Abstract
Statin therapy response is highly variable. Variants of lipid metabolism genes and statin pharmacokinetic modulators could play a role, however, the impact of most of these variants remains unconfirmed. A prospective and multicenter study included 252 patients was carried out in order to assess, according to achievement of LDL-C or non-HDL-C therapeutic targets and quantitative changes in lipid profiles, the impact of CETP, ABCA1, CYP2D6, and CYP2C9 gene candidate variants on the simvastatin, atorvastatin, and rosuvastatin response. Patients carrier ABCA1 rs2230806 and CYP2D6*3 variants are less likely to achieve therapeutic lipid targets (p = 0.020, OR = 0.59 [0.37, 0.93]; p = 0.040, OR = 0.23 [0.05, 0.93], respectively). Among CETP variants, rs708272 was linked to a 10.56% smaller reduction in LDL-C with rosuvastatin (95% CI = [1.27, 19.86] %; p = 0.028). In contrast, carriers of rs5882 had a 13.33% greater reduction in LDL-C (95% CI = [25.38, 1.28]; p = 0.032). If these findings are confirmed, ABCA1, CYP2D6, and CETP genotyping could be used to help predict which statin and dosage is appropriate in order to improve personalized medicine.
Similar content being viewed by others
Introduction
Statins are drugs that inhibit 3-hydroxy-3-methyl-glutaril-CoA reductase, a rate limiting enzyme of the cholesterol synthesis pathway. Statins are widely used for the primary and secondary prevention of atherosclerosis [1]. However, patient responses to these drugs are highly variable (20–60%) with respect to reducing plasma concentrations of LDL cholesterol (LDL-C) and increasing plasma concentrations of HDL cholesterol (HDL-C) [2]. Genetic factors contribute to this variability in therapeutic responses [3].
The results from first whole genome association study to investigate the relationship between genetics and response to statins were reported in 2009 [4]. Studies of the interactions between genes and statin efficacy have since then become more common [5, 6].
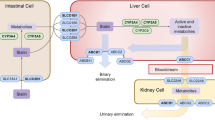
Many genes that are involved in the transport, metabolism, and elimination of statins, as well as genes involved in lipid metabolism, could influence therapeutic responses. The best-characterized candidates are the lipid metabolism genes APOE [7] and HMGCR [8] and the statin transport genes ABCB1 [9] and SLCO1B1 [10].
Other genes associated with patient response to statin treatment are SORT1/CELSR2/PRSC1 [11], KIF6 [12], and PCSK9 [13]. CETP [14] and ABCA1 [15] are known to influence lipid metabolism and modulate responses to lipid-reduction therapies, including statins. CETP mediates the transfer of cholesterol esters from HDL to VLDL and LDL [16] while ABCA1 is a transporter of intracellular cholesterol that mediates the efflux of cholesterol from the cell to the molecular acceptor apolipoprotein A1 [17]. ABCA1 therefore mediates the transport of cholesterol out of cells, a process which leads to elimination of cholesterol by hepatocytes.
Further studies of the genetic variants of the abovementioned genes are promising because conclusive information about the impact of these genes on response to statin treatment has yet to be reported.
The intronic genetic CETP variant rs708272 (Taq1B) results from the substitution of a single adenine (A) for a guanine (G). The A allele is associated with increased HDL-C and a reduced concentration of CETP protein [18, 19]. However, this allele could be responsible for a lower therapeutic response to statins, as patients carrying this allele have been reported to suffer more coronary artery disease (CAD) events [20], though the opposite has also been published [21].
The CETP variant rs5882 produces a protein with a valine substituted for isoleucine at position 405 (I405V). This amino acid change leads to downregulation of CETP [22]. The relationship of this variant with efficacy of statin treatment remains unclear [14, 23].
The ABCA1 variant rs2230806, results from a lysine substitution for arginine at position 219 (R219K). The minor genotype (TT) has been associated with a lower risk of CAD [24], a lower LDL-C, and a higher HDL-C [25]. However, other reports contradict this model and suggest that this genotype does not affect lipid profiles [26] or actually increase the risk of CAD [27]. Besides, information about the effects of this variant on statin treatment is scarce [28, 29].
The pharmacokinetics of statins are not well understood because the statin metabolic pathways have not been completely characterized. Statins are primarily metabolized by cytochrome P450 (CYP450) and CYP3A4/5 [30]. The CYP2C8 and CYP2C19 enzymes also appear to modulate these pathways, but their mechanisms of action have yet to be elucidated [31]. Meanwhile, the CYP2C9 enzyme is primarily responsible for modifying fluvastatin [32], which explains why some CYP2C9 variants are associated with changes in fluvastatin efficacy [33]. However, how these variants affect other statins is still unknown. CYP2D6 is involved in pitavastatin and rosuvastatin metabolism [34]. This gene also appears to affect statin treatment efficacy, but evidence for this model is scarce and controversial [35, 36].
In order to more deeply understand the impact of these gene variants on the efficacy of statin treatment, a prospective and multicenter study was carried out to determine the influence of the following gene variants on patient response to statins: the CETP variants rs708272 and rs5882, the ABCA1 variant rs2230806, the CYP2D6 variants rs35742686, rs3892097, and rs5030655, and the CYP2C9 variants rs1799853 and rs1057910. Three statins were examined in the study: simvastatin, atorvastatin, and rosuvastatin. Simvastatin and atorvastatin are routinely and widely administered in clinical practices. Rosuvastatin was chosen in order to remediate the lack of studies that have examined the interaction between this statin and genetic variations. The efficacy of statin treatment was determined by observing whether or not lipid targets were achieved and the resulting quantitative changes in lipid profiles.
Materials and methods
Type of study
This was a prospective, observational, and multicenter study. One primary care center and three high complexity hospitals were involved. Two of the three hospitals have a Cardiovascular Risk Unit.
Population: inclusion and exclusion criteria
Three hundred and forty-four patients were included in the study. Inclusion criteria considered patients without lipid-lowering treatment whose LDL-C was high and statin drugs, if necessary, were prescribed in conditions of usual medical practice and according solely to the patients’ physicians’ criteria.
The following exclusion criteria were applied: (1) having familial hypercholesterolemia due to mutations in the LDLR, APOB, LDLRAP1, or PSCK9 genes; (2) being a polymedicated patient (six or more drugs) or a patient being treated with immunosuppressants, antidepressants, antiretrovirals, or other lipid-reducing drugs; (3) having an autoimmune disease; (4) suffering from chronic liver disease; (5) being suspected of null or poor adherence to treatment (verified during the follow-up visit through the interview and medical criteria); (6) having hypothyroidism; (7) participation in another clinical trial; (8) intolerance to statins, defined as the inability to tolerate a dose of statin high enough to reduce cardiovascular risk, and/or suffering from side effects such as muscle symptoms, headache, sleep disorders, dyspepsia, nausea, rash, alopecia, erectile dysfunction, gynecomastia, and/or arthritis [37].
Ethical concerns and data protection
In compliance with regulation #SAS/3470/2009, this study was classified by the Division of Pharmacoepidemiology of the Department of Medicines and Health Products of Spain (ref. PR169/14). This study was also approved by the Institutional Review Boards of each participating hospital. Informed consent was obtained from each patient during their initial visit after having received a complete explanation of the study from their attending physician.
Data collection
Clinical and demographic data were collected from each patient during the initial visit. This data included: age (years), sex, prior CAD (yes/no), family history of ischemia (yes/no), hypertension (yes/no), diastolic blood pressure (mmHg), systolic blood pressure (mmHg), diabetes (yes/no), smoking history (yes/no), current tobacco use (yes/no), history of alcohol consumption (yes/no), current alcohol consumption (yes/no), body mass index, exercise (yes/no), exercise intensity (no/low/ moderate/high), initial LDL-C, HDL-C and non-HDL cholesterol (non-HDL-C), concentration of lipoprotein (a) (g/L) and carrying the SLCO1B1 gene variant rs4149056 (C allele) [38]. Treatment intensity was defined qualitatively (low, moderate, and high) according to clinical guidelines of the American College of Cardiology [39].
Data related to lipid metabolism were collected during the initial visit (prior to treatment) and during the follow-up visit (~3 months after initiation of treatment) in all treatment groups. This included plasma levels of triglycerides, total cholesterol, LDL-C, non-HDL-C, and HDL-C. Lipid profile therapy targets were established according to the clinical guidelines for statin therapies set by the National Cholesterol Education Program [40]. In the case of primary prevention, the target was an LDL-C of <3.4 mmol/L or a non-HDL-C of <4.1 mmol/L. Secondary prevention was defined as treatment subsequent to a cardiovascular event or treatment of a patient with equivalent risk factors such as diabetes. In this case, the therapeutic target was set to an LDL-C of <2.6 mmol/L or a non-HDL-C of <3.4 mmol/L. The targets for non-HDL-C were taken into account only when it was impossible to calculate an LDL-C value using the Friedewald formula [41].
Genetic analysis
The following gene variants were studied: CETP variants rs708272 (NM_000078.2:c.118 + 279G>A, known as TAQ1B) and rs5882 (NM_000078.2:c.1264A>G, NP_000069.2:p.Val422Ile, known as I405V); ABCA1 variant rs2230806 (NM_005502.3:c.656C>T, NP_005493.2:p.Arg219Lys, known as R219K), CYP2D6 variants CYP2D6*3 rs35742686 (NM_000106.5:c.775delT), CYP2D6*4 rs3892097 (NG_008376.3:g.6047C>T), and CYP2D6*6 rs5030655 (NM_000106.5:c.454delA); and CYP2C9 variants CYP2C9*2 rs1799853 (NM_000771.3:c.430C>T) and CYP2C9*3 rs1057910 (NM_000771.3:c.1075A>C).
The genotypic frequency of each variant was determined and compared with information found in the 1000 Genomes Database [42]. The PLINK program was used to predict whether or not there was a linkage disequilibrium between variants [43].
DNA was extracted from whole blood using a commercial kit (Maxwell® 16 System, Promega, Madison, USA). Genotypes were analyzed by real-time PCR using Taqman probes and a 7500 Real-Time PCR System thermocycler (Thermo Fisher Scientific, California, USA).
Importantly, copy number variations (CNVs) have been reported for CYP2D6. Therefore, some individuals may carry null alleles or extra copies. However, Taqman probes cannot detect a CNV. The frequency of variants in CNVs in the European population is 5.01% [44].
Sanger sequencing
Using a 3130 Genetic Analyzer (Applied Biosystems), PCR amplification and direct sequencing of gene variants were carried out on 24 different samples with the aim of validating the real-time PCR method of detection. Sequences were analyzed using the Sequencing Analysis v5.2 software.
Statistical analyses
Comparing averages and percentages
Relationships between genetic variant genotypes and percent changes in LDL-C, non-HDL-C, and HDL-C were analyzed using the ANOVA test once normal distribution and comparable variance was confirmed in both groups. The relationship between variant genotypes and achievement of therapy targets was analyzed using the Chi-squared test.
Multivariate analyses
A correlation matrix was constructed to find associations between control variables (criterion r > 0.4). The multivariate model was constructed using the back stepwise method.
First, a multiple logistic regression model was used to evaluate whether or not achieving the LDL-C/non-HDL-C targets depended on variant genotypes. A multiple linear regression model was then constructed to study whether or not the changes in lipid concentration that occurred between the initial and the follow-up visit were influenced by the genetic variants. Both models were constructed by grouping the statins and then stratifying them. Statistical analyses were performed using the SPSS v.17 software package (SPSS Inc., Chicago, IL, USA).
Results
Population
Three hundred and forty-four patients were prospectively included in the study between June 2014 and February 2017. Of these, 92 were later excluded for one of the following reasons:
-
Had received prior lipid-reducing therapy (33 patients).
-
Did not adhere to treatment regime (nine patients), prospectively.
-
Intolerance to statins due to muscle complications (three patients).
-
Failed to follow-up (20 patients).
-
Familial hypercholesterolemia was genetically diagnosed during the study (11 patients).
-
Hypothyroidism (two patients).
-
Autoimmune disease (one patient).
-
Other causes (13 patients).
A statistical analysis of the data provided by 252 patients was carried out. A power analyses revealed that there was a 99.95% chance that, using an ANOVA-test in the context of a multiple linear regression, a contribution of a genetic variant of as low as 5% to patient responses to statin therapies would be detected by this study.
Descriptive statistics
One hundred and six patients were treated with simvastatin, 116 were treated with atorvastatin, and 30 were treated with rosuvastatin.
Clinical and demographic data are laid out in Table 1. The genotypic frequency of the variants fulfilled the Hardy–Weinberg equilibrium principle and had a similar distribution to what had been previously reported in 1000 Genomes [42] (see Supplementary Material). No linkage disequilibrium between variants was observed (see Supplementary Material).
Comparison of means and percentages of dependent variables with respect to genotype
The effects of each genetic variant on the changes in lipid profiles after treatment with statins are displayed in Table 2. For the ABCA1 variant rs2230806, the percentage of patients who achieved therapeutic targets was lower among those carrying the T allele (p = 0.004). In the case of CYP2D6*4, carriers of the T allele had a smaller increase in HDL-C than non-carriers (p = 0.020). Regarding CYP2D6*3, the percentage of patients who achieved therapeutic targets was lower among carriers of the T deletion, a tendency which was not statistically significant (p = 0.098).
Multivariate analysis
Achievement of LDL-C/non-HDL-C targets
The impact of variant genotypes on achievement of LDL-C/non-HDL-C targets was analyzed both for statins as a whole, and for stratified groups. The models were adjusted according control variables that had made a contribution: initial non-HDL-C/LDL-C and patient age. In the case of the rosuvastatin group, the control variable was the initial non-HDL-C/LDL-C alone.
All statins:
See Fig. 1a for a summary of the results. A relationship was observed between, on the one hand, being a carrier of either the ABCA1 variant rs2230806 (the T allele) or the CYP2D6*3 variant (T deletion) and, on the other, a lower probability of achieving therapeutic targets. Carriers of either variant have a lower probability of achieving therapeutic targets than carriers of the major allele (ABCA1 rs2230806; p = 0.020; OR = 0.59; 95% CI = [0.37, 0.93] and CYP2D6*3 rs35742686; p = 0.040; OR = 0.23; 95% CI = [0.05, 0.93]).
a–c Achievement of LDL-C/non-HDL-C targets. Taking into account the minor allele with respect to the major allele, the multiple logistic regression model was adjusted according to statistically significant covariates: Initial concentration of non-HDL-C and age for all statin and simvastatin + atorvastatin study and Initial concentration of non-HDL-C for rosuvastatin study. In yellow box = Odds Ratio + 95% confidence interval. NA not applicable (n of any genotype group <3).
Simvastatin + atorvastatin:
When stratifying according to statins, simvastatin and atorvastatin were found to interact similarly with the gene variants studied. Therefore, in order to prevent the loss of statistical power, the data resulting from both treatments were grouped together (Fig. 1b). The following statistics were calculated for the variants: ABCA1 rs2230806 (T allele); p = 0.030; OR = 0.57; 95% CI = [0.35, 0.94]; and CYP2D6*3; (T deletion); p = 0.020; OR = 0.16; 95% CI = [0.03, 0.74], confirming the tendencies observed in the analysis of all three statin groups combined.
Rosuvastatin:
There was a statistically significant relationship between reaching therapeutic targets under treatment with rosuvastatin and being a carrier of the CETP variant rs708272 (A allele), see Fig. 1c. Patients carrying the minor allele were less likely to achieve therapeutic targets (p = 0.030; OR = 0.20; CI 95% = [0.05, 0.83]).
Statistically significant relationships between the achievement of lipid targets and the other variants were not detected in the case of rosuvastatin.
Changes in the plasma levels of LDL-C, non-HDL-C, and HDL-C
Changes in lipid concentrations between the initial and follow-up patient visit are laid out in Table 3. The models were adjusted according to the following control variables: initial non-HDL-C and intensity of treatment, in the case of the combined statin treatment groups. Besides, when analyzing rosuvastatin group alone, the only control variable was “prior CAD”.
All statins/simvastatin + atorvastatin:
The results from the linear multivariate regressions were consistent with the analysis of target achievement in the case of the ABCA1 variant rs2230806: for the combined statin, a statistically significant difference was observed for LDL-C and non-HDL-C (p = 0.047 and 0.050, respectively). Regarding the simvastatin + atorvastatin group, homozygotes for the minor T allele had a 7.56% (IC 95% = [0.13, 14.99]) and a 12.10% (IC 95% = [2.73, 21.46]) smaller reduction in LDL-C and non-HDL-C, (p = 0.012 and 0.046, respectively).
Rosuvastatin:
Regarding the subgroup treated with rosuvastatin, there was also a relationship between the CETP variant rs708272 and changes in LDL-C and non-HDL-C (p = 0.028 and 0.042). Carriers of the A allele had a 10.56% smaller reduction in LDL-C (IC 95% = [1.27, 19.86]) and an 8.05% smaller reduction in non-HDL-C (IC 95% = [0.32,15.77]).
No relationship was observed between the CETP variant rs5882 and achievement of therapeutic targets. Nevertheless, there was a relationship between this variant and changes in LDL-C and non-HDL-C. Carriers of the minor G allele responded better to treatment with rosuvastatin than patients carrying the major A variant. G carriers responded with a 13.33% greater reduction in LDL-C (IC 95% = [25.38, 1.28]); p = 0.0320) and a 11.61% greater reduction in non-HDL-C (IC 95% = [21.50, 2.07]; p = 0.019).
No other statistically significant relationship was observed between variants and lipid profiles.
Discussion
Statin efficacy depends in part on genetic variabilities [45]. As far as we know, this is the first observational and prospective study assessing the effectiveness of statins according to both changes in lipid profiles and achievement of therapeutic targets.
Eighty-one out of the 252 patients included in our study (32.1%) did not achieve their LDL-C/non-HDL-C targets after being treated with statins. The objective of this project was to determine whether or not the abovementioned genetic variants could be used to predicatively stratify patients according to their response to statin treatment in a way that could be used to prescribe more intensive therapies to those who are least responsive. In this way, cardiovascular risk would be reduced as soon as possible, and available resources would be used more efficiently.
The only statistically significant differences we found in the demographic variables were in age and history of alcohol consumption. The average age of the group of those who met their therapeutic targets was lower than those who did not (50 vs 55). Besides, a history of alcohol consumption is associated with a lower likelihood of meeting LDL-C/non-HDL-C targets.
It is logical to assume that the control variables initial lipid levels and statin dosage have an influence on changes in lipid profiles that occur during statin treatment. However, in the case of rosuvastatin, the control variable prior CAD event is the one with influence, a result consistent with the fact that rosuvastatin is restricted to those at high cardiovascular risk. Most patients who have suffered prior ischemic events are therefore, in this group.
Relationships between ABCA1, CYP2D6, and CETP variants and patient response to statin therapies were detected. In the case of the ABCA1 variant rs2230806, this tendency holds for the subgroup of patients who were administered simvastatin or atorvastatin.
The ABCA1 gene is located on chromosome 9 (9q31.1 region). This gene codes for the ABCA1 protein (ATP-binding cassette transporter A1). ABCA1 modulates the expulsion of intracellular cholesterol and phospholipids. Upon being expelled from inside the cell, cholesterol forms complexes with extracellular apolipoproteins to form nascent HDL. Therefore, ABCA1 forms part of an efficient cellular disposal system for excess intracellular cholesterol [46]. ABCA1 also has antiinflammatory role because it modulates the cholesterol content of membrane lipid rafts [47]. The ABCA1 variant rs2230806 results from a mutation at codon 219 that leads to a substitution of arginine for lysine in an extracellular loop known to interact with the apoA-I and to mediate cholesterol transportation [48].
Here we demonstrate that homozygous for the minor T allele of this variant respond more poorly to statin treatment, showing a 12% lower reduction in LDL-C in response to treatment with simvastatin and atorvastatin. TT carriers are also less likely to achieve their LDL-C/non-HDL-C targets than C carriers. Statins may also regulate the expression of ABCA1 [49]. The ABCA1 variant could lead to poorer treatment outcomes because of a diminished level of gene expression. However, no in vitro studies have demonstrated the specific role of this genetic variant.
In contrast with our findings, neither Akao et al. [29] nor Li et al. [28] detected a relationship between the ABCA1 polymorphism and variations in LDL-C with pravastatin, though the latter observed higher levels of HDL-C. The inconsistencies in findings from ABCA1 variant studies in relation to CAD, lipid profiles, and response to statins could be explained by the racial diversity of the populations studied, gene–gene and environmental interactions, types of statins assessed, or the differences in selection criteria. In vitro studies of the ABCA1 variant are necessary to explain its impact on statins on a molecular level.
Meanwhile, the CYP2D6*3 variant is also associated with statin treatment efficacy. Patients carrying the T deletion are less likely to achieve LDL-C/non-HDL-C targets, and therefore are more refractory to statin treatments.
CYP2D6 is located on chromosome 22 and codes for a liver enzyme that belongs to the metabolic pathway of many drugs. CYP2D6 is highly polymorphic, with over 100 allelic described [50]. The CYP2D6*3 variant is a consequence of a thymine deletion that inactivates the allele. Studies examining relationship between CYP2D6 variants and statins have been rare and inconclusive, particularly for the CYP2D6*3 variant [35, 51]. The lack of findings may be a consequence of the low allelic frequency of this variant (MAF = 0.018; European CEU) [42]. Importantly, the activity of CYP2D6 affects patient’s response to tramadol [52] and tamoxifen [53] because CYP2D6 activates these drugs. Therefore, it could be hypothesized that the CYP2D6 enzyme also could break down statins that have active metabolites, such as simvastatin and atorvastatin [54].
Finally, we have demonstrated that CETP variants rs708272 (G>A) and rs5882 (A>G) have opposing effects on rosuvastatin efficacy. While carriers of minor A allele (rs708272) respond more poorly to treatment, those with the minor G allele (rs5882) show a greater decrease in LDL-C than noncarriers. The role of these in cholesterol metabolism and the onset of cardiovascular diseases is well known [55], but their interaction with statins is not well understood [56].
The rs708272 variant is located in intron 1 and is a consequence of a guanine substitution for an adenine in position 277, which removes an endonuclease restriction site. The minor A allele is associated with lower concentrations of CETP protein and higher HDL-C [18, 19]. This relationship can be explained by the strong linkage disequilibrium between this variant and the promoter of the rs1800775 variant (known as −629A>C) [57]. Patients carrying the minor allele are less at risk for CAD, though with the caveat that this conclusion was drawn with a high cardiovascular risk population of subjects instead of general population [30].
In contrast, carriers of the minor A allele are at higher risk of CAD when treated with statins [29] in comparison with the major allele; a phenomenon which could be linked to a smaller reduction in LDL-C, in accordance with the results of our study.
The product of another CETP variant examined here, rs5882 (G), contains a single amino acid substitution on position 422(Val422Ile). This mutation downregulates CETP gene expression [31]. According to our observations, carriers of the minor G allele may benefit more from lipid-reduction therapies than carriers of the major A allele. This is the first study that has detected a relationship between the aforementioned CETP variants and the efficacy of rosuvastatin treatment. However, the small number of patients being treated by rosuvastatin in the recruitment centers limits our ability to interpret these results. Rosuvastatin is not the drug of choice in primary care settings and is reserved for patients who are at high cardiovascular risk or who suffer from other comorbidities. Therefore, naïve patients being treated with rosuvastatin are difficult to come across.
Even though large sample sizes are difficult to assemble, our results regarding rosuvastatin efficacy need to be confirmed by increasing statistical power.
Conclusion
ABCA1 variant rs2230806 and CYP2D6*3 influence patient response to treatment with statins. Variant carriers are less likely to achieve LDL-C or non-HDL-C therapeutic targets. In addition, carriers of the TT genotype of rs2230806 have a 12% lower LDL-C in response to treatment with simvastatin and atorvastatin.
CETP gene variants rs708272 and rs5882 affect patient responses to rosuvastatin. However, a larger sample size is required to corroborate these results.
If these findings are confirmed, ABCA1, CYP2D6, and CETP genotyping could be used to help predict which statin and dosage is appropriate in order to improve personalized medicine.
References
Chasman DI, Giulianini F, MacFadyen J, Barratt BJ, Nyberg F, Ridker PM. Genetic determinants of statin-induced low-density lipoprotein colesterol reduction: The Justification for the Use of Statins in Prevention: an Intervention Trial Evaluating Rosuvastatin (JUPITER) trial. Circ Cardiovasc Genet. 2012;5:257–64.
Davidson MH, Toth PP. Comparative effects of lipid-lowering therapies. Prog Cardiovasc Dis. 2004;47:73–104.
Verschuren JJW, Trompet S, Wessels JAM, Guchelaar HJ, de Maat MP, Simoons ML, et al. A systematic review on pharmacogenetics in cardiovascular disease: is it ready for clinical application? Eur Heart J. 2012;33:165–75.
Thompson JF, Hyde CL, Wood LS, Paciga SA, Hinds DA, Cox DR, et al. Comprehensive whole-genome and candidate gene analysis for response to statin therapy in the treating to new targets (TNT) cohort. Circ Cardiovasc Genet. 2009;2:173–81.
Vrablík M, Hubáček JA, Dlouhá D, Lánská V, Rynekrová J, Zlatohlávek L, et al. Impact of variants within seven candidate genes on statin treatment efficacy. Physiol Res. 2012;61:609–17.
Munshi A. Genetic variation in MDR1, LPL and eNOS genes and the response to atorvastatin treatment in ischemic stroke. Hum Genet. 2012;131:1775–81.
Deepak Voora, Svati HS, Reed CR, Zhai J, Crosslin DR, Messer C, et al. Pharmacogenetic predictors of statin-mediated low-density lipoprotein cholesterol reduction and dose response. Circ Cardiovasc Genet. 2008;1:100–6.
Chasman DI, Posada D, Subrahmanyan L, Cook NR, Stanton VP Jr, Ridker PM. Pharmacogenetic study of statin therapy and cholesterol reduction. JAMA. 2004;291:2821–7.
Becker ML, Visser LE, van Schaik RH, Hofman A, Uitterlinden AG, Stricker BH. Common genetic variation in the ABCB1 gene is associated with the cholesterol-owering effect of simvastatin in males. Pharmacogenomics. 2009;10:1743–51.
Martin NG, Li KW, Murray H, Putt W, Packard CJ, Humphries SE. The effects of a single nucleotide polymorphism in SLCO1B1 on the pharmacodynamics of pravastatin. Br J Clin Pharmacol. 2012;73:303–6.
Postmus I, Trompet S, Deshmukh HA, Barnes MR, Li X, Warren HR, et al. Pharmacogenetic meta-analysis of genome-wide association studies of LDL cholesterol response to statins. Nat Commun. 2014;5:5068.
Ruiz-Iruela C, Padró-Miquel A, Pintó-Sala X, Baena-Díez N, Caixàs-Pedragós A, Güell-Miró R, et al. KIF6 gene as a pharmacogenetic marker for lipid-lowering effect in statin treatment. PLoS ONE. 2018;13:e0205430.
Feng Q, Wei-Qi W, Chung C, Levinson R, Bastarache L, Joshua C, et al. The effect of genetic variation in PCSK9 on the LDL-cholesterol response to statin therapy. Pharmacogenomics J. 2017;17:204–8.
Ference BA, Kastelein JJP, Ginsberg HN, Chapman MJ, Nicholls SJ, Ray KK, et al. Association of genetic variants related to CETP inhibitors and statins with lipoprotein levels and cardiovascular risk. JAMA. 2017;318:947–56.
Li YY, Zhang H, Qin XY, Lu XZ, Yang B, Chen ML. ATP-binding cassette transporter A1 R219K polymorphism and coronary artery disease in Chinese population: a meta-analysis of 5,388 participants. Mol Biol Rep. 2012;39:11031–9.
Tall AR. Functions of cholesterol ester transfer protein and relationship to coronary artery disease risk. J Clin Lipidol. 2010;4:389–93.
Rocha KC, Pereira BMV, Rodrigues AC. An update on efflux and uptake transporters as determinants of statin response. Expert Opin Drug Metab Toxicol. 2018;14:613–24.
Kashani Farid MA, Azizi F, Hedayati M, Daneshpour MS, Shamshiri AR, Siassi F. Association between CETP Taq1B and LIPC −514 C/T polymor- phisms with the serum lipid levels in a group of Tehran’s population: a cross sectional study. Lipids Health Dis. 2010;9:96.
De Haan W, van der Hoogt CC, Westerterp M, Hoekstra M, Dallinga-Thie GM, Princen HM, et al. Atorvastatin increases HDL cholesterol by reducing CETP expression in cholesterol-fed APOE*3-Leiden.CETP mice. Atherosclerosis. 2008;197:57–63.
Regieli JJ, Jukema JW, Grobbee DE, Kastelein JJ, Kuivenhoven JA, Zwinderman AH, et al. CETP genotype predicts increased mortality in statin-treated men with proven cardiovascular disease: an adverse pharmacogenetic interaction. Eur Heart J. 2008;29:2792–9.
Carlquist JF, Muhlestein JB, Horne BD, Hart NI, Bair TL, Molhuizen H, et al. The cholesteryl ester transfer protein Taq1B gene polymorphism predicts clinical benefit of statin therapy in patients with significant coronary artery disease. Am Heart J. 2003;146:1007–14.
Boekholdt SM, Thompson JF. Natural genetic variation as a tool in understanding the role of CETP in lipid levels and disease. J Lipid Res. 2003;44:1080–93.
Anagnostopoulou K, Kolovou G, Kostakou P, Mihas C, Mikhailidis D, Cokkinos DV. Pharmacogenetic study of cholesteryl ester transfer protein gene and simvastatin treatment in hypercholesterolaemic subjects. Expert Opin Pharmacother. 2007;8:2459–63.
Ma XY, Liu JP, Song ZY. Associations of the ATP-binding cassette transporter A1 R219K polymorphism with HDL-C level and coronary artery disease risk: a meta-analysis. Atherosclerosis. 2011;215:428–34.
Lu Z, Luo Z, Jia A, Yu L, Muhammad I, Zeng W, et al. Associations of the ABCA1 gene polymorphisms with plasma lipid levels: a meta-analysis. Medicine. 2018;97:e13521.
Ghaznavi H, Aali E, Soltanpour MS. Association study of the atp-binding cassette transporter A1 (ABCA1) Rs2230806 genetic variation with lipid profile and coronary artery disease risk in an Iranian population. Open Access Maced J Med Sci. 2018;6:274–9.
Rejeb J, Omezzine A, Rebhi L, Boumaiza I, Kchock K, Belkahla R, et al. Associations between common polymorphisms of adenosine triphosphate-binding cassette transporter A1 and coronary artery disease in a Tunisian population. Arch Cardiovasc Dis. 2010;103:530–7.
Li J, Wang LF, Li ZQ, Pan W. Effect of R219K polymorphism of the ABCA1 gene on the lipidlowering effect of pravastatin in Chinese patients with coronary heart disease. Clin Exp Pharm Physiol. 2009;36:567–70.
Akao H, Polisecki E, Schaefer EJ, Trompet S, Robertson M, Ford I, et al. ABCA1 gene variation and heart disease risk reduction in the elderly during pravastatin treatment. Atherosclerosis. 2014;235:176–81.
Kitzmiller JP, Luzum JA, Dauki A, Krauss RM, Medina MW. Candidate-gene study of functional polymorphisms in SLCO1B1 and CYP3A4/5 and the cholesterol-lowering response to simvastatin. Clin Transl Sci. 2017;10:172–7.
Kitzmiller JP, Mikulik EB, Dauki AM, Murkherjee C, Luzum JA. Pharmacogenomics of statins: understanding susceptibility to adverse effects. Pharmacogenomics Personalized Med. 2016;9:97–106.
Neuvonen PJ, Backman JT, Niemi M. Pharmacokinetic comparison of the potential over-the-counter statins simvastatin, lovastatin, fluvastatin and pravastatin. Clin Pharmacokinet. 2008;47:463–74.
Buzková H, Pechandová K, Danzig V, Vareka T, Perlik F, Zak A, et al. Lipid-lowering effect of fluvastatin in relation to cytochrome P450 2C9 variant alleles frequently distributed in the Czech population. Med Sci Monit. 2012;18:CR512–17.
Fujino H, Saito T, Tsunenari Y, Kojima J, Sakaeda T. Metabolic properties of the acid and lactone forms of HMG-CoA reductase inhibitors. Xenobiotica. 2004;34:961–71.
Zuccaro P, Mombelli G, Calabresi L, Baldassarre D, Palmi I, Sirtori CR. Tolerability of statins is not linked to CYP450 polymorphisms, but reduced CYP2D6 metabolism improves cholesteraemic response to simvastatin and fluvastatin. Pharm Res. 2007;55:310–7.
Geisel J, Kivistö KT, Griese EU, Eichelbaum M. The efficacy of simvastatin is not influenced by CYP2D6 polymorphism. Clin Pharm Ther. 2002;72:595–6.
Banach M, Rizzo M, Toth PP, Farnier M, Davidson MH, Al-Rasadi K, et al. Position paper. Statin intolerance-an attempt at a unified definition. Position paper from an International Lipid Expert Panel. Arch Med Sci. 2015;11:1–23.
Ramsey LB, Johnson SG, Caudle KE, Haidar CE, Voora D, Wilke RA, et al. The clinical pharmacogenetics implementation consortium (CPIC) guideline for SLCO1B1 and simvastatin-induced myopathy: 2014 update. Clin Pharmacol Ther. 2014;96:423–8.
ACC/AHA Release Updated Guideline on the Treatment of Blood Cholesterol to Reduce ASCVD Risk. Am Fam Phys. 2014;90:260–5.
Third Report of the National Cholesterol Education Program (NCEP). Expert panel on detection, evaluation and treatment of high blood cholesterol in adults (adult treatment panel III) final report. Circulation. 2002;106:3143–421.
Friedewald WT, Levy RI, Fredrickson DS. Estimation of the concentration of low-density lipoprotein cholesterol in plasma, without use of the preparative ultracentrifuge. Clin Chem. 1972;18:499–502.
A global reference for human genetic variation. The 1000 Genomes Project Consortium. Nature. 2015;526:68–74.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D, et al. PLINK: a toolset for whole-genome association and population-based linkage analysis. Am J Hum Genet 2007;81:559–75. http://zzz.bwh.harvard.edu/plink/.
Crews KR, Gaedigk A, Dunnenberger HM, et al. Clinical Pharmacogenetics Implementation Consortium guidelines for cytochrome P450 2D6 genotype and codeine therapy: 2014 update. Clin Pharm Ther. 2014;95:376–82.
Barber MJ, Mangravite LM, Hyde CL, Chasman DI, Smith JD, McCarty CA, et al. Genome-wide association of lipid-lowering response to statins in combined study populations. PLoS ONE. 2010;5:e9763.
Phillips MC. Is ABCA1 a lipid transfer protein? J Lipid Res. 2018;59:749–63.
Liu Y, Tang C. Regulation of ABCA1 functions by signaling pathways. Biochim Biophys Acta. 2011;1821:522–9.
Zhao GJ, Yin K, Fu YC, Tang CK. The interaction of ApoA-I and ABCA1 triggers signal transduction pathways to mediate efflux of cellular lipids. Mol Med. 2012;18:149–58.
Niesor EJ, Schwartz GG, Perez A, Stauffer A, Durrwell A, Bucklar-Suchankova G, et al. Statin-induced decrease in ATP-binding cassette transporter A1 expression via microRNA33 induction may counteract cholesterol efflux to high-density lipoprotein. Cardiovasc Drugs Ther. 2015;29:7–14.
Gaedigk A, Ingelman-Sundberg M, Miller NA, Leeder JS, Whirl-Carrillo M, Klein TE, PharmVar Steering Committee. The pharmacogene variation (PharmVar) consortium: incorporation of the human cytochrome P450 (CYP) allele nomenclature database. Clin Pharm Ther 2018;103:399–401.
Li J, Wang X, Zhang Z, Zou J, Chen Y, Wang X, et al. Statin therapy correlated CYP2D6 gene polymorphism and hiperlipidemia. Curr Med Res Opin. 2014;30:223–8.
Dagostino C, Allegri M, Napolioni V, Bignami E, Mutti A, van Schaik RHN. CYP2D6 genotype can help to predict effectiveness and safety during opioid treatment for chronic low back pain: results from a retrospective study in an Italian cohort. Pharmgenomics Pers Med. 2018;11:179–91.
Cronin-Fenton DP, Lash TL. Clinical epidemiology and pharmacology of CYP2D6 inhibition related to breast cancer outcomes. Expert Rev Clin Pharmacol 2011;4:363–77.
Nieto-Ramirez I, Chegwin-Angarita C, Atehortúa L, Sepúlveda L. Statins: chemistry, analytical techniques, biosynthesis and pharmacokinetics. Vitae. 2013;20:49–63.
Thompson A, Di Angelantonio E, Sarwar N, Erqou S, Saleheen D, Dullaart RP, et al. Association of cholesteryl ester transfer protein genotypes with CETP mass and activity, lipid levels, and coronary risk. JAMA. 2008;299:2777–88.
Leusink M, Onland-Moret NC, de Bakker PI, de Boer A, Maitland-van der Zee AH. Seventeen years of statin pharmacogenetics: a systematic review. Pharmacogenomics. 2016;17:163–80.
Dachet C, Poirier O, Cambien F, Chapman J, Rouis M. New functional promoter polymorphism, CETP/-629, in cholesteryl ester transfer protein (CETP) gene related to CETP mass and high-density lipoprotein cholesterol levels: role of Sp1/Sp3 in transcriptional regulation. Arterioscler Thromb Vasc Biol 2000;20:507–15.
Acknowledgements
We thank Arnald Alonso-Pastor, Emili Corbella-Inglés, Ferran Trias-Vilagut, Marta Fanlo- Maresma, Hannia Elena Lafuente-González, Xavier Jusmet, and Enric Juncadella-García who provided insight and expertize that greatly assisted the research. We also would like to thank to Mollet Hospital (FSM, Mollet del Vallès, Spain) for their unstiting support in this project. The present work was performed as part of the Biochemistry, Molecular Biology and Biomedicine doctoral program of Cristina Ruiz-Iruela at Universitat Autònoma de Barcelona (Barcelona, Spain).
Fundings
This research was supported by Fundació Parc Taulí Institut Universitari UAB (Parc Taulí recerca scholarship 2012, CSPT/CIR/2012) and Fundación José Luis Castaño (FENIN scholarship 2014, JLC/FENIN/2014) from the Spanish Society of Laboratory Medicine (SEQC).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Ruiz-Iruela, C., Candás-Estébanez, B., Pintó-Sala, X. et al. Genetic contribution to lipid target achievement with statin therapy: a prospective study. Pharmacogenomics J 20, 494–504 (2020). https://doi.org/10.1038/s41397-019-0136-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41397-019-0136-7
- Springer Nature Limited