Abstract
Heterosis refers to the superior performance of F1 hybrids over their respective parental inbred lines. Although the genetic and expression basis of heterosis have been previously investigated, the metabolic basis for this phenomenon is poorly understood. In a preliminary morphological study in Brassica juncea, we observed significant heterosis at the 50% flowering stage, wherein both the growth and reproduction of F1 reciprocal hybrids were greater than that of their parents. To identify the possible metabolic causes or consequences of this heterosis, we carried out targeted LC-MS analysis of 48 primary (amino acids and sugars) and secondary metabolites (phytohormones, glucosinolates, flavonoids, and phenolic esters) in five developmental tissues at 50% flowering in hybrids and inbred parents. Principal component analysis (PCA) of metabolites clearly separated inbred lines from their hybrids, particularly in the bud tissues. In general, secondary metabolites displayed more negative heterosis values in comparison to primary metabolites. The tested primary and secondary metabolites displayed both additive and non-additive modes of inheritance in F1 hybrids, wherein the number of metabolites showing an additive mode of inheritance were higher in buds and siliques (52.77–97.14%) compared to leaf tissues (47.37–80%). Partial least regression (PLS) analysis further showed that primary metabolites, in general, displayed higher association with morphological parameters in F1 hybrids. Overall, our results are consistent with a resource-cost model for heterosis in B. juncea, where metabolite allocation in hybrids appears to favor growth, at the expense of secondary metabolism.
Similar content being viewed by others
Introduction
Heterosis (hybrid vigor) refers to the superior performance of heterozygous F1 hybrid plants compared to the average of their homozygous parental inbred lines in terms of increased biomass, size, yield, speed of development, fertility or resistance to disease, insect pests and environmental stresses (Hochholdinger and Hoecker 2007). Although the phenomenon has been exploited for centuries by various civilizations for increasing crop production, the rediscovery of Mendel’s laws in the twentieth century and accumulation of experimental data on the positive effects of cross-pollination led to the first description of the concept of heterosis (Crow 2001). Shull (1952) first recognized heterosis while working on hybrid corn, and his concept stated that different alleles exert a complementary physiological action when combined in heterozygous form, resulting in increased size, yield and vigor in hybrids.
Despite the agronomic importance of heterosis, its underlying genetic and molecular basis remains obscure. Quantitative genetic theories (dominance, overdominance and epistasis) have been used to explain the phenomenon for more than a century (reviewed by Birchler et al. 2003). Based on these theories, gene expression or protein abundance in hybrids should show a non-additive mode of inheritance in case of heterosis being equal to or greater than that of the parent with a greater value. On the other hand, if both parental alleles contribute equally to gene expression in a hybrid, they are said to exhibit additive behavior and the expression level of genes or proteins is equal to the average of both the parents referred to as the mid-parent value (MPV). High throughput methods have enabled the re-examination of the proportion of additive and non-additive expressions in heterosis, including analysis of quantitative trait loci (QTL; Li et al. 2008; Meyer et al. 2010; Shi et al. 2011; Andorf et al. 2012; Giraud et al. 2014; Liu et al. 2014a), gene expression (Song et al. 2007; Stupar et al. 2008; Ma et al. 2011; Shen et al. 2012; Qin et al. 2013), genome-wide transcriptomics (Paschold et al. 2012; Zhai et al. 2013; Ding et al. 2014; Paschold et al. 2014; Zhiguo et al. 2014; Groszmann et al. 2015, Wang et al. 2015), DNA methylation pattern analysis (Shen et al. 2012; Liu et al. 2014b), comparative proteomics (Dahal et al. 2012; Wang et al. 2014) and small and micro RNA transcriptomics (Ding et al. 2012; Shen et al. 2012; Li et al. 2014; Zhang et al. 2014; Xin et al. 2015). However, none of the current genetic models sufficiently explain heterosis in polyploid plants, which is compounded by genomic dosage, complex allelic and genic interactions, and epigenetic regulation.
With more emphasis on genetic and gene expression analysis, studies related to inheritance of metabolites have been largely under-represented in heterosis research. Metabolites are key components and regulators of biological processes, and high-throughput methods coupling gas or liquid chromatography with mass spectrometry are now able to detect a large number of plant metabolites with great accuracy (Patti et al. 2012). Metabolite analysis has provided some initial insights into the mode of inheritance in hybrids of maize (Römisch-Margl et al. 2010; Lisec et al. 2011; Meyer et al. 2012; Wang et al. 2014), rice (Ma et al. 2011) and Arabidopsis (Korn et al. 2010). However, all these studies have analyzed the metabolites of a single organ at a single developmental stage, which may not reflect the overall mode of metabolite inheritance in hybrids.
Here, we investigate the metabolic basis of heterosis in B. juncea, an economically important oilseed crop cultivated globally not only for its edible oil, but also for use as a vegetable, animal food and condiment. The crop has been frequently studied for various seed quality related traits such as oil and protein contents, fatty acid profiles and levels of secondary metabolites including glucosinolates (GSLs) (Augustine et al. 2014). In the current study, we selected a well-adapted Indian line (Varuna, V) and an early flowering east European line (Early Heera-2, EH2) and their F1 reciprocal crosses (VxEH2 and EH2xV). We conducted LC-MS based targeted analysis of key primary metabolites (amino acids and sugars) as well as secondary metabolites (phytohormones, glucosinolates, flavonoids and phenolic esters) in five different tissue types (flower buds, siliques, flag leaf, young leaf, and mature leaf), at the 50% flowering stage. Our study demonstrates that (i) the metabolic profile of B. juncea F1 hybrids are distinct from their inbred parents; (ii) some secondary metabolites (indolic glucosinolate, flavonoids and phenolic esters) display more negative heterosis (deviation from the mid-parent value) in comparison to primary metabolites (amino acids and sugars), whereas phytohormones show mixed trends of heterosis; and (iii) F1 hybrids predominantly exhibit an additive mode of metabolite inheritance, particularly in reproductive tissues.
Materials and methods
Plant material and growth conditions
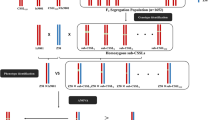
The B. juncea inbred lines Varuna (V, an Indian cultivar) and early flowering Heera-2 (EH2, an Eastern European cultivar) and their two F1 reciprocal crosses (EH2xV and VxEH2) were used in present study. The plants were grown in a greenhouse having controlled growth condition of 14 h light (24 °C), 10 h dark (20 °C) and 60% RH.
Quantitative measurement of morphological characters
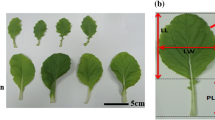
Morphological measurements were performed at two developmental stages viz., (i) 30 days after germination, and (ii) at 50% flowering stage (samples were collected for metabolite profiling at this stage). We selected morphological characters based on the recommendations of International Board for Plant Genetic Resources (IBPGR, Rome) for Brassica descriptors, which include (1) plant height, (2) leaf length, (3) petiole length, (4) leaf width, (5) number of nodes, (6) lamina length, (7) primary branches, (8) secondary branches, (9) main shoot length, (10) number of siliques on main shoot, (11) silique length on main shoot, and (12) silique density/unit length of main shoot.
Targeted metabolite analysis
A targeted metabolite analysis to quantify the levels of amino acids, sugars, phytohormones, glucosinolates, flavonoids and phenolic esters was conducted for each genotype. Five tissues namely unopened flower buds (hereafter buds), siliques, flag leaf, young leaf and mature leaf were harvested in 50% flowering stage (around 60 days post germination), from five independent biological replicates in the morning hours, snap cooled in liquid nitrogen and stored at –80 °C until further analysis was carried out.
Extraction of samples
B. juncea samples were freeze-dried and ground to fine powder. About 10 mg of the powder was extracted with 1 ml of 80% methanol solution containing 50 µM of intact 4-hydroxybenzyl-GSL and 40 ng of D6-Jasmonic acid (HPC Standards GmbH, Cunnersdorf, Germany), 40 ng D4-salicylic acid (Sigma-Aldrich, USA), 40 ng D6-abscisic acid (Santa Cruz Biotechnology, Santa Cruz, USA) and 8 ng of Jasmonic acid 13C6-isoleucine conjugate as internal standards. Jasmonic acid 13C6-isoleucine conjugate was synthesized as described by Kramell et al. (1988) using 13C6-Ile (Sigma-Aldrich, USA). Samples were centrifuged, and the supernatant was used for chemical analyses as detailed below.
Quantification of free amino acids
A 50 µl aliquot of the extract was diluted to a ratio of 1:10 (v:v) in water containing the 13C, 15N labeled algal amino acid mix (Isotec, Miamisburg, OH, USA). Amino acids in the diluted extracts were directly analysed by LC-MS/MS according to the protocol of Madsen et al. (2015) (Supplementary Table S1). Chromatography was performed on an Agilent 1200 HPLC system (Agilent Technologies, Boeblingen, Germany) using Zorbax Eclipse XDB-C18 column (50 × 4.6 mm, 1.8 µm, Agilent Technologies). The liquid chromatography was coupled to an API 5000 tandem mass spectrometer (Applied Biosystems, Darmstadt, Germany) equipped with a Turbospray ion source operated in the positive ionization mode. The instrument parameters were optimized by infusion experiments with pure standards (amino acid standard mix, Fluka, St. Louis, USA). Analyst 1.5 software (Applied Biosystems, Darmstadt, Germany) was used for data acquisition and processing.
Quantification of phytohormones
A 2 µl aliquot of the extract was subjected to LC-MS/MS analysis for the quantification of phytohormones. Chromatography was performed on an Agilent 1200 HPLC system according to the protocol described by Vahabi et al. (2015). Separation was achieved on a Zorbax Eclipse XDB-C18 column (50 × 4.6 mm, 1.8 µm, Agilent). An API 5000 tandem mass spectrometer equipped with a Turbospray ion source was operated in negative ionization mode. The instrument parameters were optimized by infusion experiments with pure standards. Multiple reaction monitoring (MRM) was used to monitor analyte parent ion → product ion, according to supplementary Table S2. For the quantification of 12-oxophytodienoic acid (cis-OPDA), D6-jasmonic acid was used as the internal standard applying an experimentally determined response factor (RF = 0.5).
Quantification of free sugars
A 50 µl aliquot of the extract was diluted in a ratio of 1:10 (v:v) in water. Sugars in the diluted extracts were directly analyzed by LC-MS/MS. Chromatography was performed on an Agilent 1200 HPLC system according to the protocol described by Madsen et al. (2015). Separation was achieved on a HILIC-HPLC-column (apHera NH2 Polymer; 15 × 4.6 mm, 5 µm, Supelco). The liquid chromatography was coupled to an API 3200 tandem mass spectrometer (Applied Biosystems, Darmstadt, Germany) equipped with a Turbospray ion source operated in negative ionization mode. The instrument parameters were optimized by infusion experiments with pure standards viz., d-(+)-glucose, d-(-)-fructose and sucrose (Sigma-Aldrich, USA). d-(+)-glucose, d-(-)-fructose and sucrose were quantified by external standard curves.
Quantification of glucosinolates
An 800 µl portion of the extract was loaded onto DEAE Sephadex A 25 columns (flow through was collected for further analysis of flavonoids and phenolic ester) and treated with aryl sulfatase for desulfation (Sigma-Aldrich) as described by Burow et al. (2006). The eluted desulfo GSL were separated using Agilent 1100 HPLC system on a reversed phase C-18 column (Nucleodur Sphinx RP, 250 × 4.6 mm, 5 µm, Machrey-Nagel, Düren, Germany) using water (solvent A)-acetonitrile (solvent B) gradient (0–1 min, 1.5% B; 1–6 min, 1.5–5% B; 6–8 min, 5–7% B; 8–18 min, 7–21% B; 18–23 min, 21–29% B; 23–23.1 min, 29–100% B; 23.1–24 min 100% B and 24.1–28 min 1.5% B; flow rate of 1.0 ml min−1). Detection was performed with a photodiode array detector and peaks were integrated at 229 nm. For the quantification of individual GSL, following response factors were used: aliphatic GSL, RF = 2; aromatic GSL, RF = 2; and indolic GSL, RF = 0.5 (Burow et al. 2006).
Quantification of flavonoids and phenolic esters
The flow-through fraction (800 µl) from DEAE Sephadex column used for GSL extraction and the fraction from washing the column with 800 µl 80% methanol (v:v) were collected and combined for flavonoid and phenolic ester analysis. Samples were separated on Agilent 1100 HPLC system equipped with a C-18 reversed phase column (Nucleodur Sphinx RP, 250 × 4.6 mm, 5 μm; Macherey-Nagel, Düren, Germany). The mobile phase consisted of 0.2% formic acid in water (v:v) (solvent A) and acetonitrile (solvent B) used in gradient mode at a flow rate of 1 ml min−1 at 25 °C. The gradient was as follows: 100% A (5 min), 0–45% B (15 min), 45–100% (0.1 min), 100% B (1.9 min), and 100% A (3.9 min). The eluent was monitored by a photodiode array detector at 330 nm. The compounds were identified based on UV visible absorption and mass spectra from LC-MS analysis on a Bruker Esquire 6000 Ion Trap mass spectrometer (LC conditions were the same as for HPLC-UV analysis) by comparing them to the identified metabolites reported in the literature (Lin and Harnly 2010; Lin et al. 2011; Table S3). Relative quantification of flavonoids and phenolic esters were performed by calculating peak area per mg of dry weight in the UV chromatograms at 330 nm. For a number of flavonoids, conversion factors for absolute quantification were determined based on an external standard curve of quercitin–3-glucoside (Sigma-Aldrich, Taufkirchen, Germany) applying a relative molar response factor (RF = 1.0); and for some phenolic esters based on an external standard curve of sinapic acid (Fluka, Buchs, Switzerland) applying a relative molar response factor (RF = 1.0, Table S3).
Statistical analysis
All the metabolites were analyzed in five biological replicates and experimental results were expressed as mean ± standard deviation (SD). Metabolic data were log transformed (log10) to better resemble a normal distribution. Principal component analysis (PCA) was performed in order to check whether data reduction obtained through a new set of variables (PCs) revealed any mode of inheritance among inbreds and hybrids or gave any indication of mode of inheritance in metabolites. When the heterosis is estimated over the mid parent (mean value of the two parents) it is known as mid parent heterosis (MPH); whereas when it is estimated over better parent it is called better or best parent heterosis (BPH). In the current study, we estimated MPH only, which is one of the most adopted ways of reporting heterosis (Melchinger et al. 2007). The MPH value was calculated using the formula MPH = 100(F1−MP/MP), where MPH = mid parent heterosis, F1 = First filial generation and MP = mean of both parents (Lisec et al. 2011). The MPH was plotted on XY axis to determine the deviation from mid-parent value. Linear contrasts in ANOVA model was used to compare hybrid vs mid-parent value (V + EH2/2 vs. EH2 × V, V + EH2/2 vs. V × EH2) for determination of heterotic mode of inheritance in metabolites (Römisch-Margl et al. 2010). To compute the mid-parent heterosis contrast, we utilized estimates of all four genotypes and specified null hypothesis as described below:
where “pq” and “qp” represents hybrids EH2 × V and V × EH2 respectively, “p’” the inbred EH2 and “q” the inbred V.
For visualization of differences between two inbred parents, we computed linear contrast for parents (EH2 vs. V) by specifying following null hypothesis:
where “p” the inbred parent EH2 and “q” the inbred parent V.
The MPH and average of inbred parent’s linear contrasts (LC) were used to decipher the heterosis effects. An additive mode of inheritance was assumed when the F1 genotype mean was not significantly different from the mean of the parents. Morphological parameters were also compared by linear contrast using one-way ANOVA (contrast: EH2 × V vs. EH2; EH2 × V vs. V; V × EH2 vs. EH2; V × EH2 vs. V; EH2 vs. V) and MPH was estimated as described above.
Partial least square (PLS) regression modeling
The PLS regression modeling was performed to study the relationship between metabolites and morphological characters. PLS regression was conducted on log transformed data with the PLS extension bundle of SPSS. Maximum numbers of latent factors were fixed as 10. A suitable model was selected based on latent factor which explained maximum variance and have the highest adjusted R2 value. For interpretation of the PLS model, VIP (variable importance projection) scores were used, where VIP scores symbolized the influence of each variable on the PLS model. VIP scores were useful in determining the metabolites that primarily influenced the morphological parameters (Hurtado et al. 2017). Therefore, VIP scores of the selected model were used to find out the most discriminatory metabolite for a specific morphological parameter. VIP score of >1 was used for selecting the discriminant metabolite for a specific trait. All statistical analyses were performed with SPSS software package v.17.0 for Windows (SPSS Corporation, Chicago, Illinois, USA).
Results
Morphological parameters of B. juncea hybrids
The two contrasting inbred lines of B. juncea used in the current study, namely Varuna and EH2, belong to two distinct and genetically diverse gene pools of B. juncea (see EH2 vs. V linear contrast in Table 1 and Table S4). The F1 hybrids obtained from these inbred lines showed heterotic potential for various agro-morphological characters during different developmental stages. Most of the morphological characters analyzed at 30 days after germination (DAG) showed positive MPH in both the F1 hybrids (Table S4). Linear contrast analysis revealed high additive inheritance in F1 (EH2 × V), whereas non-additive inheritance was observed for the F1 (V × EH2) hybrid. A high positive MPH for characters was also observed at a later 50% flowering stage (Table 1). Linear contrast of MPH showed that majority of characters in both hybrids displayed an additive mode of inheritance (8 out of 12 parameters in EH2 × V and 7 out of 12 in V × EH2 in Table 1). All the three tested reproductive characters (number of siliques on main shoot, silique length on main shoot and silique density/unit length of main shoot) showed additive inheritance. Further, pairwise linear contrast between F1 hybrids and parents at 50% flowering stage demonstrated that hybrids were significantly different from their parents. Notably, when comparison was made with the Indian parent (V) 9 out of 12 morphological characters displayed a significant difference (p ≤ 0.05). In B. juncea, 50% flowering stage was adopted in earlier studies to estimate the agronomic performances of hybrids (Pradhan et al. 1993; Ramchiary et al. 2007).
Levels of metabolites in B. juncea inbred lines and F1 hybrids
Based on the above observations, five different tissue samples representing key organs at the 50% flowering stage were taken from the two reciprocal F1 hybrids and the two inbred parents of B. juncea and used for targeted metabolite profiling. A total of 48 metabolites were quantified including 18 amino acids, 6 phytohormones, 3 sugars, 8 glucosinolates, 9 flavonoids and 4 phenolic esters. The level and distribution of these metabolites in the two inbred lines and F1 hybrids were found to vary across organs (Table S6-S10).
Amino acids
The analysis method allowed the quantification of 18 amino acids in B. juncea and the concentration of all amino acids was highest in flower buds followed by siliques and then the leaf (Table S6). Gln was found to be present in highest concentration (3012.2 to 541,753.6 nmol g−1 dry wt) followed by Glu, Pro, Ala, Ser, Asp, Thr, Val, His, Ile, Asn, Leu, Phe, Met, Tyr, Lys, Arg and Trp (57.4 to 1452.25 nmol g−1dry wt). Buds and siliques did not show any specific trend in amino acid concentrations, whereas in flag leaf, young leaf and mature leaf, a total of ten (Thr, Glu, Trp, Ser, His, Asn, Pro, Gln, Asp, Lys), nine (Glu, Trp, Ser, Asn, Pro, Leu, Gln, Asp, Lys) and eight (Glu, Ser, Ile, His, Pro, Phe, Asp, Tyr) amino acids of F1 hybrids were within parental limits, respectively. However, few amino acids in F1 hybrids also showed transgressive variation, outside the parental range.
Phytohormones
Metabolic profiling of six phytohormones viz. salicylic acid (SA), jasmonic acid (JA), abscisic acid (ABA), (-)-jasmonoyl-l-isoleucine (JA-Ile1), (+)-7-iso-jasmonoyl-L-isoleucine (JA-Ile2) and cis-(+)-12-oxo-phytodienoic acid (cis-OPDA) was carried out. The concentration of SA, JA, ABA, JA-Ile1 and JA-Ile2 was higher in reproductive tissues (floral buds and siliques), whereas cis-OPDA showed highest concentrations in vegetative tissues (flag leaf, young leaf and mature leaf) (Table S7). It was apparent that in buds, except for SA, all other phytohormones in F1 hybrids were within the parental limits, whereas, in siliques, only SA and ABA were within the parental range. No specific trend was observed in foliar tissues.
Sugars
Sugar profiling identified three major free sugars viz. fructose, glucose and sucrose, wherein, fructose was present in highest concentrations in all the plant organs studied (Table S8). Among all organs, the highest concentration of these sugars was observed in buds and siliques. The occurrence of fructose and glucose in F1 hybrids was within the parental range in young leaf, whereas other tissues exhibited transgressive variation. Sucrose was present within the parental limits across all the leaf tissues while showing transgressive variation in bud and silique of F1 hybrids.
Glucosinolates (GSL)
In total, eight GSL were detected in metabolite profiling, among which 3-butenyl GSL was found at highest concentration in all plant tissues followed by allyl GSL and 4-pentenyl GSL (Table S9). The non-aliphatic GSL namely, Indol-3-ylmethyl (I3M), 4-Hydroxyindol-3-ylmethyl (4OHI3M), 4-Methoxyindol-3-ylmethyl (4MOI3M) and 2-Phenylethyl (2-PE) were relatively low in abundance. Unlike other metabolites, most GSL, except 5-methylsulphinylpentyl (5MSOP), were not preferentially localized in any specific plant part. While Varuna accumulates the highest concentration of 3-butenyl GSL and allyl GSL in all plant parts, most GSLs were either absent or present in very low concentrations in the EH2 inbred line. The allyl GSL and 3-butenyl GSL content of F1 hybrids was within the parental ranges in the flag, young and mature leaf whereas bud and silique showed transgressive variation (Table S9).
Flavonoids and phenolic esters
We detected a total of 13 flavonoids and phenolic esters in B. juncea of which 10 compounds were identified each in flag leaf, young leaf and mature leaf, whereas only seven were identified in buds and siliques (Table S10). Two flavonoids, quercetin-glucoside-rhamnoside (QN-GR) and isorhamnetin-glucoside-rhamnoside (IS-GR) were unique to buds. Phenolic esters sinapic acid glucoside (S-G) and isorhamnetin-3-glucoside (IS-G) and a flavonoid sinapoylferuloylgentiobiose (SF-GB) occurred in all five plant organs. Five flavonoids namely, kaempferol-3-hydroxyferuloylsophoroside-7-glucoside (KM-HFSG), kaempferol-3-sophorotrioside-7-glucoside (KM-STG), kaempferol-3-glucoside-7-glucoside (KMGG), kaempferol-3-feruloylsophoroside-7-glucoside (KM-FSG) and a phenolic ester, hydroxyferulic acid glucoside (HF-G) were observed only in leaf tissues. It was notable that isorhamnetin derivatives were only found in buds and siliques, while kaempferol derivatives were present only in leaf. Interestingly, among the three leaf tissues, the flag leaf revealed increased abundance of all the kaempferol derivatives. Four compounds {isorhamnetin-3-glucoside-7-glucoside (IS-GG), S-G, IS-GR and IS-G} out of six were within parental limits in the F1 hybrids while two others showed transgressive variation. Among the four genotypes, EH2 had a higher concentration of flavonoids and phenolic esters in all plant tissues except siliques. In flag leaf, the concentration of four flavonoids (KM-HFSG, KM-GG, KM-SSG, S-G,) and a phenolic ester (KM-SSG) was within parental range while other tissues displayed transgressive variation (Table S10).
Principal component analysis of metabolic data
We performed principal component analysis (PCA) on log10 transformed metabolite data (of all 48 metabolites) from the five different tissues of B. juncea. The first two components (PC1 and PC2) explained 49.4–60.8% of the variability across all the developmental tissues (Table S11).
The PCA factor loading plot (Fig. 1a) of the first two PCs of bud samples clearly separated F1 hybrids (EH2 × V and V × EH2) from their inbred parents (EH2 and V), where both F1 hybrids were clustered together and are separated from both inbred parents, forming their own clusters. PCA factor loading revealed that Ala, Ser, Pro, Val, Thr, Ile, Leu, Met, Arg, Tyr, Lys, cis-OPDA and glucose were highly correlated with PC1; while Phe, 3-butenyl GSL, IS-GG, JA, JA-Ile1, JA-Ile2 were highly correlated with PC2 (Table S12). In the PCA loading plot (Fig. 1b) of the first two PCs of siliques, hybrids clustered together while inbred parents formed their own separate clusters. However, the separation was not as clear as for buds. The metabolites Ala, Ser, Val, Thr, Ile, Phe, Arg, Tyr, Asn, Gln, SA, fructose and glucose were highly correlated with PC1; while Glu, IS-GG, S-G, SHF-GB, JA-Ile2, 4-OHI3M GSL, 4-pentenyl GSL and I3M GSL were highly correlated with PC2 (Table S13).
PCA loading plot of metabolites in B. juncea showing separation of hybrids (EH2 × V and V × EH2) from inbreds (EH2 and Varuna) in a bud and b silique. Orange box represents metabolites and blue box represents lines (n = 5). The variability for each principal component is depicted in parenthesis and provided as supplementary Tables S12-S13. The hybrids are grouped separately from their inbreds (marked as circles)
PCA analysis of metabolites was also performed in the three leaf tissues (flag, young and mature leaf). The PCA loading plot (Fig. 2a) of the first two PCs of flag leaf clustered hybrids together, whereas inbred parents formed separate clusters. The plot showed that hybrids were located closer to the Indian parent (Varuna). PCA factor loading revealed that Ser, Pro, Asp, Glu, Trp, Asn, Gln, allyl, 3-butenyl GSL, KM-HFSG, KM-STG, KM-GG, KM-SSG, KM-FSG, S-G, SHF-GB, IS-G, SF-GB, fructose and sucrose were highly correlated with PC1; while Leu, Phe, Tyr and ABA were highly correlated with PC2 (Table S14). The PCA loading plot (Fig. 2b) of the first two PCs of young leaf revealed a clustering of hybrids and inbreds, wherein hybrids formed a distinct cluster separated from their inbred parents. The hybrids were located towards the center of inbreds, but closer to one of the parents (Varuna). A high correlation was observed with PC1 in metabolites Ser, Pro, Ile, Asp, Glu, Gln, allyl GSL, 3-butenyl GSL, I3M GSL, KM-HFSG, KM-GG, KM-SSG, S-G, SHF-GB, and IS-G, while Val, ABA and sucrose were highly correlated with PC2 (Table S15). In the PCA loading plot (Fig. 2c) of the first two PCs of mature leaf, hybrids clustered together between the two inbred lines. In mature leaf, Ser, Pro, Ile, Leu, Asp, Glu, Phe, Tyr, Gln, allyl GSL, I3M GSL, KM-SSG GSL, S-G and JA were highly correlated with PC1, while KM-STG, KM-GG and KM-FSG were highly correlated with PC2 (Table S16).
PCA loading plot of metabolites in B. juncea showing separation of hybrids (EH2 × V and V × EH2) from inbreds (EH2 and Varuna) in a flag leaf, b young leaf, and c mature leaf. Orange box represents metabolites and blue box represents lines (n = 5). The variability for each principal component is depicted in parenthesis and provided as supplementary Tables S14-16. The hybrids are grouped separately from their inbreds (marked as circles)
Mid-parent heterosis of metabolites in F1 hybrids
The mid-parent heterosis (MPH) of hybrids from parental means for all the tested metabolites was investigated across the five developmental tissues. Most metabolites showed negative MPH values in all tested plant tissue types of the two F1 hybrids of B. juncea (Fig. 3), except in buds of the V × EH2 hybrid wherein most of the identified metabolites showed positive MPH (Table S17). We further made a comparison of MPH observed for each of the primary and secondary metabolite classes across developmental tissues of the two F1 hybrids. Negative MPH values were obtained for amino acids in both F1 hybrids. Four amino acids (Val, Thr, Phe, and Tyr) in the EH2xV hybrid, and three amino acids (Asp, Glu, and Gln) in the VxEH2 hybrid showed negative MPH values in all five developmental tissues (Fig. 3; Table S17). A mixed trend of MPH (both negative and positive MPH observed) was evident for occurrence of phytohormones in both the F1 hybrids (Table S17). Interestingly, among sugars, fructose indicated negative MPH values in almost all tissues types of both the F1 hybrids. Glucose and sucrose exhibited both positive and negative trends in MPH depending upon tissue type and hybrid (Fig. 3; Table S17). A mixed pattern of MPH values was observed for GSL metabolites across all the five developmental tissues of both the F1 reciprocal hybrids (Fig. 3; Table S17). The aliphatic GSL, 3-butenyl GSL and 4-pentenyl GSL showed positive heterosis across five developmental tissues of both the F1 hybrids, whereas allyl GSL displayed positive MPH values across four developmental tissues. Out of ten flavonoids and phenolic esters detected in the leaf, seven (HF-G, KM-HFSG, KM-STG, KM-GG, KM-SSG, KM-FSG, and S-G) had negative MPH values in both F1 hybrids (Fig. 3; Table S17). However, in buds and siliques, these compounds displayed a mixed pattern of MPH values in both the crosses. S-G is a common phenolic ester that showed negative MPH values in both F1 hybrids in all developmental tissues and can be a potential candidate for detecting negative MPH in Brassica. The negative MPH values of flavonoids and phenolic esters were significantly higher (ranging from 0 to −74.848) compared to amino acids and sugars (ranging from 0 to −10.086) (Table S17).
Plot showing deviation of mid-parent heterosis (MPH) from mid-parent value at five developmental tissues namely, a buds, b siliques, c flag leaf, d young leaf, and e mature leaf. Each bar represents average deviation of hybrids from MPH value provided in Supplementary Table S18. Values within (−) 20% to (+) 20% MPH are only depicted in the figure. ABA, abscisic acid; cis-OPDA, cis-12-Oxo-Phytodienoic acid; JA, jasmonic acid; JA-Ile1, (-)-jasmonoyl-l-isoleucine; JA-Ile2, (+)-7-iso-jasmonoyl-l-isoleucine; SA, salicylic acid; Allyl, allyl isothiocyanate; I3M, indol-3-ylmethyl; 4MOI3M, 4-Methoxyindol-3-ylmethyl; 5-MSOP, 5-methylsulphinylpentyl; 4-OHI3M, 4-Hydroxyindol-3-ylmethyl; 2PE, 2-Phenylethyl; HF-G, hydroxyferuloylglucose; IS-G, isorhamnetin 3-glucoside; IS-GG, isorhamnetin 3-glucoside-7-glucoside; IS-GR, isorhamnetin glucoside-rhamnoside; KMFSG, kaempferol 3-feruloylsophoroside-7-glucoside; KMGG, kaempferol 3-glucoside-7-glucoside; KMHFSG, kaempferol 3-hydroxyferuloylsophoroside-7-glucoside; KMSSG, kaempferol 3-sinapoylsophoroside-7-glucoside; KMSTG, kaempferol 3-sophorotrioside-7-glucoside; QN-GR, quercetin glucoside-rhamnoside; SF-GB, sinapoylferuloylgentiobiose; SH-FGB, sinapoylhydroxyferuloylgentiobiose; S-G, sinapoylglucose
Mode of inheritance analysis
To dissect the mode of inheritance of metabolites in F1 hybrids, linear contrasts in ANOVA model was used to compare the levels of metabolites between hybrid and parental mean (EH2 × V vs. (V + EH2)/2; V × EH2 vs. (V + EH2)/2). Metabolites displayed both additive and non-additive mode of inheritance in B. juncea F1 hybrids (Table 2). Notably, the numbers of metabolites showing an additive mode of inheritance were higher in buds and siliques (52.77–97.14%) compared to leaf tissue (47.37–80%) in both reciprocal F1 hybrids. Additive mode of inheritance was observed in buds and siliques for more than 85% metabolites except for metabolites of V × EH2 hybrid, which revealed additive mode of inheritance in 51.42% cases. Similarly, higher additive mode of inheritance was also detected in young and mature leaf tissues (53.85–80%) compared to the flag leaf (47.3%). To visualize the differences between two inbred parents, a linear contrast in ANOVA model was also performed between two inbred lines (EH2 vs. V). It was observed that 51.28, 29.72, 63.16, 39.02, and 64.86% metabolites were significantly (p ≤ 0.05) different in the bud, silique, flag leaf, young leaf and mature leaf, respectively, between the two inbred parents (Table S18). Non-additive behavior was mainly shown by flavonoids and phenolic esters in case of foliar tissues (Table 2). Flag leaf showed non-additive mode of inheritance in both hybrids for almost all flavonoids and phenolic esters. Among GSL, 3-butenyl GSL and 4-pentenyl GSL demonstrated non-additive behavior in almost all tissues (Table 2).
Covariation of metabolites and morphological parameters by PLS regression
PLS regression was performed to investigate the covariance between the metabolites and morphological traits. The analysis revealed preferential association of primary and secondary metabolites with morphological parameters of hybrid and inbred lines, respectively (Tables 3, 4; Table S19–S28). In buds, primary metabolites (Ala, Ser, Pro, Val, Thr, Ile, His, Phe, Trp, fructose, and glucose) displayed higher positive association with morphological parameters in hybrids (Tables 3, S19), whereas secondary metabolites, particularly jasmonates and flavonoids displayed higher positive association with morphological parameters in inbreds (Table S20). The preferential association of metabolites with morphological parameters were not clearly visible in silique. Most of the secondary metabolites of inbred siliques showed positive association with morphology (Table S22).
Among foliar tissues, differential association of secondary metabolites with morphological parameters was quite visible in flag leaf (Tables 3, S23, and S24). In flag leaf of inbreds, secondary metabolites (allyl GSL, 3-butenyl GSL, 4-pentenyl GSL, KM-HFSG, KM-STG, KM-GG, KM-SSG, KM-FSG, S-G, SHFGB, ISG, and SFGB) displayed both negative and positive association with morphological parameters, in equal proportion. In young leaf, differential association of metabolites was not clearly visible (Tables 3, S25 and S26). Secondary metabolites in hybrids displayed negative relationship with morphological parameters, in contrast to inbreds that showed positive association. Mature leaf also exhibits comparatively higher and positive association of secondary metabolites with morphological parameters in inbreds compared to hybrids (Tables S27 and S28). Mature leaf displayed comparably higher association of primary metabolites with morphological parameters in hybrids.
Detailed association of morphological parameters with specific metabolites are described in Table 4. Among different class of metabolites, Ser and Pro in amino acids; ABA in phytohormones; sucrose in sugars; 3-butenyl in GSL; and KM-SSG, KM-HFSG, S-G, KM-GG, and SFGB in flavonoids and phenolic ester, displayed higher association with morphological parameters (Tables 4, S19–S28).
Discussion
Heterosis is a complex genetic phenomenon wherein hybrids show superior performance over their parental inbred lines. Indian oilseed mustard, B. juncea (AjAjBjBj, 2n = 36) is an allotetraploid species that originated from natural hybridization of two diploid progenitors, B. rapa (ArAr, 2n = 20) and B. nigra (BnBn, 2n = 16). The importance of B. juncea as an oilseed crop is growing globally due to its unique and favorable traits, such as resistance to biotic and abiotic stresses and low rate of pod shattering. Prior efforts have significantly increased the seed yield and quality traits (Augustine et al. 2014; Wei et al. 2016) but more genetic improvement is necessary (Grover and Pental 2003).
Earlier studies have provided some insights into the mode of inheritance of metabolites in hybrids of maize (Römisch-Margl et al. 2010; Lisec et al. 2011; Meyer et al. 2012; Wang et al. 2014), rice (Ma et al. 2011) and Arabidopsis (Korn et al. 2010). However, metabolite analysis in these studies was limited to a single tissue type during early stages of plant development. To get a better insight of metabolite inheritance, we conducted a detailed analysis of primary and secondary metabolites in both vegetative and reproductive tissue types during 50% flowering stage in oilseed B. juncea. Our data showed significant difference in morphological parameters for both vegetative and reproductive characters at 50% flowering (Table 1). We therefore compared the metabolite profiles of hybrids and parental lines at this stage to identify differences in a range of primary and secondary metabolites and to ascertain the predominant mode of metabolite inheritance during heterosis and dynamics involved therein. The key information obtained in this study has been summarized in Fig. 4.
Summary describing inheritance of primary and secondary metabolites in the hybrids of oilseed crop B. juncea. The figure integrates different datatypes and statistical analyses presented in the results. ABA, abscisic acid; cis-OPDA, cis-12-Oxo-phytodienoic acid; JA, jasmonic acid; JA-Ile1, (-)-jasmonoyl-l-isoleucine; JA-Ile2, (+)-7-iso-jasmonoyl-l-isoleucine; SA, salicylic acid; Allyl, allyl isothiocyanate; 5-MSOP, 5-methylsulphinylpentyl; HF-G, hydroxyferuloylglucose; IS-G, isorhamnetin 3-glucoside; IS-GG, isorhamnetin 3-glucoside-7-glucoside; IS-GR, isorhamnetin glucoside-rhamnoside; KMFSG, kaempferol 3-feruloylsophoroside-7-glucoside; KMGG, kaempferol 3-glucoside-7-glucoside; KMHFSG, kaempferol 3-hydroxyferuloylsophoroside-7-glucoside; KMSSG, kaempferol 3-sinapoylsophoroside-7-glucoside; KMSTG, kaempferol 3-sophorotrioside-7-glucoside; QN-GR, quercetin glucoside-rhamnoside; SF-GB, sinapoylferuloylgentiobiose; SH-FGB, sinapoylhydroxyferuloylgentiobiose; S-G, sinapoylglucose
Reproductive tissues accumulate higher levels of primary metabolites in B. juncea
Metabolite profiling of both inbred parents and F1 hybrids suggests that reproductive tissues, in general, displayed significantly higher level of all the investigated primary metabolites (amino acids and sugars) compared to vegetative tissues. On the other hand, secondary metabolites (phytohormones, glucosinolates, flavonoids, and phenolic esters) revealed a mixed trend of distribution across different tissue types (Fig. 4). Higher accumulation of amino acids in reproductive tissues correlates with higher nitrogen demand of developing flowers, which has a significant consequence with respect to flower setting, amount of floral abortion, pollen and embryo development and seed production. Pollen development represents a major sink for nitrogen in developing flowers, and it must be imported from source organs such as leaves (Lee and Tegeder 2004). Reproductive tissues also exhibited higher sugar accumulation, which is vital as plants undergo transition from vegetative to reproductive stage in such a manner that sufficient carbohydrate resources are available to support the energy intensive processes of flowering and seed production (Wahl et al. 2013).
Key metabolites discriminating inbreds from F1 hybrids
Some metabolites were more important than others in shaping the overall mode of metabolite inheritance in F1 hybrids. Factor loading in PCA revealed that the amino acid, Ser is the most important metabolite in all five developmental tissues (Figs. 2, 3). Ser is an important amino acid serving as an essential constituent of proteins, and a substrate for the biosynthesis of phosphatidylserine and Trp, and Trp-derived secondary metabolites such as auxin and indolic glucosinolates (Benstein et al. 2013). The amino acids Pro, Ile, Tyr, Phe, Gln, Glu, and phenolic ester S-G contribute to the differences in four developmental tissues whereas Asp, Leu, SHF-GB, KM-GG, KM-SSG, allyl GSL, 3-butenyl GSL and I3M GSL contribute in three developmental tissues. Further, PLS regression analysis (Tables 3, 4) revealed that amino acids, flavonoids and phenolic esters are important contributors in morphological trait development (Fig. 4). Amino acids Pro>Ser>Gln>Asp (in order of importance) are the most important contributors to plant morphology. Among flavonoids and phenolic ester, KM-SSG>KM-HFSG>S-G>SFGB>ISGG>ISG>KMFSG seem to have high association with plant morphology. Differential distribution of these metabolites suggests that their concentrations could play important roles in causing heterosis in hybrids. In maize, Römisch-Margl et al. (2010) reported strong heterotic mode of inheritances for Ser, Glu, Asn, and sucrose. Notably, amino acids, in general, represent a key class of primary metabolites that has a potential to be used as “heterotic biomarkers” in different crops. In previous studies, Lorenz (1972, 1975) made use of differences in concentration of seven amino acids (Asp, Glu, Gln, Gly, Ala, Ser, and sarcosine) between inbred parents as a biomarker for hybrid yield in maize. Studies over the last decade on the role of metabolic markers in plant performance have been reviewed by Fernandez et al. (2016).
Reproductive tissues predominantly display additive mode of metabolite inheritance
Using genetic analysis, Shi et al. (2011) reported that a dynamic and complex genetic network regulates yield heterosis in rapeseed (B. napus L.), shaped by additive and non-additive modes of inheritance, and epistatic interactions. Our data (Table 2) also demonstrates that both additive and non-additive modes of metabolite inheritance exist in B. juncea hybrids. Notably, the reproductive tissues, especially buds, displayed a higher preference for additivity for most of the tested metabolites (Fig. 4). Non-additive mode of inheritance was mainly restricted to secondary metabolites (GSLs and phenolic esters and flavonoids) in leaf tissues. PCA loading plots of metabolites, specifically from the buds clearly separated the parental inbred lines from their hybrids, wherein hybrids clustered together between the two parental clusters, indicating additivity (Wang et al. 2014). Since B. juncea is an oilseed crop, the higher additivity of metabolites in reproductive tissues may result in higher oilseed content of the hybrids, even under fluctuating environmental conditions. Further, our analysis also separated the two parental B. juncea inbreds, which originated from two different geographic areas (India and Europe) into two different clusters. These results demonstrate the metabolic distinctiveness of the two gene pools, which correspond well with various morphogenetic studies reported in B. juncea (Pradhan et al. 1993; Ramchiary et al. 2007).
Hybrids favor growth by modulating secondary metabolism
In the present study, the MPH values in B. juncea F1 hybrids for secondary metabolites such as indolic GSL, flavonoids and phenolic esters were found to be more negative in comparison to primary metabolites (amino acids and fructose) (Fig. 4). Phe and Tyr are the most frequent amino acids, which showed negative MPH that can be correlated with lower flavonoid levels, as deamination of Phe is a primary step in flavonoid biosynthesis pathway. PLS regression analysis further emphasized the role of secondary metabolite dynamics. Our data revealed a predominantly higher positive association of secondary metabolites in parents with morphological parameters as compared to hybrids, especially in case of buds. Among secondary metabolites, flavonoids play important role in stress response and act as negative regulators of auxin transport in vivo (Petrussa et al. 2013; Brown et al. 2001; Peer et al. 2004). Shen et al. (2012) showed that genes involved in flavonoid biosynthesis are repressed in hybrids of A. thaliana and was associated with increased DNA methylation in intraspecific hybrids. According to Miller et al. (2015), stress-responsive genes are repressed in hybrids under normal conditions but are induced to mid-parent or higher levels under stress at certain times of the day, thus balancing “trade-off” between stress response and growth. This diurnal repression of stress-responsive genes was further substantiated by repression of circadian oscillator genes CIRCADIAN CLOCK ASSOCIATED1 and LATE ELONGATED HYPOCOTYL (Shen et al. 2012; Miller et al. 2015). The higher proportion of negative MPH observed for flavonoids and phenolic esters in B. juncea hybrids might reflect a reduction in basal defense and stress responses under normal growth conditions, thereby favoring the better growth of the hybrids.
Among the glucosinolates, the aliphatic GSL (allyl GSL, 3-butenyl GSL, and 4-pentenyl GSL) displayed positive MPH values, but the indolic GSL (I3M GSL) had highly negative MPH values in most of the developmental tissues of the two F1 hybrids (Fig. 4). It has been generally observed that indolic GSL are much more stress responsive than aliphatic GSL (Textor and Gershenzon 2009). Groszmann et al. (2015) also reported down regulation of indolic GSL in A. thaliana hybrids and they concluded that blocking the production of indole-derived secondary metabolites leads to elevated levels of indole-3-acetic acid (IAA) via an accumulation of indole-3-acetaldoxime (IAOX) and subsequently an increased flow through the IAOX → indole-3-acetamide (IAM) → IAA route. Thus, the GSL composition in the F1 hybrids is also consistent with the trend that stress responsive metabolites have more negative MPH values. In addition, negative MPH for SA in bud, silique and flag leaf was also observed in this study. Groszmann et al. (2015) observed that altered expression in the hybrids leads to down regulation of SA biosynthesis pathway and increase in auxin biosynthesis. ABA, which is commonly known as the “stress hormone”, also exhibit negative MPH in all the developmental tissues. Overall, the metabolic profiles of the two reciprocal F1 hybrids appear to show that growth is favored in hybrids by altering secondary metabolite production. This underpins the importance of resource ‘trade-offs’ between growth and defense metabolites (Kleessen et al. 2014) during heterosis in B. juncea.
Overall, our study unravels the presence of a complex inheritance pattern of primary and secondary metabolites across developmental tissues in B. juncea hybrids, which is coordinately governed by additive and non-additive modes of inheritance, and alteration of secondary metabolism to favor plant growth in hybrids. However, analysis of other key metabolites such as lipids, starch, oil, and other storage substances could also be undertaken to get better insight into metabolic controls in B. juncea hybrids. In addition, information from different omics platforms, circadian rhythms, photosynthetic capacity, respiration, and flux measurements can be integrated to provide the system level information about molecular mechanisms that switch “on” and “off” growth in different tissues of the plant.
References
Andorf S, Meyer RC, Selbig J, Altmann T, Repsilber D (2012) Integration of a systems biological network analysis and QTL results for biomass heterosis in Arabidopsis thaliana. PLoS ONE 7:e49951
Augustine R, Arya GC, Nambiar DM, Kumar R, Bisht NC (2014) Translational genomics in Brassica crops: challenges, progress, and future prospects. Plant Biotechnol Rep 8:65–81
Benstein RM, Ludewig K, Wulfert S, Wittek S, Gigolashvili T, Frerigmann H et al. (2013) Arabidopsis phosphoglycerate dehydrogenase1 of the phosphoserine pathway is essential for development and required for ammonium assimilation and tryptophan biosynthesis. Plant Cell 25:5011–5029
Birchler JA, Auger DL, Riddle NC (2003) In search of the molecular basis of heterosis. Plant Cell Online 15:2236–2239
Brown DE, Rashotte AM, Murphy AS, Normanly J, Tague BW, Peer WA et al. (2001) Flavonoids act as negative regulators of auxin transport in vivo in Arabidopsis. Plant Physiol 126:524–535
Burow M, Müller R, Gershenzon J, Wittstock U (2006) Altered glucosinolate hydrolysis in genetically engineered Arabidopsis thaliana and its influence on the larval development of Spodoptera littoralis. J Chem Ecol 32:2333–2349
Crow JF (2001) Heterosis. In: Brenner S, Miller JH (eds) Encyclopedia of genetics, Vol 1. Academic Press, San Diego.
Dahal D, Mooney BP, Newton KJ (2012) Specific changes in total and mitochondrial proteomes are associated with higher levels of heterosis in maize hybrids. Plant J 72:70–83
Ding D, Wang Y, Han M, Fu Z, Li W, Liu Z et al. (2012) MicroRNA transcriptomic analysis of heterosis during maize seed germination. PLoS ONE 7:e39578
Ding H, Qin C, Luo X, Li L, Chen Z, Liu H et al. (2014) Heterosis in early maize ear inflorescence development: a genome-wide transcription analysis for two maize inbred lines and their hybrid. Int J Mol Sci 15:13892–13915
Fernandez O, Urrutia M, Bernillon S, Giauffret C, Tardieu F, Gouis JL et al. (2016) Fortune telling: metabolic markers of plant performance. Metabolomics 12:158
Giraud H, Lehermeier C, Bauer E, Falque M, Segura V, Bauland C et al. (2014) Linkage disequilibrium with linkage analysis of multiline crosses reveals different multiallelic QTL for hybrid performance in the flint and dent heterotic groups of maize. Genetics 198:1717–1734
Groszmann M, Gonzalez-Bayon R, Lyons RL, Greaves IK, Kazan K, Peacock WJ et al. (2015) Hormone-regulated defense and stress response networks contribute to heterosis in Arabidopsis F1 hybrids. Proc Natl Acad Sci USA 112:E6397–E6406
Grover A, Pental D (2003) Breeding objectives and requirements for producing transgenics for major field crops of India. Curr Sci 84:310–320
Hochholdinger F, Hoecker N (2007) Towards the molecular basis of heterosis. Trends Plant Sci 12:427–432
Hurtado C, Parastar H, Matamoros V, Piña B, Tauler R, Bayona JM (2017) Linking the morphological and metabolomic response of Lactuca sativa L exposed to emerging contaminants using GC × GC-MS and chemometric tools. Sci Rep 7:6546
Kleessen S, Laitinen R, Fusari CM, Antonio C, Sulpice R, Fernie AR et al. (2014) Metabolic efficiency underpins performance trade-offs in growth of Arabidopsis thaliana. Nat Commun 5:3537
Korn M, Gärtner T, Erban A, Kopka J, Selbig J, Hincha DK (2010) Predicting Arabidopsis freezing tolerance and heterosis in freezing tolerance from metabolite composition. Mol Plant 3:224–235
Kramell R, Schmidt J, Schneider G, Sembdner G, Schreiber K (1988) Synthesis of n-(jasmonoyl)amino acid conjugates. Tetrahedron 44:5791–5807
Lee YH, Tegeder M (2004) Selective expression of a novel high-affinity transport system for acidic and neutral amino acids in the tapetum cells of Arabidopsis flowers. Plant J 40:60–74
Li A, Liu D, Wu J, Zhao X, Hao M, Geng S et al. (2014) mRNA and small RNA transcriptomes reveal insights into dynamic homoeolog regulation of allopolyploid heterosis in nascent hexaploid wheat. Plant Cell 26:1878–1900
Li L, Lu K, Chen Z, Mu T, Hu Z, Li X (2008) Dominance, overdominance and epistasis condition the heterosis in two heterotic rice hybrids. Genetics 180:1725–1742
Lin L-Z, Harnly JM (2010) Phenolic component profiles of mustard greens, yu choy, and 15 other Brassica vegetables. J Agric Food Chem 58:6850–6857
Lin L-Z, Sun J, Chen P, Harnly J (2011) UHPLC-PDA-ESI/HRMS/MS(n) analysis of anthocyanins, flavonol glycosides, and hydroxycinnamic acid derivatives in red mustard greens (Brassicajuncea Coss variety). J Agric Food Chem 59:12059–12072
Lisec J, Römisch-Margl L, Nikoloski Z, Piepho HP, Giavalisco P, Selbig J et al. (2011) Corn hybrids display lower metabolite variability and complex metabolite inheritance patterns. Plant J 68:326–336
Liu L, Lai Y, Cheng J, Wang L, Du W, Wang Z et al. (2014a) Dynamic quantitative trait locus analysis of seed vigor at three maturity stages in rice. PLoS ONE 9:e115732
Liu TJ, Sun LF, Shan XH, Wu Y, Su SZ, Li SP et al. (2014b) Analysis of DNA methylation patterns and levels in maize hybrids and their parents. Genet Mol Res 13:8458–8468
Lorenz H (1972) Beziehungen zwischen dem Aminosaurepool von Inzuchtlinien und dem Ertrag ihrer Hybriden bei Mais. Z Pflanzenzucht 68:155–170
Lorenz H (1975) Relationships between yield and free amino acids in hybrids and inbred lines of corn (Zea mays L.). Z Pflanzenzucht 75:10–23
Ma Q, Hedden P, Zhang Q (2011) Heterosis in rice seedlings: its relationship to gibberellin content and expression of gibberellin metabolism and signaling genes. Plant Physiol 156:1905–1920
Madsen SR, Kunert G, Reichelt M, Gershenzon J, Halkier BA (2015) Feeding on leaves of the glucosinolate transporter mutant gtr1gtr2 reduces fitness of Myzus persicae. J Chem Ecol 41:975–984
Melchinger AE, Utz HF, Piepho HP, Zeng ZB, Schon CC (2007) The role of epistasis in the manifestation of heterosis: a system-oriented approach. Genetics 117:1815–1825
Meyer RC, Kusterer B, Lisec J, Steinfath M, Becher M, Scharr H et al. (2010) QTL analysis of early stage heterosis for biomass in Arabidopsis. Theor Appl Genet 120:227–237
Meyer RC, Witucka-Wall H, Becher M, Blacha A, Boudichevskaia A, Dörmann P et al. (2012) Heterosis manifestation during early Arabidopsis seedling development is characterized by intermediate gene expression and enhanced metabolic activity in the hybrids. Plant J 71:669–683
Miller M, Song Q, Shi X, Juenger TE, Chen ZJ (2015) Natural variation in timing of stress-responsive gene expression predicts heterosis in intraspecific hybrids of Arabidopsis. Nat Commun 6:7453
Paschold A, Jia Y, Marcon C, Lund S, Larson NB, Yeh CT et al. (2012) Complementation contributes to transcriptome complexity in maize (Zea mays L.) hybrids relative to their inbred parents. Genome Res 22:2445–2454
Paschold A, Larson NB, Marcon C, Schnable JC, Yeh CT, Lanz C et al. (2014) Nonsyntenic genes drive highly dynamic complementation of gene expression in maize hybrids. Plant Cell 26:3939–3948
Patti GJ, Yanes O, Siuzdak G (2012) Innovation: metabolomics: the apogee of the omics trilogy. Nat Rev Mol Cell Biol 13:263–269
Peer WA, Bandyopadhyay A, Blakeslee JJ, Makam SN, Chen RJ, Masson PH et al. (2004) Variation in expression and protein localization of the PIN family of auxin efflux facilitator proteins in flavonoid mutants with altered auxin transport in Arabidopsis thaliana. Plant Cell Online 16:1898–1911
Petrussa E, Braidot E, Zancani M, Peresson C, Bertolini A, Patui S et al. (2013) Plant flavonoids–biosynthesis, transport and involvement in stress responses. Int J Mol Sci 14:14950–14973
Pradhan AK, Sodhi YS, Mukhopadhyay A, Pental D (1993) Heterosis breeding in Indian mustard (Brassica juncea L. Czern & Coss): analysis of component characters contributing to heterosis for yield. Euphytica 69:219–229
Qin J, Scheuring CF, Wei G, Zhi H, Zhang M, Huang JJ et al. (2013) Identification and characterization of a repertoire of genes differentially expressed in developing top ear shoots between a superior hybrid and its parental inbreds in Zea mays L. Mol Genet Genom 288:691–705
Ramchiary N, Bisht NC, Gupta V, Mukhopadhyay A, Arumugam N, Sodhi YS et al. (2007) QTL analysis reveals context-dependent loci for seed glucosinolate trait in the oilseed Brassica juncea: importance of recurrent selection backcross scheme for the identification of ‘true’ QTL. Theor Appl Genet 116:77–85
Römisch-Margl L, Spielbauer G, Schützenmeister A, Schwab W, Piepho H-P, Genschel U et al. (2010) Heterotic patterns of sugar and amino acid components in developing maize kernels. Theor Appl Genet 120:369–381
Shen H, He H, Li J, Chen W, Wang X, Guo L et al. (2012) Genome-wide analysis of DNA methylation and gene expression changes in two Arabidopsis ecotypes and their reciprocal hybrids. Plant Cell 24:875–892
Shi J, Li R, Zou J, Long Y, Meng J (2011) A dynamic and complex network regulates the heterosis of yield-correlated traits in rapeseed (Brassica napus L.). PLoS ONE 6:e21645
Shull GH (1952) Beginnings of the heterosis concept. In: Gowen JW (eds) Heterosis. Iowa State College Press, Iowa city, p 14–48
Song S, Qu H, Chen C, Hu S, Yu J (2007) Differential gene expression in an elite hybrid rice cultivar (Oryza sativa, L) and its parental lines based on SAGE data. BMC Plant Biol 7:49
Stupar RM, Gardiner JM, Oldre AG, Haun WJ, Chandler VL, Springer NM (2008) Gene expression analyses in maize inbreds and hybrids with varying levels of heterosis. BMC Plant Biol 8:33
Textor S, Gershenzon J (2009) Herbivore induction of the glucosinolate–myrosinase defense system: major trends, biochemical bases and ecological significance. Phytochem Rev 8:149–170
Vahabi K, Sherameti I, Bakshi M, Mrozinska A, Ludwig A, Reichelt M et al. (2015) The interaction of Arabidopsis with Piriformospora indica shifts from initial transient stress induced by fungus-released chemical mediators to a mutualistic interaction after physical contact of the two symbionts. BMC Plant Biol 15:58
Wahl V, Ponnu J, Schlereth A, Arrivault S, Langenecker T, Franke A et al. (2013) Regulation of flowering by trehalose-6-phosphate signaling in Arabidopsis thaliana. Science 339:704–707
Wang L, Greaves IK, Groszmann M, Wu LM, Dennis ES, Peacock WJ (2015) Hybrid mimics and hybrid vigor in Arabidopsis. Proc Natl Acad Sci USA 112:E4959–E4967
Wang Z, Xue Z, Wang T (2014) Differential analysis of proteomes and metabolomes reveals additively balanced networking for metabolism in maize heterosis. J Proteome Res 13:3987–4001
Wei Z, Wang M, Chang S, Wu C, Liu P, Meng J et al. (2016) Introgressing subgenome components from Brassica rapa and B. carinata to B. juncea for broadening its genetic base and exploring intersubgenomic heterosis. Front Plant Sci 7:1677
Xin M, Yang G, Yao Y, Peng H, Hu Z, Sun Q et al. (2015) Temporal small RNA transcriptome profiling unraveled partitioned miRNA expression in developing maize endosperms between reciprocal crosses. Front Plant Sci 6:744
Zhai R, Feng Y, Wang H, Zhan X, Shen X, Wu W et al. (2013) Transcriptome analysis of rice root heterosis by RNA-Seq. BMC Genom 14:19
Zhang L, Peng Y, Wei X, Dai Y, Yuan D, Lu Y et al. (2014) Small RNAs as important regulators for the hybrid vigour of super-hybrid rice. J Exp Bot 65:5989–6002
Zhiguo E, Huang S, Zhang Y, Ge L, Wang L (2014) Genome-wide transcriptome profiles of rice hybrids and their parents. Int J Mol Sci 15:20833–20845
Acknowledgements
The work was supported by NIPGR core grant to NCB. PKB acknowledge the National Post-doctoral fellowship (NPDF grant no. PDF/2015/000391/LS) from SERB, DST (India). RA was funded with short-term research fellowship from NIPGR, India. NCB acknowledges the Max Planck-India mobility grant jointly funded by Max Planck Society (Germany) and DST (India). MR and JG acknowledge the funding by the Max Planck Society (Germany). We sincerely thank Dr. Payal Sanadhya (PS) for her help in drawing Fig. 4 and Dr. Arun Jagannath for language editing. Insightful suggestions from two anonymous reviewers are highly acknowledged.
Author information
Authors and Affiliations
Contributions
PKB and NCB planned and designed the research; PKB, MR, RA, JG, and NCB performed experiments, conducted field work, analyzed and interpreted data; PKB and NCB wrote the manuscript and all authors approved it.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Bajpai, P.K., Reichelt, M., Augustine, R. et al. Heterotic patterns of primary and secondary metabolites in the oilseed crop Brassica juncea. Heredity 123, 318–336 (2019). https://doi.org/10.1038/s41437-019-0213-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-019-0213-3
- Springer Nature Switzerland AG
This article is cited by
-
Combined transcriptome and metabolome analysis reveals the effects of light quality on maize hybrids
BMC Plant Biology (2023)
-
Heterosis for capsacinoids accumulation in chili pepper hybrids is dependent on parent-of-origin effect
Scientific Reports (2022)
-
Homoeolog gene expression analysis reveals novel expression biases in upland hybrid cotton under intraspecific hybridization
Functional & Integrative Genomics (2022)
-
A cell suspension based uptake method to study high affinity glucosinolate transporters
Plant Methods (2020)