Abstract
Multiple pharmacogenetic studies investigated the effectiveness of methotrexate. However, due to the use of nonvalidated outcomes, lack of validation or conflicting results it remains unclear if genetic markers can help to predict response to MTX treatment. Therefore, a systematic review was performed. PubMed was searched for articles reporting potential pharmacogenetic biomarkers associated (p < 0.05) with MTX efficacy using the validated endpoints DAS(28), EULAR, or ACR response criteria. The PICO method was used for study selection, and PRISMA guidelines to prepare the report. Thirty-five studies met the inclusion criteria, providing 39 potential genetic biomarkers in 19 genes. After Bonferroni correction, six genetic biomarkers were associated with the efficacy of MTX: ATIC rs7563206; SLC19A1 rs1051266; DHFR rs836788; TYMS rs2244500, rs2847153, and rs3786362 in at least one study. Only SLC19A1 rs1051266 was replicated in an independent cohort and promising for predicting methotrexate efficacy.
Similar content being viewed by others
Introduction
Low-dose methotrexate (MTX) is considered the “anchor drug” for the treatment of rheumatoid arthritis (RA). The precise mechanism of action of MTX remains to be elucidated, but it is known that MTX is transported over the membrane by multiple solute carriers (SLC) and that intracellular MTX has to be bound to polyglutames molecules by folylpolyglutamate synthase (FPGS) to exert its function. As illustrated in Fig. 1, the polyglutamated MTX affects multiple cellular pathways, e.g., adenosine, de novo purine synthesis, folate, methionine, and de novo pyrimidine synthesis.
Intracellular MTX mechanism pathway, divided into the methionine, folate, de novo pyrimidine synthesis, de novo purine synthesis, and adenosine pathway. 10-CHO-THF methylenetetrahydrofolate dehydrogenase, 5,10-CH-THF methyltetrahydrofolate cyclohydrolase, 5–10-MTHF formyltetrahydrofolate synthetase, 5-MTHF l-methylfolate, ABC ATP-binding cassette transporters, ADA adenosine deaminase, ADORA2A adenosine A2A receptor, AICAR 5-aminoimidazole-4-carboxamide ribonucleotide, AMP adenosine monophosphate, AMPD1 adenosine monophosphate deaminase 1 ATIC 5-aminoimidazole-4-carboxamide ribonucleotide formyltransferase, ATP adenosine triphosphate, cAMP cyclic adenosine monophosphate, CD37 transmembrane protein, CD39 transmembrane protein, DHF dihydrofolate, DHFR dihydrofolate reductase, DTMP deoxythymidine monophosphate, DTTP deoxythymidine triphosphate, DUMP deoxyuridine monophosphate FAICAR 5-formylaminoimidazole-4-carboxamide ribonucleotide, FPGS folylpoly-γ-glutaminase synthetase, GGH γ-glutamyl hydrolase, IL-10 interleukin-10, IMP inosine monophosphate, IPTA inosine triphosphatase, MS methionine synthase, MTHFD1 methylenetetrahydrofolate dehydrogenase 1, MTHFR methylenetetrahydrofolate reductase, MTRR methionine synthase reductase, MTX methotrexate, MTXPG methotrexate polyglutamate, NT nucleoside transporter, SHMT-1 serine hydroxymethyltransferase 1, SLC solute carrier, THF tetrahydrofolate, TYMS thymidylate synthase
In particular, an essential function of the folate pathway is to provide cofactors for key enzymes, such as dihydrofolate reductase (DHFR) that converts dihydrofolate into the folic acid derivative tetrahydrofolate (THF). THF and other derivatives are required for the purine and pyrimidine synthesis, which are important for cell proliferation and cell growth [1]. The methionine pathway is responsible for the synthesis of adenosine, which is an anti-inflammatory agent, alterated by methionine synthase and methionine synthase reductase (MTRR). Further, methionine is a precursor for S-adenosyl-methionine, which is a methyl donor that serves a variety of cellular functions, including DNA methylation [2]. The ubiquitin pathway is not directly related to the other pathways, but has an essential function in homeostasis and recognition of MHC class 1 for the cytotoxic T cells [3].
Approximately one-third of RA patients experience insufficient clinical response to MTX. Pharmacogenetics studies the impact of genetic variation to drug response and genetic variants in the MTX pathways described above may affect the potential effects of methotrexate on inflammation in RA. Indeed, multiple studies reported associations between single nucleotide polymorphisms (SNPs) and the efficacy of MTX. However, to date, none of the proposed markers are applied in clinical practice due to lack of validation or conflicting results. In addition, previous systematic reviews [4,5,6,7,8,9,10] described the effect of SNPs on the efficacy of MTX, but some included studies with MTX in different diseases such as juvenile idiopathic arthritis [10] or leukemia [5] or applied nonvalidated endpoints, such as red blood cell MTX polyglutamate concentrations [5, 11] or physicians’ assessment of patient’s response [9].
The goal of this review is to systematically explore which SNPs related to MTX pharmacology are associated with efficacy in RA by selecting only studies with the validated endpoints DAS(28), European League Against Rheumatism (EULAR), or American College of Rheumatology (ACR) response criteria [12, 13].
Methods
Data extraction and identification of eligible studies
Identification and selection of studies were performed according to the PICO method [14]. Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines were used to prepare the report [15]. PubMed was used to identify and extract all relevant articles published between April 2002 and March 2017. Search terms consisted of rheumatoid arthritis, methotrexate, pharmacogenetics, and SNP. The full search string is provided in Supplementary File S1. Also, we manually checked reference lists from reviews to identify relevant cross-references.
Records were screened on title and abstract. Comments, editorials, narrative reviews, letters (without original data), abstracts, and publications in languages other than English were excluded. Only studies utilizing the DAS(28), the response criteria of the ACR or the EULAR were eligible for inclusion. Included SNPs were analyzed under the additive, allelic, genotypic or haploid genetic model, and had at least one association with either DAS(28), ACR or EULAR response (p < 0.05, uncorrected for multiple testing).
SNPs were divided into MTX-related pathways: adenosine, de novo purine synthesis, transporters, polyglutamation, folate, methionine, de novo pyrimidine synthesis, and ubiquitin. Results from included studies were summarized, and reported odds ratio (OR) with 95% confidence interval (CI), p-value, type of association and SNP ID were collected. Finally, SNPs were checked on linkage disequilibrium by SNP Annotation and Proxy Search (SNAP, Broad Institute) [16], with the LD threshold of R2 > 0.8.
To control the risk of false positive findings, Bonferroni correction was applied when no correction for multiple testing was performed in the original study by calculating a significant cut-off p-value at α/n (p = 0.05 divided by the number of tested SNPs within each study). SNPs were significantly associated if the p-value was <0.05 after Bonferroni correction. Ultimately meta-analyses were used to support our findings of potential significant SNPs.
Results
Study selection
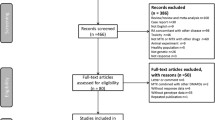
Figure 2 shows the results of the study selection. Initially, 115 publications were identified. We excluded 30 comments, editorials, letters, narrative reviews, and seven non-English written publications. Of the remaining 78 studies, 41 were excluded because none of our defined endpoints was reported and one because the report of the study could not be obtained. By cross-references, three more studies were included. In total, 35 original studies were available for analysis in this systematic review and seven meta-analyses were used to support our findings.
Study flow diagram of the systematic review inclusion [15]. MTX methotrexate, MAF minimum allele frequency, ACR American College of Rheumatology, DAS Disease Activity Score, EULAR European League Against Rheumatism
Study characteristics
Most studies (34 out of 35) were candidate gene studies investigating 1–35 polymorphisms. There was one genome-wide association study (GWAS) investigating 559,007 polymorphisms [17]. The mean study population of the studies was 197 patients (ranging from 48 to 422 patients). Most studies used the EULAR good response criteria (32%), tested <10 SNPs (76%), were conducted in Europe with RA patients of (self-)reported Caucasian origin. The average rate of good EULAR response to MTX monotherapy at t = 6 months was 55%, ranging from 23 [18] to 85% [19].
The included studies reported 39 SNPs in 20 genes associated with either DAS(28), EULAR, or ACR response with a p-value < 0.05. After Bonferroni correction, 16 SNPs in 10 genes remained significantly associated with MTX efficacy.
Adenosine pathway—ADA, ADORA2A, AMPD1, and ITPA
AMPD1 rs17602729 (allelic T) showed a significant association with DAS28 ≤ 3.2 (OR: 6.73, 95% CI: 1.74–26.01) between t = 3 and 6 months [20]. However, this was not confirmed with the genotypic CC model at t = 6 months [21]. None of the other SNPs in the adenosine pathways—ADA (rs244076), ADORA2A (rs5751876), and ITPA (rs1127354)—were significantly associated with the MTX response at t = 6 months using allelic or genotypic genetic models.
De novo purine synthesis—ATIC
Four SNPs in ATIC (rs2372536 [22], rs4673993 [23], rs7563206 [1], and rs12995526 [1]) had at least one study reporting a significant association with MTX efficacy. ATIC rs7563206 (allelic T carrier) was tested in one study, and showed an association with MTX nonresponse with the endpoint DAS28 ≤ 3.2 at t = 6 months (OR: 0.20 95% CI:0.09–0.46) [1]. At t = 6 months, ATIC rs4673993 (genotypic TT) showed a significant association with a better response (DAS28 ≤ 3.2, OR:3.86 95% CI:1.50–9.91), while rs12995526 (allelic T carriers) showed a significant association with a worse response (DAS28 ≤ 3.2, OR:0.23 95% CI:0.10–0.53) to MTX [23].
ATIC (rs2372536, genotype CC) was significantly associated with DAS ≤ 2.4 at t = 6 months, with an OR of 2.5 (95% CI: 1.3–4.8) [22]. Three other studies—using ATIC rs2372536 genotypic CC at t = 6 months—reported no significant association, of which one study reported that the CC genotype was related to MTX nonresponse with a OR below 1.0 (OR:0.27, 95% CI: 0.08–0.92) [1, 20, 24].
Transporters—ABCB1C1, ABCC1, SLC19A1 (RFC1), and SLC22A11
None of the SNPs in ABCB1 (rs1045642), ABCC1 (rs246240 and rs3784864), and SLC22A11 (rs11231809) were significantly associated with DAS28 ≤ 3.2 or EULAR good response at t = 6 months. The most studied genetic SLC19A1 SNP was rs1051266, which was investigated in 11 studies. Three studies reported a significant association with MTX efficacy at t = 6 months using ACR20 or DAS28 and different genetic models (either allelic A carriers, genotypic GG or genotypic AA). Other studies did not investigate the same genetic models, using the same efficacy endpoints with the same time evaluation point for SLC19A1 rs1051266.
Polyglutamation—FPGS and GGH
FPGS rs4451422 (allelic C carriers) was associated with MTX efficacy using EULAR good response at t = 6 months, with an OR of 0.73 (0.54–0.98) [17]. FPGS SNPs (rs1544105, rs10106, and rs10987742) and GGH SNPS (rs2305558 and rs1800909) were not significantly associated with MTX efficacy.
Folate pathway—DHFR, MTHFR, and SHMT
Both MTHFR rs1801131 (A1298C) and rs1801133 (C677T) have frequently been studied (>10 studies). One study showed a significant association with MTHFR rs1801133 CC genotype with DAS28 ≤ 3.2 at t = 6 months, with an OR of 3.4 [25]. Three other studies investigated the association of MTHFR genotypic CC at t = 6 months, and did not find an association using other endpoints (EULAR GR, ∆DAS44 < 0.6, and ACR20) [26,27,28]. For two other SNPs in MTHFR (rs17421511 and rs1476413) there was no significant association with MTX response. Also, no association was found between MTHFD1 rs17850560 or SHMT-1 rs1979277 with MTX response using DAS28(≤3.2) or EULAR GR. DHFR rs836788 was associated in one study with EULAR response at t = 6 months, with an OR of 1.44 (95% CI: 1.09–1.93) and 1.47 (95% CI: 1.09–1.96), respectively for the allelic A carriers and the genotypic AA [17].
Methionine pathway—MTR and MTRR
Six studies investigated the role of the MTR A2756G (rs1805087), of which one study reported a significant association [19]. Here, MTR rs1805087 was associated with MTX efficacy at t = 12 month, and the use of the endpoint EULAR good response with the genotypic AA (OR was not available). Other studies could not confirm the association with rs1805087, using the DAS28 with genotypic AA on t = 4 months [29], EULAR GR with the allelic G carriers on t = 4 months [30], or with the DAS28 ≤ 3.2 allelic G carriers on t = 6 months [31]. No significant association was reported with MTRR rs162040 and rs1801394.
De novo pyrimidine pathway—TYMS
TYMS rs2244500, rs2847153, and rs3786362 were all significantly associated with EULAR good response at t = 6 months and had OR of resp. 1.48. 1.92, 0.51, and 2.76 [17, 21]. No other studies investigated the effect of TYMS with MTX response.
Ubiquitin pathway—CUL1
Negi et al. investigated the association of CUL1 haplotypes with MTX efficacy using the DAS28 ≤ 3.2 at t = 6 months [32]. Here, CUL1 rs122571 haplotype A-T-T (OR: 2.83, 95% CI: 1.33–6.04) and rs243480 haplotype G-T-T (OR: 0.16, 95% CI 0.04–0.67) were significant.
KIR gene
One study tested multiple length variants of the KIR gene and showed that the full-length KIR2DS4 gene was significantly associated with DAS28 ≤2.5 (OR: 0.4344, 95% CI: 0.215, 0.987) at t = 6 months [33]. Here, possessing the KIRSDS4 gene had a lower chance of responding to MTX treatment.
Most promising genetic variants related to MTX efficacy
Table 1 lists the most promising SNPs that were significantly associated with MTX efficacy after Bonferroni correction without having conflicting results from other studies. For instance, it is ATIC rs467393 genotypic TT with better response, while allelic T carriers results in worse response or lacks validation.
The most promising SNPs were derived from the pathways de novo purine (ATIC), de novo pyrimidine (TYMS), and transporters (SLC19A1). The SNPs have a minor allele frequency > 0.2, except TYMS rs3786362 (MAF < 0.2 for all races). ATIC rs7563206 and TYMS rs2244500 were found significantly associated with an OR below 1.0, while the other eight SNPs had an OR between 1.42 and 2.83. The used genetic models were with either allelic, genotypic or haplotype. No linkage disequilibrium (R2 > 0.8) was observed for any of the SNPs in Table 2. SLC19A1 rs1051266 was tested in multiple studies and positively associated in three studies.
Of the six promising SNPs, ATIC rs7563206, TYMS rs2847153, and rs3786362 were associated with nonresponse to MTX, while SLC19A1 rs1051266, DHFR rs836788, and TYMS rs2244500 were associated with response to MTX. ORs range from 0.2 to 0.68 for MTX nonresponse and 1.42–2.76 for MTX response. The six SNPs had a MAF of >0.2 in all races except for TYMS rs3786362 which is sparse and even does not occurred in the European population.
Despite the findings of one significant association of ATIC rs473993 and rs12995526, AMPD1 rs17602729, MTHFR rs1801133, and MTR rs180508, and FPGS rs4451422, we did not mark those as promising genetic variants due to conflicting results. Also, we did not include the full-length KIR2DS4 gene as a promising genetic marker for the response to MTX, due to the complexity of the determination of the whole KIR2DS4 gene (with 15,894 bases) and the fact that it is not one SNP. This was also the case of CUL1 that was significantly associated with MTX response for two haplotypes; A-T-T (rs122571) and G-T-T (rs243480).
Discussion
This systematic review assesses the effect of genetic variation on the efficacy of MTX in RA using the validated endpoints DAS, EULAR, or ACR response criteria. After Bonferroni correction for multiple testing, we identified six genetic biomarkers related to MTX efficacy. Of these, SLC19A1 rs1051266 had the most convincing evidence with two independent studies showing significant associations. Other potentially promising SNPs are ATIC rs7563206, DHFR rs836788, TYMS rs2244500, rs2847153, and rs3786362, but these lack replication studies. The six genetic biomarkers could have clinical implications for the disease outcome of RA. In fact, SLC19A rs1051266, DHFR rs836788, and TYMS rs2244500 showed a 40% or more increased chance of the effectiveness of MTX, and ATIC rs7563206 and rs378636, and TYMS rs2847153 showed 45% or more chance of the reduced effectiveness of MTX. Still we believe that additional studies are necessary before implementing pharmacogenetic testing for these SNPs in the treatment of RA.
A limitation of the investigated studies in this systematic review is the difference in the evaluation time points for measuring MTX efficacy. MTX is a slow-acting prodrug that becomes active when polyglutamated in the cells. The process of polyglutamation is slow and takes up to 27.5 weeks (range 6.6–62.0 weeks) to reach steady state [34]. This delay in steady-state polyglutamation explains the relatively long time to clinical response, and therefore most studies had the endpoint set to 6 months after the start of MTX therapy. However, some studies evaluated response earlier than t = 6 months, while MTX may not yet have exerted its full potential. Furthermore, the genotypic or allelic genetic models were often used, when in fact the hypothesis-free driven additive genetic model seems more appropriate because the underlying genetic model is unknown.
Another limitation is that most studies tested with univariate analysis, without taking into account baseline variables (multivariate testing), such as gender, smoking status, disease severity which are known to influence response to MTX. Most drug-gene interaction studies were explorative, with the use of retrospective data and lack validation. Pharmacogenetic testing in RA remains limited mainly because the evidence for drug-gene interactions are marginal. MTX is involved in multiple pathways with different genes. Yet, most pharmacogenetic studies were candidate studies that tested only a single or a small number of SNPs, but not a combination of multiple genes or pathways [35]. To get clear evidence, additional studies with the use of a combination of multiple genes are needed. This review can show a basis, to test all suggestive SNPs together in association with the efficacy of MTX.
The strength of our study is that a systematic approach was used to identify SNPs and the selection of the articles was performed according to the PRISMA guidelines. Another strength is that only validated outcome criteria were used and that adjustment for multiple testing by Bonferroni correction was applied for the included studies. A potential weakness of this review is that only English publications were included. This results in the exclusion of seven non-English studies, and important findings could have been missed. Another weakness was the limited sample size of some studies and the lack of power analysis to check the validity of the outcomes.
Finally, a common limitation of systematic reviews is publication bias. Meaning that important—albeit negative—results were never published, which could lead to misinterpretation of the actual findings. Another limitation was that not all studies were performed with MTX monotherapy, and therefore the effect on response could be influenced by other DMARDs. Several meta-analyses have been performed on pharmacogenetics biomarkers for the efficacy or toxicity of MTX in RA. Of our promising SNPs, SLC19A1 rs1051266 with the genotypic AA (vs AG/AG) was tested in MTX efficacy in three meta-analyses. Two meta-analyses, conducted by Li et al. [50] and Chen et al. (2016), confirmed the significant association with an OR of 1.42 (95% CI: 1.04–1.93) and 1.49 (CI: 1.17–1.90), respectively. However, the third meta-analysis by Chen et al. (2016) showed substantial heterogeneity (I2) of 72% for the allelic model and thus represented inconsistencies of the pooled studies and affects the validity of the results. None of the other variants was evaluated in meta-analysis.
In summary, through the use of a systematic review and inclusion of studies with validated RA efficacy endpoints, we identified six SNPs for which there is substantial evidence for an association with MTX response in RA patients. For clinical application more evidence from prospective studies with multivariate testing is needed.
References
Lima A, Bernardes M, Azevedo R, Seabra V, Medeiros R. Moving toward personalized medicine in rheumatoid arthritis: SNPs in methotrexate intracellular pathways are associated with methotrexate therapeutic outcome. Pharmacogenomics. 2016. https://doi.org/10.2217/pgs-2016-0067.
Ulrich CM, Robien K, Sparks R. Pharmacogenetics and folate metabolism—a promising direction. Pharmacogenomics. 2002;3:299–313.
Kawaida R, Yamada R, Kobayashi K, Tokuhiro S, Suzuki A, Kochi Y, et al. CUL1, a component of E3 ubiquitin ligase, alters lymphocyte signal transduction with possible effect on rheumatoid arthritis. Genes Immun. 2005;6:194–202.
Hider SL, Bruce IN, Thomson W. The pharmacogenetics of methotrexate. Rheumatology. 2007;46:1520–4.
Malik F, Ranganathan P. Methotrexate pharmacogenetics in rheumatoid arthritis: a status report R eview. Pharmacogenomics. 2013;14:305–14.
Plant D, Wilson AG, Barton A. Genetic and epigenetic predictors of responsiveness to treatment in RA. Nat Rev Rheumatol. 2014;10:329–37.
Ranganathan P, McLeod HL. Methotrexate pharmacogenetics: the first step toward individualized therapy in rheumatoid arthritis. Arthritis Rheum. 2006;54:1366–77.
Brinker RR, Ranganathan P. Methotrexate pharmacogenetics in rheumatoid arthritis. Clin Exp Rheumatol. 2010;28. https://doi.org/10.3899/jrheum.091095.
Stamp L, Roberts R, Kennedy M, Barclay M, O’Donnell J, Chapman P. The use of low dose methotrexate in rheumatoid arthritis—are we entering a new era of therapeutic drug monitoring and pharmacogenomics? Biomed Pharmacother. 2006;60:678–87.
Zhu H, Deng F-Y, Mo X-B, Qiu Y-H, Lei S-F. Pharmacogenetics and pharmacogenomics for rheumatoid arthritis responsiveness to methotrexate treatment: the 2013 update. Pharmacogenomics. 2014;15:551–66.
Dervieux T, Furst D, Lein DO, Capps R, Smith K, Caldwell J, et al. Pharmacogenetic and metabolite measurements are associated with clinical status in patients with rheumatoid arthritis treated with methotrexate: results of a multicentred cross sectional observational study. Ann Rheum Dis. 2005;64:1180–5.
Ranganath VK, Khanna D, Paulus HE. ACR remission criteria and response criteria. Clin Exp Rheumatol. 2006;24:1909.
Fransen J, van Riel PL. The disease activity score and the EULAR response criteria. Rheum Dis Clin North Am. 2009;35:745–57.
Counsell C. Formulating questions and locating primary studies for inclusion in systematic reviews. Ann Intern Med. 1997;127:380–7.
Moher D, Shamseer L, Clarke M, Ghersi D, Liberati A, Petticrew M, et al. Preferred reporting items for systematic review and meta-analysis protocols (PRISMA-P) 2015 statement. Syst Rev. 2015;4:1.
Johnson AD, Handsaker RE, Pulit SL, Nizzari MM, O’Donnell CJ, De Bakker PIW. SNAP: a web-based tool for identification and annotation of proxy SNPs using HapMap. Bioinformatics. 2008;24:2938–9.
Senapati S, Singh S, Das M, Kumar A, Gupta R, Kumar U, et al. Genome-wide analysis of methotrexate pharmacogenomics in rheumatoid arthritis shows multiple novel risk variants and leads for TYMS regulation. Pharmacogenet Genomics. 2014;24:211–9.
Ghodke-Puranik Y, Puranik AS, Shintre P, Joshi K, Patwardhan B, Lamba J, et al. Folate metabolic pathway single nucleotide polymorphisms: a predictive pharmacogenetic marker of methotrexate response in Indian (Asian) patients with rheumatoid arthritis. Pharmacogenomics. 2015;16:2019–34.
James HM, Gillis D, Hissaria P, Lester S, Somogyi AA, Cleland LG, et al. Common polymorphisms in the folate pathway predict efficacy of combination regimens containing methotrexate and sulfasalazine in early rheumatoid arthritis. J Rheumatol. 2008;35:562–71.
Grabar PB, Rojko S, Logar D, Dolzan V. Genetic determinants of methotrexate treatment in rheumatoid arthritis patients: a study of polymorphisms in the adenosine pathway. Ann Rheum Dis. 2010;69:931–2.
Sharma Shruti, Das Mitashree, Kumar Ashok, Vishal Marwaha, Subramanian Shankar, Paramjeet Singh, et al. Purine biosynthetic pathway genes and methotrexate response in rheumatoid arthritis patients among north Indians. PubMed commons. Pharmacogenet Genomics. 2009;19:823–8.
Wessels JAM, Kooloos WM, De Jonge R, De Vries-Bouwstra JK, Allaart CF, Linssen A, et al. Relationship between genetic variants in the adenosine pathway and outcome of methotrexate treatment in patients with recent-onset rheumatoid arthritis. Arthritis Rheum. 2006;54:2830–9.
Lee YC, Cui J, Costenbader KH, Shadick NA, Weinblatt ME, Karlson EW. Investigation of candidate polymorphisms and disease activity in rheumatoid arthritis patients on methotrexate. Rheumatology. 2009;48:613–7.
Kurzawski M, Malinowski D, Szarmach N, Nowak A, Goryniak A, Pawlik A, et al. ATIC missense variant affects response to methotrexate treatment in rheumatoid arthritis patients. Pharmacogenomics. 2016;17:1971–8.
Swierkot J, Bogunia-Kubik K, Nowak B, Bialowas K, Korman L, Gebura K, et al. Analysis of associations between polymorphisms within genes coding for tumour necrosis factor (TNF)-alpha and TNF receptors and responsiveness to TNF-alpha blockers in patients with rheumatoid arthritis. Jt Bone Spine. 2015;82:94–9.
Aggarwal P, Naik S, Mishra KP, Aggarwal A, Misra R. Correlation between methotrexate efficacy & toxicity with C677T polymorphism of the methylenetetrahydrofolate gene in rheumatoid arthritis patients on folate supplementation. Indian J Med Res. 2006;124:521–6.
Wessels JAM, De Vries-Bouwstra JK, Heijmans BT, Slagboom PE, Goekoop-Ruiterman YPM, Allaart CF, et al. Efficacy and toxicity of methotrexate in early rheumatoid arthritis are associated with single-nucleotide polymorphisms in genes coding for folate pathway enzymes. Arthritis Rheum. 2006;54:1087–95.
Taraborelli M, Andreoli L, Archetti S, Ferrari M, Cattaneo R, Tincani A. Methylenetetrahydrofolate reductase polymorphisms and methotrexate: no association with response to therapy nor with drug-related adverse events in an Italian population of rheumatic patients. Clin Exp Rheumatol. 2009;27:499–502.
Dervieux T, Greenstein N, Kremer J. Pharmacogenomic and metabolic biomarkers in the folate pathway and their association with methotrexate effects during dosage escalation in rheumatoid arthritis. Arthritis Rheum. 2006;54:3095–103.
Dervieux T, Wessels JaMM, van der Straaten T, Penrod N, Moore JH, Guchelaar H-J, et al. Gene–gene interactions in folate and adenosine biosynthesis pathways affect methotrexate efficacy and tolerability in rheumatoid arthritis. Pharmacogenet Genomics. 2009;19:935–44.
Lima A, Bernardes M, Azevedo R, Medeiros R, Seabra V. Pharmacogenomics of methotrexate membrane transport pathway: can clinical response to methotrexate in rheumatoid arthritis be predicted? Int J Mol Sci. 2015;16:13760–80.
Negi S, Kumar A, Thelma BK, Juyal RC. Association of Cullin1 haplotype variants with rheumatoid arthritis and response to methotrexate. Pharmacogenet Genomics. 2011;21:590–3.
Majorczyk E, Pawlik A, Gendosz D, Kuśnierczyk P. Presence of the full-length KIR2DS4 gene reduces the chance of rheumatoid arthritis patients to respond to methotrexate treatment. BMC Musculoskelet Disord. 2014;15:256.
Dalrymple JM, Stamp LK, O’Donnell JL, Chapman PT, Zhang M, Barclay ML. Pharmacokinetics of oral methotrexate in patients with rheumatoid arthritis. Arthritis Rheum. 2008;58:3299–308.
Eektimmerman F, Swen JJ, Böhringer S, Huizinga TW, Kooloos WM, Allaart CF, et al. Pathway analysis to identify genetic variants associated with efficacy of adalimumab in rheumatoid arthritis. Pharmacogenomics. 2017;18:945–53.
Muralidharan N, Mariaselvam CM, Jain VK, Gulati R, Negi VS. ATIC 347C>G gene polymorphism may be associated with methotrexate-induced adverse events in south Indian Tamil rheumatoid arthritis. Pharmacogenomics. 2016;17:241–8.
Hayashi H, Tazoe Y, Tsuboi S, Horino M, Morishita M, Arai T, et al. A single nucleotide polymorphism of reduced folate carrier 1 predicts methotrexate efficacy in Japanese patients with rheumatoid arthritis. Drug Metab Pharmacokinet. 2013;28:164–8.
Salazar J, Moya P, Altés A, Díaz-Torné C, Casademont J, Cerdà-Gabaroi D, et al. Polymorphisms in genes involved in the mechanism of action of methotrexate: are they associated with outcome in rheumatoid arthritis patients? Pharmacogenomics. 2014;15:1079–90.
Sharma S, Das M, Kumar A, Marwaha V, Shankar S, Aneja R, et al. Interaction of genes from influx-metabolism-efflux pathway and their influence on methotrexate efficacy in rheumatoid arthritis patients among Indians. Pharmacogenet Genomics. 2008;18:1041–9.
Kooloos WM, Wessels JA, van der Straaten T, Allaart CF, Huizinga TW, Guchelaar H-J. Functional polymorphisms and methotrexate treatment outcome in recent-onset rheumatoid arthritis. Pharmacogenomics. 2010;11:163–75.
Drozdzik M, Rudas T, Pawlik A, Gornik W, Kurzawski M, Herczynska M. Reduced folate carrier-1 80G>A polymorphism affects methotrexate treatment outcome in rheumatoid arthritis. Pharmacogenomics J. 2007;7:404–7.
Muralidharan N, Mariaselvam CM, Mithun CB, Negi VS. Reduced folate carrier-1 80G>A gene polymorphism is not associated with methotrexate treatment response in South Indian Tamils with rheumatoid arthritis. Clin Rheumatol. 2016;35:879–85.
Moya P, Salazar J, Arranz MJ, Díaz-Torné C, del Río E, Casademont J, et al. Methotrexate pharmacokinetic genetic variants are associated with outcome in rheumatoid arthritis patients. Pharmacogenomics. 2016;17:25–9.
Chatzikyriakidou A, Georgiou I, Voulgari PV, Papadopoulos CG, Tzavaras T, Drosos AA. Transcription regulatory polymorphism -43T>C in the 5′-flanking region of SLC19A1 gene could affect rheumatoid arthritis patient response to methotrexate therapy. Rheumatol Int. 2007;27:1057–61.
van der Straaten R, Wessels JA, de Vries-Bouwstra JK, Goekoop-Ruiterman YP, Allaart CF, Bogaartz J, et al. Exploratory analysis of four polymorphisms in human GGH and FPGS genes and their effect in methotrexate-treated rheumatoid arthritis patients. Pharmacogenomics. 2007;8:141–50.
Wessels JAM, Van Der Kooij SM, Le Cessie S, Kievit W, Barerra P, Allaart CF, et al. A clinical pharmacogenetic model to predict the efficacy of methotrexate monotherapy in recent-onset rheumatoid arthritis. Arthritis Rheum. 2007;56:1765–75.
Milic V, Jekic B, Lukovic L, Bunjevacki V, Milasin J, Novakovic I, et al. Association of dihydrofolate reductase (DHFR) -317AA genotype with poor response to methotrexate in patients with rheumatoid arthritis. Clin Exp Rheumatol. 2012;30:178–83.
Soukup T, Dosedel M, Pavek P, Nekvindova J, Barvik I, Bubancova I, et al. The impact of C677T and A1298C MTHFR polymorphisms on methotrexate therapeutic response in East Bohemian region rheumatoid arthritis patients. Rheumatol Int. 2015;35:1149–61.
Kung TN, Dennis J, Ma Y, Xie G, Bykerk V, Pope J, et al. RFC1 80G>A is a genetic determinant of methotrexate efficacy in rheumatoid arthritis: a human genome epidemiologic review and meta-analysis of observational studies. Arthritis Rheumatol. 2014;66:1111–20.
Li X, Hu M, Li W, Gu L, Chen M, Ding H, et al. The association between reduced folate carrier-1 gene 80G/A polymorphism and methotrexate efficacy or methotrexate related-toxicity in rheumatoid arthritis: A meta-analysis. Int Immunopharmacol. 2016;38:8–15.
Acknowledgements
We are particularly grateful for the assistance for search terms given by Jan Schoones of the Walaeus library (LUMC).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Eektimmerman, F., Swen, J.J., Madhar, M.B. et al. Predictive genetic biomarkers for the efficacy of methotrexate in rheumatoid arthritis: a systematic review. Pharmacogenomics J 20, 159–168 (2020). https://doi.org/10.1038/s41397-019-0098-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41397-019-0098-9
- Springer Nature Limited