Abstract
In plants, proteolysis is emerging as an important field of study due to a growing understanding of the critical involvement of proteases in plant cell death, disease and development. Because proteases irreversibly modify the structure and function of their target substrates, proteolytic activities are stringently regulated at multiple levels. Most proteases are produced as dormant isoforms and only activated in specific conditions such as altered ion fluxes or by post-translational modifications. Some of the regulatory mechanisms initiating and modulating proteolytic activities are restricted in time and space, thereby ensuring precision activity, and minimizing unwanted side effects. Currently, the activation mechanisms and the substrates of only a few plant proteases have been studied in detail. Most studies focus on the role of proteases in pathogen perception and subsequent modulation of the plant reactions, including the hypersensitive response (HR). Proteases are also required for the maturation of coexpressed peptide hormones that lead essential processes within the immune response and development. Here, we review the known mechanisms for the activation of plant proteases, including post-translational modifications, together with the effects of proteinaceous inhibitors.

Similar content being viewed by others
Facts
-
Plant proteases are involved in various biological processes, from organ development to plant biotic and abiotic responses.
-
By processing their substrates, proteases control multiple signaling mechanisms, such as the generation of mature plant peptide hormones, relevant in plant adaptation to their environment and cell-to-cell communication.
-
Unleashed proteolysis can result in disease and cell death; therefore, proteases are tightly controlled by different means in their cellular environment.
-
Proteolysis can also be modulated from a substrate-centered point of view; e.g. post-translational modifications of the substrates can prohibit their cleavage.
-
Proteases take a prominent role in plant-pathogen interactions by modulating the plant defense response often leading to a type of cell death called hypersensitive response (HR).
Open questions
-
What are the substrate landscapes for plant proteases and what are their spatiotemporal dynamics?
-
What are the factors triggering protease activities? Extended knowledge of their regulation will help to pinpoint their roles in vivo.
-
How are the regulatory mechanisms that control plant protease activity integrated with the spatiotemporal availability of the substrates?
Introduction
Proteases are abundant enzymes present in every life kingdom with the capacity to hydrolyze peptidyl bonds between two amino acids in their substrates [1]. This irreversible post-translational modification (PTM) results in the formation of new carboxyl and amino termini in the cleaved substrate. Through this proteolytic processing, substrates may lose, gain or alter their function and, in addition to proteases roles in protein degradation, they are often key regulatory players within signaling cascades [2,3,4,5]. By cleaving a substrate, proteases can act as molecular “switches” that activate or inactivate a specific cellular process. In plants, the number of proteins with a predicted peptidase activity represents a large part of the genome. For example, in Arabidopsis thaliana (Arabidopsis) and Populus sp.(poplar) 570 and 955 proteases are annotated, respectively [6,7,8]. Plant proteases are involved in a wide pallet of processes like organellar protein import [9], programmed cell death [10, 11], growth and development [12, 13] responses to abiotic stresses, immunity and the hypersensitive response (HR) [14, 15]. However, in contrast to animal proteases, the number and identity of plant protease substrates remain largely unknown.
Proteases are often produced as zymogens, inactive proenzymes that are activated by a particular trigger in a specific cellular context. Many zymogens contain a so-called prodomain that blocks the accessibility to the active site. Removal of this prodomain generally requires limited proteolysis in cis or trans of the zymogen [16] and after cleavage, structural changes such as dimerization or complex assembly are required to permit proteolytic catalytic activity [17,18,19,20,21]. Reciprocally, proteases can be controlled by physical interaction with promiscuous protease inhibitors [22]. Sometimes, like for serpins, this inhibitor-protease interaction occurs only after activation of the protease [23, 24]. Other proteases, such as the Tobacco Etch Virus protease, are limiting their own lifetime and activity by destructive self-processing [25].
In summary, the proteases are regulated at multiple levels, starting from spatio-temporal gene expression, enzymatic activation by cellular stimuli or PTMs, and the tempering or destruction of their activity by proteinaceous inhibitors. As proteolysis is irreversible, such a multi-layered control system guards unwanted or precocious cleavage of substrates and of off-target proteins. Here, we discuss the current knowledge related to activating mechanisms and mode of action of plant proteases, together with their proteinaceous inhibitors that control proteolytic activity, mainly within the context of signal transduction events.
Protease regulation by calcium, pH and redox
Changes to the cellular environment can activate protease zymogens. For instance, most plant metacaspases depend on an elevated concentration of calcium ions for their activation [26,27,28,29,30,31]. Metacaspases have similar structural features to mammalian caspases, including a Cys-His catalytic dyad and a caspase-hemoglobinase fold [32]. However, both the activation mechanism and substrate preference of metacaspases and caspases are different. Caspases cleave their substrate after Asp residues, whereas metacaspases prefer to cleave after Arg or Lys [26, 33, 34]. This difference in cleavage specificity is explained by the conformation of the catalytic pockets; in caspases an Arg, Gln, Arg triad creates a basic pocket around the substrate binding site (S1) that efficiently binds to the Asp residue in the substrate [35]. In metacaspases, the catalytic pocket entails Asp, Asp/Glu and Asp, generating an acidic S1 microenvironment that is well suited to accept the basic residues Arg and Lys [36, 37]. Most metacaspases display the p20 and p10 conserved regions of 20- and 10-kilodalton sizes respectively, which are joined by a linker that varies in length and primary structure [32, 38,39,40]. Based on their p20 and p10 arrangement and presence of additional features, plant metacaspases are divided in two types. Type-I metacaspases contain an N-terminal prodomain, which in Arabidopsis was shown to physically interact with the zinc-finger protein LSD1, a negative regulator of cell death. In response to bacterial infection, AtMC1 acts as a positive regulator, whereas a second type-I metacaspase, AtMC2, has an antagonistic effect on cell death. Intriguingly, whereas the catalytic activity of AtMC1 is required for cell death induction, the catalytic residues in AtMC2 are not necessary for its inhibitory action [41, 42]. The trigger and mechanism leading to AtMC1 activation in response to bacteria remain unresolved, but it was demonstrated that the suicide inhibitor SERPIN1 blocks both AtMC1-mediated cell death and AtMC1 autocatalytic processing in planta [24]. Type-II metacaspases lack a prodomain, but entail a longer linker between the p20 and p10 regions. Most of the studied type-II metacaspases require Ca2+ concentrations in the millimolar range to be activated. Recently, the crystal structure of Arabidopsis AtMC4 revealed critical insights into the Ca2+-dependency of its activation mechanism [28]. A negatively charged region in the linker region proximal to an internal Lys is hindering the Cys-His catalytic pocket. Upon calcium binding, this Lys side chain approaches the catalytic cysteine, gets cleaved and initiates subsequent cleavages in other sites of the linker, thereby leading to AtMC4 activation. Within a biological context, Ca2+ levels in the millimolar range are observed during wounding stress, which can be caused by herbivory, insect chewing or stinging, and during mere physical damage. Cell rupture leads to calcium influxes and accumulation locally in the wounded and surrounding cell, where AtMC4 first self-processes and subsequently cleaves the immunomodulatory peptides PROPEPs, which are damage/danger-associated molecular patterns (DAMPs) in immunity responses [30, 43]. The mature PROPEPs, called Peps, bind to the cell-surface receptors PEPR1 and PEPR2, that with their coreceptor BRI1-ASSOCIATED RECEPTOR KINASE 1 (BAK1) activate a signaling cascade, leading to the transcriptional activation of defense/immune response genes [44]. It remains an open question whether other conditions, like bacterial or fungal infections and exposure to abiotic stresses, can elicit calcium fluxes that are sufficient to activate Ca2+ -dependent metacaspases. In Arabidopsis, AtMC9 is involved in xylem formation, a genetically controlled process in which protoxylem cells require clearance of their cellular content and thickening of their cell walls to become suitable vessels within the vascular tissue [45]. Unlike the other Arabidopsis type-II metacaspases, the activation mechanism of AtMC9 is calcium-independent and exhibits optimal processing in acidic conditions between pH 5 and 6 [26]. AtMC9 self-processing is impeded by S-nitrosylation, but this redox-dependent PTM of the catalytic cysteine is not affecting its activity towards peptidic substrates. This can be explained by the role of a second Cys that is not S-nitrosylated and that is positioned proximal to the catalytic pocket and, together with the catalytic His, can preserve AtMC9 proteolytic activity [46]. In addition, AtMC9 can be irreversibly blocked by SERPIN1 [23]. SERPINs contain a reactive center loop, which is cleaved by AtMC9, resulting in a covalent complex between inhibitor and protease at the catalytic pocket, and thereby inactivating AtMC9 [47]. Using an N-terminomic approach, a plethora of potential AtMC9 substrates was identified and revealed that the cleavage of AtMC9 after Arg or Lys, is preferentially followed by Glu or Asp residues at the P1 position in proteinaceous substrates [48]. Future structural insights into AtMC9 will be instrumental to depict a model explaining its pH-dependent activation, the mode of AtMC9-SERPIN1 interaction and the redox-dependent control of the catalytic residues. In summary, plant metacaspases remain inactive in resting conditions, can be activated by calcium or pH changes, and are regulated by PTMs or SERPINs (Fig. 1A).
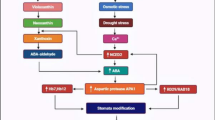
A Type-II metacaspase activation can occur via calcium increase or pH drop. Calcium-dependent metacaspases are activated through conformational changes in their structure triggered by stresses such as wounding, whereas the pH-dependent activation mechanism remains to be elucidated. Metacaspases are also regulated by nitrosylation of their catalytic cysteine, or by SERPIN after their self-processing. During wound stress, activated metacaspases cleave PROPEPs to Peps, allowing binding of their receptors PEPR1/2 to the coreceptor BAK1. B Arabidopsis legumains are able to switch from proteolytic to ligase activity. At lower pH, legumains function as peptidases, while at neutral pH, they mainly act as a ligase. Still, in an intermediate status, they can combine both activities. This flexibility allows legumains to work as different enzymes depending on the pH of the suborganelle in which they are located. C The cysteine proteases ATG4A and ATG4B control processing of ATG8, which is induced under environmental stimuli like nitrogen starvation and osmotic stress. Upon cleavage of ATG8, it can recruit adaptor and cargo proteins to the autophagosome (brown) and direct these proteins to lytic vacuoles. DArabidopsis S1P (SBT6.1) processes RALF precursors containing a dibasic motif. Processed RALF peptides act as negative regulators of immune responses binding to its receptor FERONIA (FER) and diminishing the perception and responses in conjunction with the bacteria-derived peptides elf18 and flg22.
Proteolytic activity change by mono/dimerization
Legumains, like the metacaspases, are also cysteine proteases structurally related to caspases and preferentially cleave substrates after Asn or Asp residues. In Arabidopsis, four legumains (AtLEGα, β, γ and δ) have been identified, all with both a ligase and protease activity [49]. AtLEGγ has a unique activation mechanism: in neutral pH, an alpha helix, called activation peptide, is stabilized and dimerization blocks the access to the pocket of the active site. In these conditions, AtLEGγ acts predominantly as a ligase. In protonated environments, the forces acting on the activation peptide repel it from the active site, diplace it and enables its processed. Surprisingly, after cleavage, both the catalytic domain and the C-terminal prodomain, referred to as legumain stabilization and activity modulation (LSAM) domain, remain together through disulfide bridges. At intermediate pH levels, both ligase and protease activities coexist. Such a mechanism with a two-chain intermediate state is unique to plant legumains. of which the type of enzymatic activity is controlled by the monomer-dimer protein status, which is mainly affected by the organellar pH (Fig. 1B).
Redox regulation of ATG4
Other cysteine proteases, like the autophagy-related protein 4 cysteine proteases A and B (ATG4A/B), are responsible for the processing of the C-terminal end of the ubiquitin-like protein ATG8. This cleavage event exposes a glycine residue at the neo-C-terminus that enables ATG8 cargo binding in the nascent autophagosome and their transport to the lytic vacuole (Fig. 1C). ATG4 was the first autophagy-related protein reported to be redox regulated [50,51,52]. In Arabidopsis, the in vitro proteolytic activities of both ATG4A and ATG4B on an ATG8-based synthetic substrate are reversibly inhibited by H2O2 [51]. In yeast and the green algae Chlamydomonas reinhardtii, reduction by thioredoxins of a regulatory disulfide bond outside the catalytic region was identified as the regulatory mechanism [53]. Within a more oxidized cellular context, for instance promoted by adverse environmental stress conditions, the disulfide bond is formed and inhibits the ATG4 activity. Mutation of one of the cysteines involved in the disulfide formation turns ATG4 redox-insensitive and increases its activity towards its substrate ATG8. In addition, persulfidation of a Cys residue within the catalytic Cys-His-Asp triad also inhibits its protease activity in a reversible manner [54]. Here, the regulation of ATG4 activity by its redox status shows the importance of controlled proteolysis in a conserved process such as autophagy.
Proteolysis in phytocytokine maturation
Plant peptide hormones involved in cell-to-cell communication are known as phytocytokines [55]. Many of these peptides, such as Pep1, require proteolytic processing prior to maturation before becoming bioactive [56, 57]. Phytaspase, a subtilisin-like protease, cleaves substrates after Asp residues, similar to caspases [58]. Tobacco phytaspase can self-cleave its prodomain before processing and maturation of the propeptide systemin [59]. Systemin is the first ever identified plant peptide hormone, originally discovered in Solanum lycopersicum (tomato). The active form entails 18 amino acids and is induced during wound responses [60]. Phytaspase-mediated cleavage from its 200-amino acid precursor prosystemin, releases first the intermediate Leu-Systemin. For full functional systemin, the leucine at the N-side requires to be removed, likely by a leucine aminopeptidase, which is also induced by wounding [61]. Systemin is then recognized by the systemin receptor like kinase (SYR1), inducing an oxidative burst and ethylene production [62]. An interesting regulatory aspect of tobacco phytaspases is that under specific circumstances, like cell death, it relocalizes from the apoplast into the cytoplasm to cleave endogenous substrates [63]. Methyl viologen treatment and viral infection induce remobilization of the apoplastic phytaspase to the cytoplasm in clathrin-coated vesicles. The particularity of this endocytic process is that it seems to be selective for phytaspase, while other proteases at the extracellular space, such as cathepsins, are not remobilized [63, 64]. Whether phytaspase relocalization also takes place during wound responses has not been reported.
Rapid alkalinization-like factor (RALF) peptides were first discovered in tobacco as small-size peptides proteolytically processed after a dibasic substrate motif [65]. In Arabidopsis, a third of the RALF family members present a canonical cleavage site processed by site-1 protease (S1P), also named SBT6.1 [66, 67]. Additionally, S1P processes other substrates than RALF peptides, such as the ER membrane-bound transcription factor bZIP17, which relocalizes to the nucleus after processing during the ER stress response [68], and pectin methylesterases, whose N-terminal regions need to be released from the Golgi in order to reach their final destination at the cell wall [69]. Arabidopsis RALF23 is processed by S1P and perceived by the receptor-like kinase FERONIA and LORELEI-LIKE-GPI-ANCHORED PROTEIN 1. RALF23 perception represses the complex formation of the receptors of pathogen-associated molecular patterns flg22 and elf22, FLAGELLIN-SENSITIVE 2 (FLS2) or EF-TU RECEPTOR (EFR), respectively, with their coreceptor BRI1-ASSOCIATED RECEPTOR KINASE (BAK1). By this action, S1P-processed RALF23 negatively regulates plant immunity, leading to a reduction of the reactive oxygen species production and increased bacteria colonization [70, 71] (Fig. 1D).
Sometimes, multiple proteases can act synergistically or sequentially in order to produce mature peptides. For instance, the overexpression phenotype of CLAVATA3 EMBRYO SURROUNDING REGION (CLE)-like family peptides (CLEL) is controlled by SBT6.1 (S1P) and SBT6.2 [72]. The curvy root phenotype produced by the CLEL GOLVEN1 (GLV1) is lost in the subtilase mutants. The expression patterns of the protease inhibitor SERPIN1 and SBT6.1 partially overlap and both proteins interact. Moreover, SERPIN1 reduces GLV1 processing in vitro and SERPIN1 overexpression in Arabidopsis reduces GLV1-induced hypocotyl elongation. In an independent study, CLEL peptide processing at the conserved motif by S1P/SBT6.1 was reported to occur at the Golgi [73]. After S1P processing, CLEL3 and CLEL9 are additionally processed in the apoplastic space by the subtilase SBT3.8, whose activity is regulated by the lower pH in the extracellular space [73].
The trio of subtilases SBT1.4, SBT1.7 and SBT4.13 processes the precursor of the Arabidopsis CLAVATA3/ESR-RELATED 40 (CLE40) at two different sites. Mature CLE40 contributes to root stem cell maintenance and, interestingly, the cleavage of the second site is blocked by proline hydroxylation, a common PTM in many secreted peptides, and hence modulating the peptide bioactivity [74]. This is a nice example on how protease-substrate interactions are also regulated of a substrate-centric way. Another example of substrate-centered regulation is SFH8 processing by separase [75]. Separases are cysteine proteases, reported to be part of the KINESIN-SEPARASE Complex [76]. Both separase and kinesin are corecruited to the plasma membrane by SFH8, a lipid-like transferase. Once at their final location, SFH8 is cleaved by separase. Proteolysis of SFH8 leads to the formation of filamentous states at polar domains. SFH8 localization in mature cells resembles a droplet-like structure and a reduced processing, hinting towards a restriction of proteolysis based on its subcellular localization.
CASPARIAN STRIP INTEGRITY FACTORs (CIFs) are a family of sulfated peptides, including TWISTED SEED 1 (TWS1), which are cleaved at their C-termini by the subtilisin-like serine protease ABNORMAL LEAF SHAPE 1 (ALE1) [77]. In seeds, ALE1 is expressed in the endosperm [78], and the TWS1 receptors GASSHO1 and 2 (GSO1/2) are allocated at the opposite side of the nascent cuticle [79]. The TWS1 precursor and ALE1 are expressed at different cell types but can encounter each other in a shared extracellular space, allowing TWS1 processing. Mature TWS1 initiates the process that through GSO1/2 ends up in cuticle formation, creating a hydrophobic extracellular barrier that spatially confines the previously shared space. In such manner, ALE1 and TWS1 interaction is interrupted once the cuticle is built and the process shuts down itself. Although ALE1 is necessary for TWS1 peptide maturation, it is not the only protease able to process TWS1. A recent study shows that Arabidopsis SBT1.8 transcriptionally coexpresses with ALE1 in developing seeds, and it is capable to process TWS1 at both flanking sites of the mature peptide [80]. The N-terminal processing by SBT1.8 requires a sulfotyrosine at the P2 position after cleavage, as the protease is unable to process TWS1 bearing a natural tyrosine. This PTM dependency is explained by the interaction of the negatively charged sulfotyrosine with an Arg302 in SBT1.8. Interestingly, mutation of this Arg abolishes N-terminal cleavage but does not affect the SBT1.8 capacity to process the C-terminal part of the peptide. Therefore, tyrosine sulfation ensures an appropriate docking and processing in pair with SBT1.8 and increases the processing specificity during TWS1 maturation. This case illustrates that PTMs of substrates can be critical in the regulation of their interactions, including protease accessibility to potential substrates and can condition their proteolytic hydrolysis.
Abscission of flower organs and other cell separation processes like lateral root emergence depend on the peptide hormone Inflorescence Deficient in Abscission (IDA) [81]. Mutant ida plants fail to drop their sepals and petals after fertilization and revealed a role of the processed peptide in cell wall loosening during lateral root emergence [82]. When oomycetic extracellular protease inhibitors (EPI), that inhibit apoplastic phytaspase in tomato, are transgenically expressed under control of the IDA promotor in Arabidopsis, both petal and anther detachments are impaired due to impaired subtilase activity and consequent IDA maturation (Fig. 2A). A phenotype that was reconstituted by local application of mature IDA peptides. Along the same line, gene expression patterns of a cohort of subtilases (SBT2.2, SBT2.6, SBT3.1, SBT4.6, SBT4.8, SBT4.10, SBT4.12, SBT4.13 and SBT5.2) overlap with IDA expression in the basipetal zone during flower development [83,84,85]. Timely flower abscission in tomato plants is critical for optimal fruit production and yield. Flower abscission is controlled by the drought-inducible SlPhyt2, a subtilisin-like phytaspase, with proteolytic preference after Asp residues [86]. Drought-induced abscission in tomato is independent of the plant phytohormones auxin and ethylene, but instead depends on the peptide hormone phytosulfokine (PSK). PSK is cleaved at a conserved Asp by SlPHYT2 and exogenous addition of the mature PSK to SlPHYT2-silenced tomato plants presented a normal abscission phenotype [87] (Fig. 2B). In both abscission processes for tomato flower organs and Arabidopsis petal drop, cell separation is the final causative process, showing that peptide maturation of different peptide hormones and responses through independent receptors can converge on parallel abscission mechanisms in various plant organs [88,89,90].
A A broad protein subtilase inhibitor expressed under the control of the IDA promoter abolishes dehiscence of petals and anthers during fertilization stages, marked with black arrows. Candidate subtilases were identified by expression analysis using specific promoter region fusions with the β-glucuronidase (GUS) reporter in the basal apex of Arabidopsis flowers of possible responsible executors of IDA processing. B Drought-stressed tomato plants produce lower number of flowers in the inflorescence, marked with black arrows. Drought stress is responsible for the expression of SlPhyt2 in leaves and regions proximal to flower buds during their development. SlPhyt2 cleaves PSK and initiates a series of processes leading to cell separation and flower loss. C Subtilases expressed at the contact points of the parasitic plant Phtheirospermum japonicus assist in the early events of plant interaction with its host and xylem bridge (XB) formation, through expression of PJICSL1 genes and XB formation preserving auxin levels.
Post-Translational Modifications and their spatial context affect protease activities, substrates and their relationships
DA1 is a peptidase, named after the Chinese word “big” (大: phonetically pronounced “dà”), that together with its family members DA1-related 1 and 2 affects organ growth by regulating endoreduplication [91]. Its C-terminus encodes a zinc metallopeptidase that is controlled by a “cysteine-switch”: an intramolecular complex between a cysteine in the prodomain and a zinc atom blocking the active site that can be released by mono-ubiquitination of DA1 at multiple sites by E2 and E3 ligases. Subsequently, active DA1 can process its substrates like UBIQUITIN-SPECIFIC PROTEASE 15 (UBP15), TEOSINTE BRANCHED/CYCLOIDEA/ PCF 14 and 15 (TCP14/15) and TCP22 [92] and BIG BROTHER (BB) [93]. The activity of DA1 and homologs is further controlled by the ubiquitin proteases UBP12 and UBP13 that can deubiquitinate DA1 [94]. Furthermore, in the presence of brassinosteroid phytohormones, DA1 is phosphorylated in a BRI1-BAK1-dependent manner, which deactivates its enzymatic function and stabilizes its substrates in vivo [95]. More recently, DA1 and homologs were found to cleave TRANSMEMBRANE KINASE 1 (TMK1) at its C-terminus in a motif similar to BB [96]. Accumulation of the plant growth-regulating hormone auxin leads to the cleavage of TMK1 and relocalization of the cytosolic part from the membrane to the nucleus. There it interacts with the transcriptional repressors INDOLE-3-ACETIC ACID INDUCIBLE 32 (IAA32) and IAA34, ultimately controlling the formation of the apical hook of a developing seedling [4].
The role of proteases in plant-plant and plant-pathogen interactions
Some parasitic plants have developed a dependency on other plants to initiate germination and growth [97]. After attachment and penetration to their hosts, parasitic plants develop specialized organs to access the host vasculature and nutrients (Fig. 2C). Recent studies in P. japonicus identified four subtilases expressed specifically in these intrusive cells during colonization (SBT1.1.1, SBT1.2.3, SBT1.7.2, and SBT1.7.3). Inhibition of subtilase activity by expressing EPIs at the host-parasite contact point leads to reduced levels of colonization, lower expression of the intrusive cell marker INTRUSIVE CELL-SPECIFIC LEUCINE-RICH REPEAT RECEPTOR-LIKE KINASE1 (PjICSL1) and lower auxin levels, necessary for xylem bridge formation. Hence, proteolytic activity of these subtilases is required for a successful colonization, through unidentified substrates and mechanisms facilitating the parasitism relationship [98].
Early detection of pathogen infection and an adequate response are crucial for plant survival. Plants make use of small signaling peptides to sense bacteria [55, 99, 100], and proteases, both at the plant and pathogen side, are involved in the generation of these peptides. Zip1 (Zea mays immune peptide 1) is a maize peptide that is produced after salicylic acid (SA) treatment [101]. Mutational analysis and specific inhibitor applications confirmed that the two papain-like cysteine proteases (PLCPs) CP1 and CP2 are responsible for Zip1 maturation. Zip1 application induces the expression of SA-dependent genes and promotes additional cleavage of its precursor, proZip1, by an unknown positive feedback loop. The authors suggested that Zip1 might work by activating the proteases CP1 and CP2 by binding to an exosite and promoting cleavage of proZip1. The activation of SA-like responses could be explained as the evolution of an independent parallel mechanism that results in the activation of genes involved in SA responses. In fact, SA responses are often targeted and suppressed by bacterial effectors (Fig. 3A). For example, some biotrophic pathogens, such as Ustilago maydis, impair SA accumulation by secreting a chorismate mutase that lowers SA precursor availability and thereby minimizes plant defense responses [102]. Maize PLCPs are regulated by other effectors, including the U. maydis effector Pit2, and endogenous plant protease inhibitors such as the cystatin ZmCC9 [103, 104]. The existence of multiple PLCP control mechanisms are likely reflective of their importance in plant pathogen defense.
A Plant biotrophs induce SA responses in Z. mays, thereby activating the PLCPs CP1 and CP2, which cleave the precursor of Zip1 (ProZip1). Zip1 can provoke similar responses to SA, including transcriptional induction of SA-dependent genes and activation of their processing proteases CP1 and CP2 in a positive feedback loop. Zip1-proteolytic activation remains to be confirmed by direct interaction with CP1 and CP2. A Zip1 SA-independent pathway disentangles maize plants from the necessity to generate SA, which in some situations can be dampened by bacterial effectors such as the Ustillago maydis chorismate mutase 1 (Cmu1). B PrpL and ArgC bacterial protease homologs from P. aeruginosa and X. campestris trigger phosphorylation of MAPK3 and MAPK6. This novel mechanism couples G protein interaction with RACK1C and a phosphorylation cascade after PrpL and ArgC proteolysis. C ASP1 and ASP2 process the extracellular bacterial protein MucD, which contributes to bacterial growth. By degrading MucD, plants can keep at bay the growth of colonizing bacteria in the plant apoplast, maintaining a balance between both pathogenic and commensal bacteria.
Infection with P. aeruginosa, an opportunistic pathogen of plants and animals, can induce an immune response in Arabidopsis plants, for which a Pseudomonas serine protease (PrpL) and its homolog ArgC in Xanthomonas are responsible for the induction of a set of defense genes [105]. Through a screening of a collection of P. aerigusosa mutants, PrpL was identified to trigger the oxidative burst and the RECEPTOR FOR ACTIVATED C KINASE 1 (RACK1) was discovered to work as a scaffold in the phosphorylation cascade, similar to flg22 responses, upon protease perception (Fig. 3B). How plants perceive PrpL/ArgC proteases remains an outstanding question and it is not known whether there is a direct receptor for PrpL/ArgC or rather a detection mechanism for PrpL/ArgC substrates.
Some plant proteases cleave bacterial proteins to redundantly control their growth during invasion. This is the case for the SECRETED ASPARTIC PROTEASES 1 and 2 (SAP1 and SAP2), which process MucD, a bacterial conserved HtrA-like protease required for bacterial growth [106]. Although this processing would not discriminate between pathogenic and beneficial microorganisms on the surface of the plants, the authors suggested that it could work as a surveillance mechanism to keep excessive bacterial growth at bay (Fig. 3C).
Detection of pathogen infection by proteolysis leading to the hypersensitive response
As indicated above, proteolysis during plant-pathogen interactions is used at both sides of the plant-pathogen frontline. Some bacterial effectors, once injected in the plant host environment, dampen the plant defense responses to allow the pathogen to remain unnoticed as long as possible. Undetected pathogens are more likely to spread through the plant tissues, reproduce and colonize other parts of the plant. However, plants can recognize proteolytic products of bacterial or plant origin and thereby induce a defense response. Within incompatible biotrophic plant-pathogen interactions, the plant launches an HR [15], with proteolysis being involved on both sides of the host-pathogen divide. A case in point is the recognition of pathogenic effectors via the membrane protein RPM1 INTERACTING PROTEIN 4 (RIN4). Multiple effectors target RIN4, disrupting its interaction with RPM1 and its HR induction, and reducing pathogen growth [107]. RIN4 is also a target of AvrRpt2, a bacterial cysteine protease that is injected in the plant cell through a typical type-3 secretion system. In the presence of AvrRpt2, RIN4 is degraded, resulting in the perception of processed RIN4 by the NB-LRR RESISTANT TO P. SYRINGAE 2 (RPS2), which subsequently activates the HR [108]. RIN4 degradation is caused through direct cleavage by AvrRpt2 at two conserved motifs at the N and C-terminal parts [109]. Cleavage at the C-terminal site turns out to be indispensable for RIN4 release from the membrane domain and subsequent degradation, which activates RPS2, while cleavage of the N-terminal motif has no effect in the plant response (Fig. 4A). Another effector protease from P. syringae (AvrPphB) works inside plant cells where it processes key kinases of pathogen presence like BOTRYTIS-INDUCED KINASE1 (BIK1), PBS1-LIKE 1 (PBL1) and PBL2 [110]. Another target of AvrPphB is PBS1, a plant kinase that triggers HR and whose cleavage products are perceived by RESISTANCE TO P. SYRINGAE 5 (RPS5) [111]. Immunoprecipitation experiments showed that both active and inactive versions of AvrPphB are able to bind PBS1. The cleavage site was determined using Edman sequencing and identified a glycine-aspartic-lysine (↓GDK) motif, showing possible preference for glycine after cleavage, as AvrPphB self-processing occurs between a lysine and a glycine (K ↓ G). Although the GDK motif is conserved and found in other plant kinases, its presence is necessary but not sufficient for substrate cleavage. RPS5 is a nucleotide-binding leucine-rich repeat (NLR) protein recognizing AvrPphB upon infection [112]. RPS5 contains an N-terminal coiled-coil and nucleotide-binding site domains indispensable for the HR, and a C-terminal LRR domain that inhibits HR in the absence of infection and in the presence of uncleaved PBS1. In the absence of the effector, RPS5 and PBS1 remain in a pre-assembled complex, while in the presence of the effector, RPS5 detects the complex of the cleaved PBS1 and AvrPphB (Fig. 4B). The LRR domain binds to the C-terminal part of PBS1, possibly retaining AvrPphB, which leads to a change in the structure of RPS5. That physical change switches RPS5A affinity from ADP to ATP, with subsequent changes in direct partner interactions that guides the plant towards HR [113].
A RPS2-dependent HR induction after RIN4 cleavage by AvrRpt2. RIN4 is processed by AvrRpt2 at both the RCS1 and RCS2 sites, leading to the separation of RIN4 from the membrane and subsequent degradation, and allowing the release of RPS2 to initiate an HR response. B Perception of the cysteine protease AvrPphB from P. syringae pv. phaseolicola. RPS5 and PBS1 are interacting upon normal conditions in a primed status. During infection, the effector AvrPphB cleaves PBS1, which triggers a conformational change in the decoy RPS5 and triggers HR after exchange of ADP by ATP. C Perception mechanism of the Avr2 effector of C. fulvum in tomato plants. RCR3 PLCP is secreted to the apoplast as a zymogen, where subtilases like P69B process it by release of the RCR3 prodomain, generating a mature RCR3 (mRCR3). When Avr2 is secreted to the apoplast from the fungi, it targets the active mRCR3 and forms a complex with it. This complex is recognized by Cf-2 only in the presence of both RCR3 and Avr2 and only then Cf-2 activates HR.
In tomato, the fungal effector Avr2 targets the papain-like cysteine protease REQUIRED FOR C. FULVUM RESISTANCE 3 (RCR3) [114]. Initially, the secreted RCR3 was thought to be self-activated in a pH-dependent manner. However, the RCR3 prodomain was still processed in transgenic plants expressing a catalytically inactive RCR3. A group of subtilases, including P69B, was reported to be capable of hydrolyzing the RCR3 prodomain in trans [114]. Mature RCR3 (mRCR3) is capable to interact with the fungal effector Avr2, shaping a complex that is recognized by the leucine-rich repeat receptor Cf-2 and culminates in localized cell death, a process shared in Solanaceae species [115]. Interestingly, RCR3 activity is not required to establish an Avr2-RCR3 complex, but removal of the RCR3 prodomain enables Avr2 accessibility (Fig. 4C). This pathway, including a pallet of subtilases that act as initiator proteases, described the first real proteolytic cascade in plants [114], and illustrates the evolutionary complexity of plant pathogen response.
Conclusions
Proteases are important actors in multiple cellular pathways ranging from the recognition of external signals to protein dismantlers in degradation processes. In animal systems, proteases have extensively attracted researchers’ attention, mainly due to their functions in cell death and disease, and as potential drug targets for biomedical applications. In addition, diverse proteases serve as research tools and are useful enzymes in various industrial applications [116, 117]. In contrast, our current knowledge of the mechanisms that regulate plant protease activity remains scarce, as is the functional understanding of the cleavage of their substrates. In comparison to mammalian systems, only a handful of plant proteomic studies towards an unbiased identification of protease targets are reported [48, 118,119,120] and only a few activation mechanisms are known (Table 1). Proteolysis can be validated in an ex situ framework by in vitro cleavage assays of recombinant proteins or by other means in alternative model organisms, lysates or reticulocyte systems [121]. As for some proteases recombinant production and purification might be challenging, production in cell-free systems with or without coexpression of their substrate might provide a good alternative. Although not critical, defining the preferred substrate-processing sites will facilitate to unravel the mode of action of proteases. The activation conditions should also be carefully regarded when looking at enzymatic activity. Some cues directly modify the structure of proteases, as such activating them, while other ones rather facilitate the substrate-protease encounter. PTMs are also important for the protease activation and the substrate availability for its processing. A hurdle found when working with protease families is their redundant activity towards individual substrates. In the last years, protease-class inhibitors were shown to be effective tools to surpass redundancy and discover hidden phenotypes that with other means, such as using single mutant lines, would remain undetectable [85, 87, 98]. Fortunately, nowadays genome editing tools allow to generate higher-order mutants, for instance targeting several members of the type-II metacaspase family in Arabidopsis using CRISPR, which leads to the discovery of an enhanced phenotype when tetra-mutants were challenged with pathogens [43]. Further unraveling of plant protease regulatory mechanisms will require complementary studies on the cellular conditions affecting protease structure in parallel with the spatiotemporal characterization of active proteases and their substrates.
Data availability
There are no research data associated with this Review paper
References
Rawlings ND, Barrett AJ, Thomas PD, Huang X, Bateman A, Finn RD. The MEROPS database of proteolytic enzymes, their substrates and inhibitors in 2017 and a comparison with peptidases in the PANTHER database. Nucleic Acids Res. 2018;46:D624–D32.
Jobin PG, Solis N, Machado Y, Bell PA, Rai SK, Kwon NH, et al. Moonlighting matrix metalloproteinase substrates: enhancement of proinflammatory functions of extracellular tyrosyl-tRNA synthetase upon cleavage. J Biol Chem. 2020;295:2186–202.
Paulus JK, Van der Hoorn RAL. Do proteolytic cascades exist in plants? J Exp Bot. 2019;70:1997–2002.
Cao M, Chen R, Li P, Yu Y, Zheng R, Ge D, et al. TMK1-mediated auxin signalling regulates differential growth of the apical hook. Nature. 2019;568:240–3.
Liu C, Törnkvist A, Charova S, Stael S, Moschou PN. Proteolytic proteoforms: elusive components of hormonal pathways? Trends Plant Sci. 2020;25:325–8.
van der Hoorn RAL. Plant proteases: from phenotypes to molecular mechanisms. Annu Rev Plant Biol. 2008;59:191–223.
García-Lorenzo M, Sjödin A, Jansson S, Funk C. Protease gene families in Populus and Arabidopsis. BMC Plant Biol. 2006;6:30.
Lallemand J, Bouché F, Desiron C, Stautemas J, de Lemos Esteves F, Périlleux C, et al. Extracellular peptidase hunting for improvement of protein production in plant cells and roots. Front Plant Sci. 2015;6:37.
van Wijk KJ. Protein maturation and proteolysis in plant plastids, mitochondria, and peroxisomes. Annu Rev Plant Biol. 2015;66:75–111.
Salvesen GS, Hempel A, Coll NS. Protease signaling in animal and plant-regulated cell death. FEBS J. 2016;283:2577–98.
Buono RA, Hudecek R, Nowack MK. Plant proteases during developmental programmed cell death. J Exp Bot. 2019;70:2097–112.
Schaller A. A cut above the rest: the regulatory function of plant proteases. Planta 2004;220:183–97.
Liu H, Hu M, Wang Q, Cheng L, Zhang Z. Role of papain-like cysteine proteases in plant development. Front Plant Sci. 2018;9:1717.
Balakireva AV, Zamyatnin AA. Indispensable role of proteases in plant innate immunity. Int J Mol Sci. 2018;19:629.
Salguero-Linares J, Coll NS. Plant proteases in the control of the hypersensitive response. J Exp Bot. 2019;70:2087–95.
Turk B, Turk D, Turk V. Protease signalling: the cutting edge. EMBO J. 2012;31:1630–43.
Riedl SJ, Salvesen GS. The apoptosome: signalling platform of cell death. Nat Rev Mol Cell Biol. 2007;8:405–13.
Pop C, Fitzgerald P, Green DR, Salvesen GS. Role of proteolysis in caspase-8 activation and stabilization. Biochemistry 2007;46:4398–407.
Kischkel FC, Hellbardt S, Behrmann I, Germer M, Pawlita M, Krammer PH, et al. Cytotoxicity-dependent APO-1 (Fas/CD95)-associated proteins form a death-inducing signaling complex (DISC) with the receptor. EMBO J. 1995;14:5579–88.
Zou H, Li Y, Liu X, Wang X. An APAF-1.cytochrome c multimeric complex is a functional apoptosome that activates procaspase-9. J Biol Chem. 1999;274:11549–56.
Martinon F, Burns K, Tschopp J. The inflammasome: a molecular platform triggering activation of inflammatory caspases and processing of proIL-β. Mol Cell. 2002;10:417–26.
Grosse-Holz FM, van der Hoorn RAL. Juggling jobs: roles and mechanisms of multifunctional protease inhibitors in plants. N Phytol. 2016;210:794–807.
Vercammen D, Belenghi B, van de Cotte B, Beunens T, Gavigan J-A, De Rycke R, et al. Serpin1 of Arabidopsis thaliana is a suicide inhibitor for metacaspase 9. J Mol Biol. 2006;364:625–36.
Lema Asqui S, Vercammen D, Serrano I, Valls M, Rivas S, Van Breusegem F, et al. AtSERPIN1 is an inhibitor of the metacaspase AtMC1-mediated cell death and autocatalytic processing in planta. N Phytol. 2018;218:1156–66.
Kapust RB, Tözsér J, Fox JD, Anderson DE, Cherry S, Copeland TD, et al. Tobacco etch virus protease: mechanism of autolysis and rational design of stable mutants with wild-type catalytic proficiency. Protein Eng. 2001;14:993–1000.
Vercammen D, van de Cotte B, De Jaeger G, Eeckhout D, Casteels P, Vandepoele K, et al. Type II metacaspases Atmc4 and Atmc9 of Arabidopsis thaliana cleave substrates after arginine and lysine. J Biol Chem. 2004;279:45329–36.
Watanabe N, Lam E. Calcium-dependent activation and autolysis of Arabidopsis metacaspase 2d. J Biol Chem. 2011;286:10027–40.
Zhu P, Yu X-H, Wang C, Zhang Q, Liu W, McSweeney S, et al. Structural basis for Ca2+-dependent activation of a plant metacaspase. Nat Commun. 2020;11:2249.
Wen S, Ma Q-M, Zhang Y-L, Yang J-P, Zhao G-H, Fu D-Q, et al. Biochemical evidence of key residues for the activation and autoprocessing of tomato type II metacaspase. FEBS Lett. 2013;587:2517–22.
Hander T, Fernández-Fernández AD, Kumpf RP, Willems P, Schatowitz H, Rombaut D, et al. Damage on plants activates Ca2+-dependent metacaspases for release of immunomodulatory peptides. Science 2019;363:eaar7486.
van Midden KP, Peric T, Klemenčič M. Plant type I metacaspases are proteolytically active proteases despite their hydrophobic nature. FEBS Lett. 2021;595:2237–47.
Tsiatsiani L, Van Breusegem F, Gallois P, Zavialov A, Lam E, Bozhkov PV. Metacaspases. Cell Death Differ. 2011;18:1279–88.
González IJ, Desponds C, Schaff C, Mottram JC, Fasel N. Leishmania major metacaspase can replace yeast metacaspase in programmed cell death and has arginine-specific cysteine peptidase activity. Int J Parasitol. 2007;37:161–72.
Watanabe N, Lam E. Two Arabidopsis metacaspases AtMCP1b and AtMCP2b are arginine/lysine-specific cysteine proteases and activate apoptosis-like cell death in yeast. J Biol Chem. 2005;280:14691–9.
Fuentes-Prior P, Salvesen GS. The protein structures that shape caspase activity, specificity, activation and inhibition. Biochem J. 2004;384:201–32.
Vercammen D, Declercq W, Vandenabeele P, Van Breusegem F. Are metacaspases caspases? J Cell Biol. 2007;179:375–80.
Eichinger A, Beisel H-G, Jacob U, Huber R, Medrano F-J, Banbula A, et al. Crystal structure of gingipain R: an Arg-specific bacterial cysteine proteinase with a caspase-like fold. EMBO J. 1999;18:5453–62.
Uren AG, O’Rourke K, Aravind LA, Pisabarro MT, Seshagiri S, Koonin EV, et al. Identification of paracaspases and metacaspases: two ancient families of caspase-like proteins, one of which plays a key role in MALT lymphoma. Mol Cell. 2000;6:961–7.
Minina EA, Coll NS, Tuominen H, Bozhkov PV. Metacaspases versus caspases in development and cell fate regulation. Cell Death Differ. 2017;24:1314–25.
Klemenčič M, Funk C. Evolution and structural diversity of metacaspases. J Exp Bot. 2019;70:2039–47.
Coll NS, Vercammen D, Smidler A, Clover C, Van Breusegem F, Dangl JL, et al. Arabidopsis type I metacaspases control cell death. Science. 2010;330:1393–7.
Dietrich RA, Delaney TP, Uknes SJ, Ward ER, Ryals JA, Dangl JL. Arabidopsis mutants simulating disease resistance response. Cell. 1994;77:565–77.
Shen W, Liu J, Li J-F. Type-II metacaspases mediate the processing of plant elicitor peptides in Arabidopsis. Mol Plant. 2019;12:1524–33.
Yamada K, Yamashita-Yamada M, Hirase T, Fujiwara T, Tsuda K, Hiruma K, et al. Danger peptide receptor signaling in plants ensures basal immunity upon pathogen-induced depletion of BAK1. EMBO J. 2016;35:46–61.
Bollhöner B, Zhang B, Stael S, Denance N, Overmyer K, Goffner D, et al. Post mortem function of AtMC9 in xylem vessel elements. N Phytol. 2013;200:498–510.
Belenghi B, Romero-Puertas MC, Vercammen D, Brackenier A, Inzé D, Delledonne M, et al. Metacaspase activity of Arabidopsis thaliana is regulated by S-nitrosylation of a critical cysteine residue. J Biol Chem. 2007;282:1352–8.
Roberts TH, Hejgaard J. Serpins in plants and green algae. Funct Integr Genom. 2008;8:1–27.
Tsiatsiani L, Timmerman E, De Bock P-J, Vercammen D, Stael S, van de Cotte B, et al. The Arabidopsis METACASPASE9 degradome. Plant Cell. 2013;25:2831–47.
Zauner FB, Dall E, Regl C, Grassi L, Huber CG, Cabrele C, et al. Crystal structure of plant legumain reveals a unique two-chain state with pH-dependent activity regulation. Plant Cell. 2018;30:686–99.
Scherz-Shouval R, Shvets E, Fass E, Shorer H, Gil L, Elazar Z. Reactive oxygen species are essential for autophagy and specifically regulate the activity of Atg4. EMBO J. 2007;26:1749–60.
Woo J, Park E, Dinesh-Kumar SP. Differential processing of Arabidopsis ubiquitin-like Atg8 autophagy proteins by Atg4 cysteine proteases. Proc Natl Acad Sci USA. 2014;111:863–8.
Pérez-Pérez ME, Zaffagnini M, Marchand CH, Crespo JL, Lemaire SD. The yeast autophagy protease Atg4 is regulated by thioredoxin. Autophagy. 2014;10:1953–64.
Pérez-Pérez ME, Lemaire SD, Crespo JL. Control of autophagy in Chlamydomonas is mediated through redox-dependent inactivation of the ATG4 protease. Plant Physiol. 2016;172:2219–34.
Laureano-Marín AM, Aroca A, Pérez-Pérez ME, Yruela I, Jurado-Flores A, Moreno I, et al. Abscisic acid-triggered persulfidation of the Cys protease ATG4 mediates regulation of autophagy by sulfide. Plant Cell. 2020;32:3902–20.
Luo L. Plant cytokine or phytocytokine. Plant Signal Behav. 2012;7:1513–4.
Tavormina P, De Coninck B, Nikonorova N, De Smet I, Cammue BPA. The plant peptidome: an expanding repertoire of structural features and biological functions. Plant Cell. 2015;27:2095–118.
Hou S, Liu D, He P. Phytocytokines function as immunological modulators of plant immunity. Stress Biol. 2021;1:8.
Galiullina RA, Kasperkiewicz P, Chichkova NV, Szalek A, Serebryakova MV, Poreba M, et al. Substrate specificity and possible heterologous targets of phytaspase, a plant cell death protease. J Biol Chem. 2015;290:24806–15.
Beloshistov RE, Dreizler K, Galiullina RA, Tuzhikov AI, Serebryakova MV, Reichardt S, et al. Phytaspase-mediated precursor processing and maturation of the wound hormone systemin. N Phytol. 2018;218:1167–78.
Pearce G, Strydom D, Johnson S, Ryan CA. A polypeptide from tomato leaves induces wound-inducible proteinase inhibitor proteins. Science. 1991;253:895–7.
Chao WS, Gu Y-Q, Pautot V, Bray EA, Walling LL. Leucine aminopeptidase RNAs, proteins, and activities increase in response to water deficit, salinity, and the wound signals systemin, methyl jasmonate, and abscisic acid. Plant Physiol. 1999;120:979–92.
Wang L, Einig E, Almeida-Trapp M, Albert M, Fliegmann J, Mithöfer A, et al. The systemin receptor SYR1 enhances resistance of tomato against herbivorous insects. Nat Plants. 2018;4:152–6.
Chichkova NV, Shaw J, Galiullina RA, Drury GE, Tuzhikov AI, Kim SH, et al. Phytaspase, a relocalisable cell death promoting plant protease with caspase specificity. EMBO J. 2010;29:1149–61.
Trusova SV, Teplova AD, Golyshev SA, Galiullina RA, Morozova EA, Chichkova NV, et al. Clathrin-mediated endocytosis delivers proteolytically active phytaspases into plant cells. Front Plant Sci. 2019;10:873.
Pearce G, Moura DS, Stratmann J, Ryan CA Jr. RALF, a 5-kDa ubiquitous polypeptide in plants, arrests root growth and development. Proc Natl Acad Sci USA. 2001;98:12843–7.
Abarca A, Franck CM, Zipfel C. Family-wide evaluation of RAPID ALKALINIZATION FACTOR peptides. Plant Physiol. 2021;187:996–1010.
Srivastava R, Liu J-X, Guo H, Yin Y, Howell SH. Regulation and processing of a plant peptide hormone, AtRALF23, in Arabidopsis. Plant J. 2009;59:930–9.
Liu J-X, Srivastava R, Che P, Howell SH. Salt stress responses in Arabidopsis utilize a signal transduction pathway related to endoplasmic reticulum stress signaling. Plant J. 2007;51:897–909.
Wolf S, Rausch T, Greiner S. The N-terminal pro region mediates retention of unprocessed type-I PME in the Golgi apparatus. Plant J. 2009;58:361–75.
Stegmann M, Monaghan J, Smakowska-Luzan E, Rovenich H, Lehner A, Holton N, et al. The receptor kinase FER is a RALF-regulated scaffold controlling plant immune signaling. Science 2017;355:287–9.
Xiao Y, Stegmann M, Han Z, DeFalco TA, Parys K, Xu L, et al. Mechanisms of RALF peptide perception by a heterotypic receptor complex. Nature. 2019;572:270–4.
Ghorbani S, Hoogewijs K, Pečenková T, Fernandez A, Inzé A, Eeckhout D, et al. The SBT6.1 subtilase processes the GOLVEN1 peptide controlling cell elongation. J Exp Bot. 2016;67:4877–87.
Stührwohldt N, Scholl S, Lang L, Katzenberger J, Schumacher K, Schaller A. The biogenesis of CLEL peptides involves several processing events in consecutive compartments of the secretory pathway. eLife. 2020;9:e55580.
Stührwohldt N, Ehinger A, Thellmann K, Schaller A. Processing and formation of bioactive CLE40 peptide are controlled by posttranslational proline hydroxylation. Plant Physiol. 2020;184:1573–84.
Liu C, Mentzelopoulou A, Deli A, Papagavriil F, Ramachandran P, Perraki A, et al. Phase separation of a nodulin Sec14-like protein maintains auxin efflux carrier polarity at Arabidopsis plasma membranes. bioRxiv. 2022, https://www.biorxiv.org/content/10.1101/2022.03.26.485938v1.
Moschou PN, Gutierrez-Beltran E, Bozhkov Peter V, Smertenko A. Separase promotes microtubule polymerization by activating CENP-E-related kinesin Kin7. Dev Cell. 2016;37:350–6172.
Doll NM, Royek S, Fujita S, Okuda S, Chamot S, Stintzi A, et al. A two-way molecular dialogue between embryo and endosperm is required for seed development. Science. 2020;367:431–5.
Tanaka H, Onouchi H, Kondo M, Hara-Nishimura I, Nishimura M, Machida C, et al. A subtilisin-like serine protease is required for epidermal surface formation in Arabidopsis embryos and juvenile plants. Development. 2001;128:4681–9.
Creff A, Brocard L, Joubès J, Taconnat L, Doll NM, Marsollier AC, et al. A stress-response-related inter-compartmental signalling pathway regulates embryonic cuticle integrity in Arabidopsis. PLoS Genet. 2019;15:e1007847.
Royek S, Bayer M, Pfannstiel J, Pleiss J, Ingram G, Stintzi A, et al. Processing of a plant peptide hormone precursor facilitated by posttranslational tyrosine sulfation. Proc Natl Acad Sci USA. 2022;119:e2201195119.
Shi CL, Alling RM, Hammerstad M, Aalen RB. Control of organ abscission and other cell separation processes by evolutionary conserved peptide signaling. Plants. 2019;8:225.
Kumpf RP, Shi C-L, Larrieu A, Stø IM, Butenko MA, Péret B, et al. Floral organ abscission peptide IDA and its HAE/HSL2 receptors control cell separation during lateral root emergence. Proc Natl Acad Sci USA. 2013;110:5235–40.
Tian M, Benedetti B, Kamoun S. A second Kazal-like protease inhibitor from Phytophthora infestans inhibits and interacts with the apoplastic pathogenesis-related protease P69B of tomato. Plant Physiol. 2005;138:1785–93.
Tian M, Huitema E, da Cunha L, Torto-Alalibo T, Kamoun S. A Kazal-like extracellular serine protease inhibitor from Phytophthora infestans targets the tomato pathogenesis-related protease P69B. J Biol Chem. 2004;279:26370–7.
Schardon K, Hohl M, Graff L, Pfannstiel J, Schulze W, Stintzi A, et al. Precursor processing for plant peptide hormone maturation by subtilisin-like serine proteinases. Science. 2016;354:1594–7.
Reichardt S, Repper D, Tuzhikov AI, Galiullina RA, Planas-Marquès M, Chichkova NV, et al. The tomato subtilase family includes several cell death-related proteinases with caspase specificity. Sci Rep. 2018;8:10531. [Erratum Sci Rep. 2020;10:5661]
Reichardt S, Piepho H-P, Stintzi A, Schaller A. Peptide signaling for drought-induced tomato flower drop. Science. 2020;367:1482–5.
Santiago J, Brandt B, Wildhagen M, Hohmann U, Hothorn LA, Butenko MA, et al. Mechanistic insight into a peptide hormone signaling complex mediating floral organ abscission. eLife. 2016;5:e15075.
Meng X, Zhou J, Tang J, Li B, de Oliveira MVV, Chai J, et al. Ligand-induced receptor-like kinase complex regulates floral organ abscission in Arabidopsis. Cell Rep. 2016;14:1330–8.
Zhang H, Hu Z, Lei C, Zheng C, Wang J, Shao S, et al. A plant phytosulfokine peptide initiates auxin-dependent immunity through cytosolic Ca2+ signaling in tomato. Plant Cell. 2018;30:652–67.
Peng Y, Chen L, Lu Y, Wu Y, Dumenil J, Zhu Z, et al. The ubiquitin receptors DA1, DAR1, and DAR2 redundantly regulate endoreduplication by modulating the stability of TCP14/15 in Arabidopsis. Plant Cell. 2015;27:649–62.
Dong H, Dumenil J, Lu F-H, Na L, Vanhaeren H, Naumann C, et al. Ubiquitylation activates a peptidase that promotes cleavage and destabilization of its activating E3 ligases and diverse growth regulatory proteins to limit cell proliferation in Arabidopsis. Genes Dev. 2017;31:197–208.
Vanhaeren H, Nam Y-J, De Milde L, Chae E, Storme V, Weigel D, et al. Forever young: the role of ubiquitin receptor DA1 and E3 ligase BIG BROTHER in controlling leaf growth and development. Plant Physiol. 2017;173:1269–82.
Vanhaeren H, Chen Y, Vermeersch M, De Milde L, De Vleeschhauwer V, Natran A, et al. UBP12 and UBP13 negatively regulate the activity of the ubiquitin-dependent peptidases DA1, DAR1 and DAR2. eLife. 2020;9:e52276.
Dong H, Smith C, Prior R, Carter R, Dumenil J, Saalbach G, et al. The Receptor Kinase BRI1 promotes cell proliferation in Arabidopsis by phosphorylation- mediated inhibition of the growth repressing peptidase DA1. Proc Natl Acad Sci USA. 2022;119:e2205757119.
Gu B, Dong H, Smith C, Bevan MW. Modulation of receptor-like trans-membrane kinase 1 nuclear localisation by DA1 peptidases in arabidopsis. PNAS. 2022;119:e2205757119.
Bouwmeester H, Sinha N, Scholes J. Parasitic plants: physiology, development, signaling, and ecosystem interactions. Plant Physiol. 2021;185:1267–9.
Ogawa S, Wakatake T, Spallek T, Ishida JK, Sano R, Kurata T, et al. Subtilase activity in intrusive cells mediates haustorium maturation in parasitic plants. Plant Physiol. 2021;185:1381–94.
Zipfel C, Robatzek S, Navarro L, Oakeley EJ, Jones JDG, Felix G, et al. Bacterial disease resistance in Arabidopsis through flagellin perception. Nature. 2004;428:764–7.
Zipfel C, Kunze G, Chinchilla D, Caniard A, Jones JDG, Boller T, et al. Perception of the bacterial PAMP EF-Tu by the receptor EFR restricts Agrobacterium-mediated transformation. Cell. 2006;125:749–60.
Ziemann S, van der Linde K, Lahrmann U, Acar B, Kaschani F, Colby T, et al. An apoplastic peptide activates salicylic acid signalling in maize. Nat Plants. 2018;4:172–80.
Djamei A, Schipper K, Rabe F, Ghosh A, Vincon V, Kahnt J, et al. Metabolic priming by a secreted fungal effector. Nature. 2011;478:395–8.
Mueller AN, Ziemann S, Treitschke S, Aßmann D, Doehlemann G. Compatibility in the Ustilago maydis-maize interaction requires inhibition of host cysteine proteases by the fungal effector Pit2. PLoS Pathog. 2013;9:e1003177.
van der Linde K, Hemetsberger C, Kastner C, Kaschani F, van der Hoorn RAL, Kumlehn J, et al. A maize cystatin suppresses host immunity by inhibiting apoplastic cysteine proteases. Plant Cell. 2012;24:1285–300.
Cheng Z, Li J-F, Niu Y, Zhang X-C, Woody OZ, Xiong Y, et al. Pathogen-secreted proteases activate a novel plant immune pathway. Nature. 2015;521:213–6.
Wang Y, Garrido-Oter R, Wu J, Winkelmüller TM, Agler M, Colby T, et al. Site-specific cleavage of bacterial MucD by secreted proteases mediates antibacterial resistance in Arabidopsis. Nat Commun. 2019;10:2853.
Mackey D, Holt BF 3rd, Wiig A, Dangl JL. RIN4 interacts with Pseudomonas syringae type III effector molecules and is required for RPM1-mediated resistance in Arabidopsis. Cell. 2002;108:743–54.
Mackey D, Belkhadir Y, Alonso JM, Ecker JR, Dangl JL. Arabidopsis RIN4 is a target of the type III virulence effector AvrRpt2 and modulates RPS2-mediated resistance. Cell. 2003;112:379–89.
Kim H-S, Desveaux D, Singer AU, Patel P, Sondek J, Dangl JL. The Pseudomonas syringae effector AvrRpt2 cleaves its C-terminally acylated target, RIN4, from Arabidopsis membranes to block RPM1 activation. Proc Natl Acad Sci USA. 2005;102:6496–501.
Zhang J, Li W, Xiang T, Liu Z, Laluk K, Ding X, et al. Receptor-like cytoplasmic kinases integrate signaling from multiple plant immune receptors and are targeted by a Pseudomonas syringae effector. Cell Host Microbe. 2010;7:290–301.
Shao F, Golstein C, Ade J, Stoutemyer M, Dixon JE, Innes RW. Cleavage of Arabidopsis PBS1 by a bacterial type III effector. Science. 2003;301:1230–3.
Swiderski MR, Innes RW. The Arabidopsis PBS1 resistance gene encodes a member of a novel protein kinase subfamily. Plant J. 2001;26:101–12.
Ade J, DeYoung BJ, Golstein C, Innes RW. Indirect activation of a plant nucleotide binding site-leucine-rich repeat protein by a bacterial protease. Proc Natl Acad Sci USA. 2007;104:2531–6.
Paulus JK, Kourelis J, Ramasubramanian S, Homma F, Godson A, Hörger AC, et al. Extracellular proteolytic cascade in tomato activates immune protease Rcr3. Proc Natl Acad Sci USA. 2020;117:17409–17.
Kourelis J, Malik S, Mattinson O, Krauter S, Kahlon PS, Paulus JK, et al. Evolution of a guarded decoy protease and its receptor in solanaceous plants. Nat Commun. 2020;11:4393.
Folgado A, Abranches R. Plant aspartic proteases for industrial applications: thistle get better. Plants. 2020;9:147.
Pottinger SE, Innes RW. RPS5-mediated disease resistance: fundamental insights and translational applications. Annu Rev Phytopathol. 2020;58:139–60.
Demir F, Niedermaier S, Villamor JG, Huesgen PF. Quantitative proteomics in plant protease substrate identification. N Phytol. 2018;218:936–43.
Venne AS, Solari FA, Faden F, Paretti T, Dissmeyer N, Zahedi RP. An improved workflow for quantitative N-terminal charge-based fractional diagonal chromatography (ChaFRADIC) to study proteolytic events in Arabidopsis thaliana. Proteomics. 2015;15:2458–69.
Minina EA, Stael S, Van Breusegem F, Bozhkov PV. Plant metacaspase activation and activity. Methods Mol Biol. 2014;1133:237–53.
Liu C, Stael S, Gevaert K, Van Breusegem F, Bozhkov PV, Moschou PN. The proteolytic landscape of an Arabidopsis separase-deficient mutant reveals novel substrates associated with plant development. bioRxiv. 2017, https://www.biorxiv.org/content/10.1101/140962v1.
Seo E, Woo J, Park E, Bertolani SJ, Siegel JB, Choi D, et al. Comparative analyses of ubiquitin-like ATG8 and cysteine protease ATG4 autophagy genes in the plant lineage and cross-kingdom processing of ATG8 by ATG4. Autophagy. 2016;12:2054–68.
Acknowledgements
The authors thank Dr. Martine De Cock for her assistance in the preparation of the manuscript.
Funding
This work was supported by the Research Foundation-Flanders (grant FWO14/PDO/166 to SS) and the Ghent University Special Research Fund (grant 01J11311 to FVB).
Author information
Authors and Affiliations
Contributions
ADF-F wrote the manuscript and created the figures, SS and FVB edited the manuscript. All authors approved the submitted version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fernández-Fernández, Á.D., Stael, S. & Van Breusegem, F. Mechanisms controlling plant proteases and their substrates. Cell Death Differ 30, 1047–1058 (2023). https://doi.org/10.1038/s41418-023-01120-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-023-01120-5
- Springer Nature Limited
This article is cited by
-
Dying in self-defense: cell death signaling in animals and plants
Cell Death & Differentiation (2024)