Abstract
A mosaic of cross-phylum chemical interactions occurs between all metazoans and their microbiomes. A number of molecular families that are known to be produced by the microbiome have a marked effect on the balance between health and disease1,2,3,4,5,6,7,8,9. Considering the diversity of the human microbiome (which numbers over 40,000 operational taxonomic units10), the effect of the microbiome on the chemistry of an entire animal remains underexplored. Here we use mass spectrometry informatics and data visualization approaches11,12,13 to provide an assessment of the effects of the microbiome on the chemistry of an entire mammal by comparing metabolomics data from germ-free and specific-pathogen-free mice. We found that the microbiota affects the chemistry of all organs. This included the amino acid conjugations of host bile acids that were used to produce phenylalanocholic acid, tyrosocholic acid and leucocholic acid, which have not previously been characterized despite extensive research on bile-acid chemistry14. These bile-acid conjugates were also found in humans, and were enriched in patients with inflammatory bowel disease or cystic fibrosis. These compounds agonized the farnesoid X receptor in vitro, and mice gavaged with the compounds showed reduced expression of bile-acid synthesis genes in vivo. Further studies are required to confirm whether these compounds have a physiological role in the host, and whether they contribute to gut diseases that are associated with microbiome dysbiosis.
Similar content being viewed by others
Main
In total, we analysed 768 samples from 96 sample sites of 29 different organs from 4 germ-free and 4 colonized mice by liquid chromatography–tandem mass spectrometry (LC–MS/MS) and 16S rRNA gene sequencing (Supplementary Table 1). Mapping the first principal coordinate position of each sample from specific-pathogen-free (SPF) mice onto a three-dimensional (3D) mouse model13 enabled us to visualize the similarity of the microbiome and metabolome through all organs and organ systems (Fig. 1a, b; the 3D model is available as Supplementary Data). Different sections through the gastrointestinal tract had unique microbiome and metabolome profiles. There was a distinct difference between the similarity of the two data types in mouse faecal samples. The metabolome differed between faecal samples and the distal gastrointestinal tract, whereas the microbiome was more similar between faeces and colon or caecum samples.
a, Three-dimensional model of mouse organs mapped with the mean first principal coordinate (Extended Data Fig. 1) as a heat map (according to the colour scale), from the germ-free and SPF mice (n = 4 mice each). Ad, adrenal gland; bl, bladder; br, brain; caec, caecum; col, colon; cx, cervix; duo, duodenum; er, ear; f, faeces; ft, feet; hd, hand; jej, jejunum; kd, kidney; lg, lung; lv, liver; mo, mouth; oes, oesophagus; ov, ovary; sto, stomach; tr, trachea; ut, uterus; vg, vagina. b, Mean percentage and total number of unique spectra in each organ sampled from the two mouse groups. c, Relative abundance (normalized to total ion current (TIC)) of the 30 most differential metabolites between the guts of germ-free and SPF mice. The metabolites are coloured as secondary bile acids (blue), primary bile acids (red), soyasaponins (pink), peptides (yellow) and unknown (brown). Annotations are based on spectral matching or molecular network propagation (level two or three16). Stereochemistry of the annotated molecules cannot be discerned using these methods. d, Mean and 95% confidence interval of the Shannon–Weiner diversity of the metabolomic data in each sample from the gastrointestinal tracts of germ-free and SPF mice. Statistical significance between metabolome diversity in the same sample location between germ-free and SPF mice was tested with a two-sided Mann–Whitney U-test, n = 4. *P = 0.028, #P = 0.057. e, Results of meta-mass-shift chemical profiling17 showing the spectral counts of known mass differences between unique nodes in either germ-free or SPF mice. Each mass difference corresponds to the node-to-node gain or loss of a particular chemical group.
Molecular networking of mouse data
To characterize the chemical effect of the microbiome, we subjected the mass spectrometry data to molecular networking12. The algorithm identified 7,913 spectra, of which 14.7 ± 2.2% were observed in colonized mice and 10.0 ± 0.7% were exclusive to germ-free mice (Fig. 1c, Extended Data Fig. 1). Although the overall profiles showed that the strongest differences between germ-free and SPF mice were in the gastrointestinal tract, molecular networking identified unique chemical signatures from the microbiome in all organs—ranging from 2% in the bladder to 44% in stools (Fig. 1b). The metabolome of the caecum, the main site of microbial fermentation of food, was most-markedly affected by the microbiota. Spectral library searching enabled the annotation of 8.9% of nodes in the molecular network11,15 (level two or three, according to previously published standards16). Many of the changes attributed to the microbiome were location-specific, resulting from the metabolism of plant natural products from food and bile acids (Fig. 1c, Extended Data Figs. 2–4, Supplementary Data).
In the upper gastrointestinal tract, the Shannon diversity of the metabolomes of germ-free mice mirrored those of SPF mice; in both sets of mice, diversity was low in the oesophagus and higher in the stomach and duodenum. Upon transition to the caecum, however, the diversity of the two groups of mice began to separate (Fig. 1d). The molecular diversity in the caecum and colon of colonized mice was higher than that of germ-free mice, but this was not the case in the stool samples (Fig. 1d). In the duodenum (the location at which the gallbladder adds bile to the intestine), there was a contrast in microbiome and metabolome diversity: a high metabolome diversity corresponded to a low microbial diversity (Fig. 1d, Extended Data Fig. 1).
Molecular networking enabled meta-mass-shift chemical profiling17 (an analysis of chemical transformations on the basis of parent mass shifts between related spectra without the requirement of knowing the molecular structures) of the gastrointestinal tracts of germ-free and SPF mice. In colonized mice, there was a signature for water loss in the duodenum and jejunum and the loss of H2, acetyl and methyl groups in later parts of the gastrointestinal tract (Fig. 1e). Of all the H2 shifts, 23.1% were associated with bile acids, which indicates that colonization resulted in the oxidation of bile acids (a known microbial transformation)18. Deacetylations were also prevalent in colonized mice, although the metabolites on which this occurred remain unidentified. Germ-free mice had mass gains that corresponded to saccharides in all regions of the gastrointestinal tract (Fig. 1e); these gains were primarily associated with plant natural products, such as soyasaponins and flavonoids. The absence of these sugars in SPF mice implicates the microbiome in their metabolism (Extended Data Figs. 2, 3). A unique mass gain of C4H8 was detected in the jejunum and ileum of SPF mice (Fig. 1e) and 18.2% of spectra with this mass gain were derived from an unknown molecule related to the conjugated bile acid glycocholic acid (GCA) (Fig. 2a). Overall, both germ-free and SPF mice had frequent and diverse mass losses between related molecules, but in colonized mice there were fewer molecules that gained a molecular group (Fig. 1e). This indicates that the microbiome contributed more to the catabolic breakdown of molecules, and less to anabolism. However, we found the addition of C4H8 to GCA to be a particularly interesting anabolic reaction that was dependent on the gut microbiome, and we sought to investigate this further.
a, Structures and molecular networks of newly identified microbiome-conjugated bile acids, with host-conjugated GCA shown for comparison. The molecular network is coloured by mapping to germ-free or SPF mice (according to the colour legend). Inset highlights the parent masses and mass differences between the newly discovered molecules and GCA. Each node represents a clustered tandem mass spectrum; connections between the nodes indicate relationships through the cosine score with their width scaled by the cosine size (cut-off minimum of 0.7). Circular nodes are unknown molecules, and arrowheads are spectra with matches in the GNPS libraries. b, Dot plot of the area-under-the-curve abundance of the newly identified and host-synthesized bile- acid conjugates in each SPF mouse (n = 4), through the mouse gastrointestinal tract and its subsections.
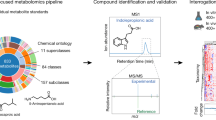
Discovery of new conjugated bile acids
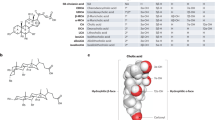
Glycine- and taurine-conjugated bile acids were detected in both germ-free and SPF mice. The glycine and taurine amino acids were removed as they passed through the gastrointestinal tract in SPF mice only, which is a known microbial transformation19 (Fig. 1b, Extended Data Fig. 4). The molecular network of conjugated bile acids had several modified forms of these compounds that were present only in colonized mice, including the C4H8 addition that was related to the tandem mass spectra of GCA (Fig. 2a). Our analysis of the tandem mass spectra of three of these SPF-mouse nodes (m/z 556.363, 572.358 and 522.379) showed the maintenance of the core cholic acid, but with a fragmentation pattern that was characteristic of the presence of phenylalanine, tyrosine or leucine through an amide bond at the conjugation site in place of glycine or taurine (Extended Data Fig. 5, Supplementary Table 2). This represents a set of unique amino acid amide conjugations to cholic acid that are mediated by the microbiome, which create the newly identified bile acids phenylalanocholic acid (Phe-chol), tyrosocholic acid (Tyr-chol) and leucocholic acid (Leu-chol). These structures were validated with synthesized standards by retention time and MS/MS matching on several instrument platforms including targeted mass spectrometry (level one matches16) (Extended Data Figs. 5, 6, Supplementary Tables 2, 5). These molecules were detected in the duodenum, jejunum and ileum of SPF mice only, with tenfold-lower levels found in the caecum and colon after targeted mass spectrometry analysis using isotopically labelled internal standards (Supplementary Table 4). The liver-synthesized glycine and taurine conjugates were not only found in these same gut locations, but were also observed in the gall bladder and liver (Fig. 2b, Extended Data Fig. 6). Phe-chol was the most abundant microbial conjugate, on average, across the gastrointestinal tract; it was present at 147.0 nmol g−1 tissue (s.d. ± 99.9) in the jejunum, 83.6 nmol g−1 tissue (s.d. ± 81.3) in the ileum, 4.7 nmol g−1 tissue (s.d. ± 3.4) in the caecum and 11.6 nmol g−1 tissue (s.d. ± 12.2) in the colon. Phe-chol was present at its highest concentration at 447.2 nmol g−1 tissue in a single sample from the jejunum (limit of detection (LOD) in Supplementary Tables 4, 6, 7).
The decreased abundance of these unique bile conjugates in the lower gastrointestinal tract prompted us to investigate whether there was reabsorption in the ileum or further metabolism by the microbiota. We collected portal and peripheral blood from an additional four SPF and six germ-free mice, and screened for the presence of conjugated bile acids. Both taurocholic acid and GCA were present in the portal and peripheral blood of colonized and sterile mice, but the newly identified amino acid amide conjugates were not detected (Extended Data Fig. 6). Furthermore, incubation of these molecules with an actively growing human faecal batch culture showed that the Tyr-, Phe- and Leu-conjugated bile acids were not deconjugated by the microbiota—even when deconjugation readily occurred on the host-synthesized GCA control, a well-known amidate hydrolase activity of bile acids that is mediated by the human microbiota20 (Extended Data Fig. 6). However, oxidation of the cholate core occurred on all three of the newly identified conjugates, which indicates that they could be modified by microbial enzymes even when no concurrent oxidation of GCA was observed (Extended Data Fig. 6).
In the extensive literature relating to bile acids (comprising more than 42,000 publication records in PubMed21,22,23,24,25,26,27), descriptions of unusual conjugations of bile acids are rare. Through 170 years of research into bile-acid chemistry, the accepted standard has been that mammalian bile acids are amide conjugated by a host liver enzyme (known as bile acid–CoA:amino acid N-acyltransferase (BAAT)) with either glycine or taurine. Here we report amide conjugations with phenylalanine, tyrosine and leucine associated with the microbiome in mice, and show that these compounds are common in humans.
Translation to humans
We performed a search using the Mass Spectrometry Search Tool (MASST) of 1,004 public datasets available in the Global Natural Products Social Molecular Networking (GNPS) database, which revealed spectral matches that correspond to Phe-chol, Tyr-chol and Leu-chol in 28 studies comprising samples from the gastrointestinal tract of both mice (3.2 to 59.4% of all samples) and humans11 (1.6 to 25.3% of all samples) (Extended Data Fig. 7). In data from faecal samples collected for the American Gut Project28, at least one of these unique bile acids was found in 1.6% of human faecal samples; Tyr-chol was the most prevalent (n = 490 samples) (Fig. 3a). These bile acids were found in higher frequency in samples from patients with inflammatory bowel disease or cystic fibrosis, or from infants, than in samples from the American gut project (Fig. 3a).
a, Percentage of samples that were positive for the newly identified bile acids from GNPS public datasets and from paediatric patients with cystic fibrosis (compared to controls without cystic fibrosis). AGP, American gut project28; CF, cystic fibrosis; IBD, inflammatory bowel disease; PS, pancreatic-sufficient; PI, pancreatic-insufficient. The colour coding of the bile acids applies to a–c. b, Abundance of the newly identified conjugates in the PRISM and iHMP (NIH Integrative Human Microbiome Project) datasets31. The statistical significance for the PRISM data was tested using the Wald’s test (Crohn’s disease (CD), n = 68 individuals; ulcerative colitis (UC), n = 53 individuals; noninflammatory bowel disease, n = 34 individuals) and for the iHMP dataset with a linear two-sided mixed-effects model. The iHMP comparisons are separated by type of inflammatory bowel disease, and by dysbiotic or nondysbiotic state (for ulcerative colitis, n = 12 dysbiotic and 110 nondysbiotic metabolomes; for Crohn’s disease, n = 48 dysbiotic, and 169 nondysbiotic metabolomes; for noninflammatory bowel disease, n = 15 dysbiotic and 107 nondysbiotic metabolomes). Significance is shown using Benjamini–Hochberg-corrected P values. Leu-chol, q = 0.031; Tyr-chol, q = 0.0074; Phe-chol, q = 0.0043. *q < 0.05, **q < 0.05. Boxes represent the interquartile range, notch is the 95% confidence interval of the mean, centre is the median and whiskers are 1.5× the interquartile range. c, Extracted ion chromatograms of Phe-chol from cultured isolates of C. bolteae compared to medium control at 0 h and 96 h (top). Experiment was performed twice. d, The ratio of 13C-Phe-chol:12C-Phe-chol in faecal samples of a mouse fed a high- fat diet with 13C-labelled phenylalanine (blue line) or unlabelled phenylalanine (black line) over time. Grey area indicates a three-day period during which a high-fat diet was fed; red indicates when the high-fat diet was supplemented with Phe. e, Quantitative PCR with reverse transcription data showing the mean and s.e.m. of the gene-expression ratio (ΔΔCt) of Fgf15, Shp, Cyp8b1 and Cyp7a1 to the 36B4 (also known as Rplp0) reference control in the ileum and/or liver of mice gavaged with different bile acids, compared to a mock control (corn oil) after 72 h. Statistical significance was tested against the mock control with a two-tailed t-test (n = 4 or 5 mice per group). CA, cholic acid.
We reanalysed data deposited in the GNPS/MassIVE repository from a previously published study of the mouse microbiome and liver cancer, which enabled us to compare the abundance of the newly identified bile acid conjugates in mice fed a high-fat diet in comparison to their abundance when the mice were fed a normal chow with or without antibiotics29 (Extended Data Fig. 7). The Phe, Tyr, and Leu amino acid conjugates were undetectable upon exposure to antibiotics, whereas GCA remained—supporting the role of the microbiome in the newly identified conjugation. In the same study29, Phe-chol and Leu-chol were more abundant in mice fed a high-fat diet, with no change observed in the host-conjugated GCA (Extended Data Fig. 7). We further validated this association in data from a separate study in which atherosclerosis-prone mice fed a high-fat diet also had increased levels of the microbial conjugates, without a corresponding change in the host-produced taurocholic acid (Extended Data Fig. 7). Cystic fibrosis is known to result in insufficient production of pancreatic lipase, microbial dysbiosis and the build-up of fat in the gut30. Reanalysis of the public data from a cohort of paediatric patients, we found that these compounds were more prevalent in patients with cystic fibrosis (particularly in those with pancreatic insufficiency) than in healthy controls (Fig. 3a). Finally, detection of the newly identified conjugates in patients with inflammatory bowel disease led us to mine metabolome data from the second stage of the human microbiome project (HMP2)31, which focused on differences between controls and patients with inflammatory bowel disease, including patients with Crohn’s disease or ulcerative colitis—subtypes of inflammatory bowel disease31 (Fig. 3b, Supplementary Table 8). All three metabolites were significantly higher in the dysbiotic state associated with patients with Crohn’s disease, but not in patients with ulcerative colitis (Fig. 3b, Supplementary Data). Our MASST-based mining of public data from the GNPS database showed that these compounds are not only found in healthy humans but are also enriched in individuals with fatty guts and inflammatory bowel disease, which suggests that these compounds may have a potential role in (or be symptoms of) gut dysbiosis and human disease.
Microorganisms make the new bile acids
There was a strong positive correlation between the presence of a species of Clostridium and all three bile acids when mice were fed a high-fat diet (Pearson’s r for Phe-chol, r = 0.73; for Tyr-chol, r = 0.50; and for Leu-chol, r = 0.74) (Extended Data Fig. 7, Supplementary Table 3). The clostridia are known to oxidize, epimerize and deconjugate bile acids32,33. We therefore cultured 20 human gut microorganisms (with an emphasis on Clostridium species) in faecal culture medium34 that contained amino acids and cholic acid precursors to screen for production of the newly identified conjugates. The Clostridium bolteae strains WAL-14578 and CC43001B both synthesized both Phe-chol and Tyr-chol (Extended Data Fig. 8). The addition of labelled 13C-phenylalanine to the medium verified that WAL-1457 could synthesize Phe-chol from the amino acid and cholate precursors (Extended Data Fig. 8). Similarly, we fed mice a high-fat diet with 13C-phenylalanine and were able to detect labelled Phe-chol in their faeces, which demonstrates microbial synthesis in vivo and shows that the amino acid precursors could come from the diet (Fig. 3d). C. bolteae is a bile-resistant gut bacterium that is more common in children with autism spectrum disorder35, is associated with abdominal infections36 and—together with Blautia producta—prevented colonization by vancomycin-resistant Enterococcus species in mice37. The production of these bile acids by C. bolteae further verifies their association with the microbiota of the mouse gut, and implicates them as potentially important for intermicrobial interactions in the gut microbiome. However, addition of the newly identified conjugates to batch cultures of human faecal samples did not affect community structure (Extended Data Fig. 8), which led us to investigate how these compounds may affect gut physiology through host receptor signalling.
New bile acids and the farnesoid X receptor
The farnesoid X receptor (FXR) is a key receptor for bile acids that is expressed in the intestine, liver and other tissues. The most-potent naturally occurring agonistic ligand of FXR is chenodeoxycholic acid, whereas tauro-β-muricholic acid is an FXR antagonist38. To assess the ability of the newly identified bile acids to affect human FXR signalling, we established a luciferase reporter assay in human embryonic kidney (HEK)293 cells39. Phe-chol and Tyr-chol were strong human-FXR agonists (Extended Data Fig. 9, Supplementary Table 9). The phenylalanine conjugate (R2 = 0.92, half maximal effective concentration (EC50) = 4.5 μM) was twice as strong of an agonist as chenodeoxycholic acid (R2 = 0.88, EC50 = 9.7 μM), and the tyrosine conjugate was the most potent of them all (R2 = 0.93, EC50 = 0.14 μM). Furthermore, gavage of mice with these compounds increased expression of the FXR effector genes Fgf15 and Shp (also known as Nr0b2) in the intestine (12.2- and 13.3-fold with Tyr-chol at 24 h, P = 0.029 and 0.009; 6.2 and 9.3-fold at 72 h, P = 0.009 and 0.019) (Fig. 3e, Extended Data Fig. 9). Although Shp expression did not change detectably in the liver at 24 h after gavage, levels were increased 2.3-fold after 72 h (P = 0.017) (Fig. 3e, Extended Data Fig. 9). Changes in expression of the bile-acid synthesis genes Cyp7a1 and Cyp8b1 also showed a time-dependent effect. Cyp7a1 was at 9% of control levels at 24 h (P = 0.001) and Cyp8b1 was at 69% (P = 0.004) (Extended Data Fig. 9). At 72 h (after 4 gavages), Cyp7a1 expression was at 8% of control levels (P = 0.004), and for Cyp8b1 the transcript was further reduced to 2% (P = 0.0002) (Fig. 3e). The strong time-dependent reduction in liver Cyp7a1 and Cyp8b1 transcripts indicates that—similar to the primary bile acid cholic acid—gavage of mice with the newly identified compounds reduced the expression of downstream FXR-target genes that are responsible for bile-acid synthesis in the liver. However, the possibility that this effect was due to FXR agonism through release of cholate from amide conjugate hydrolysis cannot be excluded.
Bile-acid metabolism by the microbiome was first described in the 1960s40. The four known mechanisms of microbial metabolism are dehydroxylation, dehydration and epimerization of the cholesterol backbone, and deconjugation of the amino acids glycine or taurine1,41,42. Here, we identify bile-acid transformation by the microbiome mediated by a fifth and completely different mechanism: amide conjugation of the cholate backbone with the amino acids phenylalanine, tyrosine and leucine. Although there are homologues of the human bile-acid-conjugation gene BAAT in clostridial genomes, the microbial enzyme in question remains unknown. Regardless of the mechanism of their synthesis, the newly identified conjugates stimulate the human FXR receptor in a cell-based system and the expression of FXR-target genes that are responsible for bile-acid production in the liver were reduced when administered to mice. Additional studies are needed to understand the health implications of bile-acid reconjugation by the human microbiome and its potential effects on FXR-related diseases.
Conclusion
This study shows that the chemistry of all organ systems is affected by the presence of the microbiome. The strongest signatures come from the gut, particularly via the breakdown of plant natural products from food and the manipulation of bile acids. The microbiome is primarily a catabolic entity, breaking down compounds through the enzymatic removal of chemical groups. However, we found an anabolic reaction that represents a fifth mechanism of bile-acid metabolism by the microbiome, which operates through unique amino acid conjugations of cholic acid. As the connections between humans and our microbial symbionts become increasingly appreciated, a combination of globally untargeted approaches and the development of tools that interlink these datasets (such as the GNPS and MASST analysis infrastructure) will enable the more-efficient characterization of microbial molecules and efficient translation between model animals and human studies, leading to a better understanding of the deep connection between our microbiota, our metabolites and our health.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
All metabolomics data that support the findings of this study are available at GNPS (https://gnps.ucsd.edu/) under MassIVE ID numbers: MSV000079949 (original germ-free and SPF mouse data), MSV000082480, MSV000082467, MSV000079134, MSV000082406, MSV000083032, MSV000083004 and MSV000083446. The sequencing data for the germ-free and SPF mouse study are available on the Qiita microbiome data analysis platform at https://qiita.ucsd.edu/ under study ID 10801 and through the European Bioinformatics Institute accession number ERP109688. Source Data for Figs. 1–3, Extended Data Fig. 7 are provided with the paper.
Code availability
MASST can be accessed at https://masst.ucsd.edu/; the development of MASST is described in ref. 43. The code for MS/MS-based MASST searching is available at https://github.com/CCMS-UCSD/GNPS_Workflows/tree/master/search_single_spectrum.
References
Ridlon, J. M., Kang, D. J., Hylemon, P. B. & Bajaj, J. S. Bile acids and the gut microbiome. Curr. Opin. Gastroenterol. 30, 332–338 (2014).
Gilbert, J. A. et al. Microbiome-wide association studies link dynamic microbial consortia to disease. Nature 535, 94–103 (2016).
Wikoff, W. R. et al. Metabolomics analysis reveals large effects of gut microflora on mammalian blood metabolites. Proc. Natl Acad. Sci. USA 106, 3698–3703 (2009).
Marcobal, A. et al. Metabolome progression during early gut microbial colonization of gnotobiotic mice. Sci. Rep. 5, 11589 (2015).
Miller, T. L. & Wolin, M. J. Pathways of acetate, propionate, and butyrate formation by the human fecal microbial flora. Appl. Environ. Microbiol. 62, 1589–1592 (1996).
Gillner, M., Bergman, J., Cambillau, C., Fernström, B. & Gustafsson, J. A. Interactions of indoles with specific binding sites for 2,3,7,8-tetrachlorodibenzo-p-dioxin in rat liver. Mol. Pharmacol. 28, 357–363 (1985).
Martin, F.-P. J. et al. A top-down systems biology view of microbiome-mammalian metabolic interactions in a mouse model. Mol. Syst. Biol. 3, 112 (2007).
Moriya, T., Satomi, Y., Murata, S., Sawada, H. & Kobayashi, H. Effect of gut microbiota on host whole metabolome. Metabolomics 13, 101 (2017).
Swann, J. R. et al. Systemic gut microbial modulation of bile acid metabolism in host tissue compartments. Proc. Natl Acad. Sci. USA 108 (Suppl 1), 4523–4530 (2011).
Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Wang, M. et al. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat. Biotechnol. 34, 828–837 (2016).
Watrous, J. et al. Mass spectral molecular networking of living microbial colonies. Proc. Natl Acad. Sci. USA 109, E1743–E1752 (2012).
Protsyuk, I. et al. 3D molecular cartography using LC-MS facilitated by Optimus and ’ili software. Nat. Protocols 13, 134–154 (2018).
Hofmann, A. F. & Hagey, L. R. Key discoveries in bile acid chemistry and biology and their clinical applications: history of the last eight decades. J. Lipid Res. 55, 1553–1595 (2014).
Yang, J. Y. et al. Molecular networking as a dereplication strategy. J. Nat. Prod. 76, 1686–1699 (2013).
Sumner, L. W. et al. Proposed minimum reporting standards for chemical analysis Chemical Analysis Working Group (CAWG) Metabolomics Standards Initiative (MSI). Metabolomics 3, 211–221 (2007).
Hartmann, A. C. et al. Meta-mass shift chemical profiling of metabolomes from coral reefs. Proc. Natl Acad. Sci. USA 114, 11685–11690 (2017).
Hirano, S. & Masuda, N. Characterization of NADP-dependent 7β-hydroxysteroid dehydrogenases from Peptostreptococcus productus and Eubacterium aerofaciens. Appl. Environ. Microbiol. 43, 1057–1063 (1982).
Wahlström, A., Sayin, S. I., Marschall, H.-U. & Bäckhed, F. Intestinal crosstalk between bile acids and microbiota and its impact on host metabolism. Cell Metab. 24, 41–50 (2016).
Huijghebaert, S. M. & Hofmann, A. F. Influence of the amino acid moiety on deconjugation of bile acid amidates by cholylglycine hydrolase or human fecal cultures. J. Lipid Res. 27, 742–752 (1986).
Myher, J. J., Marai, L., Kuksis, A., Yousef, I. M. & Fisher, M. M. Identification of ornithine and arginine conjugates of cholic acid by mass spectrometry. Can. J. Biochem. 53, 583–590 (1975).
Peric-Golia, L. & Jones, R. S. Ornithocholanic acids and cholelithiasis in man. Science 142, 245–246 (1963).
Gordon, B. A., Kuksis, A. & Beveridge, J. M. R. Separation of bile acid conjugates by ion exchange chromatography. Can. J. Biochem. Physiol. 41, 77–89 (1963).
Yousef, I. M. & Fisher, M. M. Bile acid metabolism in mammals. VIII. Biliary secretion of cholylarginine by the isolated perfused rat liver. Can. J. Physiol. Pharmacol. 53, 880–887 (1975).
Tamari, M., Ogawa, M. & Kametaka, M. A new bile acid conjugate, ciliatocholic acid, from bovine gall bladder bile. J. Biochem. 80, 371–377 (1976).
Hagey, L. R., Schteingart, C. D., Rossi, S. S., Ton-Nu, H. T. & Hofmann, A. F. An N-acyl glycyltaurine conjugate of deoxycholic acid in the biliary bile acids of the rabbit. J. Lipid Res. 39, 2119–2124 (1998).
Nair, P. P., Solomon, R., Bankoski, J. & Plapinger, R. Bile acids in tissues: binding of lithocholic acid to protein. Lipids 13, 966–970 (1978).
McDonald, D. et al. American gut: an open platform for citizen science microbiome research. mSystems 3, e00031-18 (2018).
Shalapour, S. et al. Inflammation-induced IgA+ cells dismantle anti-liver cancer immunity. Nature 551, 340–345 (2017).
Manor, O. et al. Metagenomic evidence for taxonomic dysbiosis and functional imbalance in the gastrointestinal tracts of children with cystic fibrosis. Sci. Rep. 6, 22493 (2016).
Lloyd-Price, J. et al. Multi-omics of the gut microbial ecosystem in inflammatory bowel diseases. Nature 569, 655–662 (2019).
Hirano, S., Masuda, N., Oda, H. & Mukai, H. Transformation of bile acids by Clostridium perfringens. Appl. Environ. Microbiol. 42, 394–399 (1981).
Winston, J. A. & Theriot, C. M. Impact of microbial derived secondary bile acids on colonization resistance against Clostridium difficile in the gastrointestinal tract. Anaerobe 41, 44–50 (2016).
McDonald, J. A. K. et al. Evaluation of microbial community reproducibility, stability and composition in a human distal gut chemostat model. J. Microbiol. Methods 95, 167–174 (2013).
Finegold, S. M. et al. Pyrosequencing study of fecal microflora of autistic and control children. Anaerobe 16, 444–453 (2010).
Dehoux, P. et al. Comparative genomics of Clostridium bolteae and Clostridium clostridioforme reveals species-specific genomic properties and numerous putative antibiotic resistance determinants. BMC Genomics 17, 819 (2016).
Caballero, S. et al. Cooperating commensals restore colonization resistance to vancomycin-resistant Enterococcus faecium. Cell Host Microbe 21, 592–602.e4 (2017).
Sayin, S. I. et al. Gut microbiota regulates bile acid metabolism by reducing the levels of tauro-beta-muricholic acid, a naturally occurring FXR antagonist. Cell Metab. 17, 225–235 (2013).
Downes, M. et al. A chemical, genetic, and structural analysis of the nuclear bile acid receptor FXR. Mol. Cell 11, 1079–1092 (2003).
Gustafsson, B. E., Gustafsson, J. A. & Sjövall, J. Intestinal and fecal sterols in germfree and conventional rats. Bile acids and steroids 172. Acta Chem. Scand. 20, 1827–1835 (1966).
Midtvedt, T. Microbial bile acid transformation. Am. J. Clin. Nutr. 27, 1341–1347 (1974).
Gérard, P. Metabolism of cholesterol and bile acids by the gut microbiota. Pathogens 3, 14–24 (2013).
Wang, M. et al. Mass spectrometry searches using MASST. Nat. Biotechnol. 38, 23–26 (2020).
Acknowledgements
The authors acknowledge funding from the National Institutes of Health (NIH), grants 5U01AI124316-03, 1R03CA211211-01, 1R01HL116235 U54DE023798, R24DK110499, GMS10RR029121, 1 DP1 AT010885, P30 DK120515 and R01HD084163. Additionally, B.S.B. was supported by UCSD KL2 (1KL2TR001444), T.D.A. by the National Library of Medicine Training Grant NIH grant T15LM011271. R.M.E. is an investigator of the Howard Hughes Medical Institute and March of Dimes Chair in Molecular and Developmental Biology at the Salk Institute. R.M.E. was funded by grants from the NIH (DK057978, HL105278, HL088093 and ES010337), and Samuel Waxman Cancer Research Foundation. We acknowledge G. Ackermann for her contributions. This work was also supported in part by Seed Grants from the UC San Diego Center for Microbiome Innovation. This work was funded by grants from the NIH (DK057978, HL105278 and HL088093), National Cancer Institute (CA014195), the Leona M. and Harry B. Helmsley Charitable Trust (2017PG-MED001), SWCRF Investigator Award and Ipsen/Biomeasure. J.L. is supported by grant EIA14660045, an American Heart Association Established Investigator Award. T.F. is supported by a Hewitt Medical Foundation Fellowship, a Salk Alumni Fellowship. T.F., R.K. and P.C.D. acknowledge support from the Crohn’s & Colitis Foundation (CCFA). R.M.E. and M.D. are supported in part by a Stand Up to Cancer (SU2C) - Cancer Research UK-Lustgarten Foundation Pancreatic Cancer Dream Team Research Grant (SU2C-AACR-DT-20-16). SU2C is a programme of the Entertainment Industry Foundation. Research grants are administered by the American Association for Cancer Research, the scientific partner of SU2C. Research reported in this publication was also supported by the National Institute of Environmental Health Sciences of the NIH under Award Number P42ES010337. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH.
Author information
Authors and Affiliations
Contributions
P.C.D., R.K. and R.A.Q. designed the project. P.C.D. and R.A.Q. discovered the bile acids. R.A.Q., A.A., A.V.M., F.V., J.M.G., N.G., A.T., M.P.C., L.K.C., A.T.N., M.J.M., G.H., M.P., C.C., S.B., E.G., K.C.W., P.B.-F., H.M., E.A.F., H.V., J.L.-P., T.D.A., A.C.K., J.A.-P., Z.B., E.L., M.Q., T.F., R.D.W., D.F., G.G.H. and R.B. generated data. R.A.Q., A.V., A.T., Z.B., A.V.M., R.d.S., R.J.X., T.F., R.D.W., M.D., R.M.E. and M.P.C. analysed data. R.A.Q., B.S.B., M.L., O.P., J.T.C., M.L., J.C.L., K.A.P., B.I.K., R.J., M.E., K.E.R., G.H., C.C., W.J.S. and R.B. collected samples. D.S., E.G., M.P.C. and A.T.N. were responsible for chemical synthesis. T.F., R.D.W., M.D. and R.M.E. were responsible for the FXR in vitro and in vivo functional studies and analysis, and edited the paper. P.C.D., R.K., S.K.M., V.N., C.H., R.J.X., A.C.K. and D.S. guided experimental design and analysis. M.W. converted the data in GNPS and developed MASST spectral search. T.T., V.N., M.R. and S.K.M. raised mice and guided experimental design. R.A.Q. and P.C.D. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
C.H. is on the scientific advisory board of Seres Therapeutics. M.W. is founder of, and A.A. is a consultant for, Ometa Laboratories LLC. P.C.D. and M.W. are consultants for Sirenas Therapeutics. W.J.S. consults for Abbvie, Allergan, Amgen, Arena Pharmaceuticals, Avexegen Therapeutics, BeiGene, Boehringer Ingelheim, Celgene, Celltrion, Conatus, Cosmo, Escalier Biosciences, Ferring, Forbion, Genentech, Gilead Sciences, Gossamer Bio, Incyte, Janssen, Kyowa Kirin Pharmaceutical Research, Landos Biopharma, Lilly, Oppilan Pharma, Otsuka, Pfizer, Progenity, Prometheus Biosciences (merger of Precision IBD and Prometheus Laboratories), Reistone, Ritter Pharmaceuticals, Robarts Clinical Trials (owned by Health Academic Research Trust, HART), Series Therapeutics, Shire, Sienna Biopharmaceuticals, Sigmoid Biotechnologies, Sterna Biologicals, Sublimity Therapeutics, Takeda, Theravance Biopharma, Tigenix, Tillotts Pharma, UCB Pharma, Ventyx Biosciences, Vimalan Biosciences and Vivelix Pharmaceuticals; and holds stock or stock options from BeiGene, Escalier Biosciences, Gossamer Bio, Oppilan Pharma, Prometheus Biosciences (merger of Precision IBD and Prometheus Laboratories), Progenity, Ritter Pharmaceuticals, Ventyx Biosciences and Vimalan Biosciences.
Additional information
Peer review information Nature thanks Hanns-Ulrich Marschall, Trent Northen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Microbiome and metabolome diversity in germ-free and SPF mice.
a, Principal coordinate (PC) analysis of microbiome and mass-spectrometry data highlighted by sample source as germ-free (GF) or SPF (n = 4 mice in each group). The microbial signatures from the germ-free mice are an important control, which represents background reads found in buffers, tips and tubes and other experimental materials. b, Data from a highlighted by organ source (n = 4 mice in each group). c, Bray–Curtis dissimilarities of the metabolome data collected from mouse organs. The dissimilarities are calculated within individual mice of the same group (germ-free or SPF, ‘within’) or across the germ-free and SPF groups (‘GF-SPF’) (n = 4 mice in each group). Only samples collected from exact same location (subsection) are compared. Significance was tested with a two-sided Mann–Whitney U-test. Boxes represent the interquartile range (IQR), the notch is the 95% confidence interval of the mean, the centre is the median and whiskers are 1.5× the IQR. d, Microbiome profile of the gastrointestinal tracts of SPF mice. Data were generated by sequencing 16S rRNA gene amplicons from each organ and organ section, and analysed through the Qiita Deblur pipeline as described in the Supplementary Methods. Bacterial taxa of relevance are colour-coded according to the legend. e, Molecular network of LC–MS/MS data with nodes coloured by source as germ-free, SPF, shared or detected in blanks. Molecular families with metabolites annotated by spectral matching in GNPS are listed by a number that corresponds to the molecular family. These are level-2 or -3 annotations according to the metabolomics standards consortium16. 12-OAHSA, 12-(9Z-octadecenoyloxy)-octadecanoic acid.
Extended Data Fig. 2 Microbial metabolism of soyasaponins in metabolomics data from germ-free and SPF mice.
n = 4 mice in each group. a, Molecular network cluster of soyasaponins, coloured by source of each node as germ-free, SPF or shared. Structures of corresponding molecules are shown in nodes highlighted in yellow, according to the numbering scheme. Mean total-ion-current-normalized abundance of each soyasaponin metabolite from the gastrointestinal tracts of germ-free and SPF mice. Ce, caecum; co, colon; D, duodenum; I, ileum; J, jejunum; stl, stool; sto, stomach. Boxes represent the IQR, the centre is the median and whiskers are 1.5× the IQR. n = 4 mice in each group. b, Molecular family of soyasapogenols, their structures and relative abundances in gut organs of germ-free and SPF mice (data are in the same format as in a). c, Three-dimesional model visualization (generated using ’ili) of the normalized abundance of soyasaponin I in the mouse gastrointestinal tract. The abundance of the metabolite is indicated according to the rainbow spectrum (high, red; low, blue). n = 4 mice in each group. d, Three-dimensional cartography (generated using ’ili) of the normalized abundance of soyasapogenol B onto an magnetic resonance imaging organ model of the mice. e, Mean normalized abundance of soyasaponin I through all gastrointestinal sample locations in the germ-free and SPF mice. f, Mean normalized abundance of soyasapogenol through all gastrointestinal sample locations. The annotations are level two or three16.
Extended Data Fig. 3 Microbial metabolism of plant isoflavones in metabolomics data from germ-free and SPF mice.
a, Structures, molecular network and total-ion-chromatogram-normalized abundance of glycone isoflavanoids in the mouse gastrointestinal tract. Nodes are coloured according to their source in germ-free or SPF mice (n = 4 mice each), and known library hits are shaped as arrowheads. Boxes represent the IQR, the centre is the median and whiskers are 1.5× the IQR. b, Same information as in a, for the aglycones. c, Three-dimensional molecular cartography mapping the abundance of the daidzein and glycitein glycone and sulfated forms through entire 3D mouse model. The normalized abundance of a particular molecule is indicated as a heat map. Red, most abundant; blue, least abundant. d, Three-dimesional molecular cartography mapping the abundance of the daidzein and glycitein aglycone forms through entire 3D mouse model. The gastrointestinal-tract model is inset for reference. The annotations are level two or three16.
Extended Data Fig. 4 Microbial metabolism of known bile acids in metabolomics data from germ-free and SPF mice.
n = 4 mice in each group. a, Total-ion-chromatogram-normalized abundance of taurocholic acid and secondary bile acids in gastrointestinal tract samples from germ-free and SPF mice. Gall, gall bladder; liv, liver. Boxes represent the IQR, the centre is the median and whiskers are 1.5× the IQR. b, Three-dimesional molecular cartography mapping the abundance of the same bile acids as in a through the mouse gastrointestinal-tract model; liver is separated for better visualization. The normalized abundance of a particular molecule is indicated as a heat map. Red, most abundant; blue, least abundance. The annotations are level two or three16.
Extended Data Fig. 5 Mass spectrometry analysis of newly identified conjugated bile acids.
a, Extracted-ion-chromatogram MS1 traces of Tyr-chol (m/z 572.37 ± 0.05 Da), Phe-chol (m/z 556.37 ± 0.05 Da) and Leu-chol (m/z 522.37 ± 0.05 Da). Experiments were performed four times. b, Extracted ion chromatograms for the synthetic muricholic and cholic acid versions of the Phe (m/z 556.37 ± 0.05), Tyr (572.37 ± 0.05) and Leu (522.37 ± 0.05) conjugates, showing the different retention times from the muricholic- and cholic-acid forms. c, Retention time alignments of synthetic muricholic- and cholic-acid conjugates with the newly identified conjugates found in a sample from the jejunum of a colonized mouse. The isoleucocholic- and leucocholic-acid analysis was run on a long-gradient high-performance liquid-chromatography column to separate isomeric Ile and Leu conjugates, and to compare to those detected in vivo. d, Annotation of MS/MS fragmentation patterns for the three conjugated bile acids and GCA. Structures of the immonium ions from amino acid fragmentation, whole amino acid fragments and the major sterol fragment are shown. Loss of the amino acid mass on the bile-acid steroid backbone is also highlighted.
Extended Data Fig. 6 Distribution and metabolism of newly identified conjugated bile acids.
a, Molecular network of MS/MS data from synthesized amino acid conjugated bile acids and the duodenum of SPF mice. LC–MS/MS data from synthetic standards were networked with mouse samples and spectral matching. Molecular networking is indicated by node colouring. Mirror plots show the alignment between the mouse and the synthetic standards. Nodes shaped as arrowheads had hits in the GNPS libraries, and node size is scaled to the spectral count. Tauro, taurocholic acid. These experiments were performed twice. b, Three-dimensional molecular cartography of the mean abundance of the newly discovered conjugates mapped onto a 3D-rendered model of the mouse gastrointestinal tract, as a heat map according to the colour scale. Organs are labelled as described in Fig. 1. c, Molecular network of conjugated bile acids from portal and peripheral blood of germ-free and SPF mice. Nodes are coloured by source as germ-free portal, germ-free portal and peripheral blood, SPF portal and peripheral blood, GF portal and peripheral blood and SPF peripheral blood, and all. Arrowhead nodes represent known compounds in the GNPS spectral database; circular nodes represent unknown compounds. The annotations were obtained through spectral matches against reference libraries (level two or three16). d, Mean area-under-the-curve abundance and s.d. of bile acids of interest during incubation with an actively growing batch human faecal culture for 24 h (n = 3 independent incubations). e, Molecular network of newly identified conjugated bile acids after incubation in a human faecal batch culture experiment. Each node represents a unique tandem mass spectrum; arrowhead-shaped nodes indicate known spectra in the GNPS database. The nodes are coloured by their retention time according to the legend, and the mass shifts between nodes are mapped onto the edge representing the cosine connection between related spectra. The H2 mass shift representing oxidation of the newly identified conjugates is shown. f, Mean ion intensity and s.d. of the oxidized forms of Phe-chol, Tyr-chol and Leu-chol through the 24-h batch faecal culture incubation (n = 3 independent incubations).
Extended Data Fig. 7 MASST search results and associations of newly identified conjugated bile acids with high-fat diet.
a, Proportion of samples in which Phe-chol, Tyr-chol and Leu-chol were found from a single-spectrum MASST search of publicly available data on GNPS. Massive dataset identifers are shown for each dataset, are divided into mouse (‘murine’) or human gastrointestinal samples. b, Box plots of the newly identified conjugates in a previously published mouse study, in which mice were fed high-fat diet (HFD) (n = 14 mice) or normal chow (NC) (n = 19 mice) (Gly, P = 0.72; Phe, P = 0.038; Tyr, P = 0.083; Leu P = 9.4 × 10−5) and dot plot of mice treated with (n = 27 mice) or without antibiotics (Ab) (n = 415 mice)29. Boxes represent the IQR, the line is the median and whiskers are 1.5× the IQR. Colour legend applies to both a and b. c, Mean normalized abundance of the three newly identified conjugated bile acids compared to taurocholic acid in mice (Apoe-knockout on a C57BL/6J background) fed either a high-fat diet (n = 12 mice) or normal chow (n = 12 mice) for 10 weeks. Faecal samples were collected and extracted in 50:50 methanol:water and analysed with LC–MS/MS metabolomics, as described in the Supplementary Methods. The s.d. around the mean is shown, and significance between a high-fat diet and normal chow at each time point is tested with two-sided Student’s t-test. ***P < 0.001. d, Correlations between rarefied reads of a deblurred read assigned to a Clostridium sp. from atherosclerosis-prone mice fed a high-fat diet over time (n = 12 mice). The line of best fit is plotted using the lm method in the R statistical software; grey area around the line of best fit is the 95% confidence interval.
Extended Data Fig. 8 Synthesis of newly identified conjugated bile acids by Clostridium.
a, Dot plot of the measured production of Phe-chol and Tyr-chol using a targeted liquid chromatography–mass spectrometry method for two C. bolteae strains grown in faecal culture medium (FCM) with or without labelled Phe (n = 2 independent cultures). b, The mean ratio and s.e.m. of 13C-Phe-chol:12C-phe-chol from the same C. bolteae strains when grown with faecal culture medium with 13C-labelled phenylalanine (bottom left) (n = 2 cultures). c, Mean and s.d. of the Shannon index of human faecal batch culture (n = 3 cultures) before and after 24-h growth exposed to conjugated bile acids or a mock control. NS, not significant by Mann–Whitney U-test. d, Box-and-whisker plots of concentration of Phe-chol and Tyr-chol in original samples from the gut of SPF mice. Boxes represent the IQR, the centre is the median and whiskers are 1.5× the IQR. n = 4 mice.
Extended Data Fig. 9 Effect of newly identified bile acids on FXR.
a, Mean normalized luciferase activity as a readout of human FXR stimulation when exposed to various conjugated and unconjugated bile acids, as a function of the compound dose. n = 8 measurements, ± s.e.m. DCA, deoxycholic acid; CDCA, chenodeoxycholic acid; T-βMCA, tauro-β-muricholic acid. b, Ileum mean fold expression change compared to 36B4 control of various bile acids after gavage in mice. Error bars are s.e.m. c, Liver fold expression change compared to 36B4 control of various bile acids after gavage in mice. Significance was tested with two-tailed t-test compared to the mock corn-oil control. Error bars are s.e.m.
Supplementary information
Supplementary Information
This file contains Methods, Supplementary Data, Supplementary Tables, Supplementary NMR spectra and Supplementary 3D mouse model. Supplemental 3D-Mouse model: provided in the SI are .stl files that comprise the 3D mouse model for 3D-molecular cartography mapping. These include a full mouse model, the liver only, the GI tract only, and the GI tract without the liver. Also included are x,y,z coordinates that will enable mapping of multi-omics data to locations of interest on all four .stl files.
Supplementary Table
This file contains Supplementary Table S1: Sample information from GF and SPF mice analyzed in this study.
Supplementary Table
This file contains Supplementary Table S3: Deblurred read, taxonomic assignment and Pearson’s r correlation for bacterial assigned sequence variants (ASVs) with the three novel conjugated bile acids in the HFD feeding experiment (n=12).
Rights and permissions
About this article
Cite this article
Quinn, R.A., Melnik, A.V., Vrbanac, A. et al. Global chemical effects of the microbiome include new bile-acid conjugations. Nature 579, 123–129 (2020). https://doi.org/10.1038/s41586-020-2047-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2047-9
- Springer Nature Limited
This article is cited by
-
How bile acids and the microbiota interact to shape host immunity
Nature Reviews Immunology (2024)
-
The changing metabolic landscape of bile acids – keys to metabolism and immune regulation
Nature Reviews Gastroenterology & Hepatology (2024)
-
Bile salt hydrolase acyltransferase activity expands bile acid diversity
Nature (2024)
-
Another renaissance for bile acid gastrointestinal microbiology
Nature Reviews Gastroenterology & Hepatology (2024)
-
Fast mass spectrometry search and clustering of untargeted metabolomics data
Nature Biotechnology (2024)