
Ali, S. & Coombes, R. C. Endocrine-responsive breast cancer and strategies for combating resistance. Nat. Rev. Cancer 2, 101–112 (2002).
Carroll, J. S. et al. Chromosome-wide mapping of estrogen receptor binding reveals long-range regulation requiring the forkhead protein FoxA1. Cell 122, 33–43 (2005).
Eeckhoute, J. et al. Positive cross-regulatory loop ties GATA-3 to Estrogen Receptor alpha expression in breast cancer. Cancer Res. 67, 6477–6483 (2007).
Musgrove, E. A. & Sutherland, R. L. Biological determinants of endocrine resistance in breast cancer. Nat. Rev. Cancer 9, 631–643 (2009).
Shang, Y., Hu, X., DiRenzo, J., Lazar, M. A. & Brown, M. Cofactor dynamics and sufficiency in estrogen receptor-regulated transcription. Cell 103, 843–852 (2000).
Malovannaya, A. et al. Analysis of the human endogenous coregulator complexome. Cell 145, 787–799 (2011).
Mohammed, H. et al. Endogenous purification reveals GREB1 as a key estrogen receptor regulatory factor. Cell Rep. 3, 342–349 (2013).
Fletcher, T. M. et al. ATP-dependent mobilization of the glucocorticoid receptor during chromatin remodeling. Mol. Cell. Biol. 22, 3255–3263 (2002).
John, S. et al. Interaction of the glucocorticoid receptor with the chromatin landscape. Mol. Cell 29, 611–624 (2008).
Michel, B. C. et al. A non-canonical SWI/SNF complex is a synthetic lethal target in cancers driven by BAF complex perturbation. Nat. Cell Biol. 20, 1410–1420 (2018).
Mashtalir, N. et al. Modular organization and assembly of SWI/SNF family chromatin remodeling complexes. Cell 175, 1272–1288 (2018).
Belandia, B., Orford, R. L., Hurst, H. C. & Parker, M. G. Targeting of SWI/SNF chromatin remodelling complexes to estrogen-responsive genes. EMBO J. 21, 4094–4103 (2002).
Garcia-Pedrero, J. M., Kiskinis, E., Parker, M. G. & Belandia, B. The SWI/SNF chromatin remodeling subunit BAF57 is a critical regulator of estrogen receptor function in breast cancer cells. J. Biol. Chem. 281, 22656–22664 (2006).
Jeong, K. W., Lee, Y. H. & Stallcup, M. R. Recruitment of the SWI/SNF chromatin remodeling complex to steroid hormone-regulated promoters by nuclear receptor coactivator flightless-I. J. Biol. Chem. 284, 29298–29309 (2009).
DiRenzo, J. et al. BRG-1 is recruited to estrogen-responsive promoters and cooperates with factors involved in histone acetylation. Mol. Cell. Biol. 20, 7541–7549 (2000).
Kadoch, C. & Crabtree, G. R. Mammalian SWI/SNF chromatin remodeling complexes and cancer: mechanistic insights gained from human genomics. Sci. Adv. 1, e1500447 (2015).
Kadoch, C. et al. Proteomic and bioinformatic analysis of mammalian SWI/SNF complexes identifies extensive roles in human malignancy. Nat. Genet. 45, 592–601 (2013).
Wei, Z. et al. Vitamin D switches BAF complexes to protect beta cells. Cell 173, 1135–1149 (2018).
Cho, H. D. et al. Loss of tumor suppressor ARID1A protein expression correlates with poor prognosis in patients with primary breast cancer. J. Breast Cancer 18, 339–346 (2015).
Yates, L. R. et al. Genomic evolution of breast cancer metastasis and relapse. Cancer Cell 32, 169–184 (2017).
St Pierre, R. & Kadoch, C. Mammalian SWI/SNF complexes in cancer: emerging therapeutic opportunities. Curr. Opin. Genet. Dev. 42, 56–67 (2017).
Pereira, B. et al. The somatic mutation profiles of 2,433 breast cancers refine their genomic and transcriptomic landscapes. Nat. Commun. 7, 11479 (2016).
Tzelepis, K. et al. A CRISPR dropout screen identifies genetic vulnerabilities and therapeutic targets in acute myeloid leukemia. Cell Rep. 17, 1193–1205 (2016).
Shi, J. & Vakoc, C. R. The mechanisms behind the therapeutic activity of BET bromodomain inhibition. Mol. Cell 54, 728–736 (2014).
Nagarajan, S. et al. Bromodomain protein BRD4 is required for estrogen receptor-dependent enhancer activation and gene transcription. Cell Rep. 8, 460–469 (2014).
Zhang, Y. et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. 9, R137 (2008).
Curtis, C. et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486, 346–352 (2012).
Glont, S. E. et al. Identification of ChIP-seq and RIME grade antibodies for Estrogen Receptor alpha. PLoS ONE 14, e0215340 (2019).
Papachristou, E. K. et al. A quantitative mass spectrometry-based approach to monitor the dynamics of endogenous chromatin-associated protein complexes. Nat. Commun. 9, 2311 (2018).
Bruna, A. et al. A biobank of breast cancer explants with preserved intra-tumor heterogeneity to screen anticancer compounds. Cell 167, 260–274 (2016). e22.
Filippakopoulos, P. et al. Histone recognition and large-scale structural analysis of the human bromodomain family. Cell 149, 214–231 (2012).
Johnson, T. A., Elbi, C., Parekh, B. S., Hager, G. L. & John, S. Chromatin remodeling complexes interact dynamically with a glucocorticoid receptor-regulated promoter. Mol. Biol. Cell 19, 3308–3322 (2008).
Augello, M. A., Hickey, T. E. & Knudsen, K. E. FOXA1: master of steroid receptor function in cancer. EMBO J. 30, 3885–3894 (2011).
Jozwik, K. M., Chernukhin, I., Serandour, A. A., Nagarajan, S. & Carroll, J. S. FOXA1 directs H3K4 monomethylation at enhancers via recruitment of the methyltransferase MLL3. Cell Rep. 17, 2715–2723 (2016).
Cirillo, L. A. & Zaret, K. S. An early developmental transcription factor complex that is more stable on nucleosome core particles than on free DNA. Mol. Cell 4, 961–969 (1999).
Cirillo, L. A. & Zaret, K. S. Specific interactions of the wing domains of FOXA1 transcription factor with DNA. J. Mol. Biol. 366, 720–724 (2007).
Berns, K. et al. ARID1A mutation sensitizes most ovarian clear cell carcinomas to BET inhibitors. Oncogene 37, 4611–4625 (2018).
Caumanns, J. J., Wisman, G. B. A., Berns, K., van der Zee, A. G. J. & de Jong, S. ARID1A mutant ovarian clear cell carcinoma: a clear target for synthetic lethal strategies. Biochim. Biophys. Acta Rev. Cancer 1870, 176–184 (2018).
Kent, W. J. BLAT–the BLAST-like alignment tool. Genome Res. 12, 656–664 (2002).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Corces, M. R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Bailey, T. L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Stark, R. & Brown, G. D. DiffBind: differential binding analysis of ChIP-Seq peak data v.3.10 (Bioconductor); http://bioconductor.org/packages/release/bioc/html/DiffBind.html
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Mohammed, H. et al. Progesterone receptor modulates ERalpha action in breast cancer. Nature 523, 313–317 (2015).
Centenera, M. M., Raj, G. V., Knudsen, K. E., Tilley, W. D. & Butler, L. M. Ex vivo culture of human prostate tissue and drug development. Nat. Rev. Urol. 10, 483–487 (2013).
Centenera, M. M. et al. A patient-derived explant (PDE) model of hormone-dependent cancer. Mol. Oncol. 12, 1608–1622 (2018).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Lai, Z. et al. VarDict: a novel and versatile variant caller for next-generation sequencing in cancer research. Nucleic Acids Res. 44, e108 (2016).
Acknowledgements
The authors thank Genomics, Proteomics, Bioinformatics, Preclinical genome editing, Histopathology and the Flow cytometry core facilities at CRUK Cambridge Institute. In particular, we thank C. D’Santos from Proteomics, A. Smith from Preclinical Genome Editing and M. Clayton, G. Cronshaw, R. Ellis and all the staff at CRUK. PX458 was a kind gift from F. Zhang and BRD4 antibody from C. M. Chiang. We acknowledge the suggestions from Z. Najafova in regard to mouse experiments, and A. Nicholson, M. Sen, X. Wang and S. A. Johnsen for reagents and antibodies. We thank R. Rony for his help in graphic design and models. We acknowledge the support of the University of Cambridge and Cancer Research UK. J.S.C. is supported by an ERC Consolidator award (no. 646876), a Susan G Komen leadership grant and CRUK core funding. S.N. is supported by a Komen grant. S.V.R. is funded by Innovation postdoc granted by the Norwegian Research Council. R.S. is funded by the Novo Nordisk Foundation (no. NNF15OC0014136).
Author information
Authors and Affiliations
Contributions
S.N. contributed to conceptualization, methodology, experimental work, formal analysis and figure assembly, writing of text, reviewing and advising on the manuscript. S.V.R. contributed to experimental work, analysis and figure assembly of mouse experiments and advising on the manuscript. J.S. contributed to experimental work, analysis and figure assembly of mouse experiments and advising on the manuscript. D.C. and E.A.G. contributed to experimental work and advising on the manuscript. S.D., S.-E.G., A.R. and S.K. contributed to methodology and advising on the manuscript. E.K.P. contributed to experimental work, formal analysis and figure assembly and advising on the manuscript. J.-E.G.P. contributed to experimental work and advising on the manuscript. D.-L.C. contributed to statistical analyses, figure assembly, reviewing and advising on the manuscript. K.K. and C.S.R.C. contributed to bioinformatic analyses, figure assembly and advising on the manuscript. C.B. contributed to histopathological analyses and advising on the manuscript. N.G. and R.N. performed ARID1A immunohistochemistry and advising on the manuscript. A.B. contributed to methodology regarding PDX material and advising on the manuscript. C.C. contributed to methodology regarding PDX material, funding acquisition and advising on the manuscript. R.S. contributed to methodology, supervision and advising on the manuscript. K.Y. contributed to methodology, funding acquisition and advising on the manuscript. I.C. contributed to methodology, bioinformatic analyses and figure assembly, writing of text and advising on the manuscript. J.S.C. contributed to conceptualization, supervision, funding acquisition, writing of text, reviewing and advising on the manuscript.
Corresponding author
Ethics declarations
Competing interests
J.S.C. is the founder and chief scientific officer of Azeria Therapeutics.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Enrichment of BAF and P-BAF components in the CRISPR screen.
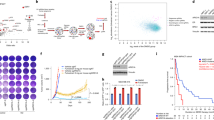
a. Scatterplot of CRISPR screening data, showing enrichment of BAF components following 26 days of different drug treatment, relative to DMSO treated control cells. n = 3 independent viral infections. b. Log2 fold changes showing gRNA enrichment/depletion against all BAF, P-BAF and ncBAF components in the CRISPR screen. Treatment conditions are compared to DMSO control. More proliferative changes represent enriched gRNA after treatment, indicating genes that contribute to drug resistance. c, e. Validation of ARID1A perturbation effect on proliferation and drug response using ARID1A siRNA on MCF7 (c) and ZR-75-1 (e), representative experiments shown from 2 similar independent experiments each cell line. p-values calculated by One way ANOVA test. * denote p < 0.05, *** denotes p < 0.001. Sample size mentioned in S4. Measure of centre represents mean ± SEM (c) and mean ± SD (e). d. Western blot of ARID1A protein levels after siRNA transfection in MCF7 cells. A representative image is shown from 3 similar independent experiments. Unprocessed Western blot in Source Data Fig. 2.
Extended Data Fig. 2 ARID1A co-binds ER and FOXA1-bound regulatory elements, but is depleted with estrogen treatment.
a-c. Single gene profiles showing the binding of ER, FOXA1 and ARID1A on overlapping sites in MCF7 cells. ChIP-seq was performed using three independent biological cell cultures. d. Overlap of binding sites for ER, FOXA1 and ARID1A binding sites in ZR-75-1 cells. e. Boxplots showing the normalized ChIP-seq tag density around 400 bp window around the center of ARID1A binding on DiffBind-defined estrogen independent (constant) and dependent (reduced with estrogen) sites in MCF7. Both classes show reduced ARID1A binding upon estrogen. p-values were calculated by Welch’s t-test, two-sided. Centre line shows the median values with bounds of box corresponding to the first and third quartiles and the upper and lower whiskers extend to the largest or the smallest value no further than 1.5 × IQR (inter-quartile range). Statistical test details are mentioned in Supplementary Table 5e.
Extended Data Fig. 3 Enrichment of SWI/SNF factors with ER and FOXA1 in RIME.
a. ARID1A and BRG1 RIME were conducted on asynchronous MCF7 cells on two biological cell cultures. Label free quantification was performed to show the log 2 scaled normalized intensities of the BAF, P-BAF, ncBAF and common subunits of SWI/SNF complex. Rabbit polyclonal IgG is used as the negative control. b. ER qPLEX-RIME was performed on five primary tumours from ER + breast cancer patients and the ER interactors are shown as enrichment over IgG vs -log10 p-value, corrected by Benjamini and Hochberg multiplicity correction, two-sided. c, d. Boxplots illustrating the more enrichment of HDAC1 (c) and less enrichment of random factors (d) in ERα RIME in five patients compared to IgG negative control in human breast tumours. The values are scaled to the median of IgG and log2 transformed. e. Boxplots illustrating the enrichment of selected known ERα interactors from the RIME experiment in MCF7 cells at a representative timepoint (4-hydroxytamoxifen- 24 hrs) comparing to IgG negative control. The values are scaled to the median of IgG and log2 transformed. n = 5 independent biological cell cultures. For all boxplots, Centre line shows the median values with bounds of box corresponding to the first and third quartiles and the upper and lower whiskers extend to the largest or the smallest value no further than 1.5 × IQR (inter-quartile range).
Extended Data Fig. 4 Enrichment of SWI/SNF factors during Tamoxifen and Fulvestrant in ChIP-seq experiments.
a-d. Asynchronous MCF7 cells were treated with vehicle or Fulvestrant, an ER degrader and ChIP-seq was conducted for ARID1A (b), BRG1 (c) or SNF5 (d). Triplicate independent cell cultures were conducted. d. Single gene profile showing the induction of SWI/SNF complex binding during Fulvestrant treatment. e. Overlap of ARID1A lost sites during estrogen treatment with gained sites during Tamoxifen and Fulvestrant from three independent biological cell cultures. f. Overlap of ARID1A gained sites during Tamoxifen treatment with Fulvestrant and Tamoxifen downregulated genes.
Extended Data Fig. 5 FOXA1 promotes the binding of ARID1A and BRG1.
Hormone-deprived ZR-75-1 cells were transfected with control or FOXA1 siRNA and ChIP-seq was conducted for ARID1A (a) and BRG1 (b). n = 3 independent biological cell cultures. MA plots are shown with the average intensity of binding vs log2 fold change with FOXA1 siRNA relative to control siRNA. c. Scatterplot showing the association of the loss of ARID1A and BRG1 binding upon FOXA1 knockdown. PCC – Pearson Correlation coefficient, two-sided. d. Heatmaps shown on ARID1A and BRG1 FOXA1 independent (common) and dependent (lost sites with FOXA1 knockdown) sites in ZR-75-1 cells. e. Boxplots showing the normalized ChIP-seq tag density around 400 bp window of ARID1A and BRG1 on FOXA1 independent (constant, n = 70,429 sites) and dependent (lost sites with siFOXA1, n = 17,357 sites) sites in ZR-75-1. p-value calculated by Welch’s test, two-sided. n = 3 independent biological cell culture samples. Centre line shows the median values with bounds of box corresponding to the first and third quartiles and the upper and lower whiskers extend to the largest or the smallest value no further than 1.5 × IQR (inter-quartile range). Statistical test details are mentioned in Supplementary Table 5f.
Extended Data Fig. 6 FOXA1 promotes the binding of ARID1A and BRG1.
Hormone-deprived MCF7 and ZR-75-1 cells were transfected with control or FOXA1 siRNA and ChIP-seq was conducted for ARID1A and BRG1. n = 3 independent biological cell cultures. (a-b) Single gene profiles of CCND1 (a) and CDH1 (b) showing the effect on SWI/SNF complex binding with FOXA1 knockdown on MCF7 and ZR-75-1 cells. ER and FOXA1 binding overlap is shown. (c-d) ChIP-qPCR analyses on specific sites (CCND1 and CDH1 ER binding sites) showing ARID1A and BRG1 binding with FOXA1 knockdown in hormone-deprived MCF7 and ZR-75-1 cells (c) or ARID1A binding following Tamoxifen treatment in asynchronous MCF7 cells (d). n = 3 independent biological cell cultures. * denotes p ≤ 0.05, ** denotes p ≤ 0.01, *** denotes p ≤ 0.001. Precise p-values are mentioned in Supplementary Fig. 10. Mean is measured as centre shown with standard deviation. Details of the statistical tests are mentioned in Supplementary Fig. 10.
Extended Data Fig. 7 ATAC-seq analyses shows a negligible regulation of ARID1A on transcription-associated chromatin opening.
a. Heatmap showing ATAC-seq analysis in ARID1A KO clones 11 and 14 following Tamoxifen treatment. Common, gained and lost sites defined by DiffBind analysis. n = 4 independent biological cell cultures. FDR ≤ 0.05 corrected by Benjamini-Hochberg multiplicity correction, two-sided. b. Association of ARID1A KO upregulated and downregulated genes with ATAC-seq gained and lost sites.
Extended Data Fig. 8 ARID1A perturbation regulates ARID2 binding.
a. ARID2 ChIP-seq was conducted in wild type cells or the two ARID1A knock-out clonal cell lines and heatmaps are shown on ARID2 binding sites after Tamoxifen treatment. Also included was ARID1A ChIP-seq from wild type cells treated with vehicle or Tamoxifen. ARID2 binding overlapped with ARID1A binding and was dependent on ARID1A. n = 3 independent biological cell cultures. b. Signal intensity plot showing changes in ARID2 binding in wild type control cells or ARID1A knock-out cells at ARID2 binding sites. n = 3 independent biological cell cultures.
Extended Data Fig. 9 ARID1A promotes BRG1 and HDAC1 binding without affecting ER and H3K27ac occupancy.
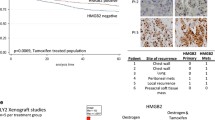
a, b. BRG1, H3K27Ac, HDAC1 and ER (b) ChIP-seq were conducted in asynchronous wild type cells treated with vehicle or tamoxifen or in the two ARID1A knock-out clones (Clones 11 and 14) following tamoxifen treatment. The binding is shown on regions where HDAC1 is lost in ARID1A knockout cells relative to wild type cells. n = 3 independent biological cell cultures. c, d. Scatterplot showing the correlation of ER (c) or H3K27Ac (d) and HDAC1 binding in ARID1A knockout clone 11 versus wild type cells. n = 3 independent biological cell cultures. PCC – Pearson Correlation coefficient. p-values were calculated by Pearson correlation test, two-sided. e. Principal Component Analysis (PCA) of normalised peptide intensities of PDX tumours after ER qPLEX-RIME. n = 2 PDX each group. f. Details of ARID1A mutations observed within ER + PDX tumours used in ER qPLEX-RIME.
Extended Data Fig. 10 ARID1A regulates histone H4 acetylation.
Upregulation of histone H4 acetylation in ARID1A knock-out clone 11 and 14 in Vehicle (a) or Tamoxifen (b) treated cells comparing to wild type cells. Heatmap representing the changes in histone H4Ac marks upon ARID1A knockout with Vehicle or Tamoxifen treatment on ER binding sites close to ARID1A repressed genes. n = 3. (c) Empirical cumulative probability distribution plots of H4K8Ac and H4K12Ac ChIP-seq signals showing upregulation in intensity (y-axis) with ARID1A knockouts clones 11 and 14. Plots were made on ER sites close to ARID1A repressed genes (n = 686 sites) with more than 75% contribution to the variance in intensity. Window – 2 kb around the center of binding.
Supplementary information
Supplementary Information
Supplementary Figs. 1–15 and Note
Supplementary Tables
Supplementary Tables 1–6
Source data
Source Data Fig. 1
Unprocessed Western Blots for Extended Data Fig. 1d.
Source Data Fig. 2.
Unprocessed Western Blots for Fig. 2a.
Rights and permissions
About this article
Cite this article
Nagarajan, S., Rao, S.V., Sutton, J. et al. ARID1A influences HDAC1/BRD4 activity, intrinsic proliferative capacity and breast cancer treatment response. Nat Genet 52, 187–197 (2020). https://doi.org/10.1038/s41588-019-0541-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-019-0541-5
- Springer Nature America, Inc.
This article is cited by
-
The androgen receptor interacts with GATA3 to transcriptionally regulate a luminal epithelial cell phenotype in breast cancer
Genome Biology (2024)
-
SMAD4 depletion contributes to endocrine resistance by integrating ER and ERBB signaling in HR + HER2− breast cancer
Cell Death & Disease (2024)
-
Prion-like domain mediated phase separation of ARID1A promotes oncogenic potential of Ewing’s sarcoma
Nature Communications (2024)
-
Lipidome and metabolome analyses reveal metabolic alterations associated with MCF-7 apoptosis upon 4-hydroxytamoxifen treatment
Scientific Reports (2023)
-
Multilevel proteomic analyses reveal molecular diversity between diffuse-type and intestinal-type gastric cancer
Nature Communications (2023)