Abstract
Altered cellular metabolism is a major mechanism by which tumours support nutrient consumption associated with increased cellular proliferation. Selective dependency on specific metabolic pathways provides a therapeutic vulnerability that can be targeted in cancer therapy. Anti-metabolites have been used clinically since the 1940s and several agents targeting nucleotide metabolism are now well established as standard of care treatment in a range of indications. However, despite great progress in our understanding of the metabolic requirements of cancer and non-cancer cells within the tumour microenvironment, there has been limited clinical success for novel agents targeting pathways outside of nucleotide metabolism. We believe that there is significant therapeutic potential in targeting metabolic processes within cancer that is yet to be fully realised. However, current approaches to identify novel targets, test novel therapies and select patient populations most likely to benefit are sub-optimal. We highlight recent advances in technologies and understanding that will support the identification and validation of novel targets, re-evaluation of existing targets and design of optimal clinical positioning strategies to deliver patient benefit.
Similar content being viewed by others
Introduction
It is well established that cells that undergo malignant transformation have altered metabolic requirements compared to non-malignant parental cells. Rewiring of cellular metabolism is required to support several processes associated with tumour progression, including increased proliferation [1] and metastasis [2] and has been recognised as one of the hallmarks of cancer [3]. Conceptually, these selective metabolic requirements provide a therapeutic vulnerability to target in cancer. Indeed, several anti-metabolite agents are approved for use in the clinic. It was in 1948 that the first use of an agent targeting metabolism, the anti-folate aminopterin, was described to induce remission in children with acute lymphocytic leukaemia (ALL) [4]. Since then, the anti-folate methotrexate has become well established as part of chemotherapy regimens for several indications including ALL [5]. Anti-folates work, in part, by inhibiting the synthesis of nucleotides from folic acid and several other agents that target nucleotide metabolism, including 5-fluorouracil [6], hydroxycarbamide [7] and gemcitabine [8] are also approved anti-cancer agents (Table 1).
Increased understanding and characterisation of the metabolic pathways linked to cancer progression in recent years [9,10,11,12,13], coupled with the patient benefit derived from the agents described above, led to significant investment in the field that was predicted to lead to a new wave of anti-cancer drugs [14]. However, despite substantial progress in understanding the requirement of key metabolic processes for tumour progression, the number of newly approved drugs in this class remains disappointingly low, with few agents acting by mechanisms other than inhibition of nucleotide metabolism (Table 1). The field has been set back in recent years by a string of disappointing outcomes associated with compounds in clinical development, including the Glutaminase inhibitor Telaglenastat (KEAPSAKE trial: NCT04265534), the TCA cycle inhibitor Devimistat (AVENGER 500 trial NCT03504423, ARMADA 2000 trial: NCT03504410), the complex I inhibitor IACS-010759 (NCT02882321, NCT03291938) [15], the tyrosine analogue SM-88 (TYME trial: NCT03512756) and the IDO1 inhibitor Epacadostat (ECHO-301/KEYNOTE-252 trial: NCT02752074) [16]. Each of these agents has demonstrated promising efficacy in pre-clinical models [17,18,19,20,21,22] that has not yet translated into clinical efficacy. With an investment of $1–2 billion [23] required to take a new agent through the drug discovery process to clinical approval, late-stage clinical failures increase the perceived risk associated with progressing novel therapeutics in this class. There are several reasons why the attrition rate of therapeutics targeting cellular metabolism may be high, including metabolic plasticity leading to resistance to single-agents, dose-limiting toxicity due to the metabolic requirements of normal tissue homoeostasis or differing metabolic dependencies between pre-clinical models and patients.
To realise the potential of therapeutic modulation of metabolism in oncology, the field needs to refine standard approaches that are sub-optimal for this complex area. Careful consideration needs to be given to ensure that optimal approaches for target identification, target validation and clinical positioning are utilised. We introduce key advances in capabilities and technology that should be applied more widely to reduce the high attrition rate for novel drug targets in this class. These recent advances allow a more accurate understanding of the metabolism of tumours in the organ- and tissue-specific microenvironment. This will allow novel targets to be rationally identified and validated in complex and clinically relevant model systems. The development of novel therapeutic modalities may also allow those targets previously considered too challenging to be re-visited.
Utilising physiologically relevant cell culture systems
The most commonly used culture media for immortalised cancer cell lines are Dulbecco’s modified Eagle’s medium (DMEM) and Roswell Park Memorial Institute (RPMI)-1640. DMEM was first described in 1959 [24] and RPMI-1640 in 1967 [25]. It is now apparent that the concentrations of many nutrients within DMEM or RPMI-1640 do not reflect the concentrations found in standard human blood plasma [26,27,28]. For example, the concentrations of glutamine and glucose in DMEM or RPMI-1640 are far in excess of those that would be considered physiologically relevant [26, 29]. Despite these discrepancies, these media are still widely used to study metabolic processes. There are numerous examples described in the literature that demonstrate how environmental nutrient concentrations can alter dependency of a cancer cell on a target of interest. Super-physiological levels of cysteine found in RPMI-1640, correlate with increased glutamine anaplerosis and sensitisation to GLS inhibition [26], dependency on methionine synthetase (MTR) for cancer cell proliferation is only apparent in physiological folate conditions [30, 31] and increased levels of uric acid are associated with resistance to 5-fluorouracil [27].
After more than 60 years since the development of DMEM and evidence demonstrating the sub-optimal nature of standard media, we strongly believe that the field should be moving to utilise more physiological conditions. Two cell culture media have been independently developed and are currently commercially available, with a nutrient concentration that reflects that found in human blood plasma, HPLM [27] and PlasmaxTM [28]. These media not only rectify the concentrations of nutrients found in standard media, but also contain nutrients that are present in human blood plasma but not found in standard media. These media represent the nutritional baseline that can be supplemented with the appropriate lipid and protein factors depending on the specific requirements of the cell type of interest. Culture of leukaemia cell lines in HPLM results in extensive metabolic changes, including altered glucose utilisation, altered redox state and a significant reduction in de novo pyrimidine synthesis [27]. Culture of breast cancer cell lines in PlasmaxTM also results in extensive phenotypic alterations, including increased colony forming ability, altered metabolite levels and reduced glutamine consumption [28]. Mass spectrometry (MS) analysis of the CAL-120 breast cancer cell line demonstrate that there are significant differences between the metabolic profiles of cells grown in PlasmaxTM relative to standard media, in both 2D and 3D growth conditions. The in vitro conditions that most closely recapitulate the metabolic profile of CAL-120 cells grown in vivo as orthotopic tumours are culture with PlasmaxTM in 3D conditions [28].
siRNA- and CRISPR-based target ID screens, such as large-scale unbiased screening datasets that are publicly available from the Cancer Dependency Map (https://depmap.org/portal), can be used for the purposes of cancer target identification, validation or clinical positioning. However, caution should be applied to the interpretation of such output for any metabolic target due to the use of non-physiologically relevant and inconsistent cell culture conditions [32]. A genome wide CRISPR screen to identify genes that reduce cell viability in a chronic myeloid leukaemia (CML) cell line cultured in either RPMI or HPLM, identified selective dependencies for each condition that span several biological processes [32]. Efforts to identify targets that influence the tumour immune response are also possible using tumour and immune co-culture systems [33] and the use of physiological culture media in these systems should be considered. Indeed, it has been demonstrated that T cells cultured in HPLM show increased levels of activation and extensive differences in gene transcription compared to T cells cultured in RPMI, driven by higher calcium levels present in HPLM [34]. This suggests that use of physiological culture media will have implications for studying both cancer cell intrinsic and extrinsic metabolic processes.
3D culture of cancer cells as spheroids or patient-derived organoids are believed to offer advantages over traditional 2D culture and may be a more physiologically relevant in vitro model system [35]. Genome wide CRIPSR screens comparing 2D versus 3D growth conditions have identified selective dependencies [36] and CRISPR screening technology is also applicable to patient-derived organoids [37]. Performing target ID screens in a 3D system with physiologically relevant media that more closely recapitulates the metabolic profile seen in vivo therefore has the potential to identify novel cancer metabolism related targets.
Moving towards greater physiological relevance of cell culture conditions is a big, and long overdue, step forward for the cancer metabolism field but there are several limitations to be aware of. Immortalised cancer cell lines have often been cultured, and therefore selected for, in far from physiological media conditions for many passages and may have undergone long-lasting adaptation to their environmental conditions. Consideration should therefore be given to the use of primary cells or organoids cultured in more physiological media. A second limitation is that cells consume, and eventually exhaust, nutrients present within the environment, resulting in non-consistent concentrations throughout the duration of an experiment. Two mitigations that can be used to delay nutrient exhaustion when using physiological media are the frequent replacing of the media, and the maximisation of the media volume to cell number ratio. A more advanced solution would be to employ a micro-fluidic system to supply the physiological concentrations of nutrients at constant rates [38, 39]. The concentrations of nutrients in HPLM and PlasmaxTM reflect the nutritional environment found in human blood plasma, however this may differ from individual tumour microenvironments (TME). Metabolomic analysis of tumour interstitial fluid of pancreatic and lung tumours from mouse models demonstrated that metabolic environments alter depending on tumour location, tissue of origin and diet [40]. Metabolic profiling on a disease specific basis to guide the design of specific media to culture patient-derived primary cells or organoids may therefore be a further step towards greater relevance to tumour biology. Implementing more physiological cell culture conditions with primary cells, frequently refreshed media, use of 3D conditions and consideration of co-cultures to understand tumour cell intrinsic and extrinsic effects, will require increased resource. However, we believe these efforts will result in more relevant target identification, target validation and clinical positioning that will consequently reduce the attrition rate at later stages of drug discovery and clinical development.
Use of clinically relevant in vivo models
Tumours exist in a complex environment containing several non-tumour cell types, including immune cells [41], cancer-associated fibroblasts (CAFs) [42] and specialised epithelial cells [43], that exhibit metabolic re-wiring during tumour development and progression. It has been shown for example that activated T cells exhibit increased nutrient uptake and increased glycolytic flux [44] whilst immunosuppressive regulatory T cells (Treg) are more dependent on oxidative phosphorylation and fatty acid oxidation to meet their energy requirements [45, 46]. Exhausted T cells also exhibit marked metabolic reprogramming, with reduced uptake of glucose and mitochondrial dysfunction being a common feature [47]. The high nutrient requirements and glycolytic activity of cancer cells result in a nutrient depleted, lactate rich microenvironment associated with immune suppression. Understanding this complex relationship may offer novel opportunities for therapeutic intervention. For example, targeting glutamine metabolism with a broad-spectrum glutamine antagonist has been shown to result in nutrient changes within the TME that promote T cell activity and improve the response to immune checkpoint blockade therapy [48]. It should also be noted that metabolic rewiring during disease progression is not limited to the TME but can be systemic in nature [49], further highlighting that metabolic dysfunction should be considered in the context of a whole body. There is therefore a need to understand the impact of novel and existing therapies on the metabolism of non-tumour cells by utilising more complex model systems. Understanding non-tumour cell metabolism also offers an opportunity to identify non-cancer autonomous therapeutic targets, both within the TME and potentially in distal organs. Co-culture experiments offer an in vitro method for target identification and validation studies, but these systems lack complexity. Studying novel targets using genetic or pharmacological methods in immune competent in vivo model systems will more readily define the impact on non-tumour tissue. The range of murine model systems available to the oncology field has been extensively reviewed previously [50], so we will concentrate on highlighting key aspects that should be considered when investigating cancer metabolism.
Transplantation models fall into two major classes: sub-cutaneous or orthotopic transplantation. Sub-cutaneous injection of human cancer cell lines is a commonly used in vivo model system for testing therapeutic agents despite studies that demonstrate they are a poor predictor for clinical activity [51, 52]. Pan-cancer analysis also demonstrate that several metabolic features associated with disease relevant characteristics, such as epithelial–mesenchymal transition, are associated with the tissue in which the tumour is formed [40, 53]. Together, this means that results obtained in sub-cutaneous transplantation models should be interpreted with caution when investigating cancer metabolism.
In orthotopic models, where cancer cells are transplanted into the appropriate tissue locations, tumours face a local metabolic environment relevant to the human pathology. Such orthotopic tumours can also be used to model spontaneous metastasis [54,55,56]. An alternative commonly used experimental murine model of metastasis is systemic introduction of tumour cells via intravascular injection [57]. Metastatic disease is a clinically unmet need and as such is often the disease setting targeted by novel therapeutic agents. Cancer cells undergo metabolic re-wiring during the metastatic cascade and encounter different metabolite environments at secondary sites [2]. Metabolic gene expression signatures differ between primary tumours and either established secondary tumours [58] or micro metastases [59]. It is therefore likely that pre-existing metabolic heterogeneity found within the primary tumour allows for selection of cells with the features required to metastasise to distal organs. Metabolic heterogeneity found within the primary tumour will be influenced by a number of factors including the extent of tumour vascularisation, the infiltration of stromal cells and contiguity to metabolically specialised epithelial cells [43]. Therefore, a fuller understanding of the metabolic differences between primary and metastatic disease and of the influence of primary tumour metabolic heterogeneity on metastatic potential is essential when considering the impact of novel therapies in the clinical setting.
Patient-derived xenograft (PDX) transplant models offer a system that more accurately recapitulate tumour heterogeneity observed in the clinic. There are PDX models available for many cancer indications and collections can be used for mouse based clinical trials to inform human clinical trial design [60]. Such drug sensitivity screens in PDX models have been shown to correlate well with patient response across multiple indications [61]. One of the limitations of PDX models, relevant to the study of metabolic pathways, is that non-tumour cells in the stroma are gradually diluted with murine cells post-transplantation. However, comparative metabolomic analysis of patient colorectal tumours and PDX models demonstrate that murine stromal cells undergo re-wiring to adopt a human-like metabolomic profile that remains stable for several generations [62]. It would be good practice to confirm this for specific PDX models of interest.
Transplantation models generally utilise mice with immune compromised backgrounds, to prevent rejection of human cancer cells. One option to overcome this limitation is to utilise murine cancer cell lines or organoid models for transplantation into syngeneic models. For example, sub-cutaneous transplantation of murine colorectal cancer cell lines have been used to understand differential response to immune checkpoint blockade therapies [63] and orthotopic transplantation of murine derived organoids have been used to model colorectal cancer metastasis in an immune competent setting [64]. This limits studies to murine-derived cancer cells which may show altered target physiology or target dependency compared to humans. Another option is to use humanised mouse models, where human haemopoietic stem cells are engrafted into immune-compromised mice, to model a human immune system. There are several well-established humanised mouse models with applications in oncology and beyond, although challenges include variable lifespans, and incomplete modelling of the human immune system [65].
A murine model system that does not require transplantation is the genetically engineered mouse model (GEMM), where germline alteration of specific cancer relevant oncogenes and/or tumour suppressors are used to generate cancer models that are driven by genetic alterations observed in the clinical setting. The use of GEMMs in oncology research are becoming increasingly popular due to the presence of a functional immune system and relevant tumour microenvironments [66]. A caveat with studying cancer metabolism in advanced murine models, is that the metabolic differences between mice and humans needs to be considered when designing and interpreting a study. The concentration of certain metabolites can differ by up to 10-fold when comparing human and mouse plasma [27], which is highly likely to alter dependency on gene–nutrient interaction and cancer target dependency. Immune cell metabolism and anticancer activity will also differ between mouse and human. This is an area in need of further investigation to better predict the differences in tumour immune response to therapy between species. Another limitation of GEMMs is that the widespread expression of oncogenic drivers can often result in the development of multi-focal tumours that differ from higher grade single tumours that are more commonly found in human cancer. This can result in differing nutrient availability and composition of the tumour microenvironment. Consideration should therefore be given to approaches that induce tumour suppressor loss and/or oncogene expression at focal regions, such as injection of tamoxifen into single sites in the colon, in tamoxifen-inducible Cre recombinase models, resulting in formation of single colonic polyps that progress into advanced carcinomas [67].
Although not the focus of this review, it would be remiss not to highlight developments of in vivo model systems for the extrapolation of drug pharmacokinetic (PK) data from mouse to human. Cytochrome P450 enzymes are the major enzymes involved in metabolism of drugs in vivo and these enzymes differ between mouse and human, that can result in species specific PK profiles. A model system, where 33 mouse P450 enzymes have been knocked out and the major human P450s and two transcription factors responsible for P450 gene expression have been introduced, has been used to accurately predict human PK from a mouse model [68].
Incorporating mitochondrial mutations into pre-clinical models
As described above, efforts to ensure clinical relevance of in vitro and in vivo model systems is of paramount importance when identifying, validating or considering clinical positioning. Recent advances in our understanding of the role of mitochondrial activity and specifically mitochondrial genome mutations coupled with advances in technology for site specific mitochondrial genome editing, allows the development of more complex pre-clinical model systems that may support patient selection for precision medicine approaches.
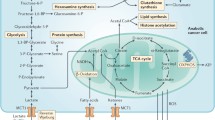
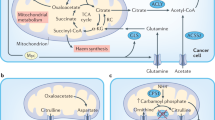
It has long been known that cancer cells upregulate glycolytic activity followed by lactate fermentation in the presence of non-limiting oxygen concentrations, known commonly as the Warburg effect [69, 70]. It has been incorrectly inferred from this observation that mitochondrial activity does not play an important role in cancer progression, but compelling findings demonstrate otherwise [71, 72]. Mitochondrial activity, namely the TCA cycle and oxidative phosphorylation (OXPHOS), are important not only for energy production but also to produce macromolecules that can support cancer anabolism, including those essential for nucleotide production and methylation reactions [71, 72]. The high dependency on mitochondrial activity has led to efforts to target TCA cycle (Devimistat), Glutaminase (Telaglenastat) and complex I (IACS-10759) activity in cancer. However, as described above, these efforts have yet to prove successful in the clinic. Cancer-associated mutations of mitochondrial enzymes have however provided an alternative therapeutic opportunity. Mutation of the TCA cycle enzymes, IDH1 and 2 induces a neomorphic activity that results in production of high levels of the oncometabolite 2-hydroxygluterate (2-HG), a reduction in α-ketoglutarate (α-KG) levels and a consequent inhibition of α-KG-dependent enzymatic processes including histone and DNA demethylation [73]. The recent approval of selective inhibitors of mutated IDH1 and IDH2 in AML [74] provides a clinical proof of concept for targeting metabolic dysfunction in cancer.
It has become apparent in recent years that somatic mutations of the mitochondrial genome may also impact upon cancer progression and response to therapy [75,76,77]. A landmark study in 2021, demonstrated somatic loss-of-function mutations in the mitochondrial genome, particularly truncating mutations impacting on the function of OXPHOS complex I, are highly prevalent across multiple cancer indications [77]. Predicted loss-of-function mutations are associated with altered transcriptional programmes, including downregulation of genes associated with innate immunity, and increased survival of colorectal cancer patients [77] that is speculated to be due to an improved response to treatment. It will be intriguing to analyse available post-clinical trial sequencing data to understand the extent to which the presence of mitochondrial genome mutations influence response to standard-of-care and novel therapies. It is possible that mitochondrial genome mutation status could be utilised as a novel patient stratification marker for specific therapies.
The finding that mitochondrial mutations can stratify patients into clinically distinct populations, demonstrate the need to incorporate cancer-associated mutations into pre-clinical models. It is unclear how stable over time are the levels of mitochondrial mutation heteroplasmy within pre-existing cancer cell lines or murine models. This uncertainty could make data interpretation and reproducibility challenging, highlighting the need to generate and characterise stable genetically engineered model systems. Mitochondrial genome editing to model mitochondrial mutations is technically challenging and conventional nuclear genome editing techniques such as CRISPR are not applicable [78]. Recent technical advancements however have described an elegant protein-based system that utilises an interbacterial toxin DddA that catalyses C:G to T:A conversions in a sequence specific manner [79]. This technique allows the impact of mitochondrial genome mutations to be investigated in vitro and in vivo using genetically engineered pre-clinical models. Utilisation of pre-clinical models that incorporate mitochondrial genome mutations may inform clinical positioning and patient stratification strategies for a broad range of therapeutics but is likely to be particularly important when evaluating dependency of a metabolic target, due to the impact of mitochondrial mutations on the metabolic landscape of the tumour [77].
Metabolic profiling technologies
Metabolomics, a relatively new addition to the omics field, involves detection of low molecular weight metabolites using methods such as nuclear magnetic resonance (NMR) or liquid chromatography mass spectrometry (LC-MS). In the context of this discussion, we consider metabolomics to include the analysis of lipids, often referred to as lipidomics and considered a specialised branch of metabolic research. Metabolomics is underpinned by two methodological approaches; untargeted analysis, a large-scale detection of identified and unidentified metabolites present in a biological sample, or targeted analysis, which is limited to the detection of a list of pre-defined metabolites that, unlike untargeted analysis, can be quantitative.
NMR based metabolomic approaches allow rapid, quantitative and reproducible analysis that preserves the original sample and can be used for liquid (body fluids, cell and tissue extracts) or solid-phase (intact cells or tissues) samples. In contrast, LC-MS analysis is sample consuming and only suitable for liquid-phase samples. LC-MS approaches are generally more sensitive, although the greatest diversity of metabolite detection is achieved when different extraction and chromatography methods are combined. For example, reversed-phase liquid chromatography-MS (RPLC-MS) is considered a standard chromatography method for metabolite detection but alternative methods such as hydrophilic interaction chromatography (HILIC)-MS are more effective for detection of highly polar metabolites [80]. Complementary analytical approaches are therefore recommended to achieve wide metabolite coverage.
The power of metabolomics to reveal tissue and cell type differences in the metabolic landscape is only beginning to be fully realised. Metabolic analysis of tumour samples can be used to detect metabolite biomarkers that are predictive of cancer risk [81, 82], aggressiveness [83, 84], response to therapy [85] and to stratify patients into metabolic based sub-groups [86]. 13C-labelling of glucose or lactate has been used in the clinical setting to assess tumour metabolism in patients, including in paediatric tumours where neuroblastoma-specific metabolic features have been identified [87]. 13C-labelling can also be combined with magnetic resonance imaging (MRI) and MR-spectroscopy (MRS) techniques to allow non-invasive visualisation of the labelled tracer in situ [88].
In addition to analysis of clinical samples, metabolomic analysis of murine models of cancer can be used to understand the differences between murine and human metabolic landscapes to inform model selection and data interpretation [89]. Despite challenges associated with current lack of widespread clinical use [90], the integration of metabolomics with other omics analysis has the potential to transform how clinicians stratify patients by identifying specific metabolic signatures predictive of treatment response.
There are, however, several limitations associated with conventional metabolomic approaches. Small molecule metabolites are not encoded by a primary sequence; therefore their MS-based identification heavily relies on a single parameter, mass-to-charge ratios. There is a lack of standardisation for novel metabolite annotation and analysis software, which together makes data analysis and comparison challenging. In addition, the complexity of the metabolome and its sensitivity to the procedure for sample preparation, means that currently no single method can detect all metabolites present in a complex biological sample.
Conventional LC-MS analysis also lack the ability to map spatial distribution of metabolites in situ. MS-based metabolomic imaging techniques overcome this limitation and have proven useful in the study of cancer metabolism. MS imaging (MSI) techniques such MALDI-MSI [91] and DESI-MSI [92] allow metabolite readout from intact tissue. MALDI-MSI uses a laser, whilst DESI-MSI utilises charged solvent droplets directed towards the sample, to ionise and desorb particles that are detected by the MS. The alternative extraction processes and differing metabolite coverage of MALDI and DESI-MSI techniques is an important factor to consider when analysing and comparing datasets.
In MSI, MS-based analysis is combined with spatial information from the input sample to generate images illustrating spatial distribution of detected metabolites. DESI-MSI has been used to visualise amino acid levels in situ from murine KRAS mutant colorectal tumours as part of a study that identified the amino acid transporter SLC7A5 as a therapeutic vulnerability of KRAS-mutant colorectal cancers [93]. MSI is a clinically applicable tool that could be used for diagnostic or patient stratification purposes when applied to tissue samples from surgically resected tumours. In support of the use of MS profiling of clinical samples, the iKnife based on a rapid evaporative ionisation mass spectrometry (REIMS) method uses MS-based quantification of metabolites in real time during surgical resection and is capable of distinguishing normal and malignant cells and of identifying tumour type [94,95,96].
Tumour heterogeneity, arising from different cell types or molecular alterations between cells of the same origin, can influence response to therapy and development of resistance [97]. Such heterogeneity will not be captured by bulk metabolic profiling of bodily fluids such as blood, urine or tumour interstitial fluid. Single-cell metabolomic analysis aims to address this by analysing metabolic profiles of individual cells within a given population [98]. This technique has the potential to identify sub-populations of tumour cells that may be dependent on specific metabolic processes or predict the emergence of treatment resistance. The high costs, low throughput and scale of data that needs to be generated and analysed, limits the accessibility of this approach. However, recent efforts to generate open-source and accessible single-cell metabolomic approach have combined MALDI-MSI and light microscopy to provide MALDI-MSI data at a cellular level [99]. This approach, named SpaceM, has several advantages over standard approaches including the possibility to be used in standard cell culture, higher throughput and ability to metabolically profile cells with specific features such as morphology or fluorescent markers.
An alternative method of metabolic profiling, that can be used to provide spatial information at the single cell level is Raman spectroscopy. Raman spectroscopy uses a monochromatic laser directed at a molecule of interest and subsequent Raman scattering produces light with different frequency, generating a Raman spectrum [100]. The unique signature of each constituent molecule can be identified from a complex Raman spectrum profile arising from a heterogeneous sample. Raman spectroscopy has multiple potential oncology applications, including in cancer diagnosis, where Raman spectrum profiles can be used to distinguish between malignant and non-malignant samples [101]. Raman imaging can also be used to identify the presence of metabolites at a sub-cellular level, aiding the understanding of organelle specific functions [102]. This technique has clear utility in determining whether therapeutic agents reach cell types and sub-cellular organelles where the disease relevant target function takes place. However, Raman spectroscopy is not as well established as MS-based methods and the range of metabolite detection is more limited. As more datasets are generated, this range will increase and establish Raman spectroscopy imaging as another powerful component in the metabolomic toolbox.
Future clinical directions
For a summary of therapeutic agents targeting metabolic processes that are in clinical development, we direct the reader to a recent comprehensive review by Lemberg and colleagues [103].
Clinical success of metabolism-targeting agents will ultimately depend on selecting a defined patient population most likely to benefit from therapy, as exemplified by the approval of mutant IDH1/2 inhibitors in patients with an IDH1/2 mutation. Conversely, patients with non-small cell lung cancer predicted to be dependent on glutaminolysis due to KEAP1 loss of function mutation [104], did not show a sufficient response to therapy with GLS inhibitors. This suggests that the success of future clinical trials targeting cancer metabolism may depend on the enrichment for responders that should be guided by patient stratification markers, such as dysregulation of nutrient transporters [105], beyond the genetic markers currently in use. As discussed in the previous sections, several metabolomic profiling technologies are now available to identify metabolic signatures from patient-derived samples. Whilst being mindful of mouse-human metabolic differences, emphasis should be placed on using clinically relevant models to identify and test patient stratification markers before moving into clinical development.
An example of a promising novel agent in clinical development that is targeting a defined patient population based on a genetically induced metabolic vulnerability is the MAT2A inhibitor AG-270 [106]. MAT2A is the first enzyme in the methionine cycle and catalyses the synthesis of the methyl-donor S-adenosylmethionine (SAM). The methionine cycle is required to replenish the pool of methionine for protein production, DNA and protein methylation, and to sustain nucleotide production via the folate cycle [107]. It has been demonstrated that MAT2A activity is selectively required for proliferation of tumours lacking expression of the methionine/adenine salvage gene MTAP [106, 108]. AG-270 is in phase 1 clinical trials, as a single agent or in combination with docetaxel in MTAP-deleted non-small cell lung cancer patients and in combination with nab (nanoparticle albumin-bound)-paclitaxel and gemcitabine in MTAP-deleted pancreatic ductal adenocarcinoma (NCT03435250).
In addition to agents in clinical or pre-clinical development, there are classes of metabolic proteins that are currently underrepresented in cancer therapy and may provide a source of novel targets going forward. One such class is the solute carrier transporter (SLC) family, comprising approximately 400 proteins that transport solutes, including amino acids, metals and other nutrients across plasma and organelle membranes [109]. There are very few examples of compounds targeting SLC family members, such as JPH-203 [110] and AZD3965 [111], that are in clinical development for oncology. The well-established differences in nutrient requirement between normal and cancer cells suggests that targeting solute uptake or efflux could be an effective strategy. A consortium between academia and pharmaceutical partners, RESOLUTE, has been formed with the specific aim to establish SLCs as a tractable class of novel drug targets [112]. Investigating this target class using physiologically relevant conditions and suitable in vivo models in defined disease settings has the potential to identify novel therapeutic opportunities within this large family [113, 114].
Although the focus of oncology drug discovery remains mainly on small molecule inhibitors, the field should also be aware of the opportunities that will arise from novel therapeutic modalities. Protein degradation approaches, such as PROTAC, are a rapidly expanding field, with several agents now in clinical development in oncology settings [115]. Degrader technology allows targets previously considered non-tractable by drug discovery to be re-evaluated in this context. For example, the RESOLUTE consortium has published a proof of concept using the first SLC PROTAC that induces selective loss of viability in cancer cell lines [116]. mRNA vaccines designed against IDH1 mutant neoepitopes, have been tested, as a single agent [117] and in combination with anti-PDL1 immune checkpoint therapy [118] in phase 1 clinical trials to induce a tumour immune response in IDH1 mutant glioma patients. Dietary manipulation is also gaining traction as a viable approach to be used alongside pharmacological approaches [119]. There are multiple studies in murine models of cancer, that demonstrate efficacy of amino acid restriction [120, 121] or dietary supplementation with sugars such as mannose [122], either alone or in a combination setting. Indeed, studies have demonstrated that cancer cells can be sensitised to radiotherapy by deprivation of specific amino acids [121, 123]. Clinical support for approaches based on amino acid restriction comes from the well-established use of asparaginases for the treatment of paediatric ALL (Table 1). In addition, promising data emerging from phase 2/3 clinical trials in mesothelioma patients has recently been shared for pegargiminase, a PEGylated arginine deiminase, which acts therapeutically by depleting arginine [124].
Metabolic plasticity of cancer, and cancer-associated cells can lead to the emergence of resistance to single agents that target metabolic processes. Combination strategies, have the potential to induce cellular dependency on specific metabolic processes and reduce the occurrence of intrinsic and acquired resistance. For example, inhibition of downstream components of the KRAS pathway in KRAS mutant cancers result in metabolic dysregulation and increased autophagic flux. Dual targeting of KRAS downstream pathway members and autophagy has proved successful in the pre-clinical setting [125, 126] and is currently being investigated in the clinic (NCT04892017). Combination approaches should therefore be considered early in the drug discovery process to both highlight specific metabolic dependencies that could be an opportunity for therapeutic intervention and mitigate the risk of resistance.
Perspectives
Although the first anti-metabolite drug, aminopterin, was approved in 1948, the modern cancer metabolism field is relatively young. The first wave of modern therapeutic agents has resulted in clinical success, with the approval of IDH mutant inhibitors, but also high-profile clinical trial failures. Consideration should be given to re-visiting existing agents to determine whether a different approach to identify defined clinical positioning offers fresh opportunities to evaluate efficacy. This involves learning from previous trial failures and designing future trials that offer the best chance of success, as modelled by the discussion on the clinical future of IDO1 inhibitors [127]. The recent development in technologies offer exciting opportunities to better understand cancer metabolism, supporting the identification and validation of novel targets and design of optimal clinical positioning strategies to expedite the delivery of patient benefit. Researchers should consider targeting not only cancer cell intrinsic metabolism but also the metabolism of immune and other non-cancer cells, that may offer unexplored therapeutic opportunities. We believe that keys to clinical success for novel metabolic interventions include biomarker-led patient selection, incorporation of metabolic targeting agents into combinatorial strategies to delay the occurrence of resistance, and utilisation of novel therapeutic modalities that expand the druggable genome (Fig. 1).
Schematic to illustrate how incorporating the technologies discussed within this review in the pre-clinical setting can impact clinical development of novel agents targeting cancer metabolism. Keys to clinical success include consideration of novel therapeutic modalities, biomarker led patient selection and incorporating combination strategies. Targets of novel therapeutic agents may include cancer cell intrinsic metabolism together with immune, epithelial and stromal cell metabolism.
Data availability
Non-applicable.
References
Keibler MA, Wasylenko TM, Kelleher JK, Iliopoulos O, Vander Heiden MG, Stephanopoulos G. Metabolic requirements for cancer cell proliferation. Cancer Metab. 2016;4:16.
Bergers G, Fendt SM. The metabolism of cancer cells during metastasis. Nat Rev Cancer. 2021;21:162–80.
Hanahan D. Hallmarks of cancer: new dimensions. Cancer Discov. 2022;12:31–46.
Farber S, Diamond LK. Temporary remissions in acute leukemia in children produced by folic acid antagonist, 4-aminopteroyl-glutamic acid. N Engl J Med. 1948;238:787–93.
Visentin M, Zhao R, Goldman ID. The antifolates. Hematol Oncol Clin North Am. 2012;26:629–48.
Vodenkova S, Buchler T, Cervena K, Veskrnova V, Vodicka P, Vymetalkova V. 5-Fluorouracil and other fluoropyrimidines in colorectal cancer: past, present and future. Pharmacol Ther. 2020;206:107447.
Madaan K, Kaushik D, Verma T. Hydroxyurea: a key player in cancer chemotherapy. Expert Rev Anticancer Ther. 2012;12:19–29.
Gesto DS, Cerqueira NM, Fernandes PA, Ramos MJ. Gemcitabine: a critical nucleoside for cancer therapy. Curr Med Chem. 2012;19:1076–87.
Harris AL. Development of cancer metabolism as a therapeutic target: new pathways, patient studies, stratification and combination therapy. Br J Cancer. 2020;122:1–3.
Luengo A, Gui DY, Vander Heiden MG. Targeting metabolism for cancer therapy. Cell Chem Biol. 2017;24:1161–80.
Seth Nanda C, Venkateswaran SV, Patani N, Yuneva M. Defining a metabolic landscape of tumours: genome meets metabolism. Br J Cancer. 2020;122:136–49.
Dang CV, Semenza GL. Oncogenic alterations of metabolism. Trends Biochem Sci. 1999;24:68–72.
Boroughs LK, DeBerardinis RJ. Metabolic pathways promoting cancer cell survival and growth. Nat Cell Biol. 2015;17:351–9.
Targeting tumour metabolism. Nat Rev Drug Discov. 2010;9:503-4.
Yap TA, Daver N, Mahendra M, Zhang J, Kamiya-Matsuoka C, Meric-Bernstam F, et al. Complex I inhibitor of oxidative phosphorylation in advanced solid tumors and acute myeloid leukemia: phase I trials. Nat Med. 2023;29:115–26.
Long GV, Dummer R, Hamid O, Gajewski TF, Caglevic C, Dalle S, et al. Epacadostat plus pembrolizumab versus placebo plus pembrolizumab in patients with unresectable or metastatic melanoma (ECHO-301/KEYNOTE-252): a phase 3, randomised, double-blind study. Lancet Oncol. 2019;20:1083–97.
Boysen G, Jamshidi-Parsian A, Davis MA, Siegel ER, Simecka CM, Kore RA, et al. Glutaminase inhibitor CB-839 increases radiation sensitivity of lung tumor cells and human lung tumor xenografts in mice. Int J Radiat Biol. 2019;95:436–42.
Hamada S, Matsumoto R, Tanaka Y, Taguchi K, Yamamoto M, Masamune A. Nrf2 activation sensitizes K-ras mutant pancreatic cancer cells to glutaminase inhibition. Int J Mol Sci. 2021;22:1870.
Gao L, Xu Z, Huang Z, Tang Y, Yang D, Huang J, et al. CPI-613 rewires lipid metabolism to enhance pancreatic cancer apoptosis via the AMPK-ACC signaling. J Exp Clin Cancer Res. 2020;39:73.
Yue EW, Sparks R, Polam P, Modi D, Douty B, Wayland B, et al. INCB24360 (Epacadostat), a highly potent and selective indoleamine-2,3-dioxygenase 1 (IDO1) inhibitor for immuno-oncology. ACS Med Chem Lett. 2017;8:486–91.
Alexander Vandell JE, Hoffman Steve, Del Priore Giuseppe, Fernandez-Zapico Martin. In vitro and in vivo anticancer effects of D/L-alpha-metyrosine (SM-88), a novel metabolism-based therapy [abstract]. Proc Annu Meet Am Assoc Cancer Res. 2020;80:Abstract nr 5998.
Molina JR, Sun Y, Protopopova M, Gera S, Bandi M, Bristow C, et al. An inhibitor of oxidative phosphorylation exploits cancer vulnerability. Nat Med. 2018;24:1036–46.
DiMasi JA, Grabowski HG, Hansen RW. Innovation in the pharmaceutical industry: new estimates of R&D costs. J Health Econ. 2016;47:20–33.
Dulbecco R, Freeman G. Plaque production by the polyoma virus. Virology. 1959;8:396–7.
Moore GE, Gerner RE, Franklin HA. Culture of normal human leukocytes. JAMA. 1967;199:519–24.
Muir A, Danai LV, Gui DY, Waingarten CY, Lewis CA, Vander Heiden MG. Environmental cystine drives glutamine anaplerosis and sensitizes cancer cells to glutaminase inhibition. Elife. 2017;6:e27713.
Cantor JR, Abu-Remaileh M, Kanarek N, Freinkman E, Gao X, Louissaint A Jr, et al. Physiologic medium rewires cellular metabolism and reveals uric acid as an endogenous inhibitor of UMP synthase. Cell. 2017;169:258.e17–72 e17.
Vande Voorde J, Ackermann T, Pfetzer N, Sumpton D, Mackay G, Kalna G, et al. Improving the metabolic fidelity of cancer models with a physiological cell culture medium. Sci Adv. 2019;5:eaau7314.
Ackermann T, Tardito S. Cell culture medium formulation and its implications in cancer metabolism. Trends Cancer. 2019;5:329–32.
Sullivan MR, Darnell AM, Reilly MF, Kunchok T, Joesch-Cohen L, Rosenberg D, et al. Methionine synthase is essential for cancer cell proliferation in physiological folate environments. Nat Metab. 2021;3:1500–11.
Ghergurovich JM, Xu X, Wang JZ, Yang L, Ryseck RP, Wang L, et al. Methionine synthase supports tumour tetrahydrofolate pools. Nat Metab. 2021;3:1512–20.
Rossiter NJ, Huggler KS, Adelmann CH, Keys HR, Soens RW, Sabatini DM, et al. CRISPR screens in physiologic medium reveal conditionally essential genes in human cells. Cell Metab. 2021;33:1248.e9–63.e9.
Gee S, Nelson N, Bornot A, Carter N, Cuomo ME, Dovedi SJ, et al. Developing an arrayed CRISPR-Cas9 co-culture screen for immuno-oncology target ID. SLAS Discov. 2020;25:581–90.
Leney-Greene MA, Boddapati AK, Su HC, Cantor JR, Lenardo MJ. Human plasma-like medium improves T lymphocyte activation. iScience. 2020;23:100759.
Farhat J, Pandey I, AlWahsh M. Transcending toward advanced 3D-cell culture modalities: a review about an emerging paradigm in translational oncology. Cells. 2021;10:1657.
Han K, Pierce SE, Li A, Spees K, Anderson GR, Seoane JA, et al. CRISPR screens in cancer spheroids identify 3D growth-specific vulnerabilities. Nature. 2020;580:136–41.
Ringel T, Frey N, Ringnalda F, Janjuha S, Cherkaoui S, Butz S, et al. Genome-scale CRISPR screening in human intestinal organoids identifies drivers of TGF-beta resistance. Cell Stem Cell. 2020;26:431.e8–40.e8.
Coluccio ML, Perozziello G, Malara N, Parrotta E, Zhang P, Gentile F, et al. Microfluidic platforms for cell cultures and investigations. Microelectron Eng. 2019;208:14–28.
Birsoy K, Possemato R, Lorbeer FK, Bayraktar EC, Thiru P, Yucel B, et al. Metabolic determinants of cancer cell sensitivity to glucose limitation and biguanides. Nature. 2014;508:108–12.
Sullivan MR, Danai LV, Lewis CA, Chan SH, Gui DY, Kunchok T, et al. Quantification of microenvironmental metabolites in murine cancers reveals determinants of tumor nutrient availability. Elife. 2019;8:e44235.
Talty R, Olino K. Metabolism of innate immune cells in cancer. Cancers. 2021;13:904.
Li Z, Sun C, Qin Z. Metabolic reprogramming of cancer-associated fibroblasts and its effect on cancer cell reprogramming. Theranostics. 2021;11:8322–36.
Altea-Manzano P, Doglioni G, Liu Y, Cuadros AM, Nolan E, Fernandez-Garcia J, et al. A palmitate-rich metastatic niche enables metastasis growth via p65 acetylation resulting in pro-metastatic NF-kappaB signaling. Nat Cancer. 2023;4:344–64.
Frauwirth KA, Riley JL, Harris MH, Parry RV, Rathmell JC, Plas DR, et al. The CD28 signaling pathway regulates glucose metabolism. Immunity. 2002;16:769–77.
Beier UH, Angelin A, Akimova T, Wang L, Liu Y, Xiao H, et al. Essential role of mitochondrial energy metabolism in Foxp3(+) T-regulatory cell function and allograft survival. FASEB J. 2015;29:2315–26.
Michalek RD, Gerriets VA, Jacobs SR, Macintyre AN, MacIver NJ, Mason EF, et al. Cutting edge: distinct glycolytic and lipid oxidative metabolic programs are essential for effector and regulatory CD4+ T cell subsets. J Immunol. 2011;186:3299–303.
Bengsch B, Johnson AL, Kurachi M, Odorizzi PM, Pauken KE, Attanasio J, et al. Bioenergetic insufficiencies due to metabolic alterations regulated by the inhibitory receptor PD-1 are an early driver of CD8(+) T cell exhaustion. Immunity. 2016;45:358–73.
Leone RD, Zhao L, Englert JM, Sun IM, Oh MH, Sun IH, et al. Glutamine blockade induces divergent metabolic programs to overcome tumor immune evasion. Science. 2019;366:1013–21.
Naser FJ, Jackstadt MM, Fowle-Grider R, Spalding JL, Cho K, Stancliffe E, et al. Isotope tracing in adult zebrafish reveals alanine cycling between melanoma and liver. Cell Metab. 2021;33:1493–504.e5.
Day CP, Merlino G, Van, Dyke T. Preclinical mouse cancer models: a maze of opportunities and challenges. Cell. 2015;163:39–53.
Voskoglou-Nomikos T, Pater JL, Seymour L. Clinical predictive value of the in vitro cell line, human xenograft, and mouse allograft preclinical cancer models. Clin Cancer Res. 2003;9:4227–39.
Johnson JI, Decker S, Zaharevitz D, Rubinstein LV, Venditti JM, Schepartz S, et al. Relationships between drug activity in NCI preclinical in vitro and in vivo models and early clinical trials. Br J Cancer. 2001;84:1424–31.
Gaude E, Frezza C. Tissue-specific and convergent metabolic transformation of cancer correlates with metastatic potential and patient survival. Nat Commun. 2016;7:13041.
Paschall AV, Liu K. An orthotopic mouse model of spontaneous breast cancer metastasis. J Vis Exp. 2016:54040.
Zhang G, Du YN. Orthotopic pancreatic tumor mouse models of liver metastasis. Methods Mol Biol. 2019;1882:309–20.
Zhang L, Bu P. Generation of an orthotopic mouse model to study colorectal cancer metastasis. STAR Protoc. 2021;2:100792.
Gomez-Cuadrado L, Tracey N, Ma R, Qian B, Brunton VG. Mouse models of metastasis: progress and prospects. Dis Model Mech. 2017;10:1061–74.
Chaika NV, Yu F, Purohit V, Mehla K, Lazenby AJ, DiMaio D, et al. Differential expression of metabolic genes in tumor and stromal components of primary and metastatic loci in pancreatic adenocarcinoma. PLoS ONE. 2012;7:e32996.
Davis RT, Blake K, Ma D, Gabra MBI, Hernandez GA, Phung AT, et al. Transcriptional diversity and bioenergetic shift in human breast cancer metastasis revealed by single-cell RNA sequencing. Nat Cell Biol. 2020;22:310–20.
Byrne AT, Alferez DG, Amant F, Annibali D, Arribas J, Biankin AV, et al. Interrogating open issues in cancer precision medicine with patient-derived xenografts. Nat Rev Cancer. 2017;17:254–68.
Gao H, Korn JM, Ferretti S, Monahan JE, Wang Y, Singh M, et al. High-throughput screening using patient-derived tumor xenografts to predict clinical trial drug response. Nat Med. 2015;21:1318–25.
Blomme A, Van Simaeys G, Doumont G, Costanza B, Bellier J, Otaka Y, et al. Murine stroma adopts a human-like metabolic phenotype in the PDX model of colorectal cancer and liver metastases. Oncogene. 2018;37:1237–50.
Sato Y, Fu Y, Liu H, Lee MY, Shaw MH. Tumor-immune profiling of CT-26 and Colon 26 syngeneic mouse models reveals mechanism of anti-PD-1 response. BMC Cancer. 2021;21:1222.
O’Rourke KP, Loizou E, Livshits G, Schatoff EM, Baslan T, Manchado E, et al. Transplantation of engineered organoids enables rapid generation of metastatic mouse models of colorectal cancer. Nat Biotechnol. 2017;35:577–82.
Allen TM, Brehm MA, Bridges S, Ferguson S, Kumar P, Mirochnitchenko O, et al. Humanized immune system mouse models: progress, challenges and opportunities. Nat Immunol. 2019;20:770–4.
Kersten K, de Visser KE, van Miltenburg MH, Jonkers J. Genetically engineered mouse models in oncology research and cancer medicine. EMBO Mol Med. 2017;9:137–53.
Knight JRP, Alexandrou C, Skalka GL, Vlahov N, Pennel K, Officer L, et al. MNK inhibition sensitizes KRAS-mutant colorectal cancer to mTORC1 inhibition by reducing eIF4E phosphorylation and c-MYC expression. Cancer Discov. 2021;11:1228–47.
Henderson CJ, Kapelyukh Y, Scheer N, Rode A, McLaren AW, MacLeod AK, et al. An extensively humanized mouse model to predict pathways of drug disposition and drug/drug interactions, and to facilitate design of clinical trials. Drug Metab Dispos. 2019;47:601–15.
Hardie DG. 100 years of the Warburg effect: a historical perspective. Endocr Relat Cancer. 2022;29:T1–T13.
Liberti MV, Locasale JW. The Warburg effect: how does it benefit cancer cells? Trends Biochem Sci. 2016;41:211–8.
Oliveira GL, Coelho AR, Marques R, Oliveira PJ. Cancer cell metabolism: rewiring the mitochondrial hub. Biochim Biophys Acta Mol Basis Dis. 2021;1867:166016.
Vasan K, Werner M, Chandel NS. Mitochondrial metabolism as a target for cancer therapy. Cell Metab. 2020;32:341–52.
Han S, Liu Y, Cai SJ, Qian M, Ding J, Larion M, et al. IDH mutation in glioma: molecular mechanisms and potential therapeutic targets. Br J Cancer. 2020;122:1580–9.
Issa GC, DiNardo CD. Acute myeloid leukemia with IDH1 and IDH2 mutations: 2021 treatment algorithm. Blood Cancer J. 2021;11:107.
Grandhi S, Bosworth C, Maddox W, Sensiba C, Akhavanfard S, Ni Y, et al. Heteroplasmic shifts in tumor mitochondrial genomes reveal tissue-specific signals of relaxed and positive selection. Hum Mol Genet. 2017;26:2912–22.
Smith AL, Whitehall JC, Bradshaw C, Gay D, Robertson F, Blain AP, et al. Age-associated mitochondrial DNA mutations cause metabolic remodelling that contributes to accelerated intestinal tumorigenesis. Nat Cancer. 2020;1:976–89.
Gorelick AN, Kim M, Chatila WK, La K, Hakimi AA, Berger MF, et al. Respiratory complex and tissue lineage drive recurrent mutations in tumour mtDNA. Nat Metab. 2021;3:558–70.
Gammage PA, Moraes CT, Minczuk M. Mitochondrial genome engineering: the revolution may not be CRISPR-ized. Trends Genet. 2018;34:101–10.
Mok BY, de Moraes MH, Zeng J, Bosch DE, Kotrys AV, Raguram A, et al. A bacterial cytidine deaminase toxin enables CRISPR-free mitochondrial base editing. Nature. 2020;583:631–7.
Harrieder EM, Kretschmer F, Bocker S, Witting M. Current state-of-the-art of separation methods used in LC-MS based metabolomics and lipidomics. J Chromatogr B Anal Technol Biomed Life Sci. 2022;1188:123069.
His M, Viallon V, Dossus L, Gicquiau A, Achaintre D, Scalbert A, et al. Prospective analysis of circulating metabolites and breast cancer in EPIC. BMC Med. 2019;17:178.
Huang J, Mondul AM, Weinstein SJ, Derkach A, Moore SC, Sampson JN, et al. Prospective serum metabolomic profiling of lethal prostate cancer. Int J Cancer. 2019;145:3231–43.
Vandergrift LA, Decelle EA, Kurth J, Wu S, Fuss TL, DeFeo EM, et al. Metabolomic prediction of human prostate cancer aggressiveness: magnetic resonance spectroscopy of histologically benign tissue. Sci Rep. 2018;8:4997.
Puchades-Carrasco L, Jantus-Lewintre E, Perez-Rambla C, Garcia-Garcia F, Lucas R, Calabuig S, et al. Serum metabolomic profiling facilitates the non-invasive identification of metabolic biomarkers associated with the onset and progression of non-small cell lung cancer. Oncotarget. 2016;7:12904–16.
He X, Gu J, Zou D, Yang H, Zhang Y, Ding Y, et al. NMR-based metabolomics analysis predicts response to neoadjuvant chemotherapy for triple-negative breast cancer. Front Mol Biosci. 2021;8:708052.
Xiao Y, Ma D, Yang YS, Yang F, Ding JH, Gong Y, et al. Comprehensive metabolomics expands precision medicine for triple-negative breast cancer. Cell Res. 2022;32:477–90.
Johnston K, Pachnis P, Tasdogan A, Faubert B, Zacharias LG, Vu HS, et al. Isotope tracing reveals glycolysis and oxidative metabolism in childhood tumors of multiple histologies. Med. 2021;2:395–410.
Woitek R, Gallagher FA. The use of hyperpolarised (13)C-MRI in clinical body imaging to probe cancer metabolism. Br J Cancer. 2021;124:1187–98.
Araujo R, Bispo D, Helguero LA, Gil AM. Metabolomic studies of breast cancer in murine models: a review. Biochim Biophys Acta Mol Basis Dis. 2020;1866:165713.
Pinu FR, Goldansaz SA, Jaine J. Translational metabolomics: current challenges and future opportunities. Metabolites 2019;9:108.
Norris JL, Caprioli RM. Analysis of tissue specimens by matrix-assisted laser desorption/ionization imaging mass spectrometry in biological and clinical research. Chem Rev. 2013;113:2309–42.
Claude E, Jones EA, Pringle SD. DESI mass spectrometry imaging (MSI). Methods Mol Biol. 2017;1618:65–75.
Najumudeen AK, Ceteci F, Fey SK, Hamm G, Steven RT, Hall H, et al. The amino acid transporter SLC7A5 is required for efficient growth of KRAS-mutant colorectal cancer. Nat Genet. 2021;53:16–26.
St John ER, Balog J, McKenzie JS, Rossi M, Covington A, Muirhead L, et al. Rapid evaporative ionisation mass spectrometry of electrosurgical vapours for the identification of breast pathology: towards an intelligent knife for breast cancer surgery. Breast Cancer Res. 2017;19:59.
Tzafetas M, Mitra A, Paraskevaidi M, Bodai Z, Kalliala I, Bowden S, et al. The intelligent knife (iKnife) and its intraoperative diagnostic advantage for the treatment of cervical disease. Proc Natl Acad Sci USA. 2020;117:7338–46.
Alexander J, Gildea L, Balog J, Speller A, McKenzie J, Muirhead L, et al. A novel methodology for in vivo endoscopic phenotyping of colorectal cancer based on real-time analysis of the mucosal lipidome: a prospective observational study of the iKnife. Surg Endosc. 2017;31:1361–70.
Dagogo-Jack I, Shaw AT. Tumour heterogeneity and resistance to cancer therapies. Nat Rev Clin Oncol. 2018;15:81–94.
Wei D, Xu M, Wang Z, Tong J. The development of single-cell metabolism and its role in studying cancer emergent properties. Front Oncol. 2021;11:814085.
Rappez L, Stadler M, Triana S, Gathungu RM, Ovchinnikova K, Phapale P, et al. SpaceM reveals metabolic states of single cells. Nat Methods. 2021;18:799–805.
Jones RR, Hooper DC, Zhang L, Wolverson D, Valev VK. Raman techniques: fundamentals and frontiers. Nanoscale Res Lett. 2019;14:231.
Auner GW, Koya SK, Huang C, Broadbent B, Trexler M, Auner Z, et al. Applications of Raman spectroscopy in cancer diagnosis. Cancer Metastasis Rev. 2018;37:691–717.
Xu J, Yu T, Zois CE, Cheng JX, Tang Y, Harris AL, et al. Unveiling cancer metabolism through spontaneous and coherent Raman spectroscopy and stable isotope probing. Cancers. 2021;13:1718.
Lemberg KM, Gori SS, Tsukamoto T, Rais R, Slusher BS. Clinical development of metabolic inhibitors for oncology. J Clin Investig. 2022;132:e148550.
Romero R, Sayin VI, Davidson SM, Bauer MR, Singh SX, LeBoeuf SE, et al. Keap1 loss promotes Kras-driven lung cancer and results in dependence on glutaminolysis. Nat Med. 2017;23:1362–8.
Zheng D, Wei Z, Guo W. Identification of a solute carrier family-based signature for predicting overall survival in osteosarcoma. Front Genet. 2022;13:849789.
Konteatis Z, Travins J, Gross S, Marjon K, Barnett A, Mandley E, et al. Discovery of AG-270, a first-in-class oral MAT2A inhibitor for the treatment of tumors with homozygous MTAP deletion. J Med Chem. 2021;64:4430–49.
Lauinger L, Kaiser P. Sensing and signaling of methionine metabolism. Metabolites. 2021;11:83.
Kalev P, Hyer ML, Gross S, Konteatis Z, Chen CC, Fletcher M, et al. MAT2A inhibition blocks the growth of MTAP-deleted cancer cells by reducing PRMT5-dependent mRNA splicing and inducing DNA damage. Cancer Cell. 2021;39:209.e11–24 e11.
Lin L, Yee SW, Kim RB, Giacomini KM. SLC transporters as therapeutic targets: emerging opportunities. Nat Rev Drug Discov. 2015;14:543–60.
Okano N, Naruge D, Kawai K, Kobayashi T, Nagashima F, Endou H, et al. First-in-human phase I study of JPH203, an L-type amino acid transporter 1 inhibitor, in patients with advanced solid tumors. Invest N Drugs. 2020;38:1495–506.
Halford S, Veal GJ, Wedge SR, Payne GS, Bacon CM, Sloan P, et al. A phase I dose-escalation study of AZD3965, an oral monocarboxylate transporter 1 inhibitor, in patients with advanced cancer. Clin Cancer Res. 2023;29:1429–39.
Superti-Furga G, Lackner D, Wiedmer T, Ingles-Prieto A, Barbosa B, Girardi E, et al. The RESOLUTE consortium: unlocking SLC transporters for drug discovery. Nat Rev Drug Discov. 2020;19:429–30.
Rebsamen M, Girardi E, Sedlyarov V, Scorzoni S, Papakostas K, Vollert M, et al. Gain-of-function genetic screens in human cells identify SLC transporters overcoming environmental nutrient restrictions. Life Sci Alliance. 2022;5:e202201404.
Chidley C, Darnell AM, Gaudio BL, Lien EC, Barbeau AM, Vander Heiden MG, et al. A CRISPRi/a screening platform to study cellular nutrient transport in diverse microenvironments. bioRxiv:2023.01.26.525375v1 [Preprint]. Cited [2023 Jan 26]: [39 p.]. Available from: https://doi.org/10.1101/2023.01.26.525375.
Bekes M, Langley DR, Crews CM. PROTAC targeted protein degraders: the past is prologue. Nat Rev Drug Discov. 2022;21:181–200.
Bensimon A, Pizzagalli MD, Kartnig F, Dvorak V, Essletzbichler P, Winter GE, et al. Targeted degradation of SLC transporters reveals amenability of multi-pass transmembrane proteins to ligand-induced proteolysis. Cell Chem Biol. 2020;27:728.e9–39.e9.
Platten M, Bunse L, Wick A, Bunse T, Le Cornet L, Harting I, et al. A vaccine targeting mutant IDH1 in newly diagnosed glioma. Nature. 2021;592:463–8.
Bunse L, Rupp AK, Poschke I, Bunse T, Lindner K, Wick A, et al. AMPLIFY-NEOVAC: a randomized, 3-arm multicenter phase I trial to assess safety, tolerability and immunogenicity of IDH1-vac combined with an immune checkpoint inhibitor targeting programmed death-ligand 1 in isocitrate dehydrogenase 1 mutant gliomas. Neurol Res Pract. 2022;4:20.
Goncalves MD, Maddocks OD. Engineered diets to improve cancer outcomes. Curr Opin Biotechnol. 2021;70:29–35.
Maddocks ODK, Athineos D, Cheung EC, Lee P, Zhang T, van den Broek NJF, et al. Modulating the therapeutic response of tumours to dietary serine and glycine starvation. Nature. 2017;544:372–6.
Gao X, Sanderson SM, Dai Z, Reid MA, Cooper DE, Lu M, et al. Dietary methionine influences therapy in mouse cancer models and alters human metabolism. Nature. 2019;572:397–401.
Gonzalez PS, O’Prey J, Cardaci S, Barthet VJA, Sakamaki JI, Beaumatin F, et al. Mannose impairs tumour growth and enhances chemotherapy. Nature. 2018;563:719–23.
Falcone M, Uribe AH, Papalazarou V, Newman AC, Athineos D, Stevenson K, et al. Sensitisation of cancer cells to radiotherapy by serine and glycine starvation. Br J Cancer. 2022;127:1773–86.
Slzlosarek WP, Creelan B, Sarkodie T, Nolan L, Taylor P, Olevsky O, et al. Phase 2-3 trial of pegargiminase plus chemotherapy versus placebo plus chemotherapy in patients with non-epithelioid pleural mesothelioma [abstract]. Proceedings of the American Association for Cancer Research Annual Meeting 2023; Part 2 (Clinical Trials and Late-Breaking Research). 2023.
Kinsey CG, Camolotto SA, Boespflug AM, Guillen KP, Foth M, Truong A, et al. Protective autophagy elicited by RAF->MEK->ERK inhibition suggests a treatment strategy for RAS-driven cancers. Nat Med. 2019;25:620–7.
Bryant KL, Stalnecker CA, Zeitouni D, Klomp JE, Peng S, Tikunov AP, et al. Combination of ERK and autophagy inhibition as a treatment approach for pancreatic cancer. Nat Med. 2019;25:628–40.
Eynde BJVD, Baren NV, Baurain J-F. Is there a clinical future for IDO1 inhibitors after the failure of epacadostat in melanoma? Annu Rev Cancer Biol. 2020;4:241–56.
Acknowledgements
We would like to acknowledge all members of Cancer Research Horizons and the Cancer Research UK Beatson Institute who were involved in relevant discussions or gave suggestions based on draft versions of this review. We particularly want to thank Neil Jones, Nathan Breeds and David Sumpton.
Funding
This work was funded by Cancer Research UK award A23982 to ST.
Author information
Authors and Affiliations
Contributions
ST drafted and revised the manuscript and approved the final version. CM conceived the review, drafted and revised the manuscript and approved the final version.
Corresponding author
Ethics declarations
Competing interests
ST is the inventor of PlasmaxTM medium.
Ethics approval and consent to participate
Non-applicable.
Consent for publication
Non-applicable.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tardito, S., MacKay, C. Rethinking our approach to cancer metabolism to deliver patient benefit. Br J Cancer 129, 406–415 (2023). https://doi.org/10.1038/s41416-023-02324-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-023-02324-9
- Springer Nature Limited