Abstract
Stimulation of the serotonin (5-HT)1A receptor (HTR1A) has been shown to contribute to the mechanism of action of some atypical antipsychotic drugs (APDs), including clozapine and lurasidone. A meta-analysis of rs6295, a functional polymorphism located at the promoter region of HTR1A, showed association with clinical response in schizophrenic patients treated with atypical APD. We have now tested whether other SNPs related to rs6295 predict response to lurasidone. We first evaluated whether rs358532 and rs6449693, tag SNPs for rs6295, predicted response to lurasidone, using data from two clinical trials of acutely psychotic schizophrenia patients with European (EUR, n = 171) or African (AFR, n = 131) ancestry; we then determined if those findings could be replicated in a third trial of lurasidone of similar design. Weekly changes (up to 6 weeks) in the Positive and Negative Syndrome Scale (PANSS) Total score and its five subscales were used to assess response. In EUR, a significant association, or trends for association, were observed for PANSS Total (p = 0.035), positive (p = 0.039), negative (p = 0.004), and disorganization (p = 0.0087) subscales, at week 1–6. There was a trend for replication with PANNS Total (p = 0.036) in the third trial. No significant association was observed in AFR or the placebo group. Meta-analysis of five studies, including the three with lurasidone, showed that rs6295 was associated with improvement in positive (p = 0.023) and negative (p ≤ 0.0001) symptoms in EUR patients with schizophrenia. This is the first study to show a significant association between functional HTR1A polymorphisms and treatment response to lurasidone. The meta-analysis provides additional evidence that rs6295 could be a race-dependent biomarker for predicting treatment response to APDs in schizophrenic patients with European Ancestry.
Similar content being viewed by others
Introduction
Schizophrenia (SCZ) is a common, phenotypically diverse, and genetically complex syndrome with devastating functional consequences. While antipsychotic drugs (APDs) treat the key aspects of the syndrome, psychosis, negative symptoms, and cognitive impairment, many patients are only partial responders. Pharmacogenomic studies attempt to identify genetic markers which can predict response to specific APDs and if successful, can lead to fewer failed treatment assignment [1,2,3,4,5].
Atypical APDs with clozapine as the prototype have important advantages for the treatment of SCZ over typical APDs in both efficacy and side effects [6]. More potent blockade of serotonin (5-HT)2A receptors (HTR2A) than dopamine (DA) D2 receptors (DRD2) is a shared property of many APDs, which lead to their advantages for efficacy and fewer side effects, such as extrapyramidal symptoms [7,8,9,10,11]. However, the atypical APDs have a variety of direct and indirect effects on other receptors, including HTR1A, HTR7, DRD1, and muscarinic receptors. SNPs for the HTR2A, HTR7 and DRD2 genes have been most intensively studied as biomarkers for overall response and response for specific domains, e.g., positive and negative symptoms [12,13,14,15,16,17,18]. However, results have been inconsistent [19].
The HTR1A in regions of the brain relevant to cognitive impairment, negative symptoms, and positive symptoms in SCZ, e.g., the prefrontal cortex (PFC), hippocampus (HIP), dorsal and ventral striatum, is a postsynaptic 5-HT heteroreceptor, whereas it is the presynaptic auto-receptor on 5-HT neurons in the dorsal raphe [20]. Blockade of HTR2A and DRD2 receptors increases cortical and striatal DA release via activation of HTR1A due to the release of 5-HT [21]. This mechanism may contribute to their efficacy for negative symptoms and fewer extrapyramidal symptoms produced by the atypical APDs [21, 22].
Altered HTR1A density has been initially identified in PFC and amygdala in postmortem specimens from SCZ patients [23, 24]. It has also been reported that tandospirone, a partial agonist of HTR1A, could ameliorate psychopathology, including negative and cognitive symptoms in patients with SCZ [25,26,27,28,29].
HTR1A is encoded at 5q11.2-13 and is made of 422 amino acids. rs6295, also known as C(−1019)G, is the most widely investigated functional variant in HTR1A associated not only with SCZ [19], but also with mood disorders [30, 31], anxiety disorders, and suicide [32, 33]. This may be related to its location at the extended promoter region where it may impact HTR1A gene expression [34, 35]. rs6295, when cloned into HEK-293 cells, was shown to exhibit higher transcriptional activity [36]. Associations between positive symptoms and rs6295 were also reported by Drago et al. [37], Kishi et al. [19], and Takekita et al. [38]. Associations between cognitive function and rs6295 were reported by Wesnes et al. [39], and Takekita et al [38]. Recently, Takekita et al. conducted a meta-analysis of the association of improvement in positive and negative symptoms of rs6295 using 1281 patients with SCZ from ten studies [40]. This association between rs6295 and treatment response to APDs was found only for SCZ patients of EUR or east Asian ancestry [40].
Lurasidone is a major atypical APD with high affinity for HTR1A with PA action [41,42,43,44]. It has been shown to improve psychopathology in SCZ in three 6-week, double-blind, randomized, placebo-controlled, phase 3 clinical trials [45,46,47]. A nonhypothesis driven GWAS based on a meta-analysis of these three clinical trials identified SNPs from ion channels, one of which reached genome wide significance, and synaptic adhesion molecules as the top predictors of efficacy [48, 49]. Here, we conducted a candidate SNP approach focused on the probable importance of HTR1A in the action of lurasidone by selection of rs358532 (C/A) and/or rs6449693 (G/A), proxy SNPs of rs6295, which are in high linkage disequilibrium (LD) with rs6295(C/G), according to the 1000Genome project. Unlike rs6295, a transverse SNP, these are transition SNPs which are tagged in widely used microarray platforms such as Affymetrix 6.0 SNP array or Illumina Beadchip. We first evaluated whether rs358532 and rs6449693, predicted treatment response to lurasidone, based on two clinical trials [49] of acutely psychotic patients with European (EUR, n = 171) or African (AFR, n = 131) ancestry. We then determined if that association could be replicated in a third trial of lurasidone with a similar design. Finally, we conducted a meta-analysis of independent cohorts from several studies, including ours to determine whether rs6295 predicted treatment response to APDs which produce HTR1A agonism at clinically effective doses.
Material and methods
Subjects
The genomic DNA was collected from SCZ patients who participated in three 6-week randomized double-blind, placebo-controlled, multicenter registration trials of lurasidone [45,46,47]. All patients were acutely psychotic and diagnosed as SCZ by Structured Clinical Interview for DSM-IV. In the first two trials, the ethnicity of 171 (EUR) and 131 (AFR) patients was verified by principal component analysis (PCA) using 1000Genome sample as the reference [48]. One hundred (EUR) and 34 (AFR) patients from the third trial with a similar design was used as a replication dataset following ethnicity verification by PCA as well [49]. The demographic features of GWAS samples from three clinical trials are described in Table 1 and Supplementary Table 1. Treatment-resistant SCZ patients were identified by evaluation of history of treatment response to two or more trials of APDs and were excluded. Both trials showed significantly greater improvement than placebo at the end of 6 weeks in the lurasidone treated patients [45, 47]. Trial 3, efficacy of treatment of 100 SCZ patients with lurasidone (80 or160 mg/day) for 12 months was compared with that of quetiapine XR (QXR) (200–800 mg/day) in outpatients with acute exacerbation. The proportions of responders with improvement greater than 20% in PANSS Total were similar in all the clinical trials. All the participants were provided written informed consent. The trials were performed in accordance with the Helsinki Declaration of 1975 (as revised in 1983).
Evaluation of treatment response
The change (Δ) and % in PANSS Total were calculated each of the first 6 weeks with last observation carried forward (LOCF) in SCZ patients with European or African ancestry from Pearl 1, 2 and 3 trials of lurasidone and placebo. We also conducted the association test on Δ and % in PANSS five subscales; positive, negative, disorganization, excitement, anxiety and depression, based on the five-factor model of SCZ psychopathology [50], at week 1, 2, 4, and 6 from Pearl 1, 2 trials of lurasidone and placebo.
Genotyping
Details have been provided previously [47, 51]. DNA samples from the first two clinical trials were genotyped using Affymetrix SNP Array 6.0 (Affymetrix, Santa Clara, CA, USA) [45, 47]. DNA samples from the third trial were genotyped with Illumina Omni5Exome-4v1beadchip [49]. The probes for rs6295 were unavailable in both array platforms. Due to the confusion brought by genotyping transverse SNP with minor allele frequency (MAF) close to 0.5 like rs6295 (C/G, 0.445) in EUR, we selected two proxy SNPs, rs358532 (C/A) and rs6449693 (G/A), which are highly in LD with rs6295 (r2 = 0.95/D′ = 1 in EUR and r2 = 0.80/D′ = 0.90 in AFR for rs358532; r2 = 1/D′ = 1 in EUR and r2 = 0.24/D′ = 1 in AFR for rs6449693), respectively, according to the report from 1000Genome (Supplementary Table 2). Genotyping data for rs6449693 or rs358532 were used for Pearl 1, 2 or 3 samples, respectively. The minor/major alleles and meta-analysis of the results from other studies used samples from EUR only. The phased reference genome data (1000Genome) in EUR were obtained to confirm that the minor A allele in rs358532 is linked to rs6295 G allele. The genotype call-rates for both proxy SNPs were 100% without significant deviation from Hardy–Weinberg Equilibrium (p = 0.0001 as a cutoff). The MAF in discovery samples were very close to the reported value from 1000Genome.
Association testing
A linear regression based on a recessive model after adjusting for age, gender, and dosage was conducted by PLINK 1.9 [52] for the association testing. Baseline psychopathology showed no significant contribution to the full model and was not included in the final model. A threshold of significance was set as 0.05 for the p-value calculated from the association between the genotype and change in PANSS Total and 0.01 for the association with PANSS five subscales, for Bonferroni correction for the multiple testing.
Association analyses by mixed model repeated measures
Multilevel multiple imputation is considered as a better approach for behavioral science data due to its flexibility to tailor the missing data handling procedure to match a specific set of analysis goals (https://stefvanbuuren.name/fimd/). Here we integrate R mice and lmer packages by conducting imputation, analysis by mixed-effects model, and pooling of the result by ‘Rubin’s rules’, to determine the association of the candidate genetic variant, rs6449693, and dynamic change of PANSS total (TOT) or five domains for up to 6 weeks. The imputation method (meth) was a two-level normal model, ‘2l.norm’, which could recover the intra-class correlation quite well, even for severe missing at random cases and high amounts of missing data in the outcome or the predictor [53]. We specified all predictors (rs6449693, PC1, PC2, PC3, gender, dose, time, baseline) as random effects. Patient ID (FID) was the class variable for ‘2l.norm’. The next step was to fit a linear model, ‘PANSS ~ rs6449693(recessive) + PC1 + PC2 + PC3 + gender + dose + (1+1|time)’ to each imputed dataset, and then to pool the results together. Time was a grouping factor and ‘1 + 1|time’ was random intercept with fixed mean. The rest of other variables in this model were fixed factors. According to the margin plot of wk2 versus wk6 data (e.g., PANSS Total) as drawn by the margin plot function in R VIM package, There were different distributions of missing and observed data from Pearl 3 but not Pearl 1 & 2 (Supplementary Fig. 1), suggesting that the missing for repeated measures in Pearl 3 data was not completely random. We only apply this two-level model to imputation of missing data from Pearl 2 trial.
Meta-analysis
AY and JL conducted a literature search independently using PubMed, Web of Science, and Google scholar to identify articles published in English until July 10, 2018. Combinations of the following MeSH terms: (‘Receptor, Serotonin, 5-HT1A’[MeSH] AND ‘Antipsychotic Agents’[MeSH] AND ‘Treatment Outcome’[MeSH] AND ‘Genetic Association Studies’[MeSH]) were used as search queries. Finally, we manually checked related articles published in English. Any disagreement was resolved before the meta-analysis.
The following inclusion criteria were used to select studies: (1) investigation of the association between HTR1A functional polymorphisms, rs6295 or its proxy SNPs, or both and improvement after treatment with APDs; (2) the majority of patients had a diagnosis of SCZ based on DSM or ICD criteria (also including first-episode patients); (3) treatment response was evaluated using a standardized rating scale, such as PANSS at baseline and endpoint; (4) genotype distribution followed Hardy–Weinberg equilibrium; (5) the study was performed by the candidate SNPs approach; (6) the study was published in English in a peer-reviewed journal; (7) the study was based on an independent sample collection (nonoverlapping with each other); (8) all the patients were European origin. Based on these inclusion criteria, we selected three studies with five independent cohorts (N = 523) [54, 55]. Among them, two studies genotyped rs6295 directly [53, 54].
Meta-analysis was performed by R ‘meta’ package. Since clinical heterogeneity among the populations and treatments was expected, a random-effects model was used to analyze the data [56, 57]. The standardized mean difference between genotypes was collected from the original articles and its 95% confidence interval and standard deviation were either calculated or imputed using a correlation coefficient (0.80, this value was calculated by other studies) [58]. Study heterogeneity was measured using the chi-square and I-square statistics, with chi-square values of p < 0.05 and I2 values of >50% indicating relevant heterogeneity. We only performed a meta-analysis separated by PANSS subscales, positive and negative symptoms since the original data from these two domains were the only ones available.
Results
HTR1A rs6295 or its related SNPs was associated with overall symptoms improvement only in SCZ patients with European ancestry
In EUR patients, the average of overall symptom improvement (Δ or % change in PANSS Total) or improvement in each subscale (Δ or % change in positive or negative symptoms) at LOCF or MMRM 6 weeks showed no significant difference between patients with AC and CC genotypes of rs358532 or between patients with GA and GG genotypes of rs6449693 (p > 0.10) (Table 2). Therefore, we conducted a recessive model of minor allelic association with phenotypes in subsequent analyses. This is also in accord with previous studies of association between rs6295 and treatment response to APDs [40].
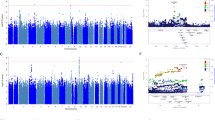
In Fig. 1, overall symptoms improvement (ΔPANSS Total) after 6 weeks of treatment were significantly predicted by rs6449693 or rs358532, proxy SNPs of rs6295 in Pearl 1,2 studies (β = 6.107, p = 0.046) for both SNPs as they are in complete LD with each other. This significant association was validated for rs358532, a proxy SNP of rs6295 in Pearl 3 study (β = 10.35, p = 0.036). In contrast, rs6295 proxy SNPs did not predict overall improvement in AFR patients (all p < 0.05, Supplementary Table 1) or patients treated with placebo (all p < 0.05, Supplementary Table 1), suggesting that this association was race-dependent and specific to the treated group. We then examined whether rs6295 proxy SNPs predicted improvement in specific domains of psychopathology, positive, negative, disorganization, excitement, and anxiety/depression, separately in EUR patients of Pearl 1,2 trials. There was a significant association between rs353832/rs6449693 and treatment response to lurasidone in positive symptoms (Δ positive, β = 1.801, p = 0.041 for Pearl 1,2), disorganization (Δ disorganization, β = 1.628; p = 0.031 for Pearl 1,2 and β = 2.754, p = 0.048 for Pearl 3) and a trend in that direction in positive symptoms (Δ positive, β = 2.003; p = 0.094 for Pearl 3) were identified in both datasets, but none of the p-value survived Bonferroni correction for multiple testing of individual domains. Similar results were found for improvement in negative symptom was observed in Pearl 1,2 (β = 1.928; p = 0.02) but not Pearl 3 (β = 1.696; p = 0.208).
Boxplots of association between rs6295, tagged by rs6449693 (Pearl 1, 2) or rs358532 (Pearl 3), and Δ change in PANSS Total and five subscales in SCZ patients with European ancestry after 6 weeks of treatment with lurasidone in Pearl 1, 2, and 3 trials. The Δ change in PANSS Total and five subscales was calculated with last observation carried forward (LOCF). Δ change = PANSSLOCF6wk – PANSSBaseline. Boxplots were created by R ‘ggplot’ package
Alternatively, we conducted imputation, analysis with a mixed-effects model, and pooling of the results to determine the association of the candidate genetic variant, rs6449693, and the changes in PANSS Total and five domains for up to 6 weeks in the Pearl 1 and 2 dataset. According to Table 2B, rs6449693, a tag SNP of rs6295, was significantly associated with the change in PANSS Negative (original p = 0.005), which survived Bonferroni correction. The negative value for the estimated fixed effect of rs6449693 indicates that the minor allele carriers (recessive mode of inheritance) had a worse response to treatment with lurasidone compared with those homozygous for major alleles. A trend of association with the same direction for the minor allele was also observed for PANSS Total and the Positive, and Disorganization domains in the Pearl 1 & 2 dataset.
rs6295 (tagged by rs6449693 or rs358532) contributes to trajectory of early response to lurasidone
It was noted that a significant association for response was identified as early as week one after initiating lurasidone (β = 3.510; p = 0.035 for PANSS Total). The association between rs6449696/rs358532 and improvement in negative symptom was significant at week one (β = 1.153; p = 0.027). This association was more robust at week 2 (β = 1.726; p = 0.004), and week 4 (β = 2.303; p = 0.003), suggesting HTR1A PA is crucial for negative symptom improvement. Major allele carriers had 2.61 in Δ change (2.58% in % change), compared with minor allele carriers, with only 0.77 in Δ change (2.58% in % change) at week 2 in negative symptoms (β = 1.726; p = 0.004, Table 2). A trend of significant association between rs6449696/rs358532 and improvement in positive symptom was also observed at week two but not in week 1 and 4. All such associations were not identified in the corresponding placebo group (p < 0.10).
Meta-analyses confirmed the association in SCZ patients with European Ancestry
Meta-analysis was used to further test the association between functional HTR1A polymorphisms and treatment response to APDs in positive and negative symptoms in SCZ patients with European ancestry from independent cohorts (Supplementary Table 3).
SCZ patients (n = 523) with European ancestry from five independent cohorts were included in this meta-analysis [54, 55]. The results from using rs6295 or its proxy SNPs were included in this analysis. All these studies showed the distribution of genotypes followed Hardy–Weinberg equilibrium. A significant association was found between rs6295 and improvement in negative symptoms (five studies, SMD = 0.56, 95%CI = 0.33 to 0.80, p = < 0.0001, Fig. 2a) and positive symptoms (five studies, SMD = 0.33, 95% CI = 0.05–0.62, p = 0.023, Fig. 2b). Regarding heterogeneity, no significant heterogeneity was detected in any of the comparisons for rs6295 (p = 0.42 or 0.17, respectively, Fig. 2a, b)
Forest plots and summary statistics of the meta-analysis of five independent datasets to show the association between HTR1A functional polymorphism, rs6295, and improvement in PANSS negative symptom (a) and positive symptom (b) after treatment with antipsychotic drugs in SCZ patients with European ancestry. Forest plots were created by ‘R meta’ package
Discussion
To our knowledge, this is the first study to evaluate the association between the tag SNPs for the functional HTR1A SNP rs6295 and treatment response to lurasidone, in acutely psychotic patients with SCZ. This is of interest because extensive evidence supports that HTR1A partial agonism is a major component of the pharmacology of lurasidone, as it is for some other atypical and APDs [42, 43, 59]. The significant association between rs358532/rs6449693, tag SNPs for rs6295, and Δ or % change in PANSS Total (LOCF) was only identified in EUR (β = 6.107, p = 0.046) but not in AFR (β = −3.147, p = 0.399) or the matched placebo groups (β = 2.205, p = 0.739), suggesting that this association is race-dependent and specific to the treated group. The direction of the association between minor allele of HTR1A rs6295 and treatment response is consistent with the result from the latest meta-analysis which reported G allele carriers of rs6295 had less improvement in response to APDs [40]. We replicated this association by using an independent dataset from the third trial of lurasidone which had a similar design to the discovery sample (β = 10.35; p = 0.036). Our subsequent analyses, separated by individual domains of psychopathology, provided further evidence of predicting symptom improvement in a domain-dependent manner (Fig. 1), suggesting that HTR1A-mediated signaling is most important in the neural circuitries related to ‘negative’, ‘disorganization’, and ‘positive’ symptoms, the three most important clinical features of SCZ and related psychotic disorders.
In addition, we presented the first evidence that the extent of this association is related to duration of the treatment, with the most significant association observed as early as one week for Disorganization, and as early as two weeks for Negative symptoms. Finally, we conducted a meta-analysis of five independent cohorts of EUR patients and showed that this functional HTR1A polymorphism or the tag SNPs were significantly associated with treatment response to APDs in positive (p = 0.0023) and negative symptom (p < 0.0001) improvement (Fig. 2a, b).
Importantly, the significant association between rs6295 and response to APDs was not restricted to negative symptom improvement. This is consistent with other research [54, 55] including a study reporting an association between rs6295 and General psychopathology subscale and total PANSS score in 63 drug-naive SCZ patients [54]. Drago et al. also found a significant association between rs6295 and improvement in positive symptoms (p = 0.003) in 96 acutely psychotic patients treated with haloperidol [60].
Hypodopaminergia has been suggested to be causally related to negative symptom [61, 62]. The association between a functional SNP of HTR1A and improvement in negative symptoms could be based on the increased release of DA in PFC induced by stimulation of HTR1A [21]. An association between HTR1A and negative symptom is consistent with an imaging study that the G allele (major allele) carriers, which is associated with increased amygdala reactivity to emotional stimuli [35], have greater improvement in negative symptoms with atypical APD treatment. Considering that disorganization is a manifestation of dysconnectivity in the brain of SCZ, and that G-carriers (major allele) of rs6295, who are predicted to have lower expression of HTR1A in PFC, leads to poor functional connectivity in DLPFC [63], and are more likely to respond to HTR1A agonism and show better response in negative symptoms and disorganization. Another possibility is that HTR1A activation promotes neurogenesis in HIP [64] ameliorate may be different cognition which is the basis of improvement in other dimensions.
The C(−1019) allele carriers have been suggested to have increased presynaptic HTR1A auto-receptors, enhancing inhibitory input to PFC 5-HT neurons [65]. This could lead to disinhibition of parvalbumin-positive or somatostatin-positive GABAergic interneuron in the PFC, increasing release of DA in PFC and greater improvement in negative symptoms in SCZ (Fig. 3). Enhanced inhibition due to increased HTR1A auto-receptors in the dorsal raphe nucleus may also increase inhibition of dopaminergic neurons in ventral tegmental area (VTA), which would be expected to diminish psychosis (Fig. 3). In addition, it might affect the balance between the pre/post HTR1A ratio, stimulating postsynaptic HTR1As to increase DA release in PFC [66].
Schematic diagram illustrated the functional impact of HTR1A polymorphism, rs6295 on the clinical response to lurasidone. SCZ patients from European ancestry harboring C allele of rs358532, corresponding to the C allele in rs6295, showed better improvement in PANSS positive, negative, and disorganization symptoms than homozygous G allele counterparts. C allele carriers of rs6295 have been shown to express more HTR1A auto-receptors in dorsal raphe nucleus (DRN), resulting in the enhanced inhibition of serotonergic firing activity at PFC (Albert, 2012). Disinhibition of parvalbumin-positive (PV+) GABAergic interneuron in PFC leads to increased release of dopamine in PFC. This may lead to a better improvement in negative symptoms in SCZ. Enhanced inhibition due to the increased HTR1A auto-receptors in DRN may also cause an enhanced inhibition of dopaminergic neuron in VTA, resulting in better improvement in positive symptoms in SCZ. Ameliorating positive and negative symptoms further supports improvement in disorganization. Transcriptional factor(s) binding is important to regulate the gene expression of HTR1A. Using mfold, we simulated the 2nd structure of transcriptional factors binding site of the palindromic sequence formed by rs6295 G allele but absence in rs6295 C allele. Having C allele in HTR1A promoter region, the transcriptional suppressor has no palindrome structure to bind to, resulting in enhanced inhibition by increased expression of presynaptic HTR1A auto-receptors
We selected HTR7, one of a small number of neurotransmitter receptors for which lurasidone has high affinity, as a candidate gene to determine if its common variants are associated with clinical response to lurasidone. However, there was no significant association with annotated functional SNPs of the HTR7 and clinical response measures after correction for multiple testing (unpublished data). However, we previously demonstrated a significant inverse correlation between expression of HTR7 and the top genes identified by GWAS which predicted clinical response to lurasidone at 6 weeks; most of these encode synaptic adhesion and scaffolding proteins 4, suggesting that the HTR7-mediated signaling pathway may contribute to the mechanisms of action of lurasidone at the transcriptional level.
There are several limitations in our study. First, the small effect-size for rs6295 in the discovery sample, with nominally significant association to treatment response to lurasidone, in addition to the high MAF, increased the challenge to achieve replication in an independent dataset of smaller sample size. Nevertheless, we showed a trend for replication. Second, in our study, rs6295 was not actually genotyped but the tag SNPs were examined. Although rs6449693 might be better in terms of avoiding the confusion due to the C/G transverse SNP, the interpretation of the results needs caution in this perspective. Third, only five studies with the varied duration of treatment of APDs were included in the meta-analysis. It is debatable whether the conclusion made by the meta-analysis can be generalized to other APDs with HTR1A pharmacodynamic feature at a distinct duration of treatment. Fourth, caution is needed about generalizing the result of meta-analysis to other APDs because of the limited number of drugs in the first studies eligible for the meta-analysis. Also, the duration of treatment was different in the meta-analysis, ranging from 6 weeks to 3 months. Only five studies with nonuniform duration of treatment with APDs were included in the meta-analysis. It uncertain as to whether the conclusion from the meta-analysis can be generalized to other APDs with HTR1A pharmacodynamic feature with varying durations of treatment. Fifth, epigenetic factors may also affect treatment response. In Han Chinese inpatients presenting with their first psychotic episode, methylation of the specific CpG site adjacent to the functional polymorphism rs6295 was shown to be correlated with improvement in negative symptoms during treatment with APDs, including risperidone and clozapine [34]. Further study of genetic and epigenetic factors impacting the relation of rs6295 and clinical response is indicated.
In conclusion, we reported here that a common functional HTR1A polymorphism, rs6295, predicted response to lurasidone, which has a relatively high affinity for HTR1A. This study included the largest samples of Caucasian patients with SCZ to date and the first study for Africans. The results indicated that the tag SNPs for HTR1A functional polymorphism were significantly associated with improvement in PANSS positive, negative, and disorganization subscales in a race-specific and a time-dependent manner. We obtained a replication of the association in meta-analysis which included two treatment groups other than risperidone, but also involved patients of EUR ancestry. We further confirmed the association of rs6295 with improvement in positive and negative symptom in Caucasian patients with SCZ.
References
McClay JL, Adkins DE, Aberg K, Stroup S, Perkins DO, Vladimirov VI, et al. Genome-wide pharmacogenomic analysis of response to treatment with antipsychotics. Mol Psychiatry. 2011;16:76–85.
McClay JL, Adkins DE, Aberg K, Bukszar J, Khachane AN, Keefe RS, et al. Genome-wide pharmacogenomic study of neurocognition as an indicator of antipsychotic treatment response in schizophrenia. Neuropsychopharmacology. 2011;36:616–26.
Arranz MJ, de Leon J. Pharmacogenetics and pharmacogenomics of schizophrenia: a review of last decade of research. Mol Psychiatry. 2007;12:707–47.
Charlab R, Zhang L. Pharmacogenomics: historical perspective and current status. In: Pharmacogenomics. Springer; 2013. pp 3–22.
Taylor DL, Tiwari AK, Lieberman JA, Potkin SG, Meltzer HY, Knight J, et al. Genetic association analysis of N-methyl-d-aspartate receptor subunit gene GRIN2B and clinical response to clozapine. Hum Psychopharmacol: Clin Exp. 2016;31:121–34.
Meltzer HY. Update on typical and atypical antipsychotic drugs. Annu Rev Med. 2013;64:393–406.
Meltzer HY. Clinical studies on the mechanism of action of clozapine: the dopamine-serotonin hypothesis of schizophrenia. Psychopharmacology. 1989;99:S18–S27.
Meltzer HY, Li Z, Kaneda Y, Ichikawa J. Serotonin receptors: their key role in drugs to treat schizophrenia. Prog Neuro-Psychopharmacol Biol Psychiatry. 2003;27:1159–72.
Kapur S, Zipursky R, Jones C, Shammi C, Remington G, Seeman P. A positron emission tomography study of quetiapine in schizophrenia: a preliminary finding of an antipsychotic effect with only transiently high dopamine D2 receptor occupancy. Arch Gen Psychiatry. 2000;57:553–9.
Meltzer HY. The mechanism of action of novel antipsychotic drugs. Schizophr Bull. 1991;17:263.
Huang M, Li Z, Ichikawa J, Dai J, Meltzer HY. Effects of divalproex and atypical antipsychotic drugs on dopamine and acetylcholine efflux in rat hippocampus and prefrontal cortex. Brain Res. 2006;1099:44–55.
Need AC, Keefe RS, Ge D, Grossman I, Dickson S, McEvoy JP, et al. Pharmacogenetics of antipsychotic response in the CATIE trial: a candidate gene analysis. Eur J Hum Genet. 2009;17:946–57.
Ikeda M, Yamanouchi Y, Kinoshita Y, Kitajima T, Yoshimura R, Hashimoto S, et al. Variants of dopamine and serotonin candidate genes as predictors of response to risperidone treatment in first-episode schizophrenia. 2008.
Hwang R, Shinkai T, De Luca V, Müller DJ, Ni X, Macciardi F, et al. Association study of 12 polymorphisms spanning the dopamine D2 receptor gene and clozapine treatment response in two treatment refractory/intolerant populations. Psychopharmacology. 2005;181:179–87.
Zhang J-P, Lencz T, Malhotra AK D2 receptor genetic variation and clinical response to antipsychotic drug treatment: a meta-analysis. Am J Psychiatry. 2010;167:763–72.
Lencz T, Robinson DG, Xu K, Ekholm J, Sevy S, Gunduz-Bruce H, et al. DRD2 promoter region variation as a predictor of sustained response to antipsychotic medication in first-episode schizophrenia patients. Am J Psychiatry. 2006;163:529–31.
Kim Y-K, Yoon H-K. Effect of serotonin-related gene polymorphisms on pathogenesis and treatment response in Korean schizophrenic patients. Behav Genet. 2011;41:709–15.
Takekita Y, Fabbri C, Kato M, Nonen S, Sakai S, Sunada N, et al. Serotonin 7 receptor variants are not associated with response to second-generation antipsychotics in Japanese schizophrenia patients. Neuropsychobiology. 2015;72:118–25.
Kishi T, Okochi T, Tsunoka T, Okumura T, Kitajima T, Kawashima K, et al. Serotonin 1A receptor gene, schizophrenia and bipolar disorder: an association study and meta-analysis. Psychiatry Res. 2011;185:20–26.
Pazos A, Palacios J. Quantitative autoradiographic mapping of serotonin receptors in the rat brain. I. Serotonin-1 receptors. Brain Res. 1985;346:205–30.
Ichikawa J, Ishii H, Bonaccorso S, Fowler WL, O'laughlin IA, Meltzer HY. 5‐HT2A and D2 receptor blockade increases cortical DA release via 5‐HT1A receptor activation: a possible mechanism of atypical antipsychotic‐induced cortical dopamine release. J Neurochem. 2001;76:1521–31.
Díaz-Mataix L, Scorza MC, Bortolozzi A, Toth M, Celada P, Artigas F. Involvement of 5-HT1A receptors in prefrontal cortex in the modulation of dopaminergic activity: role in atypical antipsychotic action. J Neurosci. 2005;25:10831–43.
Sumiyoshi T, Stockmeier CA, Overholser JC, Dilley GE, Meltzer HY. Serotonin 1A receptors are increased in postmortem prefrontal cortex in schizophrenia. Brain Res. 1996;708:209–14.
Yasuno F, Suhara T, Ichimiya T, Takano A, Ando T, Okubo Y. Decreased 5-HT 1A receptor binding in amygdala of schizophrenia. Biol Psychiatry. 2004;55:439–44.
Sumiyoshi T, Matsui M, Nohara S, Yamashita I, Kurachi M, Sumiyoshi C, et al. Enhancement of cognitive performance in schizophrenia by addition of tandospirone to neuroleptic treatment. Am J Psychiatry. 2001;158:1722–5.
Sumiyoshi T, Matsui M, Yamashita I, Nohara S, Kurachi M, Uehara T, et al. The effect of tandospirone, a serotonin 1A agonist, on memory function in schizophrenia. Biol Psychiatry. 2001;49:861–8.
Sumiyoshi T, Matsui M, Yamashita I, Nohara S, Uehara T, Kurachi M, et al. Effect of adjunctive treatment with serotonin-1A agonist tandospirone on memory functions in schizophrenia. J Clin Psychopharmacol. 2000;20:386–8.
Meltzer HY, Sumiyoshi T. Does stimulation of 5-HT 1A receptors improve cognition in schizophrenia? Behav Brain Res. 2008;195:98–102.
Sumiyoshi T, Meltzer HY. Serotonin 1A receptors in memory function. Am J Psychiatry. 2004;161:1505–1505.
Serretti A, Artioli P, Lorenzi C, Pirovano A, Tubazio V, Zanardi R. The C (−1019) G polymorphism of the 5-HT1A gene promoter and antidepressant response in mood disorders: preliminary findings. Int J Neuropsychopharmacol. 2004;7:453–60.
Kato M, Fukuda T, Wakeno M, Okugawa G, Takekita Y, Watanabe S, et al. Effect of 5‐HT1A gene polymorphisms on antidepressant response in major depressive disorder. Am J Med Genet Part B: Neuropsychiatr Genet. 2009;150:115–23.
Wasserman D, Geijer T, Sokolowski M, Rozanov V, Wasserman J. The serotonin 1A receptor C (-1019) G polymorphism in relation to suicide attempt. Behav Brain Funct. 2006;2:14.
Wang L, Chen L, Ma J, Qiao Z, Qiu X, Yang X, et al. Polymorphisms in 5-HTR1A and coupled G-proteins in association with negative life events increase susceptibility to suicide attempt. Int J Clin Exp Pathol. 2017;10:6189–97.
Tang H, Dalton CF, Srisawat U, Zhang ZJ, Reynolds GP. Methylation at a transcription factor-binding site on the 5-HT1A receptor gene correlates with negative symptom treatment response in first episode schizophrenia. Int J Neuropsychopharmacol. 2014;17:645–9.
Le François B, Czesak M, Steubl D, Albert PR. Transcriptional regulation at a HTR1A polymorphism associated with mental illness. Neuropharmacology. 2008;55:977–85.
Wu X, Yao J, Ding M, Shi Z-s, Xu F-l, Zhang J-j, et al. 5-HT1A receptor (HTR1A) 5′ region haplotypes significantly affect protein expression in vitro. Neurosci Lett. 2017;638:51–54.
Drago A, De Ronchi D, Serretti A. 5-HT1A gene variants and psychiatric disorders: a review of current literature and selection of SNPs for future studies. Int J Neuropsychopharmacol. 2008;11:701–21.
Takekita Y, Fabbri C, Kato M, Nonen S, Sakai S, Sunada N, et al. HTR1A gene polymorphisms and 5-HT1A receptor partial agonist antipsychotics efficacy in schizophrenia. J Clin Psychopharmacol. 2015;35:220–7.
Wesnes KA, Hopkins SC, Brooker HJ, Koblan KS. Differences in memory function between 5-HT1A receptor genotypes in patients with major depressive disorder. CNS Spectr. 2016;21:379–84.
Takekita Y, Fabbri C, Kato M, Koshikawa Y, Tajika A, Kinoshita T, et al. HTR1A polymorphisms and clinical efficacy of antipsychotic drug treatment in schizophrenia: a meta-analysis. Int J Neuropsychopharmacol. 2016;19:1–10.
Sanford M. Lurasidone. CNS Drugs. 2013;27:67–80.
Ishibashi T, Horisawa T, Tokuda K, Ishiyama T, Ogasa M, Tagashira R, et al. Pharmacological profile of lurasidone, a novel antipsychotic agent with potent 5-hydroxytryptamine 7 (5-HT7) and 5-HT1A receptor activity. J Pharmacol Exp Therapeutics. 2010;334:171–81.
Huang M, Horiguchi M, Felix AR, Meltzer HY. 5-HT1A and 5-HT7 receptors contribute to lurasidone-induced dopamine efflux. Neuroreport. 2012;23:436–40.
Rajagopal L, Massey BW, Michael E, Meltzer HY. Serotonin (5-HT) 1A receptor agonism and 5-HT7 receptor antagonism ameliorate the subchronic phencyclidine-induced deficit in executive functioning in mice. Psychopharmacology. 2016;233:649–60.
Nasrallah HA, Silva R, Phillips D, Cucchiaro J, Hsu J, Xu J, et al. Lurasidone for the treatment of acutely psychotic patients with schizophrenia: a 6-week, randomized, placebo-controlled study. J Psychiatr Res. 2013;47:670–7.
Loebel A, Cucchiaro J, Xu J, Sarma K, Pikalov A, Kane JM. Effectiveness of lurasidone vs. quetiapine XR for relapse prevention in schizophrenia: a 12-month, double-blind, noninferiority study. Schizophr Res. 2013;147:95–102.
Meltzer HY, Cucchiaro J, Silva R, Ogasa M, Phillips D, Xu J, et al. Lurasidone in the treatment of schizophrenia: a randomized, double-blind, placebo-and olanzapine-controlled study. Am J Psychiatry. 2011;168:957–67.
Li J, Yoshikawa A, Brennan MD, Ramsey TL, Meltzer HY Genetic predictors of antipsychotic response to lurasidone identified in a genome wide association study and by schizophrenia risk genes. Schizophr. Res. 2017.
Li J, Loebel A, Meltzer HY Identifying the genetic risk factors for treatment response to lurasidone by genome-wide association study: a meta-analysis of samples from three independent clinical trials. Schizophr Res. 2018;199:203–13.
Levine SZ, Rabinowitz J. Revisiting the 5 dimensions of the Positive and Negative Syndrome Scale. J Clin Psychopharmacol. 2007;27:431–6.
Loebel A, Cucchiaro J, Sarma K, Xu L, Hsu C, Kalali AH, et al. Efficacy and safety of lurasidone 80mg/day and 160mg/day in the treatment of schizophrenia: a randomized, double-blind, placebo-and active-controlled trial. Schizophr Res. 2013;145:101–9.
Purcell S, Chang C. PLINK 1.9. 2015. www.cog-genomicsorg/plink2
Groothuis-Oudshoorn K, Van Buuren S. Mice: multivariate imputation by chained equations in R. J Stat Softw. 2011;45:1–67.
Reynolds GP, Arranz B, Templeman LA, Fertuzinhos S, San L. Effect of 5-HT 1A receptor gene polymorphism on negative and depressive symptom response to antipsychotic treatment of drug-naive psychotic patients. Am J Psychiatry. 2006;163:1826–9.
Mössner R, Schuhmacher A, Kühn K-U, Cvetanovska G, Rujescu D, Zill P, et al. Functional serotonin 1A receptor variant influences treatment response to atypical antipsychotics in schizophrenia. Pharmacogenet Genom. 2009;19:91–94.
DerSimonian R, Laird N. Meta-analysis in clinical trials. Control Clin Trials. 1986;7:177–88.
DerSimonian R, Kacker R. Random-effects model for meta-analysis of clinical trials: an update. Contemp Clin Trials. 2007;28:105–14.
Higgins J. Cochrane handbook for systematic reviews of interventions. Version 5.1. 0. 2011. The Cochrane Collaboration. www.cochrane-handbook.org
Huang M, Kwon S, Rajagopal L, He W, Meltzer HY. 5-HT 1A parital agonism and 5-HT 7 antagonism restore episodic memory in subchronic phencyclidine-treated mice: role of brain glutamate, dopamine, acetylcholine and GABA. Psychopharmacology. 2018;235:2795–808.
Drago A, Giegling I, Schäfer M, Hartmann AM, Friedl M, Konte B, et al. AKAP13, CACNA1, GRIK4 and GRIA1 genetic variations may be associated with haloperidol efficacy during acute treatment. Eur Neuropsychopharmacol. 2013;23:887–94.
Davis KL, Kahn RS, Ko G, Davidson M. Dopamine in schizophrenia: a review and reconceptualization. Am J Psychiatry. 1991;148:1474–86.
Meltzer HY, Huang M. In vivo actions of atypical antipsychotic drug on serotonergic and dopaminergic systems. Prog Brain Res. 2008;172:177–97.
Zheng H, Onoda K, Wada Y, Mitaki S, Nabika T, Yamaguchi S. Serotonin-1A receptor C-1019G polymorphism affects brain functional networks. Sci Rep. 2017;7:12536.
Schreiber R, Newman-Tancredi A. Improving cognition in schizophrenia with antipsychotics that elicit neurogenesis through 5-HT 1A receptor activation. Neurobiol Learn Mem. 2014;110:72–80.
Albert PR. Transcriptional regulation of the 5-HT1A receptor: implications for mental illness. Philos Trans R Soc B. 2012;367:2402–15.
Sakaue M, Somboonthum P, Nishihara B, Koyama Y, Hashimoto H, Baba A, et al. Postsynaptic 5-hydroxytryptamine1A receptor activation increases in vivo dopamine release in rat prefrontal cortex. Br J Pharmacol. 2000;129:1028–34.
Acknowledgements
The authors would like to thank all the patients, healthy volunteers, and psychiatrists who intensively cooperated with us to recruit patients in this study. This work was supported by a grant awarded to HYM by Sunovion.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
HYM is a stockholder in ACADIA. HYM receives grant support from ACADIA, Allergan, Aptinyx, Eli Lilly, Janssen, Lundbeck, Neurocrine, Sumitomo Dainippon, and Sunovion. Other authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yoshikawa, A., Li, J. & Meltzer, H.Y. A functional HTR1A polymorphism, rs6295, predicts short-term response to lurasidone: confirmation with meta-analysis of other antipsychotic drugs. Pharmacogenomics J 20, 260–270 (2020). https://doi.org/10.1038/s41397-019-0101-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41397-019-0101-5
- Springer Nature Limited