Abstract
Lung cancer is a major cause accounting for cancer-related mortalities, with lung adenocarcinoma (LUAD) being the most prevalent subtype. Given the high clinical and cellular heterogeneities of LUAD, accurate diagnosis and prognosis are crucial to avoid overdiagnosis and overtreatment. Taking full advantage of scRNA-Seq data to resolve the tumor heterogeneities, we explored the overall landscape of LUAD microenvironment. Utilizing the stage-specific tumor cell markers, we have developed highly accurate diagnostic and prognostic models with elevated sensitivity and specificity. The diagnostic model, developed through random forest algorithms with a thirteen-gene signature, achieved an accuracy of 96.4% and an AUC of 0.993. These metrics were further demonstrated by benchmarking with available models and scoring systems in independent cohorts. Concurrently, the prognostic model, formulated via Cox regression with a six-gene signature, effectively predicted overall survival, with elevated risk scores associated with increased fractions of cancer-associated fibroblasts, and higher likelihood of immune escape and T-cell exclusion. Subsequently, two nomograms were developed to predict survival and drug responses, facilitating their integration into clinical practice. Overall, this study underscores the potential of our models for efficient, rapid, and cost-effective diagnosis and prognosis of LUAD, adaptable to multiple expression profiling platforms and quantification methods.

Similar content being viewed by others
Introduction
Cancer is a pervasive health challenge with increasing incidence and mortality rates worldwide [1]. Lung cancer stands as the second most diagnosed cancer and remains the major cause of cancer mortality [1]. Lung adenocarcinoma (LUAD), a main subtype of non-small cell lung cancer (NSCLC, a main subtype of lung cancer), is frequently diagnosed with high malignany and low survival probabilities [2, 3]. This complexity is partially exacerbated by its tumor heterogeneity, manifesting both within and between tumors [4]. Tumor heterogeneity, including clonal architecture, mutational burden, and TME (tumor microenvironment), contributes to diagnostic inaccuracies, treatment inefficacies, and unfavorable prognoses [3, 5].
Given the resulting poor diagnosis and prognosis of LUAD, early and precise detection is imperative for improving patient outcomes [3, 6]. The traditional histopathological assessment, however, faces several limitations due to the high intratumor and intertumor heterogeneity of LUAD, such as sampling bias from the heterogeneous tissue, the integrity of biopsy, the formation of artifacts, antibody specificity, and the subjective nature of diagnosis based on experience caused by the inter pathologist variability about the histopathology interpretations [7,8,9]. In addition to histopathology, existing models usually exhibit limited diagnostic performance or lack practical clinical applicability. For instance, in lung cancer diagnosis, a model based on circRNA and mRNA profiles achieved AUC values ranging from 0.81 to 0.92 [10], a model based on immune-related genes achieved AUC values ranging from 0.86 to 0.92 [11]. Furthermore, additional diagnostic performance leveraging various liquid biopsies were discussed, achieving AUC values ranging from 0.5 to 0.97 [12].
During cancer treatment, accurately evaluating cancer stages and the outcome of treatment strategies is critical, yet often challenging. Available prognostic models have demonstrated varying degrees of performance. A model based on alternative splicing signature achieved an AUC of approximately 0.82 [13], while another based on methylation-driven lncRNA achieved an AUC of 0.68 [14]. Additionally, a prognostic model incorporating immune gene expression profiles reported an AUC of 0.72 [15]. In addition to the limited performance, these existing models accepted input profiles from single quantification method respectively and usually lack practical applicability. These gaps underscore the need for more robust, accurate, and clinically applicable models to address the challenges.
Over the past several years, researchers have increasingly turned to scRNA-Seq to investigate tumor heterogeneity at a finer resolution [16,17,18,19]. By profiling the single-cell transcriptomes, researchers can capture the intricate TME landscape, further demonstrate the cellular reprogramming patterns of metastatic LUAD [16], and illustrate the early immune changes during multiple myeloma progression [18]. Unlike conventional bulk RNA-Seq, which provides average gene expression profiles of a sample, scRNA-Seq provides transcriptome profiles of individual cells [20]. This makes scRNA-Seq particularly adept at characterizing specific tumor cells and evaluating corresponding markers of certain subsets within the TME.
In the current study, we identified stage-specific markers of tumor cells in LUAD using scRNA-Seq. By integrating scRNA-Seq and bulk RNA-Seq data, we developed and validated both a diagnostic model and a prognostic model for LUAD, achieving high accuracy. We further evaluated their potential clinical applicability in different independent cohorts, offering a promising avenue for the efficient, rapid, and cost-effective diagnosis and prognosis of LUAD.
Materials and Methods
scRNA-Seq data processing
The LUAD single-cell transcriptome data GSE131907 [16] was obtained from GEO, including fifty-eight samples. Due to the clinical information, we divided the eleven distant normal lung tissues, eleven early-stage tumor lung tissues, and four advanced-stage tumor lung tissues for subsequent analyses. The raw UMI count matrix, representing the absolute transcript counts, was imported into the Seurat [21] package in R [22], encompassing a total of 100,217 cells. The mitochondrial percentage was quantified using the PercentageFeatureSet function. Four low-quality samples exhibiting potential doublet contamination were excluded, resulting in a refined dataset of 83,429 cells, comprising 34,669 normal tissue cells, 41,869 early-stage tissue cells, and 6891 advanced-stage tissue cells. Normalization, highly variable feature selection, and scaling were performed with the NormalizeData, FindVariableFeatures, and ScaleData function within the Seurat [21]. Subsequent dimensionality reductions were performed with the RunPCA and RunUMAP function, and cell clustering was performed with the FindClusters function. Cells were then visualized in a low-dimensional UMAP (Uniform Manifold Approximation and Projection) space.
Focusing on stage-specific markers of LUAD, we isolated epithelial components from the tumor microenvironment (TME). This subset comprised 9348 epithelial cells, with 2895 from normal tissues, 4658 from early-stage tissues, and 1795 from advanced-stage tissues. Using the FindAllMarkers function, we identified specific markers of normal, early-stage, and advanced-stage tissues. Differential expression percentages were computed for each gene across cell populations, and the top 200 markers, ranked by descending differential expression percentages, were selected and deduplicated for subsequent analyses.
Diagnostic model construction
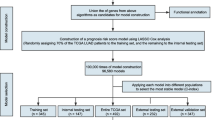
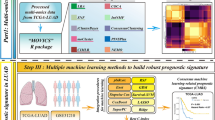
To construct the diagnostic model, we commenced by selecting the deduplicated stage-specific epithelial markers. Normalization in reference to GAPDH, a housekeeping gene, was subsequently conducted, adaptable to expression profiles across diverse platforms and methods. The formula is \({Normalized\; expression\; of\; GeneA}=\,{{Expression}}_{{GeneA}}/{{Expression}}_{{GAPDH}}\). A preliminary random forest model was then constructed using the randomForest package [23], incorporating the 9348 epithelial cells and the 502 deduplicated markers. Hyperparameter tuning was performed with the tuneRF function, resulting in the optimization of the preliminary model with 500 trees and 22 variables sampled at each split (Fig. S6A, B). Subsequently, the thirteen features exhibiting a Gini index decrease greater than 50 were identified and used to construct the final random forest model. The Gini index, obtained from the randomForest package [23], quantifies feature importance, with a greater decrease indicating a more significant feature. Following further hyperparameter tuning with the tuneRF function, the final hyperparameters for the final model were set to 350 trees and 3 variables sampled at each split (Fig. S6C, D). During the training of random forests, approximately one-third of the cases were withheld internally according to the bootstrap sampling, serving as out-of-bag data, which were employed to obtain an unbiased estimation of the model accuracy and the classification error. To validate the model’s robustness, expression profiles from multiple independent cohorts were included: TCGA-LUAD with 585 samples, GSE30219 [24] with 99 samples, GSE102287 [25] with 66 samples, GSE10072 [26] with 107 samples, GSE7670 [27] with 66 samples, and GSE19804 [28] with 120 samples.
To assess the performance and comparative effectiveness of our diagnostic model, we conducted a comprehensive benchmarking analysis against other available models and scoring systems. These included the scoring system from Evolutionary Bioinformatics 2019 [29] and models from Cancers 2022 [30], Translational Cancer Research 2021 [31], and Journal of Translational Medicine 2021 [32]. The AUC and confusion matrix were compared to determine the sensitivity, specificity, and overall accuracy.
The diagnostic risk score was calculated using the formula: \({risk\; score}=\sum ({coefficient}\times {normalized\; expression\; value})\), where the coefficient represented the weighted percentage of the decreasing Gini index for each feature. Features upregulated in tumor tissues were assigned positive coefficients, while features upregulated in normal tissues were assigned negative coefficients. The specific coefficients for each feature were as follows: −0.138 for SCGB1A1, 0.128 for IGKC, −0.086 for ADIRF, −0.083 for SFTPC, −0.081 for FABP5, 0.073 for CD24, −0.069 for SLPI, −0.066 for CYB5A, −0.066 for TPPP3, −0.065 for FABP4, 0.05 for IGHG4, −0.048 for FOLR1, and −0.045 for CLDN18, respectively.
Prognostic model construction
To establish a prognostic model, we initially identified the 355 common features between LUAD markers (both early-stage and advanced-stage markers in scRNA-Seq data) and the significant features (P-value < 0.01) derived from survival analyses with the TCGA-LUAD data. The CoxPH regression model was then employed to ascertain the effects of individual genes, obtaining corresponding HR (Hazard Ratio) and coefficients. The six features with HR greater than 1.1 or less than 0.8 were selected to construct the prognostic model.
The prognostic risk score was assessed as follows: \({risk\; score}=\sum ({coefficient}\times {normalized\; expression\; value})\), where the coefficients were derived from the CoxPH regression model, and the expression values were normalized by dividing them by the mean of the six features. The specific coefficients were as follows: 0.098 for MRPS11, 0.181 for CD3EAP, 0.123 for EMC6, 0.114 for SIX5, −0.257 for DMD, and −0.288 for STK33.
In TCGA-LUAD data, each sample was assigned a prognostic risk score, which was subsequently used to evaluate associations with clinical traits, including differential risk comparisons between different tumor stages and the pearson correlations related to new tumor events. The prognostic model’s validity was further evaluated using additional independent datasets: GSE31210 [33], GSE13213 [34], and GSE72094 [35], which collectively contain 226, 117, and 442 paired transcriptome profiles and clinical information of tumor lung tissues, respectively.
Survival analysis
Survival analyses were conducted with the survival [36] package, version 3.2. To categorize samples, cutoff thresholds were established due to the 0.25 and 0.75 quartiles. Accordingly, samples with gene expressions ranking in the top 25% were designated as the high expression group, whereas those in the bottom 25% were designated as the low expression group. This quartile-based approach was similarly applied to risk scores, where samples falling into the highest 25% were classified as high-risk, and those in the lowest 25% were classified as low-risk. Survival plots were generated with the survminer [37] package version 0.4.9.
Cell type identification and TIDE (tumor immune dysfunction and exclusion)
The immunedeconv [38] package was used to deconvolute cell types and estimate cellular fractions within bulk RNA-Seq samples. The ESTIMATE [39] was employed to estimate the total immune and stromal compositions. TIDE framework was employed to asses the potential for tumor immune escape and predict responses to immunotherapies. The T cell dysfunction scores and MDSC (myeloid-derived suppressor cell) scores were obtained from TIDE [40, 41].
Statistical analysis
All statistical analyses were conducted within R [22], version 4.1.0. Mann‒Whitney test or Student’s t-test was employed for the differential comparisons between distinct groups. Variance estimation was conducted with var.test function. Fisher’s exact test was used to compare the differential IHC staining results. Statistical significance was indicated by a P-value < 0.05 (***P-value < 0.001, **P-value < 0.01, *P-value < 0.05).
Results
The landscape of the LUAD tumor microenvironment and stage-specific marker identification
Using the LUAD (lung adenocarcinoma) scRNA-Seq dataset GSE131907 [16], we obtained the raw UMI matrix of normal lung tissues, early-stage and advanced-stage tumor lung tissues. After filtration, the expression profiles of 83,429 cells were processed using Seurat [21]. Through clustering and annotation, distinct cell types were clearly separated on the two-dimensional UMAP space, including epithelial cells, T lymphocytes, myeloid cells, B lymphocytes, and others (Fig. 1A). These cell types were distinguished through the expression of canonical specific markers, as exemplified by the specific expression of CD3D and CD3E in T lymphocytes (Fig. 1B).
A The UMAP plot demonstrates the major cell types in the LUAD tumor microenvironment. Different cell types are labeled by different colors. B The expression levels of canonical markers in specific cell types. C The compositions of the epithelial part in the tumor microenvironment, including normal epithelial cells and early-stage and advanced-stage tumor epithelial cells. D The stage-specific marker expression in epithelial cells. The color spectrum represents the expression levels of the markers.
Focusing on the epithelial components, we isolated and reprocessed the epithelial cells. By performing highly variable feature selection, scaling, and clustering, cells originating from samples at different stages could be roughly separated (Fig. 1C). Using the FindAllMarkers function, we identified the stage-specific markers of epithelial cells, as exemplified by the top five markers expressed at each stage (Fig. 1D, Figure S1; Table S1).
Construction of the diagnostic model and evaluations
To construct the diagnostic model, we commenced by ranking the stage-specific epithelial markers according to the decreasing differential expression percentages across distinct groups. The top 200 significant markers for each stage were selected and deduplicated, resulting in a curated list of 502 features. Using the epithelial components in the scRNA-Seq dataset, we normalized these features in reference to GAPDH and loaded them to train a preliminary random forest model to distinguish normal and tumor tissues, achieving an accuracy of 96.7% and an AUC of 0.994. The top thirty features, defined by the decreasing Gini index that indicates variable importance, were depicted (Fig. 2A). The thirteen features with a Gini index decrease greater than 50 were selected to reconstruct a final diagnostic model. These thirteen features exhibited distinct expression profiles across samples (Fig. 2B, C, Fig. S2). The final diagnostic model achieved an accuracy of 96.4% and an impressive AUC of 0.993 (Fig. 2D). When applied to additional independent cohorts, the model consistently achieved impressive AUCs, as exemplified by an AUC of 0.988 in the TCGA-LUAD with 585 samples (Fig. 2E), 0.996 in the GSE30219 [24] with 99 samples (Fig. 2F), 0.990 in the GSE102287 [25] with 66 samples (Fig. 2G), and three more datasets (Fig. S3A–C). Further benchmarking with other available models and scoring systems [29,30,31,32] was subsequently conducted. Our final diagnostic model achieved higher AUCs and accuracies than others across different independent cohorts (Fig. 2H–K, Fig. S3D). This finding underscored the potential of our diagnostic model to efficiently differentiate between normal and tumor tissues.
A The top thirty features ranked by the decreasing Gini index, representing variable importance. B, C Expression profiles of the potential candidates with the decreased Gini index greater than 50, as exemplified by SCGB1A1 (B) and CD24 (C). D ROC curve of the diagnostic model in the training dataset. E–G ROC curve of the diagnostic model in independent cohorts, including TCGA-LUAD (E), GSE30219 (F), and GSE102287 (G). H−K Benchmarking with other available models and scoring systems in dataset GSE102287 (H), GSE10072 (I), GSE30219 (J), and GSE7670 (K). The RFmodel stands for our diagnostic model, EvoBio2019 stands for the scoring system in Evolutionary Bioinformatics 2019 [29], Cancers2022 stands for the model in Cancers 2022 [30], TCR2021 stands for the model in Translational Cancer Research 2021 [31], and JTM2021 stands for the model in Journal of Translational Medicine 2021 [32]. L–N The risk scores of tumor tissues were significantly higher than those of normal adjacent tissues, as illustrated in independent cohorts, including TCGA-LUAD (L), GSE102287 (M), and GSE30219 (N). O Tumors from high-risk patients exhibited larger fractions of CAFs (cancer-associated fibroblasts) than those from low-risk patients. ***P-value < 0.001, by Mann-Whitney test.
The diagnostic risks were further evaluated for samples from different independent cohorts, revealing that tumor tissues consistently exhibited significantly higher risk scores compared to normal adjacent tissues (Fig. 2L–N, Figure S3E–G). To further investigate this difference, we evaluated the corresponding cellular compositions and fractions in high- and low-risk samples using the immunedeconv [38] package. Intriguingly, high-risk tissues exhibited significantly larger fractions of cancer-associated fibroblasts (CAFs) (Fig. 2O), which play pivotal roles in carcinogenesis [42], promoting immunosuppression [43], and enhancing the metastatic potential of lung cancer cells [44].
Construction of the prognostic model and validations
With the TCGA-LUAD transcriptome profiles and clinical data, survival analyses revealed that 3272 features, encompassing both coding genes and non-coding RNAs, significantly impacted patients’ overall survival (P-value < 0.05). Among these features, 1427 exhibited even greater significance with P-value < 0.01 (Fig. 3A). The common parts between these 1427 more significant features and two stage-specific markers of LUAD were identified as candidates (Fig. 3B, C), which were subsequently combined and deduplicated, resulting in a refined list of 355 genes. Univariate Cox regression was employed to evaluate the HR (Hazard Ratio) values and coefficients of these candidates. Ultimately, six candidates with an average HR greater than 1.1 or less than 0.8 were selected to construct a prognostic model (Fig. 3D).
A Survival analysis illustrated the features significantly affecting patient survival status. Dots in red represent features with P-value less than 0.01, while dots in blue represent features with P-value less than 0.05. B, C Advanced-stage (B) and early-stage (C) tumor-specific epithelial cell markers were mapped to the features with P-value less than 0.01 derived from survival analysis, indicated by red. D Cox regression analysis illustrated the ranked factors selected for constructing the prognostic model. E Cox regression analysis illustrated the risk score as a significant risk factor. F Survival analysis illustrated the significantly lower survival probabilities of the high-risk patients compared to the low-risk patients. G Validations in independent cohorts of the prognostic model using Cox regression analysis. H–J Tumors from the high-risk patients exhibited larger fractions of CAFs (H), lower fractions of endothelial cells (I) and CD4+ T cells (J) than those from the low-risk patients. ***P-value < 0.001, *P-value < 0.05, by Mann-Whitney test.
The prognostic model’s output variable, the risk score, was identified as a significant risk factor, with higher risk scores indicating a worse prognosis and a reduced survival probability in the TCGA-LUAD data (Fig. 3E, F). Furthermore, we verified the validity of this model in three additional independent datasets, including GSE13213 [34], GSE31210 [33], and GSE72094 [35] (Fig. 3G, Fig. S4A–C). Through evaluations of tumor cell types and compositions using the immunedeconv [38], tumors with high risks were significantly associated with higher CAF fractions (Fig. 3H), lower endothelial cell fractions (Fig. 3I), and lower CD4+ T cell fractions (Fig. 3J). Consistently, samples with higher risks exhibited lower stromal fractions and higher tumor purities (Fig. S4D, E). These findings collectively underscored the prognostic value of our model and its potential association with the tumor microenvironment compositions.
Associations between the prognostic risk and clinical traits
To further evaluate the prognostic model, we explored the associations between clinical traits and prognostic risks. Consistent with survival analysis, patients with high risks exhibited significantly shorter survival time (Fig. S5A). Additionally, significant differences in risk scores were observed across different tumor stages (Fig. 4A). Specifically, tumors at higher stages exhibited higher risk scores, which are typically associated with metastasis to nearby tissues or organs. Given the widespread use of the TNM staging system, with higher stages representing more deteriorating cancers, we also evaluated its association with prognostic risks. Our analysis revealed that risk scores differed significantly among distinct T stages, with higher risk scores associated with larger tumors or metastasis to nearby tissues (Fig. 4B). Similarly, we found differential risk scores among N stages, indicating more regional lymph nodes affected by cancer in high-risk patients (Fig. 4C).
A Differential prognosis risk score comparisons among different LUAD stages. Higher risk scores indicated larger tumor size and more metastasis. B, C Differential prognosis risk score comparisons among different pathologic T stages (B) and N stages (C). ***P-value < 0.001, **P-value < 0.01, *P-value < 0.05, by Mann-Whitney test. D Negative correlations between the prognosis risk scores and time to new tumor events. E Relapsed tumors were characterized by significantly higher prognosis risk scores in the independent dataset GSE31210. F Negative correlations between the prognosis risk scores and time to tumor relapse events. G, H Differential prognosis risk score comparisons between different responses to paclitaxel (G) and pemetrexed (H) treatments. For the treatment responses, complete response, partial response, and stable disease were classified as “Responses/Controlled”, while clinical progressive disease was classified as “No responses”. **P-value < 0.01, *P-value < 0.05, by Student’s t-test. I Differential prognosis risk score comparisons between predicted different responses to immunotherapy. ***P-value < 0.001, by Mann-Whitney test. J–L Positive correlations between prognosis risk scores and immunotherapy response (J), MDSC scores (K), and T-cell exclusion (L).
Beyond tumor stages, we also investigated the occurrence of the prognosis-related new tumor events. Subsequent analysis revealed the negative correlation between risk scores and time to experience new tumor events, encompassing recurrent metastasis and new primary tumors (Fig. 4D). Similarly, we also observed a higher frequency of relapsed events in high-risk patients in independent cohorts GSE13213 [34] and GSE31210 [33] (Fig. 4E, F, Fig. S5B). Intriguingly, higher risk scores were observed in samples with KRAS or TP53 mutations, with samples harboring double mutations exhibiting even higher risk scores (Fig. S5C). Considering the report that STK33, a gene in our prognostic model, functioned in a mutant KRAS-dependent manner [45], the prognostic model genes may be potentially regulated by these factors. These findings underscored the prognostic model’s clinical relevance in LUAD.
Associations between the prognostic risk and therapy response
We conducted further analysis to evaluate the predictive potential of the prognostic model for therapy responses. For paclitaxel, lower risk scores were associated with more favorable responses, including complete response, partial response, and stable disease (Fig. 4G). Similarly, for pemetrexed, lower risk scores were associated with complete response and stable disease (Fig. 4H). In the context of immunotherapy, through evaluating responses with TIDE, the non-responders exhibited significantly higher prognostic risks (Fig. 4I). Consistently, immunotherapy response scores were positively correlated with prognostic risk scores (Fig. 4J), where higher immunotherapy scores demonstrate a higher likelihood of immune escape and less potential to benefit from immunotherapy. Furthermore, both MDSC (myeloid-derived suppressor cell) scores and T cell exclusion scores showed positive correlations with prognostic risk scores (Fig. 4K, L), where increasing MDSC and T cell exclusion scores indicate an immune suppressive status. These analyses collectively underscored the prognostic model’s utility in predicting therapy responses and its potential to guide personalized treatment strategies.
Expression validation of the prognostic model genes
To elucidate the protein profiles of the prognostic model genes, we quantified the immunohistochemistry (IHC) staining results from the Human Protein Atlas (HPA) database [46,47,48]. The IHC staining were categorized into four levels: high, medium, low, and not detected. Our analysis demonstrated that the MRPS11 protein was predominantly detected at high and medium levels in lung cancer samples, whereas it was not detected in normal alveolar or endothelial cells (Fig. 5A). Similarly, the CD3EAP protein was primarily detected in lung cancer samples but not in normal alveolar cells (Fig. 5B). Representative IHC staining results for both MRPS11 and CD3EAP are presented (Fig. 5C–F). Consistently, both MRPS11 and CD3EAP exhibited significantly higher expression in lung cancer samples at the transcriptomic level (Fig. 5G, H), and elevated levels of both genes were significantly associated with reduced survival probabilities (Fig. 5I, J). These findings provided a molecular basis for the prognostic value of the model genes and underscored the importance of these genes in the pathobiology of lung cancer.
A, B Statistical analysis of MRPS11 (A) and CD3EAP (B) protein IHC staining results. C, D Representative figures illustrating MRPS11 protein IHC staining in normal lung tissue (C) and tumor lung tissue (D). E, F Representative figures illustrating CD3EAP protein IHC staining in normal lung tissue (E) and tumor lung tissue (F). G, H Differential expression levels of MRPS11 (G) and CD3EAP (H) between normal adjacent tissues and LUAD at the transcriptomic level. ***P-value < 0.001, by Mann-Whitney test. I, J Survival analysis indicated that high expression levels of MRPS11 (I) and CD3EAP (J) significantly affected patient survival probabilities.
Predictions of the outcome and drug response based on the prognostic risk model
To illustrate the practical applications of the prognostic risk model, we assessed its potential to reflect patients’ survival probabilities and drug responses. By combining the tumor stage information, the risk score were able to predict patients’ overall survival probabilities (Fig. 6A). Correspondingly, the calibration curves confirmed the model’s accuracy in predicting patient outcomes in comparison to their actual overall survival (Fig. 6B–D). Despite the inclusion of both risk score and tumor stage to predict survival probabilities, the risk score remained the most significant factor with the highest HR values (Fig. 6E), while the tumor stage aided in the predictions. Furthermore, we presented an additional nomogram to predict drug responses to pemetrexed or paclitaxel based on risk scores from the prognostic model (Fig. 6F), achieving an AUC of 0.881 (Fig. 6G). These results underscored the prognostic model’s potential as a valuable tool for personalized medicine and predicting outcomes.
A The nomogram to predict patient outcomes according to the prognostic model. Individual points derived from the risk score and tumor stage were added to predict the outcomes. B–D Calibration curves of 1-year (B), 3-year (C), and 5-year (D) survival demonstrated the high accuracy of the nomogram. E Cox regression analysis illustrated the HR values of the risk score and tumor stage. The risk score remained the most significant factor with the highest HR value. F The nomogram to predict patient responses to paclitaxel or pemetrexed treatments according to the prognostic model. G ROC curve of the nomogram to predict responses to paclitaxel or pemetrexed treatments.
Discussion
Lung cancer, particularly lung adenocarcinoma (LUAD), is the leading cause of cancer-related deaths. Traditional diagnostic methods for LUAD, including auxiliary imaging and pathological examinations, are usually accompanied by several limitations, such as sampling bias, integrity of biopsy, the formation of artifacts, antibody specificity, and subjective diagnosis based on experience caused by the inter pathologist variability [7,8,9]. Considering the tumor heterogeneity, these limitations could lead to inaccurate diagnosis. Existing models suffer from limited performance or lack practical applicability, due to their limited sensitivity or specificity, validation within the same cohort of samples, and reliance on the same quantification method.
To address these challenges, we aimed to develop transcriptomic profiling-based diagnostic and prognostic models using scRNA-Seq data of LUAD, as scRNA-Seq provides deeper insights into tumor microenvironments at single-cell resolution. By leveraging scRNA-Seq data, we trained and constructed a diagnostic model and a prognostic model with high accuracy. We also evaluated the potential applications of both models, facilitating their translation to practical applications in clinical guidance.
During the construction of the diagnostic model with high sensitivity and specificity, we delineated a stage-specific thirteen-gene signature. The final diagnostic model achieved an accuracy of 96.4% and an impressive AUC of 0.993, which was further validated across additional datasets from different platforms. Subsequent benchmarking with available models and scoring systems in independent cohorts further underscored our model’s exceptional performance. Despite the dependence on lung tissue biopsies, our model could to some extent aid in the accurate diagnosis of LUAD, mitigating several limitations in traditional diagnostic methodologies. While more peripheral blood samples are available, we would promote the accurate non-invasive and early detection of lung cancer through the integration of multi-omics data, including TCR/BCR-Seq.
During the construction of the prognostic model, we identified the combination of six genes: MRPS11, CD3EAP, EMC6, SIX5, DMD, and STK33. The functions and potential roles of these genes in LUAD have not been extensively investigated. MRPS11, as a DNA damage response gene, was potentially regulated by miR-211 in ovarian cancer [49], and its high expression level was significantly correlated with low survival probabilities in uveal melanoma [50]. Knocking down CD3EAP reduced colony formation and inhibited cell proliferation, sphere formation, and sizes in prostate cancer [51]. DMD, as a tumor suppressor, played important roles in myogenic cancers, including inhibiting myogenic sarcoma cell migration, invasion, and invadopodia formation [52]. Additionally, knocking down STK33 impaired colony formation in a mutant KRAS-dependent manner across cancers [45]. These researches provide support for the potential roles of these prognostic model genes in evaluating the outcomes of LUAD.
Given that the differential levels of the prognostic model genes were quantified and confirmed by IHC staining results in normal and tumor lung tissues, it remains unclear what exact roles they play in the development of different LUAD stages. Further study and verification of these critical genes and their functions may lead to the discovery of novel drug targets for cancer treatment, which would be a great follow-up study.
For the input of both the diagnostic and the prognostic models, we incorporated data from both high-throughput sequencing and microarray platforms. However, for the diagnostic model, the input could be expression matrices normalized in reference to GAPDH from various platforms and quantification methods. For the prognostic model, the input could be expression matrices normalized in reference to the mean expression of the six genes from multiple platforms and quantification methods. By standardizing the input in this manner, both models were able to maintain high accuracy while enabling efficient, rapid, and cost-effective diagnosis and prognosis.
Overall, we presented a diagnostic model and a prognostic model with high sensitivity and specificity, and evaluated the potential clinical applications of both models. Benchmarking in independent cohorts demonstrated their exceptional performance, underscoring their reliability and generalizability. We believe that this study could contribute to the auxiliary diagnosis and prognosis of lung adenocarcinoma, including ancillary clinical guidance, and provide deeper insights for future studies regarding the functions of pivotal factors involved in the early diagnosis, prognosis, and therapy responses of lung cancer.
Data availability
The LUAD scRNA-Seq profiles GSE131907 [16] was obtained from GEO. Expression profiles and clinical traits of TCGA-LUAD were obtained from the UCSC Xena browser (https://xenabrowser.net/datapages/). Additionally, microarray datasets GSE7670 [27], GSE102287 [25], GSE30219 [24], GSE19804 [28], GSE10072 [26], GSE31210 [33], GSE13213 [34], and GSE72094 [35] were obtained from GEO. The drug responses and corresponding clinical data were obtained from a previous study [53]. The final diagnostic model is available from the Github repository (https://github.com/univerchen/LUAD).
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin. 2021;71:209–49.
Spella M, Stathopoulos GT. Immune Resistance in Lung Adenocarcinoma. Cancers (Basel). 2021;13:384.
Senosain MF, Massion PP. Intratumor Heterogeneity in Early Lung Adenocarcinoma. Front Oncol. 2020;10:349.
Seguin L, Durandy M, Feral CC. Lung Adenocarcinoma Tumor Origin: A Guide for Personalized Medicine. Cancers (Basel). 2022;14:1759.
Diaz-Cano SJ. Tumor heterogeneity: mechanisms and bases for a reliable application of molecular marker design. Int J Mol Sci. 2012;13:1951–2011.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2016. CA Cancer J Clin. 2016;66:7–30.
Chatterjee S. Artefacts in histopathology. J Oral Maxillofac Pathol. 2014;18:S111–6.
Hillman H. Limitations of clinical and biological histology. Med Hypotheses. 2000;54:553–64.
Taqi SA, Sami SA, Sami LB, Zaki SA. A review of artifacts in histopathology. J Oral Maxillofac Pathol. 2018;22:279.
D’Ambrosi S, Giannoukakos S, Antunes-Ferreira M, Pedraz-Valdunciel C, Bracht JWP, Potie N, et al. Combinatorial Blood Platelets-Derived circRNA and mRNA Signature for Early-Stage Lung Cancer Detection. Int J Mol Sci. 2023;24:4881.
Ye XD, Zhang N, Jin YX, Xu B, Guo CY, Wang XQ, et al. Dramatically changed immune-related molecules as early diagnostic biomarkers of non-small cell lung cancer. Febs J. 2020;287:783–99.
Freitas C, Sousa C, Machado F, Serino M, Santos V, Cruz-Martins N, et al. The Role of Liquid Biopsy in Early Diagnosis of Lung Cancer. Front Oncol. 2021;11:634316.
Li Y, Sun N, Lu Z, Sun S, Huang J, Chen Z, et al. Prognostic alternative mRNA splicing signature in non-small cell lung cancer. Cancer Lett. 2017;393:40–51.
Li R, Yang YE, Yin YH, Zhang MY, Li H, Qu YQ. Methylation and transcriptome analysis reveal lung adenocarcinoma-specific diagnostic biomarkers. J Transl Med. 2019;17:324.
Sun L, Zhang Z, Yao Y, Li WY, Gu J. Analysis of expression differences of immune genes in non-small cell lung cancer based on TCGA and ImmPort data sets and the application of a prognostic model. Ann Transl Med. 2020;8:550.
Kim N, Kim HK, Lee K, Hong Y, Cho JH, Choi JW, et al. Single-cell RNA sequencing demonstrates the molecular and cellular reprogramming of metastatic lung adenocarcinoma. Nat Commun. 2020;11:2285.
Ma KY, Schonnesen AA, Brock A, Van Den Berg C, Eckhardt SG, Liu Z, et al. Single-cell RNA sequencing of lung adenocarcinoma reveals heterogeneity of immune response-related genes. JCI Insight. 2019;4:121387.
Zavidij O, Haradhvala NJ, Mouhieddine TH, Sklavenitis-Pistofidis R, Cai S, Reidy M, et al. Single-cell RNA sequencing reveals compromised immune microenvironment in precursor stages of multiple myeloma. Nat Cancer. 2020;1:493–506.
Lu T, Yang X, Shi Y, Zhao M, Bi G, Liang J, et al. Single-cell transcriptome atlas of lung adenocarcinoma featured with ground glass nodules. Cell Discov. 2020;6:69.
Olsen TK, Baryawno N. Introduction to Single-Cell RNA Sequencing. Curr Protoc Mol Biol. 2018;122:e57.
Hao Y, Hao S, Andersen-Nissen E, Mauck WM 3rd, Zheng S, et al. Integrated analysis of multimodal single-cell data. Cell. 2021;184:3573–3587.e29.
Team RC. R: A language and environment for statistical computing. 2013.
Liaw A, Wiener M. Classification and regression by randomForest. R N. 2002;2:18–22.
Rousseaux S, Debernardi A, Jacquiau B, Vitte AL, Vesin A, Nagy-Mignotte H, et al. Ectopic activation of germline and placental genes identifies aggressive metastasis-prone lung cancers. Sci Transl Med. 2013;5:186ra66.
Mitchell KA, Zingone A, Toulabi L, Boeckelman J, Ryan BM. Comparative Transcriptome Profiling Reveals Coding and Noncoding RNA Differences in NSCLC from African Americans and European Americans. Clin Cancer Res. 2017;23:7412–25.
Landi MT, Dracheva T, Rotunno M, Figueroa JD, Liu H, Dasgupta A, et al. Gene expression signature of cigarette smoking and its role in lung adenocarcinoma development and survival. PLoS One. 2008;3:e1651.
Su LJ, Chang CW, Wu YC, Chen KC, Lin CJ, Liang SC, et al. Selection of DDX5 as a novel internal control for Q-RT-PCR from microarray data using a block bootstrap re-sampling scheme. BMC Genomics. 2007;8:140.
Lu TP, Tsai MH, Lee JM, Hsu CP, Chen PC, Lin CW, et al. Identification of a novel biomarker, SEMA5A, for non-small cell lung carcinoma in nonsmoking women. Cancer Epidemiol Biomark Prev. 2010;19:2590–7.
Sheng M, Xie X, Wang J, Gu W. A Pathway-Based Strategy to Identify Biomarkers for Lung Cancer Diagnosis and Prognosis. Evol Bioinform Online. 2019;15:1176934319838494.
Zhang BZ, Wang YD, Zhou XZ, Zhang Z, Ju HY, Diao XQ, et al. Construction of a Prognostic and Early Diagnosis Model for LUAD Based on Necroptosis Gene Signature and Exploration of Immunotherapy Potential. Cancers. 2022;14:5153.
Chen Q, Wang XY, Hu J. Systematically integrative analysis identifies diagnostic and prognostic candidates and small-molecule drugs for lung adenocarcinoma. Transl Cancer Res. 2021;10:3619–46.
Cai SH, Guo XT, Huang CJ, Deng YJ, Du LD, Liu WY, et al. Integrative analysis and experiments to explore angiogenesis regulators correlated with poor prognosis, immune infiltration and cancer progression in lung adenocarcinoma. J Transl Med. 2021;19:361.
Okayama H, Kohno T, Ishii Y, Shimada Y, Shiraishi K, Iwakawa R, et al. Identification of genes upregulated in ALK-positive and EGFR/KRAS/ALK-negative lung adenocarcinomas. Cancer Res. 2012;72:100–11.
Tomida S, Takeuchi T, Shimada Y, Arima C, Matsuo K, Mitsudomi T, et al. Relapse-related molecular signature in lung adenocarcinomas identifies patients with dismal prognosis. J Clin Oncol. 2009;27:2793–9.
Schabath MB, Welsh EA, Fulp WJ, Chen L, Teer JK, Thompson ZJ, et al. Differential association of STK11 and TP53 with KRAS mutation-associated gene expression, proliferation and immune surveillance in lung adenocarcinoma. Oncogene. 2016;35:3209–16.
Therneau T. A package for survival analysis in S. R package version, 2015. 2.
Kassambara, A, Kosinski M, Biecek P, Fabian S. Survminer: Drawing Survival Curves Using Ggplot2. 2021. https://CRAN.R-project.org/package=survminer. R package version 0.4, 2021. 9.
Sturm G, Finotello F, List M. Immunedeconv: An R Package for Unified Access to Computational Methods for Estimating Immune Cell Fractions from Bulk RNA-Sequencing Data. Methods Mol Biol. 2020;2120:223–32.
Yoshihara K, Kim H, Verhaak R. estimate: Estimate of Stromal and Immune Cells in Malignant Tumor Tissues from Expression Data. R package version, 2016. 1: p. r21.
Fu J, Li K, Zhang W, Wan C, Zhang J, Jiang P, et al. Large-scale public data reuse to model immunotherapy response and resistance. Genome Med. 2020;12:21.
Jiang P, Gu S, Pan D, Fu J, Sahu A, Hu X, et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat Med. 2018;24:1550–8.
Wong KY, Cheung AH, Chen B, Chan WN, Yu J, Lo KW, et al. Cancer-associated fibroblasts in nonsmall cell lung cancer: From molecular mechanisms to clinical implications. Int J Cancer. 2022;151:1195–215.
Xiang H, Ramil CP, Hai J, Zhang C, Wang H, Watkins AA, et al. Cancer-Associated Fibroblasts Promote Immunosuppression by Inducing ROS-Generating Monocytic MDSCs in Lung Squamous Cell Carcinoma. Cancer Immunol Res. 2020;8:436–50.
Wang L, Cao L, Wang H, Liu B, Zhang Q, Meng Z, et al. Cancer-associated fibroblasts enhance metastatic potential of lung cancer cells through IL-6/STAT3 signaling pathway. Oncotarget. 2017;8:76116–28.
Scholl C, Frohling S, Dunn IF, Schinzel AC, Barbie DA, Kim SY, et al. Synthetic Lethal Interaction between Oncogenic KRAS Dependency and STK33 Suppression in Human Cancer Cells. Cell. 2009;137:821–34.
The Human Protein Atlas. 2022; Available from: http://www.proteinatlas.org.
Thul PJ, Akesson L, Wiking M, Mahdessian D, Geladaki A, Ait Blal H, et al. A subcellular map of the human proteome. Science. 2017;356:eaal3321.
Uhlen M, Fagerberg L, Hallstrom BM, Lindskog C, Oksvold P, Mardinoglu A, et al. Proteomics. Tissue-based map of the human proteome. Science. 2015;347:1260419.
Wang T, Hao D, Yang S, Ma J, Yang W, Zhu Y, et al. miR-211 facilitates platinum chemosensitivity by blocking the DNA damage response (DDR) in ovarian cancer. Cell Death Dis. 2019;10:495.
Zhao DD, Zhao X, Li WT. Identification of differentially expressed metastatic genes and their signatures to predict the overall survival of uveal melanoma patients by bioinformatics analysis. Int J Ophthalmol. 2020;13:1046–53.
Zhang D, Park D, Zhong Y, Lu Y, Rycaj K, Gong S, et al. Stem cell and neurogenic gene-expression profiles link prostate basal cells to aggressive prostate cancer. Nat Commun. 2016;7:10798.
Wang YX, Marino-Enriquez A, Bennett RR, Zhu MJ, Shen YP, Eilers G, et al. Dystrophin is a tumor suppressor in human cancers with myogenic programs. Nat Genet. 2014;46:601–6.
Ding Z, Zu S, Gu J. Evaluating the molecule-based prediction of clinical drug responses in cancer. Bioinformatics. 2016;32:2891–5.
Acknowledgements
This work was funded by the National Key R&D Program of China (2022YFA1303200 to XS), the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB29030104 to TJ), the National Natural Science Foundation of China (Grant 82272301 to TJ), and the Fundamental Research Funds for the Central Universities (WK9100000001 to TJ).
Author information
Authors and Affiliations
Contributions
Qingyu Cheng: Conceptualization, Data curation, Investigation, Visualization, Writing - original draft. Weidong Zhao: Writing - review & editing. Xiaoyuan Song: Project administration, Writing - review & editing. Tengchuan Jin: Conceptualization, Project administration, Supervision, Writing - review & editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cheng, Q., Zhao, W., Song, X. et al. Machine-learning and scRNA-Seq-based diagnostic and prognostic models illustrating survival and therapy response of lung adenocarcinoma. Genes Immun (2024). https://doi.org/10.1038/s41435-024-00289-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41435-024-00289-0
- Springer Nature Limited