Abstract
The iron chelator deferasirox is widely used in patients with iron overload. Patients with low-grade myelodysplastic syndromes (MDS) get transfusion dependency and need to be treated with deferasirox to avoid iron overload. Moreover, in some patients an increase in both erythroid and platelets have been observed after deferasirox therapy. However, the mechanisms involved in these clinical findings are poorly understood. The aim of this work was to analyze, in patients treated with deferasirox, the changes in the gene-expression profile after receiving the treatment. A total of 15 patients with the diagnosis of low-grade MDS were studied. Microarrays were carried out in RNA from peripheral blood before and after 14 weeks of deferasirox therapy. Changes in 1457 genes and 54 miRNAs were observed: deferasirox induced the downregulation of genes related to the Nf kB pathway leading of an overall inactivation of this pathway. In addition, the iron chelator also downregulated gamma interferon. Altogether these changes could be related to the improvement of erythroid response observed in these patients after therapy. Moreover, the inhibition of NFE2L2/NRF2, which was predicted in silico, could be playing a critical role in the reduction of reactive oxygen species (ROS). Of note, miR-125b, overexpressed after deferasirox treatment, could be involved in the reduced inflammation and increased hematopoiesis observed in the patients after treatment. In summary this study shows, for the first time, the mechanisms that could be governing deferasirox impact in vivo.
Similar content being viewed by others
Introduction
Myelodysplastic syndromes (MDS) are a heterogeneous group of clonal stem cell disorders, usually characterized by the presence of anemia. Red blood cell transfusion is the only therapeutic option for most patients, who usually develop transfusion dependence, which can be correlated with poorer outcomes partially caused by transfusional iron overload. The magnitude of total body iron load correlated with hepatic iron concentration (HIC) [1]. Thresholds of HIC predict the development of potentially fatal complications: [2] HIC exceeding 15 mg iron/g liver, dry weight, and sustained elevations of serum ferritin (SF) over 2500 μg/L, increase the risk of premature death. Deferoxamine has been the standard drug for iron chelation therapy over the past four decades. However, the major disadvantage is the patient’s lack of adherence, because it needs an 8–12 h parenteral administration since it has a short half-life and a very poor oral bioavailability [1,2,3,4]. Therefore, in the 1980s, the parenteral requirement for deferoxamine stimulated studies of orally active chelators. Deferasirox was licensed as first line therapy by the FDA in 2005, following randomized trials comparing this drug, and demonstrating its noninferiority, to deferoxamine [5]. The safety profile is acceptable, the drug is well tolerated and effective even in long-term follow-up [6]. Deferasirox is widely used in low-risk MDS [1] and also in high-risk MDS [7].
Changes in the gene-expression profile (GEP) of MDS have been described [8], being different between low-risk and high-risk MDS patient groups. Thus, patients with refractory anemia with ring sideroblasts showed an overexpression of mitochondrial and iron homeostasis genes [9]. Moreover, iron overload could selectively affect peripheral T lymphocytes and induce an impaired cellular immunity by increasing reactive oxygen species (ROS) level [10]. The application of system-wide “omics” technologies could provide new insights into the knowledge of pathogenic mechanisms involved after deferasirox therapy. In fact, several in vitro studies analyzed the effects of deferasirox in the blood cells, identifying pathways or cellular mechanisms such as Nf kB [11], MTOR [12], or ROS [10] as the most frequently involved during this treatment. However, only few in vivo studies have been performed to assess the main cellular mechanisms affected by deferasirox therapy in MDS patients [1].
In this work, we aimed to perform an in vivo pharmacogenomic analysis in patients diagnosed as low-risk MDS treated with deferasirox, in order to identify the changes in the GEP induced by the treatment in these patients.
Patients and methods
Patients
Fifteen patients diagnosed as low-risk MDS and treated with deferasirox were included in the study. All patients have the diagnosis of low-risk MDS according to WHO 2008 classification [13]. Blood samples were analyzed before deferasirox and 14 weeks after treatment (range 6–40 weeks). Supplementary Table 1 summarizes the main clinical characteristics and laboratory findings before and during the treatment. All patients received the treatment without additional drugs during the study. The median age was 77 years (range 62–90 years), and 36% patients were male. This study was performed in accordance with the Declaration of Helsinki guidelines, and approved by the Local Ethical Committees “Comité Ético de Investigación Clínica, Hospital Universitario de Salamanca”. All patients provided written informed consent.
Methods
PBMCs and RNA isolation
Peripheral blood mononuclear cells (PBMCs) were isolated from whole peripheral blood using Ficoll-Paque Plus gradient, snap-frozen and stored at −80 °C. Total RNA was extracted by QIAgen (Qiagen, Valencia, CA, USA), following the manufacturer’s recommendations. RNA integrity was assessed on an Agilent 2100 Bioanalyzer (Agilent technologies, Santa Clara, CA, USA) employing an RNA 6000 Nano Assay kit. All RNA samples had an A260/A280 ratio ~1.8 and RNA integrity number (RIN) ≥8.0.
Gene expression profiling
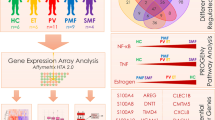
Genome-wide expression analysis of the paired samples (before and after the treatment) was performed using Human Transcriptome Array 2.0 ST (Affymetrix, Inc., Santa Clara, CA, USA) following the manufacturer’s protocols for the GeneChip platform by Affymetrix. Briefly, cDNA was regenerated through a random-primed reverse transcription (RT) using a dNTP mix containing dUTP. After cDNA was hybridized to the arrays at 60 rpm for 18 h at 45 °C, the chips were processed in a Genechip Fluidics Station 450 (Affymetrix). Microarray images were collected by Affymetrix GeneChip 3000 scanner, and data were extracted using Affymetrix GCOS Software.
Bioinformatic analysis: normalization, signal calculation, and significant differential expression
The robust microarray analysis algorithm was used for background correction, intra- and inter-microarray normalization, and expression signal calculation [14]. The absolute expression signal for each gene was calculated for each microarray. Differential gene expression before and after treatment were obtained using the MultiExperiment Viewer 4 (MeV4, TM4 software suite), using the Student’s paired t test for matched samples to analyze the statistical significance. P < 0.05 was considered statistically significant.
Functional analysis and gene annotation
The functional assignment of the genes included in the expression signature of MDS patients was carried out by the Database for Annotation, Visualization, and Integrated Discovery [15, 16] and the GeneCodis application [17] which identifies concurrent annotations in GO and KEGG, and thereby constructs several groups of genes of functional significance. The most significant biological mechanisms, pathways, and functional categories in the data sets of genes selected by statistical analysis were identified through the use of Ingenuity Pathways Analysis (Ingenuity Systems Inc., Redwood City, CA, USA).
Upstream regulator analysis
The goal of this analysis is to identify the cascade of upstream transcriptional regulators that can explain the observed gene expression changes in a dataset. This analysis is based on prior knowledge of expected effects between transcriptional regulators and their target genes stored in the Ingenuity® Knowledge Base. The analysis examines how many known targets of each transcription regulator are present in the analyzed dataset, and also compares their direction of change to what is expected from the literature in order to predict likely relevant transcriptional regulators. If the observed direction of change is mostly consistent with a particular activation state of the transcriptional regulator (“activated” or “inhibited”), then a prediction is made about that activation state.
Integrative analysis of miRNA and gene-expression profile
MiRNAs with significantly different expression (P < 0.05) before and after treatment were further analyzed to identify the networks and pathway targets. For this purpose, IPA’s microRNA Target Filter, which enables prioritization of experimentally validated and predicted mRNA targets from TargetScan Knowledge Base was used. This tool identified the putative targets for the input miRNAs and then developed the networks among the targets and identified the known and most relevant biological functions, pathways, and annotations in this enriched set of target genes. By applying the expression pairing tool, the analysis was focused on targets exhibiting altered expression in our analysis, finding miRNAs and their target genes with opposite or same expression.
Droplet digital PCR
First-stranded cDNA was synthesized from 500 ng total RNA using the SuperScript III First-Strand Synthesis SuperMix assay (Invitrogen) according to manufacturer’s instruction. Transcript expression for selected genes were quantified in MDS patients before and after treatment using QX200 ddPCR (Bio-Rad, Hercules, CA, USA). Primers are listed in Supplementary Table 2. ddPCR mix were prepared containing 10 μL of 2× QX200 ddPCR EvaGreen Supermix (Bio-Rad, cat. no. 1864034), 8 μL of nuclease-free water, 1 μL of primers 20× and 1 μL of diluted cDNA. A 20 μL aliquot was taken from each of the assembled ddPCR mixtures and pipetted into each sample well of an eight-channel disposable droplet generator cartridge (Bio-Rad, Hercules, CA, USA). A 70 μL volume of Droplet Generation Oil for EvaGreen (Bio-Rad) was then loaded into each of the eight oil wells. The cartridge was placed into the QX200 droplet generator (Bio-Rad) where a vacuum was applied to the outlet wells to simultaneously partition each 20 μL sample into nanoliter sized droplets. After ∼1.5 min, the cartridge was removed from the generator, and the droplets that had collected in each of the independent outlet wells were transferred with a multichannel pipet to a 96-well polypropylene plate (Eppendorf, Hamburg, Germany). The plate was heat-sealed with foil using a PX1 PCR Plate Sealer (Bio-Rad) and placed in a conventional thermal cycler (C1000 Touch, Bio-Rad), where an endpoint PCR run was performed with the following program: an activation period (95 °C for 10 min), followed by 40 cycles of a two-step thermal profile comprising of a denaturation step (95 °C for 30 s), and a combined annealing-extension step (60 °C for 1 min). A final signal stabilization step was also included (90 °C for 10 min) and then cooling to 4 °C. After PCR, the 96-well plate was loaded into the QX200 Droplet Reader (Bio-Rad), and the appropriate assay information was entered into the analysis software package provided (Quanta-Soft, Bio-Rad). Droplets were automatically aspirated from each well and streamed single-file past a two-color fluorescence detector and finally to waste. The quality of all droplets was analyzed and rare outliers (e.g., doublets and triplets) were gated based on detector peak width. Analysis of the ddPCR data was performed with QuantaSoft analysis software (Bio-Rad) that accompanied the QX200 Droplet Reader. Expression levels of the selected genes before and after treatment were analyzed using the Mann–Whitney U test with a two tailed value of P = 0.05 for statistical significance. All tests were performed using SPSS v19.0.
Results and discussion
GEP was carried out in paired samples before deferasirox and during the treatment. The analysis revealed changes in 1457 genes (709 overexpressed after the therapy), and 54 miRNAs (33 of them, overexpressed) (Supplementary Tables 3 and 4). The expression data are available to the public on the NIH GEO (http://www.ncbi.nlm.nih.gov/geo/) under accession number GSE141958. Table 1 shows the top-ten deregulated genes after deferasirox therapy. Of note, several genes from the Nuclear factor-κappa B, Nf kB pathway (NFBIA), and immune system (IFNG, CX3CR1) as well as miRNAs (miR-2682 and miR-147b) were deregulated. Moreover, the functional analysis of the most relevant pathways affected showed an overall downregulation of the Nf kB pathway, as well as NO and ROS and hypoxia-related genes (Table 2).
Deferasirox treatment downregulated the NF kB pathway in low-risk MDS patients
Deferasirox therapy induced an overall downregulation in several genes involved in the Nf kB pathway (Table 2). These results now detected in patients confirmed previous in vitro observations, in which deferasirox was associated to repression of Nf kB activity in samples showing high basal activity as well as in cell lines, whereas no similar behavior was observed with other iron chelators. This observation has been linked to the hemoglobin improvement showed in some patients treated with this iron chelator [18,19,20,21]. Moreover, our results pointed out the genes which could be involved in this downregulation, such as BCL10, IL1B, NFKBIA, RIPK1, RELA, BID, BIRC2, and TANK (Table 3). The interactions among these genes leading to an overall downregulation of the whole Nf kB pathway is showed in Fig. 1. Expression changes in BIRC2, TRAF6, BID, RELA, REL, TANK, and NFKBIA were validated by ddPCR (Supplementary Fig. 1).
The overall downregulation of the Nf kB pathway could be related to a decrease in levels of pro-inflammatory cytokines and also could be promoting an increased apoptosis in the cells. Altogether, these changes could be related to a decrease in the transfusion dependency of these patients, as previously reported [1, 11, 22].
Deferasirox treatment deregulated genes related to ROS in low-risk MDS patients
In silico upstream regulator analysis predicted the involvement of the transcriptional regulator nuclear factor erythroid 2 (NFE2L2/NRF2) as inhibited, pointing it as the responsible for the deregulation of a great number of genes after deferasirox treatment. NFE2L2/NRF2 is a master gene which controls the response to the oxidative stress induced by the ROS and regulates over 200 genes (Fig. 2). ROS are a group of highly reactive chemicals containing oxygen produced either exogenously or endogenously. They are related to a wide variety of human disorders, such as chronic inflammation, age-related diseases, and cancers [10, 23]. Interestingly, in MDS patients the iron overload is related to an increase of the cytoplasmic ROS. By contrast, the decreased ROS activity observed after deferasirox treatment could be related to the Nf kB activation [23] and linked to the abnormalities in T cell lymphocytes [10].
Our study confirmed that these previous results, mainly based on in vitro models, are also observed in patients during deferasirox therapy. Moreover, in a recent study, deferasirox has been reported to reduce DNA oxidation and double strand breaks in bone marrow cells from MDS patients, which would suggest beneficial effects of this therapy [24].
Erythropoiesis improvement after deferasirox therapy in low-risk MDS patients could be related to the overexpression of GFI1 and the downregulation in the interferon pathway
The present study showed an activation of the gene growth factor independent 1 (GFI1) transcriptional repressor, a transcription factor which plays a critical role in the hematopoiesis, activating both erythroid and megakaryocyte lineages [25]. Of note, several studies have reported an improvement of hemoglobin and platelet levels in MDS patients treated with deferasirox [18,19,20, 26, 27].
Moreover, the GEP of patients after deferasirox therapy showed a marked downregulation of gamma interferon, as well as other cytokines such as TRAF6, IL8, ILB, and IL1RAP. The role of IL-1 and gamma interferon as inhibitors of erythropoiesis has been already reported [28]. Thus, our results showing the downregulation of the expression of these genes could be related to an erythropoiesis activation. Taking together, several mechanisms reported in this study, including activation of GFI1 or repression of gamma interferon, could lead to an erythropoiesis stimulation, thus providing the biological basis for the clinical results observed in MDS patients treated with this iron chelator.
miRNA deregulation in MDS patients treated with deferasirox
The analysis of miRNA expression revealed that 54 miRNAs were deregulated in MDS patients after the treatment: hsa-miR-2682 was the most highly upregulated miRNA (Diff exp = 0.48), while hsa-miR-147b was the most significantly downregulated (Diff exp = −1.08) (Supplementary Table 4). Functional analysis of the deregulated miRNAs after treatment (Fig. 3) revealed that the most strongly affected cell function by these miRNAs was gene expression, with a total of seven miRNAs involved (mir-146, mir-515, mir-665, mir-548, mir-30, mir-154, and mir-130). Inflammation (inflammatory disease and inflammatory response), cancer, hematological-related functions (hematological disease and hematological system development and function), and hematopoiesis were other important functions affected by these miRNAs.
A functional enrichment analysis was performed in the dataset of 54 deregulated miRNAs. Category names are presented on the vertical axis. Of note, gene expression signaling and inflammatory signaling were among the most significant cellular functions affected. The significance of the association between the dataset and the canonical pathway was measured by the P value determining the probability of the association between the genes in the dataset and the canonical pathway, calculated by Fisher’s exact test. The horizontal axis on the top indicates the −log (P value), the higher value indicates the higher significance.
The influence of these deregulated miRNAs on deferasirox treated MDS patients was assessed. Specifically, we investigated whether observed changes in miRNAs were correlated with changes in the expression of genes. Therefore, the posttranscriptional regulatory network of miRNA and genes in MDS patients treated with deferasirox was carried out by analyzing the miRNA-mRNA relationships. Indeed, because miRNAs tend to downregulate the target genes, we focused our study on the subset of eight miRNAs selected for analysis in IPA and the genes experimentally observed or predicted with high confidence to be regulated by them and characterized by expression profiles strongly anticorrelated. MiR-125b, overexpressed after deferasirox treatment, was negatively correlated with the expression of 20% expected target genes (data not shown) demonstrating the relationship between miRNA and gene deregulation. Interestingly, the expression of miR-125b is modulated by NF kB signaling. MiR-125b targets the 3′UTR region of TNF-alpha gene to negatively regulate the inflammatory response [29]. By contrast, another study has reported that miR-125b can promote macrophage mediated inflammation and enhance antitumor activities by targeting interferon regulatory factor 4 [30]. These data suggest that miRNAs could play different roles in diverse biological contexts. Of note, a recent study [31] has shown that miR-146b, also overexpressed after deferasirox therapy (Supplementary Table 4), can regulate inflammatory responses by targeting mRNAs encoding TRAF6, a gene downregulated in our work (Table 3 and Supplementary Table 3).
Moreover, miR-125 has recently emerged as a key regulator of hematopoietic stem cells (HSCs) [32], although there are contradictory findings regarding the lineage that is expanded because of the overexpression of miR-125b, lymphoid [33], or myeloid [32]. However, the contribution of miR-125 family members in blood cell production is well recognized [32], as miR-125 has been identified in a subset of miRNAs enriched in HSCs, providing evidence that these miRNAs function to properly manage hematopoietic output.
In summary, in low-risk MDS patients, the treatment with deferasirox induced strong changes both in gene and miRNA expression of the patients. These changes mainly involved the downregulation of the Nf kB pathway, which could be related to the reduced inflammation. as well as the hemoglobin improvement observed after deferasirox treatment. Moreover, an important role regarding ROS-related genes, namely NFE2L2/NRF2, is suggested. In addition, the GFI1 overexpression and the downregulation of gamma interferon could be related to the increase in hemoglobin and platelets levels following the therapy. Besides, miRNAs may play an important role in the hematopoiesis, controlling diverse biological process.
Our in vivo work not only confirms previous results, mainly based on in vitro models, but also provides the biological basis for the clinical results observed in MDS patients treated with deferasirox, such as the erythropoiesis improvement and the decrease in the transfusion dependency of these patients, thus explaining the effects of this iron chelator and making this therapy more rational.
References
Gattermann N. Iron overload in myelodysplastic syndromes (MDS). Int J Hematol. 2018;107:55–63.
Cario H, Janka-Schaub G, Janssen G, Jarisch A, Strauss G et al. Recent developments in iron chelation therapy. Klin Padiatr. 2007;219:158–65.
Kushner JP, Porter JP, Olivieri NF. Secondary iron overload. Hematol Am Soc Hematol Educ Progr. 2001:47–61.
Olivieri NF, Brittenham GM. Iron-chelating therapy and the treatment of thalassemia. Blood. 1997;89:739–61.
Olivieri NF, Sabouhanian A, Gallie BL. Single-center retrospective study of the effectiveness and toxicity of the oral iron chelating drugs deferiprone and deferasirox. PLoS ONE. 2019;14:e0211942.
Piciocchi A, Sargentini V, Cotugno F, Bontempi K, Beltrami G, Di Tucci AA, et al. Update of the GIMEMA MDS0306 study: deferasirox for lower risk transfusion-dependent patients with myelodysplastic syndromes. Eur J Haematol. 2019;102:442–3.
Musto P, Maurillo L, Simeon V, Poloni A, Finelli C, Balleari E, et al. Iron-chelating therapy with deferasirox in transfusion-dependent, higher risk myelodysplastic syndromes: a retrospective, multicentre study. Br J Haematol. 2017;177:741–50.
Mills KI, Kohlmann A, Williams PM, Wieczorek L, Liu W, Li R, et al. Microarray-based classifiers and prognosis models identify subgroups with distinct clinical outcomes and high risk of AML transformation of myelodysplastic syndrome. Blood. 2009;114:1063–72.
del Rey M, O’Hagan K, Dellett M, Aibar S, Colyer HAA, Alonso ME, et al. Genome-wide profiling of methylation identifies novel targets with aberrant hypermethylation and reduced expression in low-risk myelodysplastic syndromes. Leukemia. 2013;27:610–8.
Chen J, Lu W-Y, Zhao M-F, Cao X-L, Jiang Y-Y, Jin X, et al. Reactive oxygen species mediated T lymphocyte abnormalities in an iron-overloaded mouse model and iron-overloaded patients with myelodysplastic syndromes. Ann Hematol. 2017;96:1085–95.
Messa E, Carturan S, Maffè C, Pautasso M, Bracco E, Roetto A, et al. Deferasirox is a powerful NF-kappaB inhibitor in myelodysplastic cells and in leukemia cell lines acting independently from cell iron deprivation by chelation and reactive oxygen species scavenging. Haematologica. 2010;95:1308–16.
Ohyashiki JH, Kobayashi C, Hamamura R, Okabe S, Tauchi T, Ohyashiki K. The oral iron chelator deferasirox represses signaling through the mTOR in myeloid leukemia cells by enhancing expression of REDD1. Cancer Sci. 2009;100:970–7.
Vardiman JW, Thiele J, Arber DA, Brunning RD, Borowitz MJ, Porwit A, et al. The 2008 revision of the World Health Organization (WHO) classification of myeloid neoplasms and acute leukemia: rationale and important changes. Blood. 2009;114:937–51.
Bolstad BM, Irizarry RA, Astrand M, Speed TP. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics. 2003;19:185–93.
Huang DW, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4:44–57.
Huang DW, Sherman BT, Lempicki RA. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009;37:1–13.
Carmona-Saez P, Chagoyen M, Tirado F, Carazo JM, Pascual-Montano A. GENECODIS: a web-based tool for finding significant concurrent annotations in gene lists. Genome Biol. 2007;8:R3.
List AF, Baer MR, Steensma DP, Raza A, Esposito J, Martinez-Lopez N, et al. Deferasirox reduces serum ferritin and labile plasma iron in RBC transfusion-dependent patients with myelodysplastic syndrome. J Clin Oncol. 2012;30:2134–9.
Gattermann N, Finelli C, Della Porta M, Fenaux P, Stadler M, Guerci-Bresler A, et al. Hematologic responses to deferasirox therapy in transfusion-dependent patients with myelodysplastic syndromes. Haematologica. 2012;97:1364–71.
Angelucci E, Santini V, Di Tucci AA, Quaresmini G, Finelli C, Volpe A, et al. Deferasirox for transfusion-dependent patients with myelodysplastic syndromes: safety, efficacy, and beyond (GIMEMA MDS0306 Trial). Eur J Haematol. 2014;92:527–36.
Banerjee A, Mifsud NA, Bird R, Forsyth C, Szer J, Tam C, et al. The oral iron chelator deferasirox inhibits NF-κB mediated gene expression without impacting on proximal activation: implications for myelodysplasia and aplastic anaemia. Br J Haematol. 2015;168:576–82.
Lyle L, Hirose A. Iron overload in myelodysplastic syndromes: pathophysiology, consequences, diagnosis, and treatment. J Adv Pract Oncol. 2018;9:392–405.
Meunier M, Ancelet S, Lefebvre C, Arnaud J, Garrel C, Pezet M, et al. Reactive oxygen species levels control NF-κB activation by low dose deferasirox in erythroid progenitors of low risk myelodysplastic syndromes. Oncotarget. 2017;8:105510–24.
Jiménez‐Solas T, López‐Cadenas F, Aires‐Mejía I, Caballero‐Berrocal JC, Ortega R, Redondo AM, et al. Deferasirox reduces oxidative DNA damage in bone marrow cells from myelodysplastic patients and improves their differentiation capacity. Br J Haematol. 2019;187:93–104.
Anguita E, Candel FJ, Chaparro A, Roldán-Etcheverry JJ. Transcription factor GFI1B in health and disease. Front Oncol. 2017;7:54.
Nolte F, Höchsmann B, Giagounidis A, Lübbert M, Platzbecker U, Haase D, et al. Results from a 1-year, open-label, single arm, multi-center trial evaluating the efficacy and safety of oral Deferasirox in patients diagnosed with low and int-1 risk myelodysplastic syndrome (MDS) and transfusion-dependent iron overload. Ann Hematol. 2013;92:191–8.
Molteni A, Riva M, Pellizzari A, Borin L, Freyrie A, Freyre A, et al. Hematological improvement during iron-chelation therapy in myelodysplastic syndromes: the experience of the «Rete Ematologica Lombarda». Leuk Res. 2013;37:1233–40.
Morales-Mantilla DE, King KY. The role of interferon-gamma in hematopoietic stem cell development, homeostasis, and disease. Curr Stem Cell Rep. 2018;4:264–71.
Tili E, Michaille J-J, Cimino A, Costinean S, Dumitru CD, Adair B, et al. Modulation of miR-155 and miR-125b levels following lipopolysaccharide/TNF-alpha stimulation and their possible roles in regulating the response to endotoxin shock. J Immunol. 2007;179:5082–9.
Chaudhuri AA, So AY-L, Sinha N, Gibson WSJ, Taganov KD, O’Connell RM, et al. MicroRNA-125b potentiates macrophage activation. J Immunol. 2011;187:5062–8.
Park H, Huang X, Lu C, Cairo MS, Zhou X. MicroRNA-146a and microRNA-146b regulate human dendritic cell apoptosis and cytokine production by targeting TRAF6 and IRAK1 proteins. J Biol Chem. 2015;290:2831–41.
O’Connell RM, Chaudhuri AA, Rao DS, Gibson WSJ, Balazs AB, Baltimore D. MicroRNAs enriched in hematopoietic stem cells differentially regulate long-term hematopoietic output. Proc Natl Acad Sci USA. 2010;107:14235–40.
Ooi AGL, Sahoo D, Adorno M, Wang Y, Weissman IL, Park CY. MicroRNA-125b expands hematopoietic stem cells and enriches for the lymphoid-balanced and lymphoid-biased subsets. Proc Natl Acad Sci USA. 2010;107:21505–10.
Acknowledgements
This work was partially supported by grants from the Spanish Fondo de Investigaciones Sanitarias PI15/01471, PI17/01741, PI18/01500, Instituto de Salud Carlos III (ISCIII), European Regional Development Fund (ERDF) “Una manera de hacer Europa”, “Consejería de Educación, Junta de Castilla y León” (SA271P18), “Proyectos de Investigación del SACYL”, Spain: GRS 1847/A/18, GRS 1653/A17, GRS 1850/A/18, “Fundación Memoria Don Samuel Solórzano Barruso”, by grants (RD12/0036/0069) from Red Temática de Investigación Cooperativa en Cáncer (RTICC) and Centro de Investigación Biomédica en Red de Cáncer (CIBERONC CB16/12/00233). JMHS and AERV are supported by a research grant by FEHH (“Fundación Española de Hematología y Hemoterapia”). We are grateful to I. Rodríguez, S. González, T. Prieto, M. Á. Ramos, AM, A. Díaz, A. Simón, M. del Pozo, V. Gutiérrez, and S. Pujante, Sandra Santos and Cristina Miguel from Cancer research Center of Salamanca, Salamanca, for their technical assistance.
Author information
Authors and Affiliations
Contributions
JMHS designed the experiment, interpreted the results and performed bioinformatic analysis; DAL performed bioinformatic analysis; EL, MA, and MDR contributed to the interpretation of the results; TG, MDC, AAM, RP, SE, BA, and JMHR performed patient selection and provided clinical data; JMHR and AERV contributed to the interpretation of the results and wrote the paper. All authors revised the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Sánchez, J.M.H., Lumbreras, E., Díez-Campelo, M. et al. Genome-wide transcriptomics leads to the identification of deregulated genes after deferasirox therapy in low-risk MDS patients. Pharmacogenomics J 20, 664–671 (2020). https://doi.org/10.1038/s41397-020-0154-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41397-020-0154-5
- Springer Nature Limited