Abstract
Background
Leaf morphology influences plant growth and productivity and is controlled by genetic and environmental cues. The various morphotypes of Brassica rapa provide an excellent resource for genetic and molecular studies of morphological traits.
Objective
This study aimed to identify genes regulating leaf morphology using segregating B. rapa p F2 population.
Methods
Phenotyping and transcriptomic analyses were performed on an F2 population derived from a cross between Rapid cycling B. rapa (RCBr) and B. rapa ssp. penkinensis, inbred line Kenshin. Analyses focused on four target traits: lamina (leaf) length (LL), lamina width (LW), petiole length (PL), and leaf margin (LM).
Results
All four traits were controlled by multiple QTLs, and expression of 466 and 602 genes showed positive and negative correlation with leaf phenotypes, respectively. From this microarray analysis, large numbers of genes were putatively identified as leaf morphology-related genes. The Gene Ontology (GO) category containing the highest number of differentially expressed genes (DEGs) was “phytohormones”. The sets of genes enriched in the four leaf phenotypes did not overlap, indicating that each phenotype was regulated by a different set of genes. The expression of BrAS2, BrAN3, BrCYCB1;2, BrCYCB2;1,4, BrCYCB3;1, CrCYCBD3;2, BrULT1, and BrANT seemed to be related to leaf size traits (LL and LW), whereas BrCUC1, BrCUC2, and BrCUC3 expression for LM trait.
Conclusion
An analysis integrating the results of the current study with previously published data revealed that Kenshin alleles largely determined LL and LW but LM resulted from RCBr alleles. Genes identified in this study could be used to develop molecular markers for use in Brassica breeding projects and for the dissection of gene function.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Leaves are the major biosynthetic organs of plants. As the function of a leaf is associated with its structure, leaf morphology influences plant growth and development (Tsukaya 2005; Fambrini and Pugliesi 2013). Leaf morphology varies between plant species and is under strong genetic and environmental control. The crop species Brassica rapa displays extreme morphological diversity, encompassing leafy vegetables, turnips, and oilseed rape. This diversity results from both genetic and epigenetic variation, and includes important traits for Brassica breeders (Zhao et al. 2005; Bonnema et al. 2011). Understanding morphological variation and development of leaves can help the production of novel leafy crops that attract consumers (Xiao et al. 2014) and provides insight into the adaptation of leaves to diverse environments (Kalve et al. 2014).
The determination of leaf shape and size is primarily controlled by meristem genes (including CLAVATA1, CLAVATA3, WUSCHEL, KNOTTED1, and PHANTASTICA), which are involved in leaf initiation; initiation is also regulated by hormone levels (Tsukaya 2005; Kessler and Sinha 2004). Final leaf size, however, is determined by the complex coordination of cell division and expansion (Kim and Cho 2006; Tsukaya 2006; Barkoulas et al. 2007; Micol 2009; Gonzalez et al. 2010, 2012).
A leaf is composed of the blade and the petiole, which differ from each other in terms of structure and physiology (Tsukaya 2006). The petiole supports the leaf blade and functions in a similar manner to the stem (Tsukaya et al. 2002; Kozuka et al. 2005, 2010). Petiole development is stimulated by shade-avoidance conditions (Tsukaya et al. 2002) and controlled by photoreceptors, such as phytochrome or cryptochrome (Kozuka et al. 2005). In addition, petiole development is regulated by hormones such as auxin, gibberellin, brassinosteroids (BRs), and ethylene (Kim et al. 2005). The proliferative zone, the region of the junction between the leaf blade and leaf petiole, produces both leaf blade and petiole cells and appears to be controlled by ANGUSTIFOLIA3 (AN3) (Ichihashi et al. 2011). Leaf blade development, by contrast, is controlled by two major genes, the auxin transport-related PINFORMED1 (PIN1) and a growth repressor CUP-SHAPED COTYLEDON2 (CUC2) (Hay et al. 2006; Nikovics et al. 2006; Bilsborough et al. 2011), and a plant hormone, cytokinin (Shani et al. 2010). Several additional genes are known to be involved in blade growth: BIG BROTHER (BB), ARP [ASYMMETRIC LEAVES1 (AS1), ROUGH SHEATH2 (RS2), PHANTASTICA], ASYMMETRIC LEAVES 2 (AS2), CINCINNATA-TEOSINTE BRANCHED/CYCLOIDEA/PCF (CIN-TCP), WOX (WUSCHEL-RELATED HOMEOBOX), and JAGGED (JAG) (Bar and Ori 2014). The genes NO APICAL MERISTEM (NAM)/CUP-SHAPED COTYLEDON (CUC), and REDUCED COMPLEXITY (RCO), together with auxin, are involved in the development of the leaf margin (Bar and Ori 2014; Vlad et al. 2014).
Leaf morphology, especially leaf size and shape, is a quantitative trait controlled by many genes. Quantitative trait loci (QTLs) are chromosomal regions that contain a gene or genes affecting a quantitative trait (Geldermann 1975). The most efficient method of analyzing QTLs is to examine transgression in an F2 population derived from intra- or interspecific crosses (Tanksley 1993; Pérez-Pérez et al. 2010). Since the publication of the first linkage map constructed from a cross between Chinese cabbage and broccoli (Song et al. 1995), several groups have identified QTLs in Chinese cabbages and used them to unravel the genetic regulation of leaf architecture. Lou et al. (2007) identified ten leaf trait-related QTLs using an F2/3 population derived from crosses between Chiifu, rapid cycling Brassica rapa-144 (RCBr-144), yellow sarson (oilseed rape), pak choi, and turnip. Li et al. (2009) identified candidate genes for control of the leaf lobe shape using an expressed sequence tagged (EST)-based single nucleotide polymorphism (SNP) marker linkage map and progenies derived from a cross between yellow sarson and B. rapa cv. Osome (heading type) (Li et al. 2009). Lim and colleagues generated large numbers of transgression lines from several crosses and identified several candidate genes that co-localized with QTLs in subsequent populations; these studies involved crosses between Chiifu and Kenshin (two inbred lines of B. rapa ssp. pekinensis) (Choi et al. 2017), Chiifu and Caixin (Li et al. 2015), and Chiifu and RCBr (Li et al. 2013). They found that several QTLs controlling leaf size co-localized with the genes CYCLIN D3;1 (CYCD3;1), CYCLIN B2;4 (CYCB2;4), AN3, ULTRAPETALA1 (ULT1), and AINTEGUMENTA (ANT)(Choi et al. 2017). Xiao et al. (2014) identified a large number of genes regulating leaf development in Brassica species by integrating phenotypic QTLs and transcriptomic data of segregating populations derived from a cross between yellow sarson and pak choi. They established expression QTLs (eQTLs) by microarray analysis followed by qRT-PCR analysis (Xiao et al. 2014). More recently, restriction site-associated DNA sequencing (RAD-seq) of F2 populations generated by a cross between B. rapa ssp. chinensis and pekinensis was used to generate genome-wide SNP-based genetic and QTL maps for Chinese cabbage (Huang et al. 2017).
We used an F2 population derived from a cross between rapid cycling Brassica rapa (RCBr) and Kenshin, an inbred line of B. rapa ssp. pekinensis. RCBr is one of the Wisconsin fast plants (Goldman 1999; Slankster et al. 2012). A heading-type Kenshin was a parental line of a Chiifu and Kenshin double haploid (CKDH) mapping population used to construct a reference genetic map for sequencing the Brassica rapa genome (Vanjildorj et al. 2009). The parental lines and the F2 population were subjected to phenotypic analysis to identify genes controlling leaf morphology. Next, the parental lines and four groups from the F2 population, selected using the traits lamina length (LL), lamina width (LW), leaf margin (LM), and petiole length (PL), were subjected to microarray and quantitative reverse transcriptase polymerase chain reaction (qRT-PCR) analyses. Candidate genes considered likely to control leaf morphology in B. rapa were further investigated by comparisons with previous studies.
Materials and methods
Plant materials
F1 plants derived from a cross between a RCBr inbred line (#24001) and the Brassica rapa ssp. pekinensis inbred line Kenshin were self-hybridized to generate an F2 population. For phenotypic analysis, 15 seeds from each parental line and 200 F2 seeds, which eventually gave more than 180 individuals, were sown on pots (15 cm diameter × 9 cm height) and grown in a greenhouse at Chungnam National University for 6 weeks. Leaf phenotype was analyzed using the fifth leaf. The seventh and ninth leaves were sampled from at least three individuals showing similar phenotypes, and the combined samples were used in microarray and qRT-PCR analyses. Experiments were conducted twice a year, in the spring (April to May) and fall (October to November). Samples were always collected 4 h after dawn to minimize the effect of the diurnal cycle on gene expression.
Leaf morphology
Four leaf morphology traits were analyzed: lamina (leaf) length (LL), lamina width (LW), petiole length (PL), and leaf margin (LM). The first three traits were graded into 1 cm intervals. LM was expressed as six grades (I: very little serration; II: slightly serrated; III: medium serrated; IV: highly serrated; V: slightly lobed; VI: lobed) according to the relative serration ratio and lobe phenotype.
Microarray experiment
Frozen samples (consisting of the seventh and ninth leaves from RCBr, Kenshin, and four F2 groups with similar phenotypes: F2-19, F2-33, F2-93, and F2-100 types) were ground under liquid nitrogen. Total RNA was extracted from each sample with TRIzol reagent (Invitrogen, USA) and cleaned using a NucleoSpin® RNA Clean-up kit (Macherey–Nagel, Germany). The Br135K microarray (Brapa_V3_microarray, 3′-Tiling microarray) was used for transcriptome analysis. This is a high-density DNA array prepared by NimbleGen (http://www.nimblegen.com/) using Maskless Array Synthesizer (MAS) technology, as described in Jung et al. (2014). Probes were designed from 41,173 genes of Brassica rapa accession Chiifu-401-42 (http://brassicadb.org/brad/). Each length of three probes was 60 mers designed by 30 bp overlapped in 120 bp (60 bp coding sequence plus 60 bp 3′UTR of each gene), representing 123,647 features. Fifty features were also deposited from five markers (GUS, GFP, Bar, Kan, Hyg). Labeling, data processing, and background correction were performed as described previously (Jung et al. 2014). The raw data (.pair file) was subjected to RMA (Robust Multi-Array Analysis) (Irizarry et al. 2003), quantile normalization (Bolstad et al. 2003), and background correction as implemented in the NimbleScan software package, version 2.4.27 (Roche NimbleGen, Inc.). To assess the reproducibility of the microarray analysis, the experiment was repeated using independently prepared total RNA from two biological replicates. The microarray data, was deposited in The National Center for Biotechnology Information (NCBI, https://www.ncbi.nlm.nih.gov/) in USA, with the ID of (GSE114479, http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE114479). To understand the putative biological functions and biochemical pathways of differentially expressed genes (DEGs), enrichment analyses were carried out by searching Gene Ontology (GO) (Ashburner et al. 2000), agriGO (Du et al. 2010), and the Kyoto Encyclopedia of Genes and Genomes (Kanehisa et al. 2008).
Quantitative RT-PCR (qRT-PCR)
Cleaned RNA samples were subjected to first-strand cDNA synthesis using the Ace-α kit with Oligo-dT primers (TOYOBO, Japan). Target genes for qRT-PCR were selected following consultation of a previous report by Kalve et al. (2014) and the PhenoLeaf database (http://genetics.umh.es/phenoleaf/) (Wilson-Sánchez et al. 2014). Primers were designed against sequences in the Brassica database (BRAD, http://brassicadb.org/brad/). PCR, using SYBR® Green Realtime PCR Master Mix-Plus (TOYOBO, Japan) as a SYBR green dye, was performed as follows: 30 s at 95 °C, followed by 40 cycles of 95 °C for 5 s, 58 °C for 10 s, and 72 °C for 15 s.
Results and discussion
Analyses of leaf phenotypes
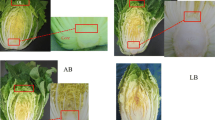
Trait analysis showed that Kenshin leaves were longer and wider, with shorter petioles and more rounded LMs, than leaves of RCBr (Fig. 1a). The F2 population derived from an F1 cross between the RCBr and Kenshin inbred lines displayed various morphological phenotypes (Table 1). The morphological characteristics of the fifth leaf of 6-week-old plants were analyzed (Fig. 1b). For phenotyping, 150 individuals were randomly selected from 180 F2 plants (Fig. 2). The trait LL (length of leaf from apex to leaf base) was measured, and the values were grouped into 1 cm intervals (Fig. 2a). The mean values of LL for RCBr and Kenshin were less than 5 cm and 9–10 cm, respectively (Table 1). The most frequently observed values of LL were between 8 and 9 cm (34 plants; Fig. 2a). Generally, LL appeared to be a quantitative trait, but slightly shifted toward the Kenshin phenotype.
Overall leaf morphology and specific leaf traits in RCBr and Kenshin lines. The fifth leaf from 6-week-old plants was measured. a Representative leaf morphologies of the fifth leaves from the parental lines RCBr and Kenshin. b The four leaf traits used in analyses of leaf phenotypes. LL lamina length (cm) from apex to base of blade, LW lamina width (cm) at the widest region of the lamina, PL petiole length (cm), LM leaf margin, scored from I to VI (I: very little serration; II: slightly serrated; III: medium serrated; IV: highly serrated; V: slightly lobed; VI: lobed)
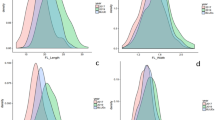
Leaf phenotypes observed in the fifth leaf (left) and distribution of trait values for each character in the F2 population (right). a Observed values of lamina length (LL) were classified into ten groups ranging from < 5 to > 13 cm. b Observed values of lamina width (LW) were classified into seven groups ranging from < 3 to > 8 cm. c Observed values of petiole length (PL) were classified into nine groups ranging from < 2 to > 9 cm. d Observed values for the leaf margin (LM) were classified into six groups, labeled I–VI. N normal distribution. Red arrows indicate the representative phenotype selected for microarray analysis. Horizontal red line indicated the base of leaf. (Color figure online)
The values obtained for LW ranged from less than 3 cm to more than 8 cm (Fig. 2b). The mean values of LW for RCBr and Kenshin were less than 3 cm and 5–6 cm, respectively (Table 1). When grouped into 1 cm intervals, the most frequently observed values of LW in the F2 population were between 5 and 6 cm (47 plants), indicating that LW was a quantitative trait and again shifted toward the Kenshin phenotype (Fig. 2b).
Both RCBr and Kenshin had mean values of PL of approximately 3 cm: 2.6 cm for RCBr and 2.8 cm for Kenshin (Table 1). The values obtained for PL in the F2 population ranged from less than 2 cm to more than 9 cm. When these results were grouped into 1 cm intervals, 51 plants had PL values of between 4 and 5 cm (Fig. 2c). PL appeared to be a quantitative trait, but the PLs of most F2 plants were longer than those of either parental line (Fig. 2c).
The LM trait was scored using a six-point scale ranging from I (very little leaf serration) to VI (leaf lobes present) (Fig. 2d). LMs of RCBr and Kenshin were scored as VI and I, respectively, and the F2 population showed a left-shifted normal distribution (Fig. 2d). The most frequent LM score was II (45 plants), implying that a round margin was semi-dominant.
Most leaf phenotypes were normally distributed, implying regulation by QTLs. The values observed for LL, LW, and LM in the F2 population were slightly shifted toward the Kenshin phenotype, but values of PL exceeded those of the parent phenotypes. This suggested that leaf morphology in F2 individuals was controlled by alleles from Kenshin but PL was additive. The cauliflower Orange (Or) gene enhances petiole elongation by suppressing eukaryotic release factor 1 (Zhou et al. 2011). Expression of its orthologue BrOr (Bra035916) was elevated slightly in RCBr but did not correlate with PL in F2 individuals. ROTUNDIFOLIA3 (ROT3) encodes CYP90C1, which is involved in the late stages of BR biosynthesis and controls leaf petiole growth in the dark (Kim et al. 2005). A BrROT3 (Bra017757) expression was 3.4-fold higher in Kenshin than in RCBr, and no correlation was found between Bra017757 expression and phenotype in F2 individuals. This suggests that regulation of PL is likely to be complex in B. rapa and differ from the mechanism known from Arabidopsis thaliana.
Transcriptome analysis using the Br135K microarray
To identify genes whose expression were correlated with leaf phenotype values, six types of plants with different leaf characteristics were selected: the two parental lines (RCBr and Kenshin inbred lines) and four types of F2 plants represented by F2-93, F2-33, F2-100, and F2-19 types. In the analyses, one representative plant plus two additional plants showing comparable phenotypes were pooled for each type (Fig. 3; Table 1). Although we have used a representative plant plus two additional individuals for experiments, only representative plant names were used to indicate each type. To identify genes involved in the determination of leaf morphology, the seventh and ninth leaves were harvested from representative plants and from two additional plants showing comparable phenotypes, while leaf phenotypes were determined with 5th leaves. The three samples were combined prior to extraction of RNA for use in the microarray experiment and real-time PCR analysis.
Schematic explanation of RCBr, Kenshin, and F2 leaves used in the microarray and qRT-PCR analyses. The trait values used for classification were obtained from measurements made on the fifth leaf of plants from the two parental lines and four F2 plants. The seventh and ninth leaves were harvested from plants showing the phenotypes shown above, and RNA was extracted from these samples. Red and blue text indicates RCBr-like and Kenshin-like traits, respectively. (Color figure online)
Data obtained from the Br135K microarray were expressed as probe intensity (PI) values for expression levels (Table S1) rather than log values to distinguish small differences in expression levels. Of the 41,173 genes present on the Br135K microarray, 5349 genes did not have a counterpart in Arabidopsis (i.e., could not be annotated using TAIR 10). A total of 28,679 genes showed PI values over 500 in at least one sample (Table S1); these levels of expression mean that transcript can be easily detected by 25 cycles of RT-PCR. Genes whose expression correlated with phenotype were selected for further characterization (Figs. 4, 5, 6, 7).
Genes whose level of expression correlated with the extent of the phenotypic trait LL. Candidate genes were selected on the basis of (1) expression PI value > 500; and (2) similarity of patterns between gene expression and LL trait value. a Hierarchical clustering of 96 and 53 genes showing positive and negative correlations between expression levels and phenotype, respectively. b, c K-means clustering and expression patterns. Gray lines: Expression of individual genes. Pink lines: Mean expression of all genes. d Plot of lamina length against genotype. RCBr: 4.7 cm; F2-100: 7.9 cm; Kenshin: 9.3 cm; F2-93: 13.7 cm. (Color figure online)
Genes with expression levels correlated with the value of the phenotypic trait LW. Genes were selected on the basis of (1) expression PI value > 500; and (2) similarity of patterns between gene expression and LW trait value. a Hierarchical clustering of 142 and 82 genes showing positive and negative correlation between expression levels and phenotype, respectively. b, c K-means clustering and expression patterns. Gray lines: Expression of individual genes. Pink lines: Mean expression of all genes. d Plot of lamina width against genotype. RCBr: 2.8 cm; Kenshin: 5 cm; F2-19: 6.2 cm; F2-93: 8.5 cm. (Color figure online)
Genes with expression levels correlated with the value of the phenotypic trait PL. Genes were selected on the basis of (1) expression PI value > 500; and (2) similarity of patterns between gene expression and PL trait value. a Hierarchical clustering of 174 and 381 genes showing positive and negative correlation between expression levels and phenotype, respectively. b, c K-means clustering and expression patterns. Gray lines: Expression of individual genes. Pink lines: Mean expression of all genes. d Plot of petiole length against genotype. RCBr: 2.6 cm; F2-93: 2.7 cm; Kenshin: 2.8 cm; F2-100: 9.8 cm. (Color figure online)
Genes with expression levels correlated with the classification of the phenotypic trait LM. Genes were selected on the basis of (1) expression PI value > 500; and (2) similarity of patterns between gene expression and LM trait value. a Hierarchical clustering of 74 and 86 genes showing positive and negative correlation between expression levels and phenotype, respectively. b, c K-means clustering and expression patterns. Grey lines: Expression of individual genes. Pink lines: Mean expression of all genes. d Plot of leaf margin classification against genotype. Kenshin: I; F2-33 and F2-93: II; F2-19: V; RCBr: VI. (Color figure online)
The order of LL values, ranked from lowest to highest, was RCBr (4.7 cm), F2-100 (7.9 cm), Kenshin (9.3 cm), and F2-93 (13.7 cm) (Fig. 3; Table 1). Transcript levels of 96 genes were positively correlated with LL, and expression of 53 genes was negatively correlated with LL (Fig. 4; Table S2). Expression of most of these genes matched the LL phenotypes well. Several positively or negatively correlated genes were recognizable; for example, BrGH3.5 (Bra019060), BrAS2 (Bra039733), and Bra002068, encoding the auxin efflux carrier family protein, were positively correlated with LL, whereas Bra031986, encoding the tetratricopepetide repeat (TPR)-like superfamily protein, and Bra019610, encoding transducin/WD40 repeat-like superfamily protein, were negatively correlated. ARABIDOPSIS AUXIN-RESPONSIVE GH3 FAMILY 5 (AtGH3.5) increases leaf size and trichome number (Kryvych et al. 2008), Arabidopsis AS2-like/lateral organ boundary (LOB) domain family protein increases leaf size (Li et al. 2016), and Arabidopsis auxin efflux carrier family protein (AT2G17500) regulates auxin homeostasis. Orthologues of all three genes appeared to be associated with leaf size, especially LL, in B. rapa. On the other hand, expression of two negatively correlated genes might inhibit leaf growth. The Arabidopsis orthologue of Bra031986 is important for RNA editing of a mitochondrial respiration gene (Arenas-M et al. 2014), and Bra031986 encodes a protein possibly involved in a COMPASS-like complex, which was initially identified in Caenorhabditis elegans and is involved in the epigenetic regulation of development (Fisher et al. 2010). Moreover, AT3G49660, the Arabidopsis orthologue of Bra031986, interacts with transcription factors to regulate expression of specific genes (Song et al. 2015). Thus, these genes might be involved in determining leaf length in B. rapa.
To identify genes involved in determining LW, four plants were compared: RCBr (LW: 2.8 cm), Kenshin (LW: 5.0 cm), F2-19 (LW: 6.2 cm), and F2-93 (LW: 8.5 cm) (Fig. 3; Table 1). Transcript levels of 142 genes were proportional to LW values, and transcript levels of 82 genes were inversely proportional to LW values (Fig. 5; Table S3). Most genes whose expression positively associated with LW values were stress-responsive genes. Sixteen transcription factors, including BrSPL11 (Bra030040), a SQUAMOSA PROMOTER-BINDING PROTEIN (SBP)-box family-like (SPL) gene, belonged to this category. In Arabidopsis, SPL10, SPL11, and SPL2 affect lamina shape and trichome formation by changing shoot maturation (Shikata et al. 2009). Expression of GIBBERELLIN 20-OXIDASE (BrGA20OX3, Bra009285, Bra0033189) and GA-RESPONSIVE GATS1 HOMOLOGUE 1 (BrGASA1, Bra029227) was negatively correlated with LW. Arabidopsis GA20OX1, -2, and -3 are involved in growth and fertility, and mutations of these genes lead to dwarfism (Plackett et al. 2012).
The same four plants used in the association analysis of PL were used to identify genes involved in determining PL: RCBr (PL: 2.6 cm), F2-93 (PL: 2.7 cm), Kenshin (PL: 2.8 cm), and F2-100 (PL: 9.8 cm) (Fig. 3; Table 1). Transcript levels of 174 genes were positively correlated with PL values, and expression of 381 genes was negatively correlated (Fig. 6; Table S4). Expression of only 25 and 36 genes, however, was exactly positively and negatively proportional, respectively, to the PL phenotypes; other genes showed a clear difference only between RCBr (short PL) and F2-100 (long PL). BrOPT6 (Bra026311), an orthologue of ARABIDOPSIS OLIGOPEPTIDE TRANSPORTER 6 (AtOPT6), was a likely candidate gene for involvement in determining the PL phenotype; AtOPT6 is expressed in vascular tissue and has broad substrate specificity (Pike et al. 2009). The functions of Bra0296512, encoding a Fe-2S ferredoxin-like superfamily protein, Bra013540, encoding an oxidoreductase, zinc-binding dehydrogenase family protein, and Bra040419, encoding a NPL4-like protein 1 (BrNPL4L), whose expression all showed a negative correlation with the PL phenotype, have not yet been identified.
To uncover genes involved in determining the LM phenotype, five plants were selected: Kenshin (class I), F2-33 and F2-93 (both class II), F2-19 (class V), and RCBr (class VI) (Fig. 3; Table 1). The numbers of genes whose transcript levels were proportional and inversely proportional to the LM classification were 74 and 86, respectively (Fig. 7; Table S5). The most dramatic differences in expression levels were found between the parental lines, Kenshin, which showed no serration, and RCBr, whose leaves showed a high level of serration and lobing (Table S5). Expression of three genes showed a strong, positive correlation with the LM phenotype, Bra016851, encoding LYSINE KETOGLUTARATE REDUCTASE TRANS-SPLICING-LIKE PROTEIN, Bra007991, encoding SUCROSE-PROTON SYMPORTER 1 (BrSUC1), and Bra007197, encoding RECEPTOR-LIKE PROTEIN 47 (BrRLP47); the functions of these genes have not yet been studied. The expression of ten genes (Bra012716, Bra012818, Bra020690, Bra008167, Bra020091, Bra033287, Bra029776, Bra001726, Bra037638, and Bra000189) showed a strong negative correlation with the LM phenotype. Moreover, expression of LOB DOMAIN-CONTAINING PROTEIN 37/ASYMMETRIC LEAVES2-LIKE (ASL) (BrLBD37, Bra012164), which is involved in leaf morphogenesis (Albinsky et al. 2010), was negatively correlated with the degree of the LM phenotype across all the genotypes, with the exception of F2-33. BrLBD37 is thus a good candidate for a gene determining the LM phenotype.
These microarray data are summarized in Table 2. Individuals resembling F2-93 showed superior leaf phenotypes with respect to the traits LL and LW, indicating that they might be useful for generating molecular markers for breeding leafy vegetables. A previous study (Xiao et al. 2014) identified novel genes likely to be involved in leaf development, in addition to several known genes. That study used Agilent 105K Brassica species oligoarrays, which are not specific for B. rapa and do not contain a suitable number of B. rapa genes. Our microarray analysis used a chip specifically designed for B. rapa as well as 3′-specificity and revealed many genes whose expression correlated strongly with four selected leaf phenotypes. This has provided a large number of candidate genes likely to be important in determining leaf morphology.
Analysis of selected genes by functional category
The DEGs associated with each leaf phenotype were annotated using GO in Biological Process in TAIR (The Arabidopsis Information Resource; http://arabidopsis.org/) (Table 3). DEGs were declared if expression levels in PI values between the lowest- and highest-ranked samples in Table 2 differed by more than 1.5-fold. A large number of genes were classified as unknown in all phenotypes. After this, “phytohormone” genes were found to be the most abundant (9.3%) GO category. The second to sixth categories by gene number were “developmental process”, “enzyme activity”, “transport”, “cell division and elongation, differentiation”, and “cell wall and cytoskeleton”. The finding that phytohormone-associated genes formed the largest category of DEGs across all leaf phenotypes agreed with previous studies showing that phytohormones are involved in leaf development (Kim et al. 2006; Barkoulas et al. 2007; Cha et al. 2007; Micol 2009; Gonzalez et al. 2010; Kalve et al. 2014; Xiao et al. 2014). Other enriched genes also appear to play important roles in leaf growth and differentiation, as they are involved in developmental processes, cell division and elongation, differentiation, and cell wall and cytoskeleton (Cosgrove 2005; Wolf et al. 2012; Kalve et al. 2014).
Using our data and the findings of a previous report (Kalve et al. 2014), the genes involved in four biological process categories (“phytohormone”, “cell division and elongation, differentiation”, “developmental process”, and “cell wall and cytoskeleton”) were further characterized with respect to the four leaf traits (Tables S6–S9). These analyses were supplemented with information from the PhenoLeaf database (http://genetics.umh.es/phenoleaf/) and the phenotypes of Salk mutant lines (indicated by underlining and an asterisk in the Supplementary Tables).
First, DEGs associated with the phenotypic trait LL were considered (Table S6). Some important up-regulated genes shared a high level of sequence identity with IAA-AMIDO SYNTHASE (WES1), PIN-LIKE 5 (PILS5), and CYP74A, while the down-regulated genes included homologues of ABI FIVE-BINDING PROTEIN 2 (AFP2) and STRICTOSIDINE SYNTHASE 2 (SS2), which are associated with phytohormone signaling and biosynthesis. Other up-regulated genes associated with LL were involved in cell division, elongation, and differentiation; these included ARABIDOPSIS THALIANA HOMEOBOX 1 (ATHB-1), KIP-RELATED PROTEIN 5 (KRP5), and CHROMATIN REMODELING FACTOR 1 (CHC1/SWP73B). HB-1 and AS2, two genes involved in developmental processes, were also up-regulated; however, another gene in this category, TOPLESS/WUS-INTERACTING PROTEIN 1 (TPL/WSIP1), was down-regulated. Only one gene in the “cell wall and cytoskeleton” category, XYLOSYL TRANSFERASE 1 (XT1), was associated with LL; this remains to be studied.
In the analysis of DEGs associated with the LW phenotype (Table S7), four genes related to well-known phytohormone-associated genes [AFP1, GIBBERELLIN 2-XODASE 4 (GA2OX4), ETHYLENE RESPONSE ELEMENT-BINDING FACTOR 5 (ERF5), and RING DOMAIN LIGASE 1 (RGLG1)] were up-regulated. Three genes from the same category were down-regulated [two GA2OX3 genes and CYTOKININ OXIDASE 5 (CKX6)]. Additional well-characterized genes from other processes were identified as DEGs in the LW analysis. Genes associated with “cell division and elongation, differentiation” included 3x HIGH MOBILITY GROUP-BOX 2 (3xHMG-BOX2), which was up-regulated, and GA-RESPONSIVE GAST1 HOMOLOGUE1 (GASA1), which was down-regulated. Other DEGs associated with the LW phenotypic trait were from the “developmental process” category. These included five up-regulated genes [HB-1, UBIQUITIN-SPECIFIC PROTEASE 15 (UBP15), CYTOCHROME C LYASE (CCMH/CycI), and ABL INTERACTOR-LIKE PROTEIN 3 (ABIL3)], and three down-regulated genes [TCP family transcription factor 4, EMBRYONIC DEFECTIVE 1075 (EMB1075), and EMBRYONIC DEFECTIVE 24 (EMB24)]. Genes from the “cell wall and cytoskeleton” category were also differentially expressed; XYLOGLUCAN ENDOTRANSGLUCOSYLASE/HYDROLASE 22 (XTH22) and XYG XYLOSYLTRANSFERASE 1 (XT1) were up-regulated, and REVERSIBLY GLYCOSYLATED POLYPEPTIDE 1 (RGP1) and IRREGULAR XYLEM 10 (IRX10) were down-regulated.
The PL phenotype was associated with the largest number of DEGs. The full list is given in Table S8, but a few genes are described here. In the “phytohormone” category, SLOW MOTION (SLOMO) and RUB1 CONJUGATING ENZYME 1 (RCE1) were up-regulated, whereas PIL6, NUCLEAR FACTOR Y SUBUNIT 4 (NF-YA4), SMALL AUXIN UPREGULATED RNA32 (SAUR32), ABERRANT LATERAL ROOT FORMATION 1 (ALF1), BR-INSENSITIVE 1 (BRI1), BRZ-INSENSITIVE-LONG HYPOCOTYLS 4 (BIL4), HISTIDINE-CONTAINING PHOSPHOTRANSMITTER 1 (HP1), and GALACTURONOSYLTRANSFERASE-LIKE 2 (GATL2) were all down-regulated. Several genes from the “developmental process” category were associated with PL, up-regulated genes included PIN1, ACAULIS 5 (ACL5), DIACYLGLYCEROL KINASE 7 (DGK7), DsRNA-BINDING PROTEIN 1 (DRB1), CONSTITUTIVE MORPHOGENESIS 2 (COP2), and PIGMENT DEFECTIVE 191 (PDF191), whereas down-regulated genes included BELL-LIKE HOMEODOMAIN 9 (BLH9), ENDOPLASMIC RETICULUM-ADENINE NUCLEOTIDE TRANSPORTER 1 (ER-ANT1), ARABIDOPSIS 6B-INTERACTING PROTEIN 1-LIKE 2 (ASIL2), and LON PROTEASE 1 (LON1). All the genes associated with PL in the category of “cell division and elongation, differentiation” were down-regulated. These included RETARDED ROOT GROWTH (RRG), MEMBRANE-RELATED BIGGER1 (MRB1), AS1/2 ENHANCER 7 (AE7), E2F TRANSCRIPTION FACTOR 1 (E2F1/E2FB), COP9 SIGNALOSOME SUBUNIT 5 = 6B (CSN6B), RAN GTPASE ACTIVATING PROTEIN 2 (RANGAP2), LONG AFTER FAR-RED 3 (LAF3), LEUCINE-RICH REPEAT/EXTENSION 1 (LRX1), and ARPC4. In the “cell wall and cytoskeleton” category, REF4-RELATED 1 (RFR1) was up-regulated while CELLULOSE SYNTHASE-LIKE D3 (CSLD3), GLUCURONIC ACID SUBSTITUTION OF XYLAN 3 (GUX3), CINNAMYL ALCOHOL DEHYDROGENASE 5 (CAD5), and an unknown gene (possibly a Golgin family A protein) were down-regulated. Although many DEGs were associated with PL, no key transcription factors or core genes were identified.
DEGs associated with the LM trait also included large numbers of genes associated with the GO categories “phytohormone”, “cell division and differentiation”, “developmental process”, and “cell wall and cytoskeleton” (Table S9). Some of these were examined with respect to their function. In the “phytohormone” category, CHLOROPHYLLASE 1 (CLH1) was up-regulated and G-PROTEIN GAMMA SUBUNITS 2 (GG2) was down-regulated. Up-regulated genes from the “developmental process” category included CINNAMATE 4-HYDROXYLASE (C4H) and FLOWERING LOCUS T (FT); down-regulated genes from the same category included GATA TRANSCRIPTION FACTOR 21 (GATA21), ASYMMETRIC LEAVES 2-LIKE 39 (ASL39), and RESPONSE REGULATOR 1 (RR1). Only one gene from the “cell wall and cytoskeleton” category, MYOSIN 1, was down-regulated.
The analysis of four leaf phenotypes focused on four biological process categories has identified 33 up-regulated and 42 down-regulated genes (Kalve et al. 2014 and websites). These genes were also identified in the present study, demonstrating that we had used an appropriate sampling regime (parents and F2 population) and methodology (Br135K microarray). The genes enriched in each leaf morphology phenotype rarely overlapped, suggesting that each phenotype considered in the present study was regulated by a different set of genes.
Confirmation of microarray data with qRT-PCR analysis
To confirm the associations between identified genes and leaf morphology, transcript levels of 77 genes, selected from our microarray experiments, previous reports, and the PhenoLeaf database, were examined using qRT-PCR with gene-specific primers designed against sequences in BRAD (http://brassicadb.org/) (Table S10). A ribosomal protein gene from B. rapa, BrRPL22, was used as a control. Approximately half of the tested genes, including those presented in Fig. 8, showed the same or similar patterns of expression in both the microarray and qRT-PCR analyses.
qRT-PCR analysis to confirm the expression levels of selected genes. Genes positively correlated with a phenotypic trait are shown in the left-hand panel, and genes negatively correlated with a trait are shown in the right-hand panel. Expression levels of each gene were normalized against BrRPL22 expression. Blue bars: Mean RNA expression levels measured using qRT-PCR. Orange line: Probe intensity (PI) values obtained from Br135K microarray analysis. Error bars: Standard error. K: Kenshin inbred line. (Color figure online)
Expression levels of BrAS2, TERPENE SYNTHASE 10 (BrTPS10), and XYLOTRANSFERASE1 (BrXT1) were positively correlated with LL, while BrSS2 and BrTPL expression levels were negatively correlated with LL (Fig. 8, top panel). AS2 is known to be involved in leaf morphology and venation (Rédei and Hirono 1964; Semiarti et al. 2001; Iwakawa et al. 2007). The function of TPS10 is not yet known. XT1 is essential for cell wall biosynthesis (Park and Cosgrove 2012). An xxt1/xxt2 double mutant shows a reduction in cell wall xyloglucan (Cavalier et al. 2008; Park and Cosgrove 2012) and a reduction in rosette leaves and petiole size (Park and Cosgrove 2012). The expression of SS2 is controlled by the phytohormones auxin and jasmonic acid (Menke et al. 1999). TPL is involved in embryo formation, and in cotyledon and apex development (Szemenyei et al. 2008). Results obtained from Arabidopsis suggest that these genes are also likely to perform similar functions in leaf morphogenesis in B. rapa.
Expression of four genes (BrAFP1, BrXTH22, BrERF5, and BrRGLG1) was positively correlated with LW, whereas expression of two genes (BrGA20OX3 and BrIRX10) was negatively correlated (Fig. 8, second panel from top). XTH22 modifies cell wall structure (Cosgrove 2005; Sasidharan et al. 2014). Although ERF5 inhibits leaf growth (Dubois et al. 2013), it showed elevated expression as LW values increased. Mutations in RGLG1 change leaf morphology (Yin et al. 2007). GA20OX3 reduces gibberellin levels (Thomas et al. 1999), and overexpression reduces rice growth (Lo et al. 2008). IRX10, a member of the glycosyltransferase family, is involved in xylan biosynthesis (Brown et al. 2009). These observations all suggest that these genes are likely to affect LW.
The qRT-PCR analysis of genes associated with PL found similar patterns of up-regulation of BrPIN1 and down-regulation of BrE2FB and BrLAF3 to those seen in the microarray data (Fig. 8, second panel from bottom). The auxin transporter PIN1 is associated with leaf morphology and pattern formation of vascular tissues (Scarpella et al. 2006). E2FB (or E2F1) is the key target of auxin and regulates cell division for cell growth and differentiation (Magyar et al. 2005). LAF3 functions in phytochrome A signaling and affects photomorphogenesis (Hare et al. 2003). The expression patterns of these genes imply that determination of PL involves a complex mechanism. In Arabidopsis, PHYTOCHROME B (PhyB), ROT3, ACAULIS 2 (ACL2), and GA-INSENSITIVE (GAI) are known to control PL (Tsukaya et al. 2002), but regulation of petiole development in B. rapa may differ from that in Arabidopsis.
Expression of ISOPENTENYLTRANSFERASE 5 (BrIPT5), BrCYP73A5, and CHLOROPHYLLASE 1 (BrCLH1), genes associated with the LM phenotype, increased in tandem with leaf serration, while ASYMMETRIC LEAVES2-LIKE 39 (BrASL39) and EXPANSIN 8A (BrEXPA8) showed the opposite pattern (Fig. 8, bottom panel). IPT5 controls cytokinin biosynthesis (Miyawaki et al. 2006), and CYP73A5 is associated with lignin biosynthesis (Sundin et al. 2014). ASL39 affects leaf morphology, nitrogen transport, and anthocyanin biosynthesis (Rubin et al. 2009). EXPANSIN 8A (EXP8A) is associated with cell wall elongation (Esmon et al. 2005). These results suggest that these genes are involved in determining leaf shape.
Integrative analysis of leaf morphology-related genes
To narrow down the numbers of genes involved in leaf morphology in B. rapa, we compared our results with previously published data (Table 4). This table, which uses the same classification as Xiao et al. (2014), includes 24 genes up-regulated in Kenshin and 14 genes up-regulated in RCBr in our microarray data. The trends in gene expression were consistent with the observation that LL and LW in the F2 population were slightly shifted toward the values of the Kenshin parental line (Fig. 2a, b). Genes associated with serration and lobe phenotypes were more likely to be up-regulated in RCBr. Expression of many of the listed genes did not differ between Kenshin and RCBr; however, ten genes mentioned by Choi et al. (2017) were up-regulated in Kenshin: ARL, CLF, CycB2.3, CycB2.4, CycD3.1, ROT3, AN3, COW1, ULT1, and ANT. Of the other genes examined, expression of AS2 was strongly correlated with LL (Fig. 8) and up-regulated in Kenshin (Table 4). AS2 and AS1 regulate leaf development by repressing BREVIPEDICELLUS (BP/KNAT1) and KNAT2, members of the class I KNOTTED1-like homeobox (KNOX) family (Li et al. 2016). KNOX genes maintain shoot apical meristem activity but repress leaf development by inhibiting leaf initiation (Hay and Tsiantis 2010; Sluis 2015). The final leaf size depends on primodium size, the cell division or expansion rate, the duration of cell division or expansion, and meristemoid division (Bar and Ori wo14; Kalve et al. 2014). In particular, two overlapping phases of leaf development, cell proliferation and cell expansion, are closely related to leaf size (Golzalez et al. 2012). The increased expression of BrAS2 (Bra039733) associated with Kenshin may reflect the fact that AS2 stimulates leaf initiation, which is thereby increased in LL. A high level expression of BrAS2 gene may be responsible for the increased leaf size observed in the F2 population, as they increase leaf size in Arabidopsis (Bar and Ori 2014).
BrCUC2 was highly expressed in RCBr (Table 4; Table S1), an inbred line that has a serrated and lobed LM. BrCUC1 (Bra021592, Bra001586), BrCUC2 (Bra003023, Bra022685), and BrCUC3 (Bra008259, Bra015750) were much more highly expressed in RCBr than in Kenshin, with increases in expression of between 3- and eightfold. Increases in the expression levels of these genes may lead to more serration and lobed margins in F2 populations, as described in Arabidopsis (Nikovics et al. 2006; Bar and Oir 2014; Vlad et al. 2014). Otherwise, it is possible that other regulatory factors are responsible for CUC2 expression variation in B. rapa.
It is worth noting that the cyclin genes, AN3, ULT1, and ANT, were reported to overlap with leaf size QTLs by Choi et al. (2017) and that these genes were more highly expressed in Kenshin than in RCBr. In contrast to LM, values of LL and LW in the F2 population resembled those of Kenshin (Fig. 2; Table 4); this observation is supported by the expression levels of these genes (Table S1). Kenshin plants showed high expression levels of BrAN3 (Bra010002, Bra020616, Bra036131), increased by 1.8- to 2.2-fold; of BrCYCB1;2 (Bra005880, Bra028741), increased by 9- to 12-fold; of BrCYCB2;1 (Bra024504), increased by eightfold; of BrCYCB2;4 (Bra003727, Bra015762), increased by 9- to 28-fold; of BrCYCD3;1 (Bra034612, Bra017629, Bra011501), increased by 10- to 15-fold; of BrCYCD3;2 (Bra012146), increased by 15-fold; of BrULT1 (Bra0024219, Bra026276), increased by Four- to fivefold; and of BrANT 9 (Bra011782, Bra017851), increased by 2- to 2.5-fold. All these data suggest that these genes are essential for controlling leaf size in B. rapa, especially increases in leaf size.
Conclusion
To identify genes associated with leaf morphology in B. rapa, we generated an F2 population from a cross between the inbred lines RCBr and Kenshin. We analyzed four leaf phenotypes, examined gene expression associated with phenotypic variation using the Br135K microarray, and finally confirmed these analyses by qRT-PCR. In addition, we determined the importance of the genes identified in this study by comparing them with genes known from QTL and eQTL mapping. All four phenotypes were quantitative traits in the F2 population. Gene expression analysis usig the Br135K microarray identified 466 genes positively associated with the leaf phenotypes and 602 genes negatively correlated with the leaf phenotypes. Many of these DEGs had been previously identified as involved in leaf development. The GO categories containing the highest numbers of DEGs were, from highest to lowest, “phytohormone”, “developmental process”, “enzyme activity”, “transport”, “cell division and elongation, differentiation”, and “cell wall and cytoskeleton”. A qRT-PCR analysis further suggested that many selected DEGs had putative functions related to leaf morphology. An integrative analysis combining our results with previously published data revealed that the values of LL and LW in our RCBr/Kenshin-derived F2 population were largely determined by alleles derived from Kenshin, but values for LM were determined by RCBr alleles. Genes previously identified as important determinants of leaf phenotypes were more highly expressed in Kenshin than in RCBr plants, suggesting that alleles from Kenshin determine large leaf size. Further research to dissect the roles of the candidate genes identified in this study in leaf development is necessary for the development of molecular markers associated with leaf phenotype that can be used in molecular approaches to plant breeding.
References
Albinsky D, Kusano M, Higuchi M, Hayashi N, Kobayashi M, Fukushima A, Mori M, Ichikawa T, Matsui K, Kuroda H, Horii Y, Tsumoto Y, Sakakibara H, Hirochika H, Matsui M, Saito K (2010) Metabolomic screening applied to rice FOX Arabidopsis lines leads to the identification of a gene-changing nitrogen metabolism. Mol Plant 3:125–142
Arenas-M A, Zehrmann A, Moreno S, Takenaka M, Jordana X (2014) The pentatricopeptide repeat protein MEF26 participates in RNA editing in mitochondrial cox3 and nad4 transcripts. Mitochondrion 19(Pt B):126–134
Ashburner M. Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, Matese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock G (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25:25–29
Bar M, Ori N (2014) Leaf development and morphogenesis. Development 141:4219–4230
Barkoulas M, Galinha C, Grigg SP, Tsiantis M (2007) From genes to shape: regulatory interactions in leaf development. Curr Opin Plant Biol 10:660–666
Bilsborough GD, Runions A, Barkoulas M, Jenkins HW, Hasson A, Galinha C, Laufs P, Hay A, Prusinkiewicz P, Tsiantis M (2011) Model for the regulation of Arabidopsis thaliana leaf margin development. Proc Natl Acad Sci USA 108:3424–3429
Bolstad BM, Irizarry RA, Ǻstrand M, Speed TP (2003) A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19:185–193
Bonnema G, Carpio DPD, Zhao JJ (2011) Diversity analysis and molecular taxonomy of Brassica vegetable crops. In: Kole C, Sadowski J (eds) Genetics, genomics and breeding of crop plants. Science Publishers, pp 81–124
Brown DM, Zhang Z, Stephens E, Dupree P, Turner SR (2009) Characterization of IRX10 and IRX10-like reveals an essential role in glucuronoxylan biosynthesis in Arabidopsis. Plant J 57:732–746
Cavalier DM, Lerouxel O, Neumetzler L, Yamauchi K, Reinecke A, Freshour G, Zabotina OA, Hahn MG, Burgert I, Pauly M, Raikhel NV, Keegstra K (2008) Disrupting two Arabidopsis thaliana xylosyltransferase genes results in plants deficient in xyloglucan, a major primary cell wall component. Plant Cell 20:1519–1537
Cha YK, Liu L, Kim JY (2007) Signaling network for stem cell maintenance and functioning in Arabidopsis shoot apical meristem. J Plant Biol 50:274–281
Choi SR, Yu X, Dhandapani V, Li X, Wang Z, Lee SY, Oh SH, Pang W, Ramchiary N, Hong CP, Park S, Piao Z, Kim H, Lim YP (2017) Integrated analysis of leaf morphological and color traits in different populations of Chinese cabbage (Brassica rapa ssp. pekinensis). Theor Appl Genet 130:1617–1634
Cosgrove DJ (2005) Growth of the plant cell wall. Nat Rev Mol Cell Biol 6:850–861
Du Z, Zhou X, Ling Y, Zhang Z, Su Z (2010) agriGO: A GO analysis toolkit for the agricultural community. Nucleic Acids Res 38:W64–W70
Dubois M, Skirycz A, Claeys H, Maleux K, Dhondt S, Bodt SD, Bossche RV, Milde LD, Yoshizumi T, Matsui M, Inzé D (2013) ETHYLENE RESPONSE FACTOR6 acts as a central regulator of leaf growth under water-limiting conditions in Arabidopsis. Plant Physiol 162:319–332
Esmon CA, Tinsley AG, Ljung K, Sandberg G, Hearne LB, Liscum E (2005) A gradient of auxin and auxin-dependent transcription precedes tropic growth responses. Proc Natl Acad Sci USA 103:236–241
Fambrini M, Pugliesi C (2013) Usual and unusual development of the dicot leaf: involvement of transcription factors and hormones. Plant Cell Rep 32:899–922
Fisher K, Southall SM, Wilson JR, Poulin GB (2010) Methylation and demethylation activities of a C. elegans MLL-like complex attenuate RAS signalling. Dev Biol 341:142–153
Geldermann H (1975) Investigations on inheritance of quantitative characters in animals by gene markers. Theor Appl Genet 46:319–330
Goldman IL (1999) Teaching recurrent selection in the classroom with Wisconsin fast plants. Hort Technol 9:579–584
Gonzalez N, De Bodt S, Sulpice R, Jikumaru Y, Chae E, Dhondt S, Van Daele T, De Milde L, Weigel D, Kamiya Y, Stitt M, Beemster GT, Inzé D (2010) Increased leaf size: Different means to an end. Plant Physiol 153:1261–1279
Gonzalez N, Vanhaeren H, Inzé D (2012) Leaf size control: complex coordination of cell division and expansion. Trends Plant Sci 17:332–340
Hare PD, Moller SG, Huang LF, Chua NH (2003) LAF3, a novel factor required for normal phytochrome A signaling. Plant Physiol 133:1592–1604
Hay A, Tsiantis M (2010) KNOX genes: Versatile regulators of plant development and diversity. Development 137:3153–3165
Hay A, Barkoulas M, Tsiantis M (2006) ASYMMETRIC LEAVES1 and auxin activities converge to repress BREVIPEDICELLUS expression and promote leaf development in Arabidopsis. Development 133:3955–3961
Huang L, Yang Y, Zhang F, Cao J (2017) A genome-wide SNP-based genetic map and QTL mapping for agronomic traits in Chinese cabbage. Sci Rep 7:46305
Ichihashi Y, Kawade K, Usami T, Horiguchi G, Takahashi T, Tsukaya H (2011) Key proliferative activity in the junction between the leaf blade and leaf petiole of Arabidopsis. Plant Physiol 157:1151–1162
Irizarry TA, Bolstad BM, Collin F, Cope RM, Hobbs B, Speed TP (2003) Summaries of Affymetrix GeneChip prove level data. Nucleic Acids Res 31:e15
Iwakawa H, Iwasaki M, Kojima S, Ueno Y, Soma T, Tanaka H, Semiarti E, Machida Y, Machida C (2007) Expression of the ASYMMETRIC LEAVES2 gene in the adaxial domain of Arabidopsis leaves represses cell proliferation in this domain and is critical for the development of properly expanded leaves. Plant J 51:173–184
Jung HJ, Dong X, Park JI, Thamilarasan SK, Lee SS, Kim YK, Lim YP, Nou IS, Hur Y (2014) Genome-wide transcriptome analysis of two contrasting Brassica rapa doubled haploid lines under cold-stresses using Br135K oligomeric chip. PLoS One 9:e106069
Kalve S, De Vos D, Beemster GT (2014) Leaf development: a cellular perspective. Front Plant Sci 5:1–25
Kanehisa M, Araki M, Goto S, Hattori M, Hirakawa M, Itoh M, Katayama T, Kawashima S, Okuda S, Tokimatsu T, Yamanishi Y (2008) KEGG for linking genomes to life and the environment. Nucleic Acids Res 36:D480–D484
Kessler S, Sinha N (2004) Shaping up: the genetic control of leaf shape. Curr Opin Plant Biol 7:65–72
Kim GT, Cho KH (2006) Recent advances in the genetic regulation of the shape of simple leaves. Physiol Plant 126:494–502
Kim GT, Fujioka S, Kozuka T, Tax FE, Takatsuto S, Yoshida S, Tsukaya H (2005) CYP90C1 and CYP90D1 are involved in different steps in the brassinosteroid biosynthesis pathway in Arabidopsis thaliana. Plant J 41:710–721
Kozuka T, Horiguchi G, Kim GT, Ohgishi M, Sakai T, Tsukaya H (2005) The different growth responses of the Arabidopsis thaliana leaf blade and the petiole during shade avoidance are regulated by photoreceptors and sugar. Plant Cell Physiol 46:213–223
Kozuka T, Kobayashi J, Horiguchi G, Demura T, Sakakibara H, Tsukaya H, Nagatani A (2010) Involvement of auxin and brassinosteroid in the regulation of petiole elongation under the shade. Plant Physiol 153:1608–1618
Kryvych S, Nikiforova V, Herzog M, Perazza D, Fisahn J (2008) Gene expression profiling of the different stages of Arabidopsis thaliana trichome development on the single cell level. Plant Physiol Biochem 46:160–173
Li F, Kitashiba H, Inaba K, Nishio T (2009) A Brassica rapa linkage map of EST-based SNP markers for identification of candidate genes controlling flowering time and leaf morphological traits. DNA Res 16:311–323
Li X, Ramchiary N, Dhandapani V, Choi SR, Hur Y, Nou IS, Yoon MK, Lim YP (2013) Quantitative trait loci mapping in Brassica rapa revealed the structural and functional conservation of genetic loci governing morphological and yield component traits in the A, B, and C subgenomes of Brassica species. DNA Res 20:1–16
Li X, Wang W, Wang Z, Li K, Lim YP, Piao Z (2015) Construction of chromosome segment substitution lines enables QTL mapping for flowering and morphological traits in Brassica rapa. Front Plant Sci 6:432
Li Z, Li B, Liu J, Guo Z, Liu Y, Li Y, Shen WH, Huang Y, Huang H, Zhang Y, Dong A (2016) Transcription factors AS1 and AS2 interact with LHP1 to repress KNOX genes in Arabidopsis. J Integr Plant Biol 58:959–970
Lo SF, Yang SY, Chen KT, Hsing YI, Zeevaart JA, Chen LJ, Yub SM (2008) A Novel class of gibberellin 2-Oxidases control semidwarfism, tillering, and root development in rice. Plant Cell 20:2603–2618
Lou P, Zhao J, Kim JS, Shen S, Del Carpio DP, Song X, Jin M, Vreugdenhil D, Wang X, Koornneef M, Bonnema G (2007) Quantitative trait loci for flowering time and morphological traits in multiple populations of Brassica rapa. J Exp Bot 58:4005–4016
Magyar Z, De Veylder L, Atanassova A, Bakó L, Inzé D, Bögre L (2005) The role of the Arabidopsis E2FB transcription factor in regulating auxin-dependent cell division. Plant Cell 17:2527–2541
Menke FL, Champion A, Kijne JW, Memelink J (1999) A novel jasmonate- and elicitor-responsive element in the periwinkle secondary metabolite biosynthetic gene Str interacts with a jasmonate- and elicitor-inducible AP2-domain transcription. EMBO J 18:4455–4463
Micol JL (2009) Leaf development: time to turn over a new leaf? Curr Opin Plant Biol 12:9–16
Miyawaki K, Tarkowski P, Matsumoto-Kitano M, Kato T, Sato S, Tarkowska D, Tabata S, Sandberg G, Kakimoto T (2006) Roles of Arabidopsis ATP/ADP isopentenyltransferases and tRNA isopentenyltransferases in cytokinin biosynthesis. Proc Natl Acad Sci USA 103:16598–16603
Nikovics K, Blein T, Peaucelle A, Ishida T, Morin H, Aida M, Laufs P (2006) The balance between the MIR164A and CUC2 genes controls leaf margin serration in Arabidopsis. Plant Cell 18:2929–2945
Park YB, Cosgrove DJ (2012) Changes in cell wall biomechanical properties in the xyloglucan-deficient xxt1/xxt2 mutant of Arabidopsis. Plant Physiol 158:465–475
Pérez-Pérez JM, Esteve-Bruna D, Micol JL (2010) QTL analysis of leaf architecture. J Plant Res 123:15–23
Pike S, Patel A, Stacey G, Gassmann W (2009) Arabidopsis OPT6 is an oligopeptide transporter with exceptionally broad substrate specificity. Plant Cell Physiol 50:1923–1932
Plackett AR, Powers SJ, Fernandez-Garcia N, Urbanova T, Takebayashi Y, Seo M, Jikumaru Y, Benlloch R, Nilsson O, Ruiz-Rivero O, Phillips AL, Wilson ZA, Thomas SG, Hedden P (2012) Analysis of the developmental roles of the Arabidopsis gibberellin 20-oxidases demonstrates that GA20ox1, -2, and -3 are the dominant paralogs. Plant Cell 24:941–960
Rédei GP, Hirono Y (1964) Linkage studies. Arabidopsis Inf Serv 1:9–10
Rubin G, Tohge T, Matsuda F, Saito K, Scheiblea WR (2009) Members of the LBD Family of transcription factors repress anthocyanin synthesis and affect additional nitrogen responses in Arabidopsis. Plant Cell 21:3567–3584
Sasidharan R, Keuskamp DH, Kooke R, Voesenek LA, Pierik R (2014) Interactions between auxin, microtubules and XTHs mediate green shade-induced petiole elongation in Arabidopsis. PLoS One 9:e90587
Scarpella E, Marcos D, Friml J, Berleth T (2006) Control of leaf vascular patterning by polar auxin transport. Genes Dev 20:1015–1027
Semiarti E, Ueno Y, Tsukaya H, Iwakawa H, Machida C, Machida Y (2001) The ASYMMETRIC LEAVES2 gene of Arabidopsis thaliana regulates formation of a symmetric lamina, establishment of venation and repression of meristem-related homeobox genes in leaves. Development 128:1771–1783
Shani E, Ben-Gera H, Shleizer-Burko S, Burko Y, Weiss D, Ori N (2010) Cytokinin regulates compound leaf development in tomato. Plant Cell 22:3206–3217
Shikata M, Koyama T, Mitsuda N, Ohme-Takagi M (2009) Arabidopsis SBP-box genes SPL10, SPL11 and SPL2 control morphological change in association with shoot maturation in the reproductive phase. Plant Cell Physiol 50:2133–2145
Slankster EE, Chase JM, Jones LA, Wendell DL (2012) DNA-based genetic markers for rapid cycling Brassica rapa (Fast Plants Type) designed for the teaching laboratory. Front Plant Sci 3:118
Sluis A, Hake S (2015) Organogenesis in plants: initiation and elaboration of leaves. Trends Genet 31:300–306
Song K, Slocum MK, Osborn TC (1995) Molecular marker analysis of genes controlling morphological variation in Brassica rapa (syn. campestris). Theor Appl Genet 90:1–10
Song ZT, Sun L, Lu SJ, Tian Y, Ding Y, Liu JX (2015) Transcription factor interaction with COMPASS-like complex regulates histone H3K4 trimethylation for specific gene expression in plants. Proc Natl Acad Sci USA 112:2900–2905
Sundin L, Vanholme R, Geerinck J, Goeminne G, Höfer R, Kim H, Ralph J, Boerjan W (2014) Mutation of the inducible ARABIDOPSIS THALIANA CYTOCHROME P450 REDUCTASE 2 alters lignin composition and improves saccharification. Plant Physiol 166:1956–1971
Szemenyei H, Hannon M, Long JA (2008) TOPLESS mediates auxin-dependent transcriptional repression during Arabidopsis embryogenesis. Science 319:1384–1386
Tanksley SD (1993) Mapping polygenes. Ann Rev Genet 27:205–233
Thomas SG, Phillips AL, Hedden P (1999) Molecular cloning and functional expression of gibberellin 2-oxidases, multifunctional enzymes involved in gibberellin deactivation. Proc Natl Acad Sci USA 96:4698–4703
Tsukaya H (2005) Leaf shape: genetic controls and environmental factors. Int J Dev Biol 49:547–555
Tsukaya H (2006) Mechanism of leaf-shape determination. Ann Rev Plant Biol 57:477–496
Tsukaya H, Kozuka T, Kim GT (2002) Genetic control of petiole length in Arabidopsis thaliana. Plant Cell Physiol 43:1221–1228
Ueno Y, Springer P (2011) Molecular genetic studies on ASYMMETRIC LEAVES2 (AS2) of Arabidopsis: Insight into the function of the AS2 protein. Plant Morphol 23:71–80
Vanjildorj E, Song SY, Yang ZH, Choi JE, Noh YS, Park S, Lim WJ, Cho KM, Yun HD, Lim YP (2009) Enhancement of tolerance to soft rot disease in the transgenic Chinese cabbage (Brassica rapa L. ssp. pekinensis) inbred line, Kenshin. Plant Cell Rep 28:1581–1591
Vlad D, Kierzkowski D, Rast MI, Vuolo F, Dello Ioio R, Galinha C, Gan X, Hajheidari M, Hay A, Smith RS, Huijser P, Bailey CD, Tsiantis M (2014) Leaf shape evolution through duplication, regulatory diversification, and loss of a homeobox gene. Science 343:780–783
Wilson-Sánchez D, Rubio-Díaz S, Muñoz-Viana R, Pérez-Pérez J, Jover-Gil S, Ponce M, Micol J (2014) Leaf phenomics: a systematic reverse genetic screen for Arabidopsis leaf mutants. Plant J 79:878–891
Wolf S, Hematy K, Hofte H (2012) Growth control and cell wall signaling in plants. Annu Rev Plant Biol 63:381–407
Xiao D, Wang H, Basnet RK, Zhao J, Lin K, Hou X, Bonnema G (2014) Genetic dissection of leaf development in Brassica rapa using a ‘genetical genomics’ approach. Plant Physiol 164:1309–1325
Yin XJ, Volk S, Ljung K, Mehlmer N, Dolezal K, Ditengou F, Hanano S, Davis SJ, Schmelzer E, Sandberg G, Teige M, Palme K, Pickart C, Bachmair A (2007) Ubiquitin lysine 63 chain forming ligases regulate apical dominance in Arabidopsis. Plant Cell 19:1898–1911
Zhao J, Wang X, Deng B, Lou P, Wu J, Sun R, Xu Z, Vromans J, Koornneef M, Bonnema G (2005) Genetic relationships within Brassica rapa as inferred from AFLP fingerprints. Theor Appl Genet 110:1301–1314
Zhou X, Sun TH, Wang N, Ling HQ, Lu S, Li L (2011) The cauliflower Orange gene enhances petiole elongation by suppressing expression of eukaryotic release factor 1. New Phytol 190:89–100
Acknowledgements
This work was supported by a grant from the Technology Development Program for Agriculture and Forestry, Ministry for Food, Agriculture, Forestry, and Fisheries (Grant 213007-05-2-SB620), Republic of Korea.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest, financial or otherwise.
Human and animal rights
No animals/humans were used for studies that are base of this research.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lee, SM., Song, H., Yi, H. et al. Transcriptomic analysis of contrasting inbred lines and F2 segregant of Chinese cabbage provides valuable information on leaf morphology. Genes Genom 41, 811–829 (2019). https://doi.org/10.1007/s13258-019-00809-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-019-00809-7