Abstract
We have previously recognized the genotypic and prognostic heterogeneity of U2AF1 mutations (MT) in myelofibrosis (MF) and myelodysplastic syndromes (MDS). In the current study, we considered 179 U2AF1-mutated patients with clonal cytopenia of undetermined significance (CCUS; n = 22), MDS (n = 108), MDS/acute myeloid leukemia (AML; n = 18) and AML (n = 31). U2AF1 variants included S34 (60%), Q157 (35%), and others (5%): corresponding mutational frequencies were 45%, 55%, and 0% in CCUS; 57%, 39%, and 4% in MDS; 61%, 33%, and 6% in MDS/AML; and 55%, 35% and 10% in AML (P = 0.17, 0.36 and 0.09), respectively. Concurrent mutations included ASXL1 (37%), BCOR (19%), RUNX1 (14%), TET2 (15%), DNMT3A (10%), NRAS/KRAS (8%), TP53 (8%), JAK2 (5.5%) and SETBP1 (5%). The two most frequent U2AF1 MT were S34F (n = 97) and Q157P (n = 46); concurrent MT were more likely to be seen with the latter (91% vs 74%; P = 0.01) and abnormal karyotype with the former (70% vs 62%; P = 0.05). U2AF1 S34F MT clustered with BCOR (P = 0.04) and Q157P MT with ASXL1 (P = 0.01) and TP53 (P = 0.03). The median overall survival (OS) in months was significantly worse in AML (14.2) vs MDS/AML (27.3) vs MDS (33.7; P = 0.001); the latter had similar OS with CCUS (30.0). In morphologically high-risk disease (n = 49), defined by ≥10% blood or bone marrow blasts (i.e., AML or MDS/AML), median OS was 14.2 with Q157P vs 37.1 months in the presence of S34F (P = 0.008); transplant-adjusted multivariable analysis confirmed the detrimental impact of Q157P (P = 0.01) on survival and also identified JAK2 MT as an additional risk factor (P = 0.02). OS was favorably affected by allogeneic hematopoietic stem cell transplantation (HR: 0.16, 95% CI; 0.04-0.61, P = 0.007). The current study defines the prevalence and co-mutational profiles of U2AF1 pathogenic variants in AML, MDS/AML, MDS, and CCUS, and suggests prognostic heterogeneity in patients with ≥10% blasts.
Similar content being viewed by others
Introduction
Myelodysplastic neoplasm/syndrome (MDS) and acute myeloid leukemia (AML) are heterogeneous diseases, with variable outcomes, largely driven by chromosomal alterations and somatic mutations [1, 2]. Mutations in genes encoding components of the spliceosome complex (SRSF2, U2AF1, SF3B1, ZRSR2) are observed in approximately one-third of the patients with MDS and nearly half of the patients with MDS transforming to AML (secondary [s] AML) [3,4,5,6]. Historically, AML patients with gene mutations related to spliceosome complex, including U2AF1 were considered as intermediate risk, but with recently updated European LeukemiaNet (ELN) 2022 recommendations, these patients are now considered adverse risk based on associated inferior outcomes [7]. U2AF1 is an important component of the spliceosome complex required for pre-mRNA splicing [8, 9]. Mutations in U2AF1 have been described in myeloid neoplasms, with variants causing specific alterations in 3′ splice site recognition [5, 10, 11]. U2AF1 mutations (MT) are typically acquired later in life and associated with rapid rates of progression to MDS and AML [12,13,14,15,16,17] U2AF1 MT predominantly occur at two hotspots (S34F, Q157) located in zinc finger regions [5]. An earlier report evaluating clinical outcomes of 78 MDS patients with U2AF1 MT demonstrated that transcriptional factor and epigenetic regulator genes (e.g., ASXL1 [26%], DNMT3A/PHF6 [12%], BCOR [15%], TET2 [13%], RUNX1/STAG2 [9%], SETBP1[8%]) were predominantly co-mutated in U2AF1 MT myeloid neoplasms. Furthermore, analysis showed that ancestral U2AF1S34F MT were associated with inferior outcomes in comparison to ancestral U2AF1Q157 MT, while no differences in outcomes were observed if the mutations were secondary/subclonal in nature [15]. In another study by Tefferi et al. [17], 52 MDS patients with U2AF1 MT were evaluated, S34F and Q157 hotspots were commonly observed. However, a cytogenetic-independent prognostic impact was not evident for either one of the two commonly observed hotspot mutations. Unlike in MDS, the mutational spectrum of U2AF1 MT in myelofibrosis is contrastingly different with higher occurrence of Q157 (50/77 [65%]) compared to S34 (26/77 [34%]) hotspots [10]. To the best of our knowledge apart from the aforementioned studies, limited information is available on the molecular profile, myeloid co-mutation pattern and survival outcomes with unique U2AF1 MT in patients with clonal cytopenia of undetermined significance (CCUS), MDS, MDS/AML and AML. In this report we have analyzed the genomic profile and clinical relevance of U2AF1 MT in a larger cohort of patients with myeloid neoplasms.
Patient and methods
We reviewed the Mayo Clinic database of patients with myeloid neoplasms who underwent next-generation genomic sequencing (NGS) between January 2015 and July 2021. We evaluated the molecular profile and outcomes in 179 patients with precursor myeloid neoplasms (clonal cytopenias of undetermined significance [CCUS; n = 22]) and myeloid neoplasms (MDS [n = 108], MDS/AML [n = 18] and AML [n = 31]) harboring U2AF1 MT. Clinical NGS testing was performed on DNA extracted from fresh bone marrow aspirates. The Mayo Clinic NGS panel included 42 genes (Supplementary Material) and has an accuracy of >99% and reproducibility of 100% for single base substitutions and insertion/deletion events. The panel has a variant sensitivity of ≥2% VAF with a minimum depth coverage of 250x. CCUS was defined according to the 2022 WHO (World Health Organization) criteria; MDS, MDS/AML and AML were defined as per International Consensus Classification (ICC) 2022 of myeloid neoplasms and acute leukemia [18, 19]. For this analysis, we operationally defined high-risk myeloid neoplasms as myeloid neoplasms with ≥10% myeloid blasts in the peripheral blood and/or bone marrow. Treatment responses in MDS and AML were assessed according to the International Working Group (IWG) MDS response criteria (2006) and the 2017 ELN AML response criteria, respectively [20, 21].
Statistical analysis
Continuous variables summarized as medians (range), while categorical variables reported as frequencies (percentage). Unadjusted comparisons of patient characteristics and outcomes among patients with different myeloid neoplasms and U2AF1 MT were made using the Wilcoxon rank sum test (continuous variables) or Fisher’s exact test (categorical variables). We derived the cut-offs for U2AF1 VAF by using Receiver Operating Characteristics (ROC) analysis to assess values that correlated with OS. The Kaplan–Meier method was used to estimate overall survival (OS). All tests were two-sided with P value < 0.05 considered statistically significant. Cox proportional hazards regression model was used to determine the univariate and multivariate predictors of overall survival in patients with high-risk myeloid neoplasm. Multivariable models included all significant univariate predictors with P = ≤0.05. We also performed landmark analysis for OS among responding patients from time of response till last follow up or death and evaluated the impact of allogeneic stem cell transplantation (allo-HCT) in these patients.
Results
Baseline characteristics
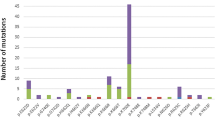
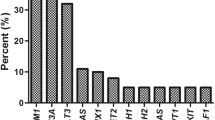
The baseline characteristics for this cohort are summarized in Table 1. Overall, the median age of the cohort was 72 years (range, 19–92), with a male preponderance (83%), and similar distributions in patients with CCUS, MDS, MDS/AML and AML. Twenty-two (12%), 108 (60%), 18 (10%) and 31 (17%) patients met criteria for CCUS, MDS, MDS/AML and AML, respectively. There was no significant difference in median white blood cell (WBC) count (P = 0.49), hemoglobin (P = 0.23) and platelets (P = 0.62) between CCUS, MDS, MDS/AML and AML patients. Sixty-seven % (n = 118) of these patients had cytogenetic (CG) abnormalities; CCUS (n = 17 [77%]), MDS (n = 70 [65%]), MDS/AML (n = 12 [70%]) and AML (n = 19 [61%]), P = 0.43. Overall, the most common CG abnormalities were del 20q (18%), complex CG (12%), trisomy 8 (9%) and monosomy 7/del 7q (8%) as outlined in Table 1. In MDS patients, 23% (n = 28), 54% (n = 64), and 23% (n = 28) had very good/good, intermediate risk and poor/very poor risk CG, respectively, as per IPSS-R (Revised International Prognostic Scoring System) criteria [22]. In the AML group, 20/31 (64.5%) had CG abnormalities, 50% (n = 10) each had ELN 2022 intermediate and adverse risk CG [7].
Somatic mutational profile and co-mutational patterns
The median U2AF1 variant allele frequency (VAF) was 35% (range [R], 5–51). The median U2AF1 VAF was 33% (R, 6–51), 36% (R, 5–49), 33% (R, 5–44) and 35% [R, 10–46] in patients with CCUS, MDS, MDS/AML and AML, respectively (P = 0.77). U2AF1 MT locations included S34 (60%), Q157 (35%), and others (5%). The corresponding mutational frequencies were 45%, 55%, and 0% in CCUS, 57%, 39%, and 4% in MDS, 61%, 33%, and 6%, in MDS/AML, 55%, 35% and 10% in AML (P = 0.17, P = 0.36 and P = 0.09, respectively). In the U2AF1 protein the S34 hotspot is in the zinc finger protein 1 region (ZF1), while the Q157 hotspot is located in the ZF2 region (Fig. 1). Concurrent myeloid MT observed in ≥ 5% of patients were ASXL1 (37%), BCOR (19%), RUNX1 (14%), TET2 (15%), DNMT3A (10%), RAS (NRAS or KRAS) (8%), TP53 (8%; 9/14 TP53 MT were “multi-hit” as per ICC 2022 [accompanying CG or loss of heterozygosity, multiple TP53 MT, single TP53 MT with VAF > 50% or 17p deletion]), JAK2 (5.5%) and SETBP1 (5%), respectively. We did not observe co-occurrence of splicing factor (SRSF2, SF3B1 and ZRSR2) MT with U2AF1 MT. Illustrations depicting concurrent myeloid co-mutations with U2AF1 MT and their corresponding VAF % are provided in Supplementary Figs. 1–3.
a Overview of U2AF1 domains, structures, and distribution of U2AF1 mutations detected, positioned on the U2AF1 protein. Protein Sequence of ZF1 (hotspots at codon 34; S34F and S34Y) and ZF2 (hotspots at codon 157; Q157R and Q157P) domains, where all U2AF1 mutations clustered. b Patterns of the co-mutations identified in the U2AF1 cohort with respective mutations. NTD N-terminal domain, ZF zinc finger domain, RRM RNA recognition motifs, RS The C-terminal Arg-Ser rich domain, CTD C-terminal domain.
We analyzed myeloid co-mutation patterns between the two most frequent U2AF1 MT (S34F and Q157P) (Table 2). Concurrent MT were more likely to be seen with Q157P compared to S34F (91% vs 74%; P = 0.01) MT, respectively. Cytogenetic abnormalities were more frequently seen with S34F compared to Q157P (70% vs 62%; P = 0.05). U2AF1 S34F MT clustered with BCOR (P = 0.04) MT, while Q157P MT clustered with ASXL1 (P = 0.01) and TP53 (P = 0.03) MT (P = 0.02), respectively. We did not observe significant differences in median WBC (P = 0.23), hemoglobin (P = 0.82) and platelet counts (P = 0.72) between S34F and Q157P MT. We then looked at differences in hemoglobin level ≤10 g/dl (P = 0.16) or ≤8 g/dl (P = 0.51) between S34F and Q157P U2AF1 MT and did not find statistically significant differences. Similarly, there were no differences in platelet counts between the two groups.
Treatment and responses
One hundred twenty-five (70%) patients received disease directed treatment. The number of patients in each group that received treatment included 6/22 (27%) patients with CCUS, 75/108 (69%) patients with MDS, 17/18 patients with MDS/AML (94%) and 27/31 (87%) patients with AML. Four of 31 (13%) AML patients did not receive leukemia directed therapy due to co-morbidities and advanced age. Two of six (33%) CCUS patients received hypomethylating agents (HMA=off label use) and the remaining 4/6 (67%) received supportive care (e.g., erythropoietin stimulating agents and/or growth factor support). Amongst MDS patients, 48 (64%) received HMA, 17 (23%) received supportive care, 6 (8%) received AML-like intensive chemotherapy and 4 (5%) patients received HMA plus venetoclax combination therapy. In the MDS/AML cohort, 10 (59%) patients received HMA, 3 (18%) received intensive chemotherapy, 3 (18%) patients other low-intensity/supportive care therapy and 1 (6%) patient received HMA plus venetoclax combination therapy. In the AML cohort 14 (52%) received intensive chemotherapy, 6 (22%) patients received HMA, 5 (18%) patients received HMA plus venetoclax combination therapy and 1 (4%) received other low-intensity/supportive care therapy (Table 1).
None of the patients treated in the CCUS group met criteria for an objective response (as adjudicated by MDS response criteria, given that CCUS response criteria do not exist). The complete remission [CR] rates were 25% in the MDS cohort, 18% in MDS/AML cohort and 26% in AML cohort. Details on responses with regards to diagnostic categories and types of therapies used summarized in Table 1. Overall, 31 (18%) patients underwent allo-HCT; 19 (17.5%) patients with MDS, 4 (22%) patients with MDS/AML and 8 (26%) patients with AML. Treatment patterns, responses and proportion of patients receiving allo-HCT were not significantly different amongst the two common U2AF1 MT (S34F and Q157P) (Table 2).
Survival outcomes
The median OS of the entire cohort was 26.5 months. The median OS among patients with CCUS, MDS, MDS/AML and AML was 29.2, 33.7, 27.3 and 14.3 months, respectively (P = 0.01; Fig. 2a). The median OS in patients with high-risk myeloid neoplasm was 26.6 months. We then performed a subset analysis among patients with high-risk myeloid neoplasms harboring; S34F and Q157P MT. The median OS were 37.1 vs 14.2 months with U2AF1S34F and U2AF1Q157P MT, respectively (P = 0.008; Fig. 2b). The median OS was better in patients with <10% myeloid blasts compared to those with ≥10% myeloid blasts (32.7 vs 21.5 months, P = 0.009; Fig. 2c). We used ROC derived U2AF1 VAF cut off for OS (VAF ≥ 25% vs <25%), however VAF as a continuous variable did not achieve statistical significance (52.3 vs 50.9 months, P = 0.48). Similarly, we also performed landmark analysis for OS among responding patients from time of response till last follow up or death and evaluated outcome with allo-HCT in these patients. The median OS was significantly better with allo-HCT (53.0 months) compared to without allo-HCT (22.8 months), P = 0.04 (Fig. 3).
Univariate and multivariate analysis for OS in high-risk myeloid neoplasms with U2AF1S34F or U2AF1157P
In univariate analysis for OS in patients with high-risk myeloid neoplasms, U2AF1Q157P compared to U2AF1S34F showed significantly inferior outcome (14.2 vs 37.1, P = 0.008). Patients with concurrent ASXL1 MT (10.77 vs 37.1 months, P = 0.008), and JAK2 MT (5.8 vs 23.2 months, P = 0.01) had inferior survival outcomes. Patients with bone marrow (BM) blast percentage ≥20% had inferior OS (14.3 months) compared to BM blast percentage between 10 and 19% (37.3 months, P = 0.03). Allo-HCT was associated with favorable OS in univariate analysis (40.0 vs 15.5 months, P = 0.04). Using predictors that demonstrated significance or trended towards significance (P = ≤ 0.1) in univariate analysis, a group of variables was assembled for multivariate OS analysis. Concurrent JAK2 MT (HR: 8.12, 95% CI; 1.39–47.31, P = 0.02) and Q157P vs S34F (HR: 4.37, 95% CI; 1.31–14.11, P = 0.01) and retained significance for inferior OS, while allo-HCT (HR: 0.71, 95% CI; 0.09–0.57, P = 0.01) retained significance for better OS in multivariate analysis (Table 3).
Discussion
We present data on the molecular profile and survival outcomes of patients with precursor myeloid neoplasms and myeloid neoplasms harboring U2AF1 MT. We observed distinct myeloid co-mutation profiles and survival outcomes associated with different mutant U2AF1 hotspot regions and amino acid changes. The most frequently observed MT were S34 (60%), Q157 (35%), followed by others (5%), in alignment with prior published data [15]. In high-risk myeloid neoplasm patients, U2AF1Q157P was associated with inferior outcome compared to U2AF1S34F. U2AF1Q157P MT significantly clustered with TP53 (15% vs 4%) and ASXL1 (50% vs 28%) mutations and had a higher percentage of co-mutations (91% vs 74%), in comparison to U2AF1S34F MT. U2AF1S34F MT on the other hand significantly clustered with BCOR MT (23% vs 0%). Cytogenetic abnormalities were more commonly seen with U2AF1S34F MT compared to U2AF1Q157P MT. While the observation that U2AF1S34F and U2AF1Q157P are the most frequent U2AF1 MT in myeloid neoplasms is not new [5, 23,24,25], our study is the first to demonstrate their unique co-mutational spectrum and differential prognostic effect.
In our cohort, aligned with most previous observations, we did not observe co-occurrence of other splicing factor MT with U2AF1 MT [26]. It is believed that concomitant splicing factor MT in the same cell could be incompatible with survival of the cell. In the new risk stratification schema for MDS and AML, U2AF1 MT are now considered high risk [7, 27]; given that 81% of patients in our cohort had concurrent myeloid MT, we asked the question as to whether or not these accompanying MT were acting as confounding factors, adversely influencing U2AF1 MT related outcomes. In our analysis, the OS of the entire cohort including patients with CCUS, MDS, MDS/AML and AML was sub-optimal at 26.5 months, with a differential prognostic impact imparted by the two U2AF1 hotspot regions; 37.1 vs 14.2 months (P = 0.008) in patients with high-risk myeloid neoplasm harboring U2AF1S34F and U2AF1Q157PMT, respectively. Earlier reports suggested differential degree of anemia and thrombocytopenia with different U2AF1 MT. In MDS, thrombocytopenia was specifically associated with U2AF1S34F MT and anemia with U2AF1Q157 MT [24]. In patients with myelofibrosis, both mutation types were associated with anemia and the association with thrombocytopenia was most evident with U2AF1Q157 MT [10]. In our cohort of patients with CCUS, MDS, MDS/AML and AML, we did not observe significant differences in anemia or thrombocytopenia among different U2AF1 MT. Interestingly, in the current study we observe a relatively higher rate of CG abnormalities and a shorter OS in U2AF1 MT CCUS patients, in comparison to patients with MDS and AML. Our group has recently reported on clinical outcomes of patients with U2AF1 MT clonal hematopoiesis (CHIP [clonal hematopoiesis of indeterminate potential] and CCUS) [16]. In that study, we observed a high rate (25%) and a short latency (17.5 months) towards progression to myeloid neoplasms. We acknowledge that the higher incidence of CG abnormalities in the CCUS group could have been due to selection bias, inherent to the structure of retrospective studies.
Current literature suggest sub-optimal responses with hypomethylating agent therapies in U2AF1-mutated myeloid neoplasms [28], and better responses with HMA plus venetoclax-based therapies [6]. Similarly, we observed lower CR rates with HMA therapy alone (12%) and relatively better responses with HMA plus venetoclax (45%) or intensive chemotherapies (65%). However, our sample size was small and larger prospective studies are needed to gauge responses with different treatment regimens. Similar to earlier reports, our study suggests benefit from allo-HCT in improving survival outcome among patients with U2AF1 MT myeloid neoplasms [29].
We acknowledge the limitations of our analysis including inherent selection bias and lack of sequential mutation testing in this cohort of patients with myeloid neoplasms. Nevertheless, our study highlights the unique co-mutation patterns and survival outcomes in CCUS, MDS and AML patients with different U2AF1 MT, underscoring the need for accurate mutational assessment and reporting.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Badar T, Szabo A, Sallman D, Komrojki R, Lancet J, Padron E, et al. Interrogation of molecular profiles can help in differentiating between MDS and AML with MDS-related changes. Leuk Lymphoma. 2020;61:1418–27.
Lindsley RC, Mar BG, Mazzola E, Grauman PV, Shareef S, Allen SL, et al. Acute myeloid leukemia ontogeny is defined by distinct somatic mutations. Blood. 2015;125:1367–76.
Reinig E, Yang F, Traer E, Arora R, Brown S, Rattray R, et al. Targeted next-generation sequencing in myelodysplastic syndrome and chronic myelomonocytic leukemia aids diagnosis in challenging cases and identifies frequent spliceosome mutations in transformed acute myeloid leukemia. Am J Clin Pathol. 2016;145:497–506.
Thol F, Kade S, Schlarmann C, Löffeld P, Morgan M, Krauter J, et al. Frequency and prognostic impact of mutations in SRSF2, U2AF1, and ZRSR2 in patients with myelodysplastic syndromes. Blood. 2012;119:3578–84.
Yoshida K, Sanada M, Shiraishi Y, Nowak D, Nagata Y, Yamamoto R, et al. Frequent pathway mutations of splicing machinery in myelodysplasia. Nature. 2011;478:64–9.
Lachowiez CA, Loghavi S, Furudate K, Montalban-Bravo G, Maiti A, Kadia T, et al. Impact of splicing mutations in acute myeloid leukemia treated with hypomethylating agents combined with venetoclax. Blood Adv. 2021;5:2173–83.
Döhner H, Wei AH, Appelbaum FR, Craddock C, DiNardo CD, Dombret H, et al. Diagnosis and management of AML in adults: 2022 recommendations from an international expert panel on behalf of the ELN. Blood. 2022;140:1345–77.
Pellagatti A, Armstrong RN, Steeples V, Sharma E, Repapi E, Singh S, et al. Impact of spliceosome mutations on RNA splicing in myelodysplasia: dysregulated genes/pathways and clinical associations. Blood. 2018;132:1225–40.
Zhao Y, Cai W, Hua Y, Yang X, Zhou J. The biological and clinical consequences of RNA splicing factor U2AF1 mutation in myeloid malignancies. Cancers. 2022;14:4406.
Tefferi A, Finke CM, Lasho TL, Hanson CA, Ketterling RP, Gangat N, et al. U2AF1 mutation types in primary myelofibrosis: phenotypic and prognostic distinctions. Leukemia. 2018;32:2274–8.
Barraco D, Elala YC, Lasho TL, Begna KH, Gangat N, Finke C, et al. Molecular correlates of anemia in primary myelofibrosis: a significant and independent association with U2AF1 mutations. Blood Cancer J. 2016;6:e415.
Abelson S, Collord G, Ng SWK, Weissbrod O, Mendelson Cohen N, Niemeyer E, et al. Prediction of acute myeloid leukaemia risk in healthy individuals. Nature. 2018;559:400–4.
Robertson NA, Latorre-Crespo E, Terradas-Terradas M, Lemos-Portela J, Purcell AC, Livesey BJ, et al. Longitudinal dynamics of clonal hematopoiesis identifies gene-specific fitness effects. Nat Med. 2022;28:1439–46.
Malcovati L, Galli A, Travaglino E, Ambaglio I, Rizzo E, Molteni E, et al. Clinical significance of somatic mutation in unexplained blood cytopenia. Blood. 2017;129:3371–8.
Adema V, Hirsch CM, Przychodzen BP, Nazha A, Kuzmanovic T, Negoro E. et al. U2AF1 mutations in S34 and Q157 create distinct molecular and clinical contexts. Blood.2016;128:3155
Pritzl SL, Gurney M, Badar T, Ferrer A, Lasho T, Finke C, et al. Clinical and molecular spectrum and prognostic outcomes of U2AF1 mutant clonal hematopoiesis—a prospective Mayo Clinic cohort study. Leuk Res. 2023;125:107007.
Tefferi A, Mudireddy M, Finke CM, Nicolosi M, Lasho TL, Hanson CA, et al. U2AF1 mutation variants in myelodysplastic syndromes and their clinical correlates. Am J Hematol. 2018;93:E146–E8.
Arber DA, Orazi A, Hasserjian RP, Borowitz MJ, Calvo KR, Kvasnicka H-M, et al. International consensus classification of myeloid neoplasms and acute leukemias: integrating morphologic, clinical, and genomic data. Blood. 2022;140:1200–28.
Khoury JD, Solary E, Abla O, Akkari Y, Alaggio R, Apperley JF, et al. The 5th edition of the World Health Organization classification of haematolymphoid tumours: myeloid and histiocytic/dendritic neoplasms. Leukemia. 2022;36:1703–19.
Döhner H, Estey E, Grimwade D, Amadori S, Appelbaum FR, Büchner T, et al. Diagnosis and management of AML in adults: 2017 ELN recommendations from an international expert panel. Blood. 2017;129:424–47.
Cheson BD, Greenberg PL, Bennett JM, Lowenberg B, Wijermans PW, Nimer SD, et al. Clinical application and proposal for modification of the International Working Group (IWG) response criteria in myelodysplasia. Blood. 2006;108:419–25.
Greenberg PL, Tuechler H, Schanz J, Sanz G, Garcia-Manero G, Solé F, et al. Revised international prognostic scoring system for myelodysplastic syndromes. Blood. 2012;120:2454–65.
Graubert TA, Shen D, Ding L, Okeyo-Owuor T, Lunn CL, Shao J, et al. Recurrent mutations in the U2AF1 splicing factor in myelodysplastic syndromes. Nat Genet. 2012;44:53–7.
Li B, Liu J, Jia Y, Wang J, Xu Z, Qin T, et al. Clinical features and biological implications of different U2AF1 mutation types in myelodysplastic syndromes. Genes Chromosomes Cancer. 2018;57:80–8.
Wu S-J, Tang J-L, Lin C-T, Kuo Y-Y, Li L-Y, Tseng M-H, et al. Clinical implications of U2AF1 mutation in patients with myelodysplastic syndrome and its stability during disease progression. Am J Hematol. 2013;88:E277–82.
Bamopoulos SA, Batcha AMN, Jurinovic V, Rothenberg-Thurley M, Janke H, Ksienzyk B, et al. Clinical presentation and differential splicing of SRSF2, U2AF1 and SF3B1 mutations in patients with acute myeloid leukemia. Leukemia. 2020;34:2621–34.
Bernard E, Tuechler H, Greenberg PL, Hasserjian RP, Ossa JEA, Nannya Y, et al. Molecular international prognostic scoring system for myelodysplastic syndromes. NEJM Evidence. 2022;1:EVIDoa2200008.
Jung SH, Kim YJ, Yim SH, Kim HJ, Kwon YR, Hur EH, et al. Somatic mutations predict outcomes of hypomethylating therapy in patients with myelodysplastic syndrome. Oncotarget. 2016;7:55264–75.
Song G-Y, Kim T, Ahn S-Y, Jung S-H, Kim M, Yang D-H, et al. Allogeneic hematopoietic cell transplantation can overcome the adverse prognosis indicated by secondary-type mutations in de novo acute myeloid leukemia. Bone Marrow Transplant. 2022;57:1810–9.
Author information
Authors and Affiliations
Contributions
TB, AT and MP designed the study, collected data, performed analysis, and co-wrote the paper. YMV, AN, JF, AAK, AM, HM, HA, DV, RH, MS, CAY, MRL and NG contributed patients, reviewed, and approved the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Badar, T., Vanegas, Y.A.M., Nanaa, A. et al. U2AF1 pathogenic variants in myeloid neoplasms and precursor states: distribution of co-mutations and prognostic heterogeneity. Blood Cancer J. 13, 149 (2023). https://doi.org/10.1038/s41408-023-00922-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41408-023-00922-7
- Springer Nature Limited
This article is cited by
-
Real world predictors of response and 24-month survival in high-grade TP53-mutated myeloid neoplasms
Blood Cancer Journal (2024)