Abstract
Medulloblastoma (MB) is the most frequent malignant brain tumor in children and it is subgrouped into 4 entities (SHH, WNT, Group 3, and Group 4). Molecular pathways involved in these different subgroups still are evolving and can be of clinical relevance to therapy. The YAP1-CTGF axis is known to regulate cell proliferation, differentiation, and cell death; however, its role in MB is poorly explored. We aimed to investigate the role of YAP1 gene in the MB SHH cell line DAOY and evaluate cell proliferation, doubling time and 3D spheroids invasion and its consequence on CTGF regulation. We assessed CTGF expression from 22 children with MB. Lastly, we validated our findings through in silico analysis in large cohorts dataset of patients. We observed an increased invasion rate of DAOY cells and CTGF downregulation under YAP1 knockdown (p < 0.0001). Additionally CTGF is overexpressed in MB with extensive nodularity subtype and an indicative of higher survival rates in pediatric MB (p < 0.05). Interestingly, no difference of CTGF expression was observed between molecular subgroups. These results provide new evidence ofCTGF as a potential prognostic marker for MB, corroborating to the role of YAP1 in restricting MB cell.
Similar content being viewed by others
Medulloblastoma, YAP1, and CTGF
Medulloblastoma (MB) is the most common malignant brain tumor of childhood. Actual consensus considers MB as four subgroups with distinct clinical and molecular features [1]. Although most patients show good response to treatment, therapy may cause long-term side effects due to the lack of personalized target therapies [2]. YAP1 (Yes Associated protein 1) is a transcriptional co-activator protein that belongs to the Hippo signaling pathway and controls organ size during embryonic development [3,4,5]. In MB patients, YAP1 is associated with poor prognosis and is considered a potential target to therapy [3, 4]. YAP1 acts as a mediator of cell proliferation in cerebellar granule neural precursors (GNPs) and drives resistance to radiation when expressed [3, 4]. YAP1 is amplified in some MB patients, particularly, in the SHH subtypes;[6] however, the clinical implications and pathway downstream activation that drives various biological processes remain elusive [3, 4, 6]. Interestingly, YAP1 plays a role in tissue regeneration and cell matrix stiffness [7]. These features place YAP1 as a candidate protein for molecular medicine regeneration due to its role in organ size control through CTGF (connective tissue growth factor), restricting migration and stimulating adhesion of various cell types [7, 8]. The dual role of this molecule highlights YAP1 as an interesting candidate to be exploited [9]. We found that YAP1 upregulates CTGF mRNA levels, and that CTGF overexpression is an indicative of superior survival rates in pediatric patients with MB. Further, we validated our findings in a larger dataset of MB patients. This rapid communication shed new lights onCTGF, a downstream target of YAP1, as a potential prognosticator marker for MB.
Depletion of YAP1 augments invasion rates of MB cell line and downregulatesCTGF, an interesting gene which expression indicates better survival rates
The efficiency of YAP1 depletion in DAOY cells stably transduced with either shScrambled (shSCR) or (shYAP_#1 or shYAP_#2) was evaluated by Immunoblotting. Both shYAP inducible vectors reduced YAP1 expression after treatment with Dox (150 and 200 ng/ml) (Fig. 1a, b). To assess cell viability and proliferation under YAP1 depletion, CCK8 viability assay and doubling time were performed. Interestingly, YAP1 depletion leads to significant decrease in cell proliferation and cell viability (Fig. 1c, d). Similarly, authors found YAP1 protein as a driver of proliferation in GNP’s, cells with the same origin as DAOY [3, 4]. Moreover, ectopic expression of YAP1 in GNP’s triggered oncogenic signaling pathways such as AKT/mTOR and IGF2, key proteins related to cell cycle progression and DNA repair mechanisms [3, 4]. Remarkably, depletion of YAP1 enhances cell invasion as shown by 3D spheroids assay (Fig. 1e, f) (p < 0.0001). YAP1 might restrict cell migration and invasion due to its interdependence of mechanotransducers, mechanical cues, cell matrix stiffness, and cell–cell contact [9]. In epithelial cells, activation of the tumor suppressor Hippo Pathway leads to phosphorylation and degradation of YAP1 and cell invasion promotion via downregulation of E-cadherin and laminin, proteins implicated in the Epithelial Mesenchymal Transition (EMT) [10]. These dual function of action warrant precaution regarding therapeutic interventions involving inhibition of YAP1, such as the use of Verteporfin in breast cancer, skin cancers, and pancreatic cancer (Clinical trials number: NCT02872064, NCT00049959, and NCT02939274) [11, 12].
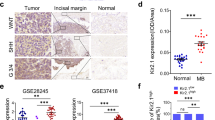
a Immunoblot showing the expression of YAP1 in DAOY transduced with SCR vector, shYAP1 #1 and YAP1 #2 lentiviral vector when incubated with 200 ng/ml of DOX [* < 0.05]. b Representing graph of a, c Knockdown of YAP1 after 24, 48, and 72 h leads to decrease in cell viability. d Knockdown of YAP1 after 24, 48, and 72 h leads to decrease in cell.e 3D spheroid invasion assay shows significant increasing of migration/invasion process in DAOY cell line after inhibition of YAP1 [* < 0.05, ** < 0.01, *** < 0.001]. f Quantification of 3D spheroid assay using aCDc in three independent experiments.g Decrease of CTGF after YAP1 knockdown in DAOY cells [** < 0.01]. h Kaplan–Meier curve according to the CTGF gene expression in 22 MB patients samples. The group with low levels ofCTGF presented significant lower event-free survival rates (Log-rant test; P = 0.04) i Kaplan–Meier curve according to the CTGF gene expression in 172 MB patients samples. The group with low levels of CTGF presented a trend to lower event-free survival rates (log-rant test; P = 0.058).j Expression of CTGF in groups of patients (dead versus alive). l Expression of CTGF in groups of patients (LCA versus MBEN versus desmoplastic versus classic histological subtype)
YAP1 triggers cytokines, mitogens, and downstream proteins (MAPKs, Akt/mTOR, VEGF, and IGF2) leading to aggressiveness and unfavorable outcomes in MBs [3, 4]. In order to investigate a potential tumor suppressor feature of YAP1 and to better understand the results obtained in the invasion assay, we elected to study CTGF, a canonical downstream effector of YAP1 and promoter of adhesion in cells [12]. We performed RT-PCR comparing YAP1 depleted cells with SCR cells (Control) and found a significant decrease in fold-change expression (p > 0.001) (Fig. 1g), indicating a potential upregulation of CTGF by YAP1. Interestingly, a YAP1/TEAD1 complex is recognized, which binds to the site ofCTGF gene and activates its transcription [15,16,17,18,19,20]. This feature corroborates with our hypothesis that YAP1 acts directly upregulating CTGF [13,14,15,16,17,18]. Looking more closely to the potential role of CTGF as a tumor suppressor factor upregulated by YAP1, we evaluated in 22 MB samples the expression levels of CTGF gene and assessed event-free survival (EFS) rates by Kaplan–Meier curve and log-rank test, considering as unfavorable events relapse or death. We observed that patients showing CTGF low expression presented inferior EFS rates when compared to patients which high expression levels (27.3% ± 13.4% versus 71.6% ± 14%; P = 0.04) (Fig. 1h). Next, we decided to validate our findings in a larger cohort utilizing a data set from GSE85217 study in 172 MB patients whose complete clinical data were publicly available. A similar survival trend was observed, although statistical significance for the comparison resulted only in a borderline survival difference (P = 0.056) (Fig. 1i). Additionally, we compared the expression levels of CTGF regarding patient status, and the histological subtypes LCA (large cell anaplastic) versus MBEN (medulloblastoma with extensive nodularity) (Fig. 1j–l). Notably, the group of patients alive and classified as MBEN subtypes had higher levels of CTGF (p > 0.001). Patients classified as MBEN are associated with good prognosis and a low frequency of metastasis compared to LCA and desmoplastic subtypes [19, 20] (Complete Method description available at Additional supplementary file 1 and [21,22,23]). Additionally, we sought to evaluateCTGF expression in MB molecular subgroups, however, we did not find any significant differences of CTGF mRNA levels comparing molecular subgroups in two large MB cohort studies: (1) Microarray data from Cavalli and colleagues (2017) (Fig. S1a and S2a), (2) RNAseq from Pfister and colleagues (2017) (Fig. S2b and Fig. S2c). Our findings suggest that CTGF is a marker for medulloblastoma independent of molecular subgroups and its transcriptional program is active at the microenvironment niche of tumors in CNS (Fig. S3).
Conclusion
In conclusion, our data shed new light on a dual role of YAP1 in MBs and highlights the need for caution when exploiting YAP1 as a therapeutic target. Likewise, YAP1 potentially drives proliferation and can restrict migration through CTGF. Moreover, decreasing CTGF as a downstream consequence of YAP1 inhibition might trigger cell migration, potentially facilitating CNS tumor dissemination, which is, one of the leading causes of death in MBs [24]. Although CTGF was identified here as potential prognostic marker, more studies, particularly in animal models are needed to elucidate its biological role in MBs.
References
Cavalli FMG, Remke M, Rampasek L, Peacock J, Shih DJH, Luu B. et al. Intertumoral heterogeneity within medulloblastoma subgroups. Cancer Cell. 2017;31(Issue 6):737–754.
Martirosian V, Chen TC, Lin M, Neman J. Medulloblastoma initiation and spread: where neurodevelopment, microenvironment and cancer cross pathways. J Neurosci Res. 2016;94:1511–9.
Fernandez-L A, Northcott PA, Dalton J, Fraga C, Ellison D, Angers S, et al. YAP1 is amplified and up-regulated in hedgehog-associated medulloblastomas and mediates Sonic hedgehog-driven neural precursor proliferation. Genes Dev. 2009;23:2729–41.
Fernandez-L A, Squatrito M, Northcott P, Awan A, Holland EC, Taylor MD, et al. Oncogenic YAP promotes radioresistance and genomic instability inmedulloblastoma through IGF2-mediated Akt activation. Oncogene. 2012;31:1923–37.
Zhao B, Li L, Lei Q, Guan KL. The Hippo-YAP pathway in organ size control and tumorigenesis: an updated version. Genes Dev. 2010;24:862–74.
Schwalbe EC, Lindsey JC, Nakjang S, Crosier S, Smith AJ, Hicks D, et al. Novel molecular subgroups for clinical classification and outcome prediction in childhood medulloblastoma: a cohortstudy. Lancet Oncol. 2017;18:958–71.
Juan WC, Hong W. Targeting the hippo signaling pathway for tissue regeneration and cancer therapy. Genes. 2016;7:pii: E55.
Low BC, Pan CQ, Shivashankar GV, Bershadsky A, Sudol M, Sheetz M, et al. YAP/TAZ as mechanosensors and mechanotransducers in regulating organ size and tumor growth. FEBS Lett. 2014;588:2663–70.
Imajo M, Ebisuya M, Nishida E. Dual role of YAP and TAZ in renewal of the intestinal epithelium. Nat Cell Biol. 2015;17:7–19.
Ma X, Wang H, Ji J, Xu W, Sun Y, Li W, et al. Hippo signaling promotes JNK-dependent cell migration. Proc Natl Acad Sci USA. 2017;114:1934–9.
Huggett MT, Jermyn M, Gillams A, Illing R, Mosse S, Novelli M. Phase I/II study of verteporfin photodynamic therapy in locally advanced pancreatic cancer. Br J Cancer. 2014;110:1698–704.
Wang LH, Tsai HC, Cheng YC, Lin CY, Huang YL, Tsai CH, et al. CTGF promotes osteosarcoma angiogenesis by regulating miR-543/angiopoietin 2 signaling. Cancer Lett. 2017;391:28–37.
Zhao B, Ye X, Yu J, Li L, Li W, Li S, et al. TEAD mediates YAP-dependent gene induction and growth control. Genes Dev. 2008;22:1962–71.
Hong W, Guan KL. The YAP and TAZ transcription co-activators: key downstream effectors of the mammalian Hippo pathway. Semin Cell Dev Biol. 2012;23:785–93.
Li Z, Zhao B, Wang P, Chen F, Dong Z, Yang H, et al. Structural insights into the YAP and TEAD complex. Genes Dev. 2010;24:235–40.
Cairns L, Tran T, Kavran JM. Structural insights into the regulation of hippo signaling. ACS Chem Biol. 2017;12:601–10.
Chen L, Loh PG, Song H. Structural and functional insights into the TEAD-YAP complex in the Hippo signaling pathway. Protein Cell. 2010;1:1073–1083.
Mesrouze Y, Bokhovchuk F, Meyerhofer M, Fontana P, Zimmermann C, Martin T, et al. Dissection of the interaction between the intrinsically disordered YAP protein and the transcription fator TEAD. Elife. 2017;6:e25068.
Jóźwiak J, Sontowska I, Bikowska B, Grajkowska W, Galus R, Roszkowski M, et al. Favourable prognosis in medulloblastoma with extensive nodularity is associatedwith mitogen-activated protein kinase upregulation. Folia Neuropathol. 2011;49:257–61.
Korshunov A, Sahm F, Stichel D, Schrimpf D, Ryzhova M, Zheludkova O, et al. Molecularcharacterization of medulloblastomas with extensive nodularity (MBEN). Acta Neuropathol. 2018;136:303–13.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25:402–8.
Kumar KS, Pillong M, Kunze J, Burghardt I, Weller M, Grotzer MA, et al. Corrigendum: computer-assisted quantification of motile and invasive capabilities of cancer cells. Sci Rep. 2018;8:46996.
Cruzeiro GAV, Salomão KB, de Biagi CAO Jr, Baumgartner M, Sturm D, Lira RCP, et al. A simplified approach using Taqman low-density array for medulloblastoma subgrouping. Acta Neuropathol Commun. 2019;7:33.
Garzia L, Kijima N, Morrissy AS, De Antonellis P, Guerreiro-Stucklin A, Holgado BL, et al. A hematogenous route for medulloblastoma leptomeningeal metastases. Cell. 2018;172:1050–1062.
Funding
This work was supported by FAPESP grant number 2017/06511–8, 2014/19976–0 and 2014/20341–0.
Author information
Authors and Affiliations
Contributions
G.A.V.C. planned and conducted all experiments, performed the in silico analysis, drafted, and critically read the manuscript. R.C.P.L conducted the overall survival analysis, helped to design the study, and critically read the manuscript. T.A.M., helped to review the design of the study, performed the in silico analysis, and critically read the manuscript. C.A.S. critically read the manuscript, M.B. critically read the manuscript and helped to design the study. L.G.T. provided patient samples and critically read the manuscript. E.T.V. critically read the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. This research was submitted to and approved by the HC/FMRP-USP Research Ethics Committee (CAAE no 37206114.1.0000.5440) no 15509/2016.
Informed consent
All samples were obtained after receiving informed consent from all participants included in the study.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Cruzeiro, G.A.V., Lira, R.C.P., de Almeida Magalhães, T. et al. CTGF expression is indicative of better survival rates in patients with medulloblastoma. Cancer Gene Ther 27, 378–382 (2020). https://doi.org/10.1038/s41417-019-0100-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41417-019-0100-3
- Springer Nature America, Inc.