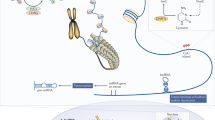
Abstract
Systemic sclerosis (SSc) is a prototypical inflammatory fibrotic disease involving inflammation, vascular abnormalities and fibrosis that primarily affect the skin and lungs. The aetiology of SSc is unknown and its pathogenesis is only partially understood. Of all the rheumatic diseases, SSc carries the highest all-cause mortality rate and represents an unmet medical need. A growing body of evidence implicates epigenetic aberrations in this intractable disease, including specific modifications affecting the three main cell types involved in SSc pathogenesis: immune cells, endothelial cells and fibroblasts. In this Review, we discuss the latest insights into the role of DNA methylation, histone modifications and non-coding RNAs in SSc and how these epigenetic alterations affect disease features. In particular, histone modifications have a role in the regulation of gene expression pertinent to activation of fibroblasts to myofibroblasts, governing their fate. DNA methyltransferases are crucial in disease pathogenesis by mediating methylation of DNA in specific promoters, regulating expression of specific pathways. We discuss targeting of these enzymes for therapeutic gain. Innovative epigenetic therapy could be targeted to treat the disease in a precision epigenetics approach.
Key points
-
In systemic sclerosis (SSc), epigenetic aberrations are prominent in the main cell types involved in the disease pathogenesis.
-
DNA in SSc fibroblasts seems to be hypermethylated, leading to repression of gene expression of negative regulators such as SOCS3.
-
Studies of open regions of chromatin using ATAC sequencing have identified multiple regions of transcriptionally active genes, although their function (or functions) needs further investigation in understanding the role in SSc pathogenesis.
-
Non-coding RNAs, including long non-coding RNAs and microRNAs, have been linked to SSc in the past few years and might be targets for anti-fibrotic therapy through alteration of their levels.
-
Epigenetic drugs already in use for other indications, such as decitabine, could be repurposed for SSc.


Similar content being viewed by others
References
Denton, C. P. & Khanna, D. Systemic sclerosis. Lancet 390, 1685–1699 (2017).
Hinchcliff, M. & O’Reilly, S. Current and potential new targets in systemic sclerosis therapy: a new hope. Curr. Rheumatol. Rep. 22, 42 (2020).
Simeón-Aznar, C. P. et al. Registry of the Spanish network for systemic sclerosis: clinical pattern according to cutaneous subsets and immunological status. Semin. Arthritis Rheum. 41, 789–800 (2012).
Vonk, M. C. et al. Systemic sclerosis and its pulmonary complications in The Netherlands: an epidemiological study. Ann. Rheum. Dis. 68, 961–965 (2009).
Allanore, Y. et al. Systemic sclerosis. Nat. Rev. Dis. Primers 1, 15002 (2015).
Varga, J. & Abraham, D. Systemic sclerosis: a prototypic multisystem fibrotic disorder. J. Clin. Invest. 117, 557–567 (2007).
Luo, Y., Wang, Y., Wang, Q., Xiao, R. & Lu, Q. Systemic sclerosis: genetics and epigenetics. J. Autoimmun. 41, 161–167 (2013).
Gladman, D. D. et al. HLA markers for susceptibility and expression in scleroderma. J. Rheumatol. 32, 1481 (2005).
Beretta, L. et al. Analysis of Class II human leucocyte antigens in Italian and Spanish systemic sclerosis. Rheumatology 51, 52–59 (2012).
Patel, S. et al. Occupational silica exposure in an Australian systemic sclerosis cohort. Rheumatology 59, 3900–3905 (2020).
Dostert, C. et al. Innate immune activation through Nalp3 inflammasome sensing of asbestos and silica. Science 320, 674 (2008).
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007).
Lyko, F. The DNA methyltransferase family: a versatile toolkit for epigenetic regulation. Nat. Rev. Genet. 19, 81–92 (2018).
Bostick, M. et al. UHRF1 plays a role in maintaining DNA methylation in mammalian cells. Science 317, 1760–1764 (2007).
Tahiliani, M. et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–935 (2009).
He, Y. F. et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 333, 1303–1307 (2011).
Shen, L. et al. Genome-wide analysis reveals TET- and TDG-dependent 5-methylcytosine oxidation dynamics. Cell 153, 692–706 (2013).
Delatte, B., Deplus, R. & Fuks, F. Playing TETris with DNA modifications. EMBO J. 33, 1198–1211 (2014).
Dees, C. et al. TGF-β–induced epigenetic deregulation of SOCS3 facilitates STAT3 signaling to promote fibrosis. J. Clin. Invest. 130, 2347–2363 (2020).
Henderson, J., Distler, J. & O’Reilly, S. The role of epigenetic modifications in systemic sclerosis: a druggable target. Trends Mol. Med. 25, 395–411 (2019).
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Horsburgh, S. et al. MicroRNAs in the skin: role in development, homoeostasis and regeneration. Clin. Sci. 131, 1923–1940 (2017).
Ozsolak, F. et al. Chromatin structure analyses identify miRNA promoters. Genes Dev. 22, 3172–3183 (2008).
Han, J. et al. Molecular basis for the recognition of primary microRNAs by the Drosha-DGCR8 complex. Cell 125, 887–901 (2006).
Denli, A. M., Tops, B. B., Plasterk, R. H., Ketting, R. F. & Hannon, G. J. Processing of primary microRNAs by the microprocessor complex. Nature 432, 231–235 (2004).
Lund, E., Güttinger, S., Calado, A., Dahlberg, J. E. & Kutay, U. Nuclear export of microRNA precursors. Science 303, 95 (2004).
Hutvágner, G. et al. A cellular function for the RNA-interference enzyme Dicer in the maturation of the let-7 small temporal RNA. Science 293, 834 (2001).
Uszczynska-Ratajczak, B., Lagarde, J., Frankish, A., Guigó, R. & Johnson, R. Towards a complete map of the human long non-coding RNA transcriptome. Nat. Rev. Genet. 19, 535–548 (2018).
Xiang, J. F. et al. Human colorectal cancer-specific CCAT1-L lncRNA regulates long-range chromatin interactions at the MYC locus. Cell Res. 24, 513–531 (2014).
He, X. et al. C-Myc-activated long noncoding RNA CCAT1 promotes colon cancer cell proliferation and invasion. Tumour Biol. 35, 12181–12188 (2014).
Zhao, J., Sun, B. K., Erwin, J. A., Song, J. J. & Lee, J. T. Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science 322, 750–756 (2008).
O’Leary, V. B. et al. PARTICLE, a triplex-forming long ncRNA, regulates locus-specific methylation in response to low-dose irradiation. Cell Rep. 11, 474–485 (2015).
Mondal, T. et al. MEG3 long noncoding RNA regulates the TGF-β pathway genes through formation of RNA-DNA triplex structures. Nat. Commun. 6, 7743 (2015).
Thomson, D. W. & Dinger, M. E. Endogenous microRNA sponges: evidence and controversy. Nat. Rev. Genet. 17, 272–283 (2016).
Piwecka, M. et al. Loss of a mammalian circular RNA locus causes miRNA deregulation and affects brain function. Science 357, eaam8526 (2017).
Luger, K., Dechassa, M. L. & Tremethick, D. J. New insights into nucleosome and chromatin structure: an ordered state or a disordered affair? Nat. Rev. Mol. Cell Biol. 13, 436–447 (2012).
Tessarz, P. & Kouzarides, T. Histone core modifications regulating nucleosome structure and dynamics. Nat. Rev. Mol. Cell Biol. 15, 703–708 (2014).
Zhang, D. et al. Metabolic regulation of gene expression by histone lactylation. Nature 574, 575–580 (2019).
Wang, Y. et al. Human PAD4 regulates histone arginine methylation levels via demethylimination. Science 306, 279–283 (2004).
Shi, Y. et al. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 119, 941–953 (2004).
Tsukada, Y. et al. Histone demethylation by a family of JmjC domain-containing proteins. Nature 439, 811–816 (2006).
Dowson, C., Simpson, N., Duffy, L. & O’Reilly, S. Innate immunity in systemic sclerosis. Curr. Rheumatol. Rep. 19, 2 (2017).
Lei, W. et al. Abnormal DNA methylation in CD4+ T cells from patients with systemic lupus erythematosus, systemic sclerosis, and dermatomyositis. Scand. J. Rheumatol. 38, 369–374 (2009).
Lian, X. et al. DNA demethylation of CD40L in CD4+ T cells from women with systemic sclerosis: A possible explanation for female susceptibility. Arthritis Rheum. 64, 2338–2345 (2012).
Elgueta, R. et al. Molecular mechanism and function of CD40/CD40L engagement in the immune system. Immunol. Rev. 229, 152–172 (2009).
Jiang, H. et al. Demethylation of TNFSF7 contributes to CD70 overexpression in CD4+ T cells from patients with systemic sclerosis. Clin. Immunol. 143, 39–44 (2012).
Wang, Y. et al. Hypomethylation and overexpression of ITGAL (CD11a) in CD4+ T cells in systemic sclerosis. Clin. Epigenetics 6, 25 (2014).
Lu, T. et al. Whole-genome bisulfite sequencing in systemic sclerosis provides novel targets to understand disease pathogenesis. BMC Med. Genomics 12, 144 (2019).
Wei, J. et al. Wnt/β-catenin signaling is hyperactivated in systemic sclerosis and induces Smad-dependent fibrotic responses in mesenchymal cells. Arthritis Rheum. 64, 2734–2745 (2012).
Ding, W. et al. Genome-wide DNA methylation analysis in systemic sclerosis reveals hypomethylation of IFN-associated genes in CD4+ and CD8+ T cells. J. Invest. Dermatol. 138, 1069–1077 (2018).
Li, T. et al. Epigenomics and transcriptomics of systemic sclerosis CD4+ T cells reveal long-range dysregulation of key inflammatory pathways mediated by disease-associated susceptibility loci. Genome Med. 12, 81 (2020).
Qiu, Y. & Huang, S. CTCF-mediated genome organization and leukemogenesis. Leukemia 34, 2295–2304 (2020).
Fullard, N. & O’Reilly, S. Role of innate immune system in systemic sclerosis. Semin. Immunopathol. 37, 511–517 (2015).
van der Kroef, M. et al. Histone modifications underlie monocyte dysregulation in patients with systemic sclerosis, underlining the treatment potential of epigenetic targeting. Ann. Rheum. Dis. 78, 529–538 (2019).
Ciechomska, M. et al. Histone demethylation and Toll-like receptor 8-dependent cross-talk in monocytes promotes transdifferentiation of fibroblasts in systemic sclerosis via Fra-2. Arthritis Rheumatol. 68, 1493–1504 (2016).
Mariotti, B. et al. The long non-coding RNA NRIR drives IFN-response in monocytes: implication for systemic sclerosis. Front. Immunol. 10, 100 (2019).
Ciechomska, M. et al. Global miRNA and mRNA expression profiles identify miRNA-26a-2-3p-dependent repression of IFN signature in systemic sclerosis human monocytes. Eur. J. Immunol. 50, 1057–1066 (2020).
Rossato, M. et al. Association of microRNA-618 expression with altered frequency and activation of plasmacytoid dendritic cells in patients with systemic sclerosis. Arthritis Rheumatol. 69, 1891–1902 (2017).
Chouri, E. et al. Implication of miR-126 and miR-139-5p in plasmacytoid dendritic cell dysregulation in systemic sclerosis. J. Clin. Med. 10, 491 (2021).
Liu, Q. et al. Chromatin accessibility landscapes of skin cells in systemic sclerosis nominate dendritic cells in disease pathogenesis. Nat. Commun. 11, 5843 (2020).
Tsou, P. S., Palisoc, P. J., Flavahan, N. A. & Khanna, D. Dissecting the cellular mechanism of prostacyclin analogue iloprost in reversing vascular dysfunction in scleroderma. Arthritis Rheumatol. 73, 520–529 (2021).
Manetti, M. et al. Overexpression of VEGF165b, an inhibitory splice variant of vascular endothelial growth factor, leads to insufficient angiogenesis in patients with systemic sclerosis. Circ. Res. 109, e14–e26 (2011).
Wang, Y. & Kahaleh, B. Epigenetic repression of bone morphogenetic protein receptor II expression in scleroderma. J. Cell Mol. Med. 17, 1291–1299 (2013).
Tsou, P. S. et al. Histone deacetylase 5 is overexpressed in scleroderma endothelial cells and impairs angiogenesis via repression of proangiogenic factors. Arthritis Rheumatol. 68, 2975–2985 (2016).
Tsou, P. S. et al. Inhibition of EZH2 prevents fibrosis and restores normal angiogenesis in scleroderma. Proc. Natl Acad. Sci. USA 116, 3695–3702 (2019).
Tsou, P. S., Palisoc, P. J., Ali, M., Khanna, D. & Sawalha, A. H. Genome-wide reduction in chromatin accessibility and unique transcription factor footprints in endothelial cells and fibroblasts in scleroderma skin. Arthritis Rheumatol. 73, 1501–1513 (2021).
Hinz, B. & Lagares, D. Evasion of apoptosis by myofibroblasts: a hallmark of fibrotic diseases. Nat. Rev. Rheumatol. 16, 11–31 (2020).
Altorok, N., Tsou, P. S., Coit, P., Khanna, D. & Sawalha, A. H. Genome-wide DNA methylation analysis in dermal fibroblasts from patients with diffuse and limited systemic sclerosis reveals common and subset-specific DNA methylation aberrancies. Ann. Rheum. Dis. 74, 1612–1620 (2015).
Baker Frost, D. et al. Differential DNA methylation landscape in skin fibroblasts from African americans with systemic sclerosis. Genes (Basel) 12, 129 (2021).
Wang, X.-F., Zhang, B.-H., Lu, X.-Q. & Wang, R.-Q. DLX5 gene regulates the Notch signaling pathway to promote glomerulosclerosis and interstitial fibrosis in uremic rats. J. Cell. Physiol. 234, 21825–21837 (2019).
Henderson, J. et al. Methyl cap binding protein 2: a key epigenetic protein in systemic sclerosis. Rheumatology 58, 527–535 (2019).
He, Y., Tsou, P. S., Khanna, D. & Sawalha, A. H. Methyl-CpG-binding protein 2 mediates antifibrotic effects in scleroderma fibroblasts. Ann. Rheum. Dis. 77, 1208–1218 (2018).
Wang, Y. et al. MBD2 serves as a viable target against pulmonary fibrosis by inhibiting macrophage M2 program. Sci. Adv. 7, eabb6075 (2021).
O’Reilly, S., Ciechomska, M., Cant, R., Hügle, T. & van Laar, J. M. Interleukin-6, its role in fibrosing conditions. Cytokine Growth Factor. Rev. 23, 99–107 (2012).
O’Reilly, S., Ciechomska, M., Cant, R. & van Laar, J. M. Interleukin-6 (IL-6) trans signaling drives a STAT3-dependent pathway that leads to hyperactive transforming growth factor-β (TGF-β) signaling promoting SMAD3 activation and fibrosis via Gremlin protein. J. Biol. Chem. 289, 9952–9960 (2014).
Shin, J. Y. et al. Epigenetic activation and memory at a TGFB2 enhancer in systemic sclerosis. Sci. Transl. Med. 11, eaaw0790 (2019).
Vichaikul, S. et al. Inhibition of histone readers bromodomain and extraterminal domain proteins alleviates scleroderma fibrosis. Arthritis Rheumatol. https://acrabstracts.org/abstract/inhibition-of-histone-readers-bromodomain-and-extraterminal-domain-proteins-alleviates-scleroderma-fibrosis/ (2019).
Stock, C. J. W. et al. Bromodomain and extraterminal (BET) protein inhibition restores redox balance and inhibits myofibroblast activation. Biomed. Res. Int. 2019, 1484736 (2019).
Sanders, Y. Y. et al. Brd4-p300 inhibition downregulates Nox4 and accelerates lung fibrosis resolution in aged mice. JCI Insight 5, e137127 (2020).
Wasson, C. W. et al. Long non-coding RNA HOTAIR drives EZH2-dependent myofibroblast activation in systemic sclerosis through miRNA 34a-dependent activation of NOTCH. Ann. Rheum. Dis. 79, 507–517 (2020).
Wasson, C. W. et al. Long non-coding RNA HOTAIR induces GLI2 expression through Notch signalling in systemic sclerosis dermal fibroblasts. Arthritis Res. Ther. 22, 286 (2020).
Lin, X., Li, J. & Xing, Y. Q. Geniposide, a sonic hedgehog signaling inhibitor, inhibits the activation of hepatic stellate cell. Int. Immunopharmacol. 72, 330–338 (2019).
Kugler, M. C. et al. Sonic hedgehog signaling regulates myofibroblast function during alveolar septum formation in murine postnatal lung. Am. J. Respir. Cell Mol. Biol. 57, 280–293 (2017).
Pachera, E. et al. Long noncoding RNA H19X is a key mediator of TGF-β-driven fibrosis. J. Clin. Invest. 130, 4888–4905 (2020).
Forrester, H. B., Li, J., Leong, T., McKay, M. J. & Sprung, C. N. Identification of a radiation sensitivity gene expression profile in primary fibroblasts derived from patients who developed radiotherapy-induced fibrosis. Radiother. Oncol. 111, 186–193 (2014).
Henderson, J., Wilkinson, S., Przyborski, S., Stratton, R. & O’Reilly, S. microRNA27a-3p mediates reduction of the Wnt antagonist sFRP-1 in systemic sclerosis. Epigenetics 16, 808–817 (2020).
Yaseen, B. et al. Interleukin-31 promotes pathogenic mechanisms underlying skin and lung fibrosis in scleroderma. Rheumatology 59, 2625–2636 (2020).
Yao, Q. et al. MiR-16-5p suppresses myofibroblast activation in systemic sclerosis by inhibiting NOTCH signaling. Aging 13, 2640–2654 (2020).
Feng, S. & De Carvalho, D. D. Clinical advances in targeting epigenetics for cancer therapy. FEBS J. 29, 375–381 (2021).
Wang, Y., Fan, P. S. & Kahaleh, B. Association between enhanced type I collagen expression and epigenetic repression of the FLI1 gene in scleroderma fibroblasts. Arthritis Rheum. 54, 2271–2279 (2006).
Dees, C. et al. The Wnt antagonists DKK1 and SFRP1 are downregulated by promoter hypermethylation in systemic sclerosis. Ann. Rheum. Dis. 73, 1232–1239 (2014).
Wang, Y. Y. et al. DNA hypermethylation of the forkhead box protein 3 (FOXP3) promoter in CD4+ T cells of patients with systemic sclerosis. Br. J. Dermatol. 171, 39–47 (2014).
Noda, S. et al. Simultaneous downregulation of KLF5 and Fli1 is a key feature underlying systemic sclerosis. Nat. Commun. 5, 5797 (2014).
Zhang, Y. et al. Poly(ADP-ribose) polymerase-1 regulates fibroblast activation in systemic sclerosis. Ann. Rheum. Dis. 77, 744–751 (2018).
Daver, N. et al. Efficacy, safety, and biomarkers of response to azacitidine and nivolumab in relapsed/refractory acute myeloid leukemia: a nonrandomized, open-label, phase II study. Cancer Discov. 9, 370–383 (2019).
Xu, X. et al. A CRISPR-based approach for targeted DNA demethylation. Cell Discov. 2, 16009 (2016).
Huber, L. C. et al. Trichostatin A prevents the accumulation of extracellular matrix in a mouse model of bleomycin-induced skin fibrosis. Arthritis Rheum. 56, 2755–2764 (2007).
Svegliati, S. et al. Oxidative DNA damage induces the ATM-mediated transcriptional suppression of the Wnt inhibitor WIF-1 in systemic sclerosis and fibrosis. Sci. Signal. 7, ra84 (2014).
Palumbo-Zerr, K. et al. Orphan nuclear receptor NR4A1 regulates transforming growth factor-β signaling and fibrosis. Nat. Med. 21, 150–158 (2015).
Grabiec, A. M., Korchynskyi, O., Tak, P. P. & Reedquist, K. A. Histone deacetylase inhibitors suppress rheumatoid arthritis fibroblast-like synoviocyte and macrophage IL-6 production by accelerating mRNA decay. Ann. Rheum. Dis. 71, 424–431 (2012).
Wei, J. et al. The histone deacetylase sirtuin 1 is reduced in systemic sclerosis and abrogates fibrotic responses by targeting transforming growth factor beta signaling. Arthritis Rheumatol. 67, 1323–1334 (2015).
Zhu, X. et al. Sirt1 ameliorates systemic sclerosis by targeting the mTOR pathway. J. Dermatol. Sci. 87, 149–158 (2017).
Akamata, K. et al. SIRT3 is attenuated in systemic sclerosis skin and lungs, and its pharmacologic activation mitigates organ fibrosis. Oncotarget 7, 69321–69336 (2016).
Chu, H. et al. Sirtuin1 protects against systemic sclerosis-related pulmonary fibrosis by decreasing proinflammatory and profibrotic processes. Am. J. Respir. Cell Mol. Biol. 58, 28–39 (2018).
Wyman, A. E. et al. Sirtuin 7 is decreased in pulmonary fibrosis and regulates the fibrotic phenotype of lung fibroblasts. Am. J. Physiol. Lung Cell Mol. Physiol. 312, L945–L958 (2017).
Sosulski, M. L., Gongora, R., Feghali-Bostwick, C., Lasky, J. A. & Sanchez, C. G. Sirtuin 3 deregulation promotes pulmonary fibrosis. J. Gerontol. A Biol. Sci. Med. Sci. 72, 595–602 (2017).
Rehan, M. et al. Restoration of SIRT3 gene expression by airway delivery resolves age-associated persistent lung fibrosis in mice. Nat. Aging 1, 205–217 (2021).
Zhu, L., Mou, Q., Wang, Y., Zhu, Z. & Cheng, M. Resveratrol contributes to the inhibition of liver fibrosis by inducing autophagy via the microRNA‑20a‑mediated activation of the PTEN/PI3K/AKT signaling pathway. Int. J. Mol. Med. 46, 2035–2046 (2020).
Bergmann, C. et al. The histone demethylase Jumonji domain-containing protein 3 (JMJD3) regulates fibroblast activation in systemic sclerosis. Ann. Rheum. Dis. 77, 150–158 (2018).
Martin-Mateos, R. et al. Enhancer of Zeste Homologue 2 inhibition attenuates TGF-β dependent hepatic stellate cell activation and liver fibrosis. Cell Mol. Gastroenterol. Hepatol. 7, 197–209 (2019).
Ligresti, G. et al. CBX5/G9a/H3K9me-mediated gene repression is essential to fibroblast activation during lung fibrosis. JCI Insight 5, e127111 (2019).
Ghosh, A. K. et al. p300 is elevated in systemic sclerosis and its expression is positively regulated by TGF-β: epigenetic feed-forward amplification of fibrosis. J. Invest. Dermatol. 133, 1302–1310 (2013).
Welti, J. et al. Targeting the p300/CBP axis in lethal prostate cancer. Cancer Discov. 11, 1118–1137 (2021).
Yan, Q., Chen, J., Li, W., Bao, C. & Fu, Q. Targeting miR-155 to treat experimental scleroderma. Sci. Rep. 6, 20314 (2016).
Peng, W. J. et al. MicroRNA-29: a potential therapeutic target for systemic sclerosis. Expert Opin. Ther. Targets 16, 875–879 (2012).
Gallant-Behm, C. L. et al. A microRNA-29 mimic (remlarsen) represses extracellular matrix expression and fibroplasia in the skin. J. Invest. Dermatol. 139, 1073–1081 (2019).
Makino, K. et al. The downregulation of microRNA let-7a contributes to the excessive expression of type I collagen in systemic and localized scleroderma. J. Immunol. 190, 3905–3915 (2013).
Krützfeldt, J. et al. Silencing of microRNAs in vivo with ‘antagomirs’. Nature 438, 685–689 (2005).
Zerr, P. et al. Sirt1 regulates canonical TGF-β signalling to control fibroblast activation and tissue fibrosis. Ann. Rheum. Dis. 75, 226–233 (2016).
Kramer, M. et al. Inhibition of H3K27 histone trimethylation activates fibroblasts and induces fibrosis. Ann. Rheum. Dis. 72, 614–620 (2013).
Hardy, T. et al. Plasma DNA methylation: a potential biomarker for stratification of liver fibrosis in non-alcoholic fatty liver disease. Gut 66, 1321 (2017).
Wielscher, M. et al. Diagnostic performance of plasma DNA methylation profiles in lung cancer, pulmonary fibrosis and COPD. EBioMedicine 2, 929–936 (2015).
Author information
Authors and Affiliations
Contributions
S.O. researched data for the article. All authors made substantial contributions to discussions of the content and contributed to writing the article and reviewing/editing of the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
S.O. became a full-time employee of Stipe Therapeutics after submission of this manuscript. The other authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Rheumatology thanks Y. Asano, B. Kahaleh and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Methyl binding domain
-
A family of methyl-CpG-binding domain proteins that translate the DNA methylation signal and that work in concert with other proteins such as histone deacetyl transferases to facilitate gene repression.
- Histone tails
-
Flexible regions that flank both ends of the histone fold and that can be modified by a plethora of modifications that impact chromatin dynamics and gene expression.
- Lactylation
-
An epigenetic modification whereby the metabolite lactate is deposited on histone lysine residues.
- Histone acetyl transferases
-
(HATs). A group of enzymes that mediate the addition of an acetyl group onto lysine residues on histones to modulate gene expression.
- Histone deacetyl transferases
-
(HDACs). A group of enzymes that mediate the removal of acetyl groups from lysine residues on histones, positively regulating gene expression.
- Stress fibres
-
Contractile actin bundles found in non-muscle cells, composed of actin and non-muscle myosin II.
Rights and permissions
About this article
Cite this article
Tsou, PS., Varga, J. & O’Reilly, S. Advances in epigenetics in systemic sclerosis: molecular mechanisms and therapeutic potential. Nat Rev Rheumatol 17, 596–607 (2021). https://doi.org/10.1038/s41584-021-00683-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41584-021-00683-2
- Springer Nature Limited
This article is cited by
-
Epigenetics as a versatile regulator of fibrosis
Journal of Translational Medicine (2023)
-
Distinct genome-wide DNA methylation and gene expression signatures in classical monocytes from African American patients with systemic sclerosis
Clinical Epigenetics (2023)
-
Enrichr in silico analysis of MS-based extracted candidate proteomic biomarkers highlights pathogenic pathways in systemic sclerosis
Scientific Reports (2023)
-
Back to the future: targeting the extracellular matrix to treat systemic sclerosis
Nature Reviews Rheumatology (2023)
-
The critical importance of epigenetics in autoimmune-related skin diseases
Frontiers of Medicine (2023)