Abstract
Lung adenocarcinoma (LUAD) remains the most common deadly disease and has a poor prognosis. Pyroptosis could regulate tumour cell proliferation, invasion, and metastasis, thereby affecting the prognosis of cancer patients. However, the role of pyroptosis-related genes (PRGs) in LUAD remains unclear. In our study, comprehensive bioinformatics analysis was performed to construct a prognostic gene model and ceRNA network. The correlations between PRGs and tumour-immune infiltration, tumour mutation burden, and microsatellite instability were evaluated using Pearson’s correlation analysis. A total of 23 PRGs were upregulated or downregulated in LUAD. The genetic mutation variation landscape of PRG in LUAD was also summarised. Functional enrichment analysis revealed that these 33 PRGs were mainly involved in pyroptosis, the NOD-like receptor signalling pathway, and the Toll-like receptor signalling pathway. Prognosis analysis indicated a poor survival rate in LUAD patients with low expression of NLRP7, NLRP1, NLRP2, and NOD1 and high CASP6 expression. A prognostic PRG model constructed using the above five prognostic genes could predict the overall survival of LUAD patients with medium-to-high accuracy. Significant correlation was observed between prognostic PRGs and immune-cell infiltration, tumour mutation burden, and microsatellite instability. A ceRNA network was constructed to identify a lncRNA KCNQ1OT1/miR-335-5p/NLRP1/NLRP7 regulatory axis in LUAD. In conclusion, we performed a comprehensive bioinformatics analysis and identified a prognostic PRG signature containing five genes (NLRP7, NLRP1, NLRP2, NOD1, and CASP6) for LUAD patients. Our results also identified a lncRNA KCNQ1OT1/miR-335-5p/NLRP1/NLRP7 regulatory axis, which may also play an important role in the progression of LUAD. Further study needs to be conducted to verify this result.
Similar content being viewed by others
Introduction
Lung cancer remains the most common deadly disease, with an estimated 2.09 million new cases and 1.76 million deaths each year [1]. Worse still, the incidence and mortality of lung cancer are rising [1]. Lung adenocarcinoma (LUAD) is the most common histologic subtype of lung cancer, accounting for approximately 40% of all cases [2]. Despite surgery, chemoradiotherapy, targeted therapy, and immunotherapy being used in the treatment of lung cancer, the prognosis remains disheartening [3], and 5-year survival ranges from 4 to 17%, depending on disease and treatment differences [4].
Although many biomarkers or gene signatures have been found to have the potential to predict the prognosis of LUAD, they are still in the molecular research phase and have not yet been applied in clinical practice. Thus, uncovering prognostic gene signatures for the prognosis of LUAD would be of great significance.
Pyroptosis, referred to as cellular inflammatory necrosis, is considered to be gasdermin-mediated programmed necrotic cell death [5]. Triggered by certain inflammasomes, pyroptosis relies on the cleavage of gasdermin D (GSDMD) and activation of inactive cytokines [6]. The correlation between pyroptosis and cancer is extremely complicated. Although pyroptosis can inhibit the oncogenesis and progression of tumours, it also develops a microenvironment delivering nutrients for cancer and accelerating cancer growth [7]. Increasing studies have demonstrated the effect of pyroptosis on tumour cell proliferation, invasion, and metastasis, thus affecting the prognosis of cancer [8, 9]. For example, a recent study identified a novel pyroptosis-related gene signature for the prognosis of ovarian cancer [10]. In lung cancer, the pyroptosis gene GSDMD can inhibit tumour proliferation by regulating the intrinsic mitochondrial apoptotic pathway and EGFR/Akt signalling [11]. The prognostic value of pyroptosis-related genes (PRGs) in LUAD has not yet been elucidated.
In the current research, bioinformatics analysis was performed to investigate PRG expression profiles and their prognostic significance as well as the associated regulatory axis in LUAD. Our data may provide additional evidence for prognostic biomarkers and therapeutic targets for LUAD.
Results
Defining of the expression of PRGs in LUAD
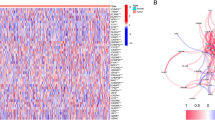
We first explored the expression of the 33 PRGs in LUAD and normal lung tissues using the TCGA LUAD dataset. A total of 23 PRGs were either upregulated or downregulated in LUAD (Fig. 1A). More specifically, the expression of PRKACA, NOD1, NLRP1, ELANE, TNF, IL1B, IL18, PYCARD, CASP5, NLRC4, NLRP3, IL6, and CASP1 was increased, while the expression of GSDMB, PJVK, CASP4, NLRP7, CASP3, CASP6, CASP8, GSDMA, GSDMC, and AIM2 was decreased in LUAD compared with normal tissues (Fig. 1A, all <0.001). A protein–protein interaction (PPI) analysis with the minimum required interaction score of 0.9 was constructed to detect the interactions of these PRGs, which revealed that CASP1, CASP5, CASP8, NLRP1, NLRP3, and PYCARD were hub genes (Fig. S1A). Supplementary Fig. S1B shows the correlation network containing all PRGs.
A The expression of 33 PRG in LUAD and lung tissues, Tumour, red; Normal, blue. The upper and lower ends of the boxes represented the interquartile range of values. The lines in the boxes represented median value. B, C The mutation frequency and classification of 33 PRG in LUAD. D The location of CNV alteration of 33 PRG on 23 chromosomes in the LUAD cohort. E The CNV variation frequency of 33 PRG in the LUAD cohort. The height of the column represented the alteration frequency. ***P < 0.001, PRG pyroptosis‑related gene, LUAD lung adenocarcinoma, SNP single nucleotide polymorphism, INS insertion, DEL deletion.
Landscape of genetic variation of PRGs in LUAD
We then summarised the incidence of copy number variations and somatic mutations of 33 PRGs in LUAD. As shown in Fig. 1B and Fig. 1C, 116 of 257 (64.59%) LUAD samples demonstrated genetic mutations. Missense mutation was the most common variant classification (Fig. 1B). SNPs were the most common variant type, and C > A ranked as the top SNV class. The results also demonstrated NLRP3 as the gene with the highest mutation frequency, followed by NLRP7 and NLRP2, among the 33 PRGs (Fig. 1C). Figure 1D presents the location of CNV alterations of these 33 PRGs on chromosomes. We also investigated CNV alteration frequency, which revealed that these 33 PRGs showed prevalent CNV alterations. More than half of the 33 PRGs had copy number amplification, while the CNV deletion frequencies of CASP9, GPX4, NLRP7, NLRP2, IL18, ELANE, NLRP6, PLCG1, CASP6, CASP3, NLRP1, and PRKACA were widespread (Fig. 1E).
Functional enrichment analysis of PRGs
To clarify the function of PRGs, the pathways were analysed using GO and KEGG databases. We found that these 33 PRGs were mainly involved in the positive regulation of cytokine production, interleukin-1 production, regulation of inflammatory response, pyroptosis, inflammasome complex, cysteine-type endopeptidase activity involved in apoptotic process, cysteine-type endopeptidase activity, and cytokine receptor binding in GO analysis (Fig. 2A). Moreover, KEGG pathway analysis suggested that 33 PRGs were mainly involved in the NOD-like receptor signalling pathway, salmonelia infection, cytosolic DNA-sensing pathway, TNF signalling pathway, Toll-like receptor signalling pathway, and apoptosis (Fig. 2B).
Construction of a pyroptosis-related prognostic gene model
To construct a prognostic gene model, univariate Cox regression analysis was performed to screen those PRG with a prognostic value. As a result, a total of five genes with a prognostic value were identified, and the Kaplan–Meier survival curves are shown in Fig. 3. The results suggested a poor survival rate in LUAD patients with low expression of NLRP7 (Fig. 3A, p = 0.021), NLRP1 (Fig. 3B, p = 0.008), NLRP2 (Fig. 3C, p = 0.014), and NOD1 (Fig. 3D, p = 0.047) and high CASP6 expression (Fig. 3E, p = 0.048). LASSO Cox regression analysis was performed to construct a prognostic gene model based on these five prognostic PRGs (Fig. 4A, B). The risk score = (0.0946) * CASP6 + (0.1573) * NLRP7 + (−0.124) * NOD1 + (−0.1627) * NLRP1 + (−0.0262) * NLRP2. Based on the risk score, LUAD patients were separated into two groups. The risk score distribution, survival status, and the expression of these five genes are presented in Fig. 4C. As the risk score increased, the patients’ risk of death increased, and the survival time decreased (Fig. 4C). The Kaplan–Meier curve revealed that LUAD patients with high-risk scores had a worse overall survival probability than those with low-risk scores (median time = 3.3 years vs. 4.9 years, p = 0.00083, Fig. 4D), with AUCs of 0.668, 0.591, and 0.612 in the 1-year, 3-year, and 5-year ROC curves, respectively (Fig. 4E).
A LASSO coefficient profiles of the five PRGs. B Plots of the ten-fold cross-validation error rates. C Distribution of risk score, survival status, and the expression of five prognostic PRGs in LUAD. D, E Overall survival curves for LUAD patients in the high-/low-risk group and the ROC curve of measuring the predictive value. PRG, pyroptosis‑related gene; LUAD, lung adenocarcinoma.
Building a predictive nomogram
Considering the clinicopathologic features and these five prognostic PRGs, we also built a predictive nomogram to predict the survival probability. Univariate and multivariate analyses revealed that NOD1 expression and pT stage, pN stage, and pM stage were independent factors affecting the prognosis of LUAD patients (Fig. 5A, B). The predictive nomogram suggested that the 3‐year and 5‐year overall survival rates could be predicted relatively well compared with an ideal model in the entire cohort (Fig. 5C, D).
A, B Hazard ratio and P‐value of the constituents involved in univariate and multivariate Cox regression considering clinical the parameters and five prognostic PRG in LUAD. C, D Nomogram to predict the 1-year, 3-year, and 5-year overall survival rate of LUAD patients. Calibration curve for the overall survival nomogram model in the discovery group. A dashed diagonal line represents the ideal nomogram. PRG pyroptosis‑related gene, LUAD lung adenocarcinoma.
PRGs were associated with tumour immune infiltration in LUAD
Pyroptosis plays a vital role in the development of the tumour-immune microenvironment. In our study, we also clarified the correlation of the expression of prognostic PRGs (NOD1, CASP6, NLRP1, NLRP2, and NLRP7) and immune infiltration in LUAD using the TIMER database. The data demonstrated a negative correlation between CASP6 expression and the abundance of B cells (Fig. 6A, p = 6.6e−4) and CD4 + T cells (Fig. 6A, p = 0.0157). Moreover, there was a positive association between NLRP7 expression and the immune infiltration level of B cells (p = 6.43e−6), CD4 + T cells (p = 1.21e−7), macrophages (p = 0.0184), neutrophils (p = 4.67e−4), and dendritic cells (p = 9.9e−5) (Fig. 6B). NLRP2 expression showed a positive association with the abundance of CD4 + T cells (p = 2.56e−4) and dendritic cells (p = 0.0426) (Fig. 6C). Figure 6D shows the correlation between NOD1 expression and the abundance of immune cells, which revealed a positive correlation between NOD1 expression and the abundance of B cells (p = 5.53e−10), CD8 + T cells (p = 0.0329), CD4 + T cells (p = 1.73e−15), neutrophils (p = 0.0151) and dendritic cells (p = 3.74e−11). We also found that NLRP1 expression was positively correlated with the abundance of B cells (p = 4.98e−23), CD8 + T cells (p = 0.0158), CD4 + T cells (p = 4.28e−48), macrophages (p = 2.03e−5), neutrophils (p = 2e−14), and dendritic cells (p = 1.47e−19) (Fig. 6E). This evidence suggests a significant correlation between PRG and tumour-immune infiltration.
TMB, MSI, and drug-sensitivity analysis of PRGs
TMB can be used as a biomarker to predict the efficacy of immunotherapy for lung cancer [12, 13]. Microsatellite instability (MSI) was also suggested as a predictive biomarker for cancer immunotherapy [14]. The above results revealed that the PRG was significantly correlated with tumour immune infiltration. To clarify whether these PRGs could also serve as biomarkers for drug screening, we then analysed the correlation between PRGs and TMB as well as MSI in LUAD. The results revealed a negative correlation between TMB and NOD1 (Fig. 7A, p = 1.39e−7) and NLRP1 (Fig. 7C, p = 1.88e−6) and a positive correlation between TMB and CASP6 (Fig. 7B, p = 0.023). However, there was no significant correlation between TMB and NLRP2 and NLRP7 (Fig. 7D, E). In MSI analysis, MSI was negatively correlated with CASP6 expression (Fig. 7G, p = 0.022), and was positively correlated with the expression of NLRP1 (Fig. 7H, p = 0.001) and NLRP2 (Fig. 7I, p = 0.044). There was no significant correlation between MSI and NOD1 or NLRP7 (Fig. 7F, J). To develop a therapy target, it is important to analyse the correlation between gene expression and existing drugs. In our study, drug-sensitivity analysis revealed that the expression of NLRP7, NLRP2, NOD1, and CASP6 was positively correlated with some or most drugs in the cancer therapeutic response portal database (Fig. 7K).
A–E The correlation between five prognostic PRG and TMB in LUAD. F–J The correlation between five prognostic PRG and MSI in LUAD. K The correlation between five prognostic PRG and CTRP drug sensitivity in LUAD. TMB tumour mutation burden, MSI microsatellite instability, LUAD lung adenocarcinoma, PRG pyroptosis‑related gene, CTRP cancer therapeutics response portal.
Construction of a network of mRNA–miRNA–lncRNA
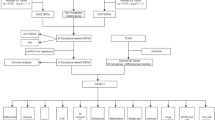
We also clarified the correlation between prognostic PRG and clinical stage, which revealed that NLRP1 (Fig. S2C, p = 0.0105) and NLRP7 (Fig. S2E, p = 0.00211) were correlated with clinical stage. However, there was no significant correlation between NOD1 (Fig. S2A, p = 0.339), CASP6 (Fig. S2B, p = 0.232) and NLRP2 (Fig. S2D, p = 0.645), and clinical stage. This suggested that NLRP1 and NLRP7 may be involved in tumour progression in LUAD. To clarify the potential molecular mechanism of NLRP1 and NLRP7 in LUAD, we then constructed a network of mRNA–miRNA–lncRNA interactions. The data identified miR-335-5p as the targeting mRNA binding to NLRP1 and NLRP7 according to mirTarBase and TarBase V.8 (Fig. 8A). Further analysis revealed that miR-335-5p was downregulated in LUAD (Fig. 8B, p = 0.00016), and LUAD patients with high miR-335-5p levels experienced better overall survival (Fig. 8C, p = 0.0328). According to this result, we also explored its upstream lncRNA targets to construct the miRNA–lncRNA axis. As shown in Fig. 8D, three lncRNAs, lncRNA XIST, lncRNA FTX, and lncRNA KCNQ1OT1, were identified as targets. The ceRNA network is shown in Fig. 8E. The expression of lncRNA targets was also detected, which revealed downregulation of lncRNA FTX (Fig. 8F, p = 5.9e−5) and upregulation of lncRNA KCNQ1OT1 (Fig. 8G, p = 7e−6) in LUAD compared with normal tissues. However, only lncRNA KCNQ1OT1 could reduce the LUAD patients’ survival probability (Fig. 8H, p = 0.0361). Thus, the lncRNA KCNQ1OT1/miR-335-5p/NLRP1/NLRP7 regulatory axis may play a vital role in the progression of LUAD.
A Results of miRNA target predicted by mirTarBase and TarBase V.8. The expression B and prognostic value C of miR-335-5p in LUAD. D Results of lncRNA targets predicted by lncBase predicted V.2 and StarBase V2.0. E The network of lncRNA–miRNA–mRNA. F-G The expression lncRNA FTX and lncRNA KCNQ1OT1 in LUAD. H The prognosis value of lncRNA KCNQ1OT1 in LUAD. LUAD lung adenocarcinoma; CeRNA competing endogenous RNA.
Discussion
Pyroptosis is a newly recognised type of programmed cell death that exerts a dual function in cancer progression and treatment mechanisms. Pyroptosis can release inflammatory factors and stimulate normal cells, resulting in transformation into tumour cells [15]. However, pyroptosis can promote tumour cell death, making pyrolysis a potential prognostic and therapeutic target for cancer [16]. In ovarian cancer, a novel PRG signature has been identified to predict prognosis [10]. However, the role of PRG in LUAD has not yet been elucidated, and our study was performed to clarify this role.
We first clarified the expression and prognostic value of PRGs in LUAD. We found that the expression of PRKACA, NOD1, NLRP1, ELANE, TNF, IL1B, IL18, PYCARD, CASP5, NLRC4, NLRP3, IL6, and CASP1 was increased, while the expression of GSDMB, PJVK, CASP4, NLRP7, CASP3, CASP6, CASP8, GSDMA, GSDMC, and AIM2 was decreased in LUAD compared with normal tissues. Prognosis analysis suggested a poor survival rate in LUAD patients with low expression of NLRP7, NLRP1, NLRP2, and NOD1 and high CASP6 expression. These data were consistent with prior results. Edward et al. suggested that low expression of NLRP1 was linked to a poor prognosis and immune infiltration in LUAD [17].
We also performed functional enrichment analysis of PRGs, which revealed that these 33 PRGs were mainly involved in the regulation of the inflammatory response, pyroptosis, NOD-like receptor signalling pathway, TNF signalling pathway, Toll-like receptor signalling pathway, and apoptosis. Interestingly, these functions or pathways were correlated with the oncogenesis and progression of LUAD. The induction of Th1-like and cytotoxic immunity by the TLR signalling pathway could result in lung cancer regression or arrest [18]. Moreover, a previous study showed that potentially functional genetic variants in TNF/TNFR signalling pathway genes were associated with prognosis in LUAD [19]. These results suggested that these 33 PRGs may also play a vital role in the oncogenesis and progression of LUAD.
LASSO Cox regression analysis was performed to construct a prognostic gene model based on five prognostic PRGs (NLRP7, NLRP1, NLRP2, NOD1, and CASP6), which could predict the overall survival of LUAD patients with medium-to-high accuracy. A predictive nomogram suggested that the 3‐year and 5‐year overall survival rates could be predicted relatively well compared with an ideal model in the entire cohort. A previous study identified several prognostic signatures for LUAD. A study performed by Sijin developed and validated an immune-related prognostic signature in LUAD [20]. Another glycolysis-related gene signature could predict metastasis and survival in LUAD patients [21]. Moreover, an autophagy-related prognostic signature showed good performance in LUAD patient prognosis prediction [22]. In our study, we first identified a pyroptosis-related prognostic gene signature for LUAD, which provides more choices for prognostic prediction in LUAD.
In our study, CASP6 was found to be one of the gene signatures. Although a previous study revealed that CASP6 could facilitate the activation of programmed cell death pathways, including pyroptosis, apoptosis, and necroptosis, CASP6 is typically not associated with pyroptosis [23]. CASP6 is generally considered to be a vital regulator of innate immunity, inflammasome activation, and host defence [23]. Increasing evidence has revealed that CASP6 is involved in carcinogenesis and progression by regulating the apoptosis and metastasis of tumours [24]. Moreover, active CASP6 is thought to be a potential therapeutic target against Alzheimer’s disease [25, 26]. This combined evidence suggested a broad role for CASP6. However, studies on the role of CASP6 in pyroptosis are limited. In our study, we found that CASP6 was one of the pyroptosis-related prognostic biomarkers in LUAD. Further in vivo and in vitro studies should be performed to verify whether CASP6 is involved in pyroptosis in LUAD.
Another important finding of our study revealed that the above five pyroptosis-related prognostic genes were significantly correlated with immune infiltration, which further confirmed the fact that pyroptosis plays a vital role in the tumour immune microenvironment. BRAF mutations could regulate the tumour immune microenvironment by regulating the pyroptosis-related signalling pathway [27]. A previous study also found that the pyroptosis gene NLRP1 is correlated with immune infiltration in LUAD [17].
We also constructed a mRNA–miRNA–lncRNA network, which identified a lncRNA KCNQ1OT1/miR-335-5p/NLRP1/NLRP7 regulatory axis. In fact, miR-335-5p could regulate the LUAD cell cycle and metastasis [28]. Moreover, miR-335-5p could suppress TGF-β1-induced EMT in lung cancer [29]. Interestingly, lncRNA KCNQ1OT1 could accelerate LUAD cell proliferation, migration, and invasion [30]. In our study, we also found that miR-335-5p and lncRNA KCNQ1OT1 were linked to the prognosis of LUAD patients. All this evidence suggests that the lncRNA KCNQ1OT1/miR-335-5p/NLRP1/NLRP7 regulatory axis may also play an important role in the progression of LUAD. Further study should be conducted to verify this result.
Our study has some limitations. All analyses were conducted using the TCGA LUAD cohort, and it would be better to verify them using the GEO cohort. Moreover, in vivo and in vitro experiments should be performed to further confirm our results.
In conclusion, we performed a comprehensive and systematic bioinformatics analysis and identified the pyroptosis‑related prognostic gene signature containing five genes (NLRP7, NLRP1, NLRP2, NOD1, and CASP6) for LUAD patients. Our results also identified a lncRNA KCNQ1OT1/miR-335-5p/NLRP1/NLRP7 regulatory axis, which may also play an important role in the progression of LUAD. Further study should be conducted to verify this result.
Materials and methods
Datasets and preprocessing
The RNA-sequencing (RNA-seq) data of 486 LUAD patients and the corresponding clinical information were obtained using The Cancer Genome Atlas (TCGA) database on April 1, 2021. The clinical information of the LUAD patients is shown in Table S1. Moreover, somatic datasets and copy number variation (CNV) data for LUAD were also downloaded from TCGA and the University of California, Santa Cruz (UCSC) Xena website, respectively. Data analysis was performed with the R (version 4.0.5) and R Bioconductor packages. The expression data were normalised to transcripts per kilobase million (TPM) values before further analysis.
Identification of differentially expressed PRGs
A total of 33 PRGs were obtained from prior reviews [10, 31], which are shown in Table S2. The difference in PRG expression in LUAD and normal tissues was identified using the “limma” and “reshape2” packages. We then constructed a protein–protein interaction (PPI) network for 33 PRGs using the Search Tool for the Retrieval of Interacting Genes (STRING).
Mutation analysis of PRGs
The mutation frequency and oncoplot waterfall plot of 33 PRGs in LUAD patients were generated by the “maftools” package. The location of CNV alteration of 33 PRGs on 23 chromosomes was drawn using the “RCircos” package in R.
Functional enrichment analysis
Gene Ontology (GO), including the biological process (BP), cellular component (CC), and molecular function (MF) categories, was conducted with the “ggplot2” package in R software. Similarly, this package was also utilised to perform Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis.
Development of the pyroptosis-related gene prognostic model
Cox regression analysis was performed to evaluate the prognostic significance of the PRGs. For Kaplan–Meier curves, p-values and hazard ratios (HRs) with 95% confidence intervals (CIs) were generated by log-rank tests and univariate Cox proportional hazard regression. PRGs with a significant prognostic value were selected for further analysis. Based on these prognostic PRGs, LASSO Cox regression analysis was then used to construct the prognostic model. The TCGA LUAD patients were divided into low- and high-risk subgroups according to the median risk score, and the overall survival (OS) time was compared between the two subgroups via Kaplan–Meier analysis. The predictive accuracy of each gene and the risk score were evaluated by performing time receiver-operating characteristic (ROC) analysis. Considering the clinical characteristics, a predicted nomogram was developed to predict the 1-, 3-, and 5-year overall survival. A forest was used to show the P-value, HR and 95% CI of each variable through the “forestplot” R package.
Immune infiltration, tumour mutation burden, and microsatellite-instability analysis
We then analysed the correlation between prognostic PRG and immune infiltration using the Tumour IMmune Estimation Resource (TIMER, https://cistrome.shinyapps.io/timer/), a web portal for comprehensive analysis of tumour-infiltrating immune cells. The “Gene” module of TIMER could visualise the correlation of gene expression with the immune infiltration level in LUAD. In tumour mutation burden (TMB) and microsatellite-instability (MSI) analysis, Spearman’s correlation analysis was performed to calculate the correlation between gene expression and TMB and MSI score. A p-value of less than 0.05 was considered statistically significant.
Competing endogenous RNA network construction
To clarify the potential function of PRG in LUAD, we then constructed a competing endogenous RNA (ceRNA) network. miRTarBase (http://mirtarbase.cuhk.edu.cn/) and TarBase V.8 (https://carolina.imis.athena-innovation.gr/diana_tools/web/index.php?r=tarbasev8%2Findex) were utilised to predict the miRNA targets binding to the PRGs. Based on the miRNAs identified, StarBase (http://starbase.sysu.edu.cn/) and LncBase Predicted v.2 (https://carolina.imis.athena-innovation.gr/diana_tools/web/index.php?r=lncbasev2/index-predicted) were utilised to predict lncRNA targets interacting with miRNAs. We also explored the expression and prognostic value of these miRNA and lncRNA targets using the TCGA LUAD dataset. All analyses were considered statistically significant at P < 0.05.
Data availability
The analysed data sets generated during the study are available from the corresponding author on reasonable request.
References
Bade BC, Dela Cruz CS. Lung cancer 2020: epidemiology, etiology, and prevention. Clin Chest Med. 2020;41:1–24.
Denisenko TV, Budkevich IN, Zhivotovsky B. Cell death-based treatment of lung adenocarcinoma. Cell Death Dis. 2018;9:117.
Greenawalt EJ, Edmonds MD, Jain N, Adams CM, Mitra R, Eischen CM. Targeting of SGK1 by miR-576-3p inhibits lung adenocarcinoma migration and invasion. Mol Cancer Res. 2019;17:289–98.
Hirsch FR, Scagliotti GV, Mulshine JL, Kwon R, Curran WJ Jr, Wu YL, et al. Lung cancer: current therapies and new targeted treatments. Lancet. 2017;389:299–311.
Shi J, Gao W, Shao F. Pyroptosis: Gasdermin-mediated programmed necrotic cell death. Trends Biochem Sci. 2017;42:245–54.
Schneider KS, Groß CJ, Dreier RF, Saller BS, Mishra R, Gorka O, et al. The inflammasome drives GSDMD-independent secondary pyroptosis and IL-1 release in the absence of caspase-1 protease activity. Cell Rep. 2017;21:3846–59.
Xia X, Wang X, Cheng Z, Qin W, Lei L, Jiang J, et al. The role of pyroptosis in cancer: pro-cancer or pro-“host”? Cell Death Dis. 2019;10:650.
Fang Y, Tian S, Pan Y, Li W, Wang Q, Tang Y, et al. Pyroptosis: a new frontier in cancer. Biomed Pharmacother. 2020;121:109595.
Tan YF, Wang M, Chen ZY, Wang L, Liu XH. Inhibition of BRD4 prevents proliferation and epithelial-mesenchymal transition in renal cell carcinoma via NLRP3 inflammasome-induced pyroptosis. Cell Death Dis. 2020;11:239.
Ye Y, Dai Q, Qi H. A novel defined pyroptosis-related gene signature for predicting the prognosis of ovarian cancer. Cell Death Disco. 2021;7:71.
Gao J, Qiu X, Xi G, Liu H, Zhang F, Lv T, et al. Downregulation of GSDMD attenuates tumor proliferation via the intrinsic mitochondrial apoptotic pathway and inhibition of EGFR/Akt signaling and predicts a good prognosis in non‑small cell lung cancer. Oncol Rep. 2018;40:1971–84.
Liu L, Bai X, Wang J, Tang XR, Wu DH, Du SS, et al. Combination of TMB and CNA stratifies prognostic and predictive responses to immunotherapy across metastatic cancer. Clin Cancer Res. 2019;25:7413–23.
Samstein RM, Lee CH, Shoushtari AN, Hellmann MD, Shen R, Janjigian YY, et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat Genet. 2019;51:202–6.
Chang L, Chang M, Chang HM, Chang F. Microsatellite instability: a predictive biomarker for cancer immunotherapy. Appl Immunohistochem Mol Morphol. 2018;26:e15–21.
Karki R, Kanneganti TD. Diverging inflammasome signals in tumorigenesis and potential targeting. Nat Rev Cancer. 2019;19:197–214.
Ruan J, Wang S, Wang J. Mechanism and regulation of pyroptosis-mediated in cancer cell death. Chem Biol Interact. 2020;323:109052.
Shen E, Han Y, Cai C, Liu P, Chen Y, Gao L, et al. Low expression of NLRP1 is associated with a poor prognosis and immune infiltration in lung adenocarcinoma patients. Aging. 2021;13:7570–88.
Pinto A, Morello S, Sorrentino R. Lung cancer and Toll-like receptors. Cancer Immunol Immunother. 2011;60:1211–20.
Guo Y, Feng Y, Liu H, Luo S, Clarke JW, Moorman PG, et al. Potentially functional genetic variants in the TNF/TNFR signaling pathway genes predict survival of patients with non-small cell lung cancer in the PLCO cancer screening trial. Mol Carcinog. 2019;58:1094–104.
Sun S, Guo W, Wang Z, Wang X, Zhang G, Zhang H, et al. Development and validation of an immune-related prognostic signature in lung adenocarcinoma. Cancer Med. 2020;9:5960–75.
Zhang L, Zhang Z, Yu Z. Identification of a novel glycolysis-related gene signature for predicting metastasis and survival in patients with lung adenocarcinoma. J Transl Med. 2019;17:423.
Zhu J, Wang M, Hu D. Development of an autophagy-related gene prognostic signature in lung adenocarcinoma and lung squamous cell carcinoma. PeerJ. 2020;8:e8288.
Zheng M, Karki R, Vogel P, Kanneganti TD. Caspase-6 is a key regulator of innate immunity, inflammasome activation, and host defense. Cell. 2020;181:674–e613.
Foveau B, Van Der Kraak L, Beauchemin N, Albrecht S, LeBlanc AC. Inflammation-induced tumorigenesis in mouse colon is caspase-6 independent. PLoS ONE. 2014;9:e114270.
Angel A, Volkman R, Royal TG, Offen D. Caspase-6 knockout in the 5xFAD model of Alzheimer’s disease reveals favorable outcome on memory and neurological hallmarks. Int J Mol Sci. 2020;21:1144.
Klaiman G, Petzke TL, Hammond J, Leblanc AC. Targets of caspase-6 activity in human neurons and Alzheimer disease. Mol Cell Proteom. 2008;7:1541–55.
Erkes DA, Cai W, Sanchez IM, Purwin TJ, Rogers C, Field CO, et al. Mutant BRAF and MEK inhibitors regulate the tumor immune microenvironment via pyroptosis. Cancer Disco. 2020;10:254–69.
Wang X, Xiao H, Wu D, Zhang D, Zhang Z. miR-335-5p regulates cell cycle and metastasis in lung adenocarcinoma by targeting CCNB2. Onco Targets Ther. 2020;13:6255–63.
Du W, Tang H, Lei Z, Zhu J, Zeng Y, Liu Z, et al. miR-335-5p inhibits TGF-β1-induced epithelial-mesenchymal transition in non-small cell lung cancer via ROCK1. Respir Res. 2019;20:225.
Wang Y, Zhang L, Yang J, Sun R. LncRNA KCNQ1OT1 promotes cell proliferation, migration and invasion via regulating miR-129-5p/JAG1 axis in non-small cell lung cancer. Cancer Cell Int. 2020;20:144.
Latz E, Xiao TS, Stutz A. Activation and regulation of the inflammasomes. Nat Rev Immunol. 2013;13:397–411.
Acknowledgements
This study was funded by the Science and Technology Special Fund of Maoming (2020KJZX016), the Science and Technology Project of Guangdong Esophageal Cancer Institute (M201914) and the Science and Technology Special Fund of Guangdong (2020S00061).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by I. Lavrik
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lin, W., Chen, Y., Wu, B. et al. Identification of the pyroptosis‑related prognostic gene signature and the associated regulation axis in lung adenocarcinoma. Cell Death Discov. 7, 161 (2021). https://doi.org/10.1038/s41420-021-00557-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-021-00557-2
- Springer Nature Limited
This article is cited by
-
Prognostic model revealing pyroptosis-related signatures in oral squamous cell carcinoma based on bioinformatics analysis
Scientific Reports (2024)
-
Machine learning developed a CD8+ exhausted T cells signature for predicting prognosis, immune infiltration and drug sensitivity in ovarian cancer
Scientific Reports (2024)
-
CRTAC1 identified as a promising diagnosis and prognostic biomarker in lung adenocarcinoma
Scientific Reports (2024)
-
Identification and bioinformatics analysis of genes associated with pyroptosis in spinal cord injury of rat and mouse
Scientific Reports (2024)
-
Gefitinib metabolism-related lncRNAs for the prediction of prognosis, tumor microenvironment and drug sensitivity in lung adenocarcinoma
Scientific Reports (2024)