Abstract
Human parainfluenza virus type 3 (hPIV3) is a respiratory pathogen that can cause severe disease in older people and infants. Currently, vaccines against hPIV3 are in clinical trials but none have been approved yet. The haemagglutinin-neuraminidase (HN) and fusion (F) surface glycoproteins of hPIV3 are major antigenic determinants. Here we describe naturally occurring potently neutralizing human antibodies directed against both surface glycoproteins of hPIV3. We isolated seven neutralizing HN-reactive antibodies and a pre-fusion conformation F-reactive antibody from human memory B cells. One HN-binding monoclonal antibody (mAb), designated PIV3-23, exhibited functional attributes including haemagglutination and neuraminidase inhibition. We also delineated the structural basis of neutralization for two HN and one F mAbs. MAbs that neutralized hPIV3 in vitro protected against infection and disease in vivo in a cotton rat model of hPIV3 infection, suggesting correlates of protection for hPIV3 and the potential clinical utility of these mAbs.






Similar content being viewed by others
Data availability
All data needed to evaluate the conclusions in the paper are included in the paper or the Supplementary Information. Sequencing reads and consensus sequences have been deposited in NCBI under BioProject PRJNA992650. The atomic models used to determine and interpret the two protein structures detailed in this manuscript are available in the Protein Data Bank under the following accession codes: PDB 1V2I, PDB 4WEF, PDB 5I19, PDB 7BEL, PDB 3Q6G, PDB 5F9O, PBD 7DPM, PDB 6MJZ, PDB 7DPM. The cryo-EM structures and volume for the hPIV3 HN and hPIV3 F in complex with neutralizing Fabs have been deposited in the Protein Data Bank with the accession codes 8TQI (hPIV3 HN complex) and 8TQK (hPIV3 F complex). The electron density volumes are also available in the Electron Microscopy Data Bank under the codes EMD-41505 (hPIV3 HN complex) and EMD-41506 (hPIV3 F complex). Source data are provided with this paper.
Code availability
No new code was generated in this study. See the source data provided with this study and the deposited data.
References
Chanock, R. M. et al. Newly recognized myxoviruses from children with respiratory disease. N. Engl. J. Med. 258, 207–213 (1958).
Shah, D. P., Shah, P. K., Azzi, J. M. & Chemaly, R. F. Parainfluenza virus infections in hematopoietic cell transplant recipients and hematologic malignancy patients: a systematic review. Cancer Lett. 370, 358–364 (2016).
Fontana, L. & Strasfeld, L. Respiratory virus infections of the stem cell transplant recipient and the hematologic malignancy patient. Infect. Dis. Clin. North Am. 33, 523–544 (2019).
Moscona, A. & Peluso, R. W. Fusion properties of cells persistently infected with human parainfluenza virus type 3: participation of hemagglutinin-neuraminidase in membrane fusion. J. Virol. 65, 2773–2777 (1991).
Porotto, M. et al. Regulation of paramyxovirus fusion activation: the hemagglutinin-neuraminidase protein stabilizes the fusion protein in a pretriggered state. J. Virol. 86, 12838–12848 (2012).
Farzan, S. F. et al. Premature activation of the paramyxovirus fusion protein before target cell attachment with corruption of the viral fusion machinery. J. Biol. Chem. 286, 37945–37954 (2011).
Bose, S., Jardetzky, T. S. & Lamb, R. A. Timing is everything: fine-tuned molecular machines orchestrate paramyxovirus entry. Virology 479, 518–531 (2015).
Porotto, M., Murrell, M., Greengard, O. & Moscona, A. Triggering of human parainfluenza virus 3 fusion protein (F) by the hemagglutinin-neuraminidase (HN) protein: an HN mutation diminishes the rate of F activation and fusion. J. Virol. 77, 3647–3654 (2003).
Marcink, T. C. et al. Subnanometer structure of an enveloped virus fusion complex on viral surface reveals new entry mechanisms. Sci. Adv. 9, eade2727 (2023).
Porotto, M., Murrell, M., Greengard, O., Doctor, L. & Moscona, A. Influence of the human parainfluenza virus 3 attachment protein’s neuraminidase activity on its capacity to activate the fusion protein. J. Virol. 79, 2383–2392 (2005).
Huberman, K., Peluso, R. W. & Moscona, A. Hemagglutinin-neuraminidase of human parainfluenza-3: role of the neuraminidase in the viral life cycle. Virology 214, 294–300 (1995).
Porotto, M., Fornabaio, M., Kellogg, G. E. & Moscona, A. A second receptor binding site on human parainfluenza virus type 3 hemagglutinin-neuraminidase contributes to activation of the fusion mechanism. J. Virol. 81, 3216–3228 (2007).
Xu, R. et al. Interaction between the hemagglutinin-neuraminidase and fusion glycoproteins of human parainfluenza virus type III regulates viral growth. mBio 4, e00803-13 (2013).
Greninger, A. L. et al. Human parainfluenza virus evolution during lung infection of immunocompromised individuals promotes viral persistence. J. Clin. Invest. https://doi.org/10.1172/JCI150506 (2021).
Iketani, S. et al. Viral entry properties required for fitness in humans are lost through rapid genomic change during viral isolation. mBio 9, e00898-18 (2018).
Porotto, M., Palmer, S. G., Palermo, L. M. & Moscona, A. Mechanism of fusion triggering by human parainfluenza virus type III. J. Biol. Chem. 287, 778–793 (2012).
Palmer, S. G. et al. Adaptation of human parainfluenza virus to airway epithelium reveals fusion properties required for growth in host tissue. mBio 3, e00137-12 (2012).
Mishin, V. P. et al. N-linked glycan at residue 523 of human parainfluenza virus type 3 hemagglutinin-neuraminidase masks a second receptor-binding site. J. Virol. 84, 3094–3100 (2010).
Lawrence, M. C. et al. Structure of the haemagglutinin-neuraminidase from human parainfluenza virus type III. J. Mol. Biol. 335, 1343–1357 (2004).
Streltsov, V. A., Pilling, P., Barrett, S. & McKimm-Breschkin, J. L. Catalytic mechanism and novel receptor binding sites of human parainfluenza virus type 3 hemagglutinin-neuraminidase (hPIV3 HN). Antiviral Res. 123, 216–223 (2015).
Dirr, L., El-Deeb, I. M., Chavas, L. M. G., Guillon, P. & Itzstein, M. V. The impact of the butterfly effect on human parainfluenza virus haemagglutinin-neuraminidase inhibitor design. Sci. Rep. 7, 4507 (2017).
Pascolutti, M. et al. Structural insights into human parainfluenza virus 3 hemagglutinin-neuraminidase using unsaturated 3-substituted sialic acids as probes. ACS Chem. Biol. 13, 1544–1550 (2018).
Dirr, L. et al. The catalytic mechanism of human parainfluenza virus type 3 haemagglutinin-neuraminidase revealed. Angew. Chem. Int. Ed. 54, 2936–2940 (2015).
Yin, H. S., Paterson, R. G., Wen, X., Lamb, R. A. & Jardetzky, T. S. Structure of the uncleaved ectodomain of the paramyxovirus (hPIV3) fusion protein. Proc. Natl Acad. Sci. USA 102, 9288–9293 (2005).
Stewart-Jones, G. B. E. et al. Structure-based design of a quadrivalent fusion glycoprotein vaccine for human parainfluenza virus types 1-4. Proc. Natl Acad. Sci. USA 115, 12265–12270 (2018).
Habibi, M. S. et al. Impaired antibody-mediated protection and defective IgA B-cell memory in experimental infection of adults with respiratory syncytial virus. Am. J. Respir. Crit. Care Med. 191, 1040–1049 (2015).
Ngwuta, J. O. et al. Prefusion F-specific antibodies determine the magnitude of RSV neutralizing activity in human sera. Sci. Transl. Med. 7, 309ra162 (2015).
Gilman, M. S. et al. Rapid profiling of RSV antibody repertoires from the memory B cells of naturally infected adult donors. Sci. Immunol. 1, eaaj1879 (2016).
van Wyke Coelingh, K. L. & Tierney, E. L. Identification of amino acids recognized by syncytium-inhibiting and neutralizing monoclonal antibodies to the human parainfluenza type 3 virus fusion protein. J. Virol. 63, 3755–3760 (1989).
van Wyke Coelingh, K. L., Winter, C. & Murphy, B. R. Antigenic variation in the hemagglutinin-neuraminidase protein of human parainfluenza type 3 virus. Virology 143, 569–582 (1985).
van Wyke Coelingh, K. L., Winter, C. C., Jorgensen, E. D. & Murphy, B. R. Antigenic and structural properties of the hemagglutinin-neuraminidase glycoprotein of human parainfluenza virus type-3: sequence analysis of variants selected with monoclonal antibodies which inhibit infectivity, hemagglutination, and neuraminidase activities. J. Virol. 61, 1473–1477 (1987).
Henrickson, K. J. & Portner, A. Antibody response in children to antigen sites on human PIV-3 HN: correlation with known epitopes mapped by monoclonal antibodies. Vaccine 8, 75–80 (1990).
van Wyke Coelingh, K. L. & Tierney, E. L. Antigenic and functional organization of human parainfluenza virus type 3 fusion glycoprotein. J. Virol. 63, 375–382 (1989).
van Wyke Coelingh, K. L. & Winter, C. C. Naturally occurring human parainfluenza type 3 viruses exhibit divergence in amino acid sequence of their fusion protein neutralization epitopes and cleavage sites. J. Virol. 64, 1329–1334 (1990).
Boonyaratanakornkit, J. et al. Protective antibodies against human parainfluenza virus type 3 infection. MAbs 13, 1912884 (2021).
Caban, M. et al. Cross-protective antibodies against common endemic respiratory viruses. Nat. Commun. 14, 798 (2023).
Madhi, S. A. et al. Transmissibility, infectivity and immunogenicity of a live human parainfluenza type 3 virus vaccine (HP1V345) among susceptible infants and toddlers. Vaccine 24, 2432–2439 (2006).
Bernstein, D. I., Falloon, J. & Yi, T. T. A randomized, double-blind, placebo-controlled, phase 1/2a study of the safety and immunogenicity of a live, attenuated human parainfluenza virus type 3 vaccine in healthy infants. Vaccine 29, 7042–7048 (2011).
Karron, R. A. et al. Evaluation of a live attenuated bovine parainfluenza type 3 vaccine in two- to six-month-old infants. Pediatr. Infect. Dis. J. 15, 650–654 (1996).
Gomez, M. et al. Phase-I study MEDI-534, of a live, attenuated intranasal vaccine against respiratory syncytial virus and parainfluenza-3 virus in seropositive children. Pediatr. Infect. Dis. J. 28, 655–658 (2009).
Greninger, A. L. A decade of RNA virus metagenomics is (not) enough. Virus Res. 244, 218–229 (2018).
Palermo, L. M. et al. Features of circulating parainfluenza virus required for growth in human airway. mBio 7, e00235 (2016).
Krammer, F. et al. NAction! How can neuraminidase-based immunity contribute to better influenza virus vaccines? mBio 9, e02332-17 (2018).
Monto, A. S. et al. Antibody to influenza virus neuraminidase: an independent correlate of protection. J. Infect. Dis. 212, 1191–1199 (2015).
Yin, H. S., Wen, X., Paterson, R. G., Lamb, R. A. & Jardetzky, T. S. Structure of the parainfluenza virus 5 F protein in its metastable, prefusion conformation. Nature 439, 38–44 (2006).
White, J. M., Delos, S. E., Brecher, M. & Schornberg, K. Structures and mechanisms of viral membrane fusion proteins: multiple variations on a common theme. Crit. Rev. Biochem. Mol. Biol. 43, 189–219 (2008).
Harrison, S. C. Viral membrane fusion. Nat. Struct. Mol. Biol. 15, 690–698 (2008).
Wen, X. et al. Structure of the human metapneumovirus fusion protein with neutralizing antibody identifies a pneumovirus antigenic site. Nat. Struct. Mol. Biol. 19, 461–463 (2012).
Jiang, J. J. et al. Functional analysis of amino acids at stalk/head interface of human parainfluenza virus type 3 hemagglutinin-neuraminidase protein in the membrane fusion process. Virus Genes 54, 333–342 (2018).
Li, L. et al. AbRSA: a robust tool for antibody numbering. Protein Sci. 28, 1524–1531 (2019).
Pascolutti, M. et al. Structural insights into human parainfluenza virus 3 hemagglutinin-neuraminidase using unsaturated 3- N-substituted sialic acids as probes. ACS Chem. Biol. 13, 1544–1550 (2018).
Taylor, G. Sialidases: structures, biological significance and therapeutic potential. Curr. Opin. Struct. Biol. 6, 830–837 (1996).
Winger, M. & von Itzstein, M. Exposing the flexibility of human parainfluenza virus hemagglutinin-neuraminidase. J. Am. Chem. Soc. 134, 18447–18452 (2012).
Bailly, B. et al. A dual drug regimen synergistically blocks human parainfluenza virus infection. Sci. Rep. 6, 24138 (2016).
Yuan, P., Paterson, R. G., Leser, G. P., Lamb, R. A. & Jardetzky, T. S. Structure of the Ulster strain Newcastle disease virus hemagglutinin-neuraminidase reveals auto-inhibitory interactions associated with low virulence. PLoS Pathog. 8, e1002855 (2012).
Zaitsev, V. et al. Second sialic acid binding site in Newcastle disease virus hemagglutinin-neuraminidase: implications for fusion. J. Virol. 78, 3733–3741 (2004).
Welch, S. R. et al. Inhibition of Nipah virus by defective interfering particles. J. Infect. Dis. 221, S460–S470 (2020).
Borisevich, V. et al. Escape from monoclonal antibody neutralization affects henipavirus fitness in vitro and in vivo. J. Infect. Dis. 213, 448–455 (2016).
Chou, T. C. Drug combination studies and their synergy quantification using the Chou–Talalay method. Cancer Res. 70, 440–446 (2010).
Ashton, J. C. Drug combination studies and their synergy quantification using the Chou–Talalay method—letter. Cancer Res. 75, 2400 (2015).
Gilchuk, P. et al. Analysis of a therapeutic antibody cocktail reveals determinants for cooperative and broad ebolavirus neutralization. Immunity 52, 388–403.e12 (2020).
Bernstein, D. I. et al. Phase 1 study of the safety and immunogenicity of a live, attenuated respiratory syncytial virus and parainfluenza virus type 3 vaccine in seronegative children. Pediatr. Infect. Dis. J. 31, 109–114 (2012).
Madhi, S. A. et al. Transmissibility, infectivity and immunogenicity of a live human parainfluenza type 3 virus vaccine (HP1V3cp45) among susceptible infants and toddlers. Vaccine 24, 2432–2439 (2006).
Skiadopoulos, M. H. et al. Evaluation of the replication and immunogenicity of recombinant human parainfluenza virus type 3 vectors expressing up to three foreign glycoproteins. Virology 297, 136–152 (2002).
Karron, R. A. et al. A live human parainfluenza type 3 virus vaccine is attenuated and immunogenic in young infants. Pediatr. Infect. Dis. J. 22, 394–405 (2003).
Domachowske, J. et al. Safety of nirsevimab for RSV in infants with heart or lung disease or prematurity. N. Engl. J. Med. 386, 892–894 (2022).
Ginsburg, A. S. & Srikantiah, P. Respiratory syncytial virus: promising progress against a leading cause of pneumonia. Lancet Glob. Health 9, e1644–e1645 (2021).
Fernandez, P. et al. A phase 2, randomized, double-blind safety and pharmacokinetic assessment of respiratory syncytial virus (RSV) prophylaxis with motavizumab and palivizumab administered in the same season. BMC Pediatr. 10, 38 (2010).
Edupuganti, S. et al. Feasibility and successful enrollment in a proof-of-concept HIV prevention trial of VRC01, a broadly neutralizing HIV-1 monoclonal antibody. J. Acquir. Immune Defic. Syndr. 87, 671–679 (2021).
Mgodi, N. M. et al. A phase 2b study to evaluate the safety and efficacy of VRC01 broadly neutralizing monoclonal antibody in reducing acquisition of HIV-1 infection in women in sub-saharan Africa: baseline findings. J. Acquir. Immune Defic. Syndr. 87, 680–687 (2021).
Corey, L. et al. Two randomized trials of neutralizing antibodies to prevent HIV-1 acquisition. N. Engl. J. Med. 384, 1003–1014 (2021).
Han, A. et al. Safety and efficacy of CR6261 in an influenza A H1N1 healthy human challenge model. Clin. Infect. Dis. 73, e4260–e4268 (2021).
Ali, S. O. et al. Evaluation of MEDI8852, an anti-influenza A monoclonal antibody, in treating acute uncomplicated influenza. Antimicrob. Agents Chemother. https://doi.org/10.1128/AAC.00694-18 (2018).
Johnson, S. et al. Development of a humanized monoclonal antibody (MEDI-493) with potent in vitro and in vivo activity against respiratory syncytial virus. J. Infect. Dis. 176, 1215–1224 (1997).
Guillon, P. et al. Structure-guided discovery of potent and dual-acting human parainfluenza virus haemagglutinin-neuraminidase inhibitors. Nat. Commun. 5, 5268 (2014).
Xu, R. et al. A recurring motif for antibody recognition of the receptor-binding site of influenza hemagglutinin. Nat. Struct. Mol. Biol. 20, 363–370 (2013).
Hong, M. et al. Antibody recognition of the pandemic H1N1 Influenza virus hemagglutinin receptor binding site. J. Virol. 87, 12471–12480 (2013).
Palermo, L. M., Porotto, M., Greengard, O. & Moscona, A. Fusion promotion by a paramyxovirus hemagglutinin-neuraminidase protein: pH modulation of receptor avidity of binding sites I and II. J. Virol. 81, 9152–9161 (2007).
Chng, J. et al. Cleavage efficient 2A peptides for high level monoclonal antibody expression in CHO cells. MAbs 7, 403–412 (2015).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Sanchez-Garcia, R. et al. DeepEMhancer: a deep learning solution for cryo-EM volume post-processing. Commun. Biol. 4, 874 (2021).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Teplyakov, A. et al. Structural diversity in a human antibody germline library. MAbs 8, 1045–1063 (2016).
Dejnirattisai, W. et al. The antigenic anatomy of SARS-CoV-2 receptor binding domain. Cell 184, 2183–2200.e22 (2021).
Spurrier, B. et al. Structural analysis of human and macaque mAbs 2909 and 2.5B: implications for the configuration of the quaternary neutralizing epitope of HIV-1 gp120. Structure 19, 691–699 (2011).
Bonsignori, M. et al. Maturation pathway from germline to broad HIV-1 neutralizer of a CD4-mimic antibody. Cell 165, 449–463 (2016).
Jiang, W. et al. Characterization of MW06, a human monoclonal antibody with cross-neutralization activity against both SARS-CoV-2 and SARS-CoV. MAbs 13, 1953683 (2021).
Pettersen, E. F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D 75, 861–877 (2019).
Schiffrin, B., Radford, S. E., Brockwell, D. J. & Calabrese, A. N. PyXlinkViewer: a flexible tool for visualization of protein chemical crosslinking data within the PyMOL molecular graphics system. Protein Sci. 29, 1851–1857 (2020).
Suryadevara, N. et al. Real-time cell analysis: a high-throughput approach for testing SARS-CoV-2 antibody neutralization and escape. STAR Protoc. 3, 101387 (2022).
Greninger, A. L. et al. Rapid metagenomic next-generation sequencing during an investigation of hospital-acquired human parainfluenza virus 3 infections. J. Clin. Microbiol. 55, 177–182 (2017).
Saitou, N. & Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987).
Tamura, K., Stecher, G. & Kumar, S. MEGA11: molecular evolutionary genetics analysis version 11. Mol. Biol. Evol. 38, 3022–3027 (2021).
Acknowledgements
We thank members of the Jardetzky, Moscona and Crowe laboratories. The work was supported by NIAID/NIH grants R01 AI13752 and R01 AI114736. Some of this work was performed at the Stanford-SLAC Cryo-EM Center (S2C2), which is supported by the National Institutes of Health Common Fund Transformative High-Resolution Cryo-Electron Microscopy programme (U24 GM129541). The authors thank H. A. Khant of the S2C2 for invaluable support and assistance. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. Special acknowledgements to R. Irving for helping to procure cotton rats and helping with animal studies. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
T.S.J. and J.E.C. conceived the project. N.S., A.R.O.-C., N.K., Y.-X.H., E.B., R.M.W., A.L.G. and L.S.H. performed laboratory experiments. A.L.G. and A.M. provided reagents. T.S.J., A.M. and J.E.C. obtained funding. R.H.C., T.S.J., A.M. and J.E.C. supervised the research. N.S., A.R.O., J.E.C. and T.S.J. wrote the first drafts of the paper. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
J.E.C. has served as a consultant for Luna Labs USA, Merck Sharp & Dohme Corporation, Emergent Biosolutions, and BTG International Inc., is a member of the Scientific Advisory Board of Meissa Vaccines, a former member of the Scientific Advisory Board of Gigagen (Grifols) and is founder of IDBiologics. The laboratory of J.E.C. received unrelated sponsored research agreements from AstraZeneca, Takeda Vaccines, and IDBiologics during the conduct of the study. T.S.J. has served as a consultant for Pfizer. A.L.G reports contract testing from Abbott, Cepheid, Novavax, Pfizer, Janssen and Hologic, and research support from Gilead, outside of the described work. A.M. reports potential future financial interests in Thylacine Biotherapeutics Inc. unrelated to the present study. All other authors declare no competing interests. Vanderbilt University has applied for a patent for some of the antibodies in this paper (2024 International Patent Application No. PCT/US2024/028239 based on US Serial No. 63/504,549).
Peer review
Peer review information
Nature Microbiology thanks Michael Hoffmann and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
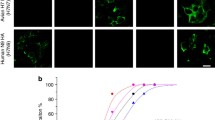
Extended Data Fig. 1 Haemagglutination Inhibition assay and Fab neutralization.
1a. Haemagglutination Inhibition assay (Highlighted in Red- IgG, Blue -Fab, Green- RBCs only, Yellow - RBCs + PIV3 virus) 1b. Neutralization assay of PIV-3 virus using equimolar ratio IgG (30 µg/mL starting concentration) and Fabs (10 µg/mL starting concentration) of the PIV3 mAbs. Error bars indicate mean ± S.D.; data are representative of n = 2 independent experiments performed in technical triplicates.
Extended Data Fig. 2 Overview of the HPIV3 F and rPIV3-18 structure determination.
A. Comparison of the gel filtration profile for the HPIV3 F and the HPIV3 F: r-PIV3-18 complex. The complex was purified at least three times independently with consistent results. B. Motion corrected micrograph for HPIV3 F:rPIV3-18 complex (n = 11194). Two independent datasets were collected with similar results. C. Representative 2D class averages for HPIV3 F:rPIV3-18. D. FSC curves. E. Cryo-EM image processing flowchart. F. Local resolution map.
Extended Data Fig. 3 Overview of the HPIV3 HN and rPIV3-23:rPIV3-28 structure determination.
A. Comparison of the gel filtration profile for the HPIV3 HN and the HPIV3 HN: r-PIV3-23:rPIV3-28 complex. The complex was purified at least three times independently with consistent results. B. Motion-corrected micrograph for HPIV3 HN:rPIV3-23:rPIV3-28 complex (n = 7242). C. Representative 2D class averages for HPIV3 HN:rPIV3-23:rPIV3-28. D. FSC curves. E. Cryo-EM image processing flowchart. F. Local resolution map.
Extended Data Fig. 4 Overview of the HPIV3 HN and rPIV3-23 structure determination.
A. Gel filtration profile for the HPIV3 HN: r-PIV3-23 complex. The complex was purified at least two times independently with consistent results. B. Motion-corrected micrograph for HPIV3 HN:rPIV3-23 complex (n = 7524). C. Representative 2D class averages for HPIV3 HN:rPIV3-23. D. FSC curves. E. Comparison between the HPIV3 HN:rPIV3-23 density map (blue), the HPIV3 HN:rPIV3-23:rPIV3-28 atomic model (beige) and the HPIV3 HN:rPIV3-23:rPIV3-28 density map (gray).
Extended Data Fig. 5 Superimposition with HPIV3 HN dimer and tetramer structures.
A. Stereoview with superimposition of the HN complex with other paramyxovirus HN dimer structures (PDB IDs: 4WEF, 4MZA, 4MZE, 1V2I, 1V3B, 1V3D, 1V3C, 1V3E, 5KV9, 4XJR, 6C0M, 5KV8, 4XJQ and 1USR). B. Superimposition of the HPIV3 HN (light pink) +rPIV3-23 (purple) +rPIV3-28 (yellow) complex with the ‘4 heads down’ NDV HN tetramer (green) (PDB ID: 3T1E). C. Superimposition of the HPIV3 HN (light pink) +rPIV3-23 (purple) +rPIV3-28 (yellow) complex with the ‘2 heads up 2 heads down’ PIV5 HN tetramer (green) (PDB ID: 4JF7).
Extended Data Fig. 6 Comparison between the HPIV3 HN binding site with rPIV3-23 HCDR3 and the Neu5Ac.
A. Comparison between the HPIV3 HN binding site with rPIV3-23 (purple) HCDR3 and the Neu5Ac (PDB model ID 1V3C) bound. B. Comparison of Ulster strain NDV HN (old rose) with HPIV3 HN (light rose):rPIV3-23 (purple). Relevant residues are displayed as sticks. C. Superimposition of NDV HN dimer with Neu5Acen (yellow) and Neu5Ac (green) displayed as spheres (PDB ID 1USR). Glycan represented as sticks. A single rPIV3-28 Fab (yellow) is shown binding to the HN dimer for clarity.
Supplementary information
Supplementary Tables 1–3
Antibody variable gene usage and HCDR3 amino acid sequences. Inhibitory concentration50 (IC50, in ng ml−1) of PIV3 mAbs against clinical isolates. DRI value was estimated by CompuSyn.
Supplementary Table 4
Cryo-EM data collection and model refinement.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Suryadevara, N., Otrelo-Cardoso, A.R., Kose, N. et al. Functional and structural basis of human parainfluenza virus type 3 neutralization with human monoclonal antibodies. Nat Microbiol 9, 2128–2143 (2024). https://doi.org/10.1038/s41564-024-01722-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-024-01722-w
- Springer Nature Limited