Abstract
Breast cancer is the second leading cause of cancer-related deaths among women, largely due to the progression of a significant fraction of primary tumours to the metastatic stage. Here, we show that zinc-finger protein 750 (ZNF750) opposes the migration and invasion of breast cancer cells by repressing a prometastatic transcriptional programme, which includes genes involved in focal adhesion and extracellular matrix interactions, such as LAMB3 and CTNNAL1. Mechanistically, ZNF750 recruits the epigenetic modifiers KDM1A and HDAC1 to the promoter regions of LAMB3 and CTNNAL1, influencing histone marks and transactivating these genomic sites. Gene expression analysis in cancer patient datasets indicated that ZNF750 and its targets were negative prognostic factors in breast cancer. Together, our findings shed light on the molecular mechanism by which ZNF750 regulates cell migration and invasion, suggesting a role in breast cancer metastasis.
Similar content being viewed by others
Introduction
Breast cancer is the second leading cause of cancer-related deaths among women worldwide. It is a complex heterogeneous disease that can be classified either histologically according to the expression of oestrogen receptor, progesterone receptor, and ERBB2 receptor or by gene expression profiling into at least five distinct subtypes (luminal A, luminal B, HER2-enriched, basal-like and normal-like) [1, 2]. While surgical resection is generally effective in resolving primary breast cancers, disease progression to metastatic stages results in incurable conditions and is therefore responsible for mortality. Understanding the molecular basis for the acquisition of metastatic properties by breast cancers is therefore of central importance.
ZNF750 is a zinc-finger transcription factor that plays a key role in the regulation of the homeostasis of epithelia [3, 4]; ZNF750 deregulation can indeed result in neoplastic transformation [5]. The structural organization of ZNF750 consists of an atypical C2H2 zinc-finger motif in the amino-terminal domain and two highly conserved PLNLS sequences that are involved in DNA binding and protein–protein interactions, respectively. ZNF750 is required for epidermal differentiation. In particular, ZNF750 interacts with RCOR1 and CTBP1/2 to exert dual gene regulatory impacts: namely, activating the expression of differentiation-associated genes in concert with KLF4 and repressing progenitor gene expression in association with KDM1A [6]. More recently, ZNF750 has been associated with tumour suppression in squamous cell carcinomas of the oesophagus (ESCCs), lung and cervix. Mutations in ZNF750 have been reported in oesophageal squamous cell carcinoma [5] in association with higher cytoplasmic protein levels, while ZNF750 has nuclear localization in normal tissues.
Here, we show that ZNF750 represses the migration and invasion of breast cancer cells. Our findings reveal that ZNF750 recruits the epigenetic platform KDM1A/HDAC1 to the genetic loci of LAMB3 and CTNNAL1, repressing their expression. Depletion of ZNF750 expression results in an increase in the repressive histone acetylation and methylation marks on both LAMB3 and CTNNAL1 promoters. Depletion of LAMB3 or CTNNAL1 reverses the promigratory effect induced by ZNF750 silencing in breast cancer cells.
Results
ZNF750 is deregulated in breast cancer
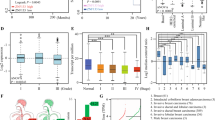
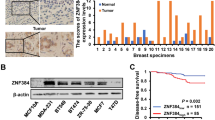
ZNF750 is a lineage-specific tumour suppressor gene in SSC [7]; however, little is known about its general tumour-suppressive role in other cancer types. Using cBioPortal for Cancer Genomics (http://www.cbioportal.org) [8, 9], we performed a comprehensive analysis of genetic alterations of ZNF750 across several tumours. Interestingly, breast and prostate cancer [10,11,12] showed the highest alteration frequency (Supplementary Fig. 1a). In particular, ZNF750 was frequently amplified in breast cancer, while deletions and mutations were less common (Supplementary Fig. 1b). As shown in Supplementary Fig. 1c, the four mutations detected thus far do not involve the biologically relevant zinc-finger domain (required for the interaction with DNA) or the PLNLS motifs (responsible for protein–protein interactions) located at the C terminus. Despite being amplified in breast cancer, expression analysis revealed that ZNF750 expression in tumours is reduced when compared with expression in normal tissue and negatively correlates with tumour stage (Fig. 1a, b), suggesting that amplification is a passenger event that is possibly counteracted by epigenetic silencing. To further confirm the expression data, we performed an immunohistochemistry analysis in our small cohort of breast cancer patients. In normal tissue, ZNF750 was mainly expressed in myoepithelial cells with a prevalent nuclear localization (Fig. 1c), and weak staining was observed in luminal cells. In contrast, ZNF750 expression was reduced in tumour tissue classified as triple-negative breast cancer (TNBC), luminal A breast cancer, luminal B breast cancer or Her2+ breast cancer (Fig. 1c and Supplementary Fig. 1d). In particular, ZNF750 expression was almost absent at the nuclear level, while weak staining was observed in the cytoplasm. We then queried the Human Protein Atlas dataset (https://www.proteinatlas.org); in agreement with our data, ZNF750 expression was found to be reduced in cancer tissues when compared with that in normal tissues (Supplementary Fig. 1e). Remarkably, reduced ZNF750 levels were associated with worse survival in several cancer datasets (Fig. 1d–f). Since ZNF750 has been previously described as a key transcriptional target of p63 [3] and considering the crucial role of p63 in the biology of mammary cancer stem cells [11, 13], we asked whether the p63/ZNF750 axis is conserved in breast cancer. To this end, we first correlated the expression of ZNF750 and p63 across several breast cancer cell lines at both the mRNA and protein levels. MDA-MB-453 and MDA-MB-468 cell lines had the highest expression of ZNF750 at both the mRNA and protein levels (Fig. 1g, h). In contrast, MDA-MB-231 and BT-549 cell lines did not express ZNF750. However, no correlation between p63 isoforms and ZNF750 was observed in the four breast cancer cell lines (Supplementary Fig. 2a–c) and in three breast cancer datasets (luminal A, luminal B and basal types) (Supplementary Fig. 2d). Next, we verified that p63 silencing did not affect ZNF750 expression in breast cancer cell lines (Supplementary Fig. 2e) despite ΔNp63 binding to the ZNF750 promoter (Supplementary Fig. 2f) and strongly regulating its expression (Supplementary Fig. 2g) in epidermal-derived models, such as head and neck SCC cell lines.
a Bioinformatics analysis showed that ZNF750 mRNA expression was reduced in breast cancer tissues when compared with that in normal tissues. b The expression of ZNF750 mRNA negatively correlated with the tumour stage. The analysis was carried out using the UALCAN website. The dataset analysed was from TCGA (see “Materials and methods”). c Expression of ZNF750 protein in normal tissues and in the indicated breast cancer subtypes was evaluated by IHC. A representative case is shown. Bar = 100 μm. d–f Patients with low expression of ZNF750 showed a shorter relapse-free survival than patients with high expression of ZNF750. The analysis was carried out using the R2 Genomic Platform (https://r2.amc.nl) and kmplot.com. The datasets analysed were GSE69031 and GSE12093. g Real-time PCR showing the expression of ZNF750 mRNA across the indicated breast cancer cell lines. Data were normalized to TBP and relative to MCF-10A, an immortalized breast epithelial cell line. Bars represent the mean ± SD of two independent biological replicates. h Western blot showing ZNF750 expression across several breast cancer cell lines. A representative western blot of two different experiments is shown.
Together, these results indicate that ZNF750 is deregulated in breast cancer and that its expression is independent of p63.
ZNF750 regulates cell migration/invasion in breast cancer cell lines
To explore the biological role of ZNF750, we first correlated the levels of ZNF750 protein in four different cell lines with their migratory capacity. Cell lines expressing high levels of ZNF750, MDA-MB-468 and MDA-MB-453, showed less migratory capacity than their low-expressing counterparts, MDA-MB-231 and BT-549 (Fig. 2a, b). Then, to confirm the possible effect of ZNF750 on cell migration in breast cancer, its expression was depleted by siRNA in MDA-MB-468 and HCC1143 cell lines (Supplementary Fig. 3a, b). Interestingly, Fig. 2c shows that the silencing of ZNF750 markedly increased the migratory potential of both cell lines. In addition, as shown in Fig. 2d and Supplementary Fig. 3c, depletion of ZNF750 resulted in an increase in invasiveness. To corroborate our findings, we overexpressed either wild-type or DNA binding domain-mutant ZNF750 in the MDA-MB-231 cell line. The migration of the breast cancer cell line was significantly inhibited by wild-type but not mutant ZNF750 (Fig. 2e).
a, b The migratory capacity of the indicated breast cancer cell lines inversely correlated with ZNF750 expression. Western blot analysis of ZNF750 levels in the indicated breast cancer cell lines. A representative micrograph of three independent experiments is shown. Bar = 100 μm. The graph depicts the quantification of the migrated cells. Bars represent the mean ± SD of three independent experiments. c, d Inhibition of ZNF750 increased the migration and invasion capabilities of the indicated cell lines (transwell assay). Bars represent the mean ± SD of three independent experiments. A representative micrograph of three independent experiments is shown. Bar = 100 μm. e Ectopic expression of ZNF750 repressed cell migration. MDA-MB-231 cells were transfected with the indicated plasmid and subjected to transwell assays. f, g ZNF750 regulated Golgi positioning and cytoskeleton remodelling. Inhibition of ZNF750 expression increased the number of cells with their Golgi apparatuses oriented towards the wound (bar = 10 μm). Bars represent the mean ± SD of three independent experiments. Statistical significance was calculated with a two-sided Student’s t test. h Actin filament polymerization (asterisk) and focal adhesion formation (arrow) increased after ZNF750 silencing (bar = 25 μm). A representative micrograph of three independent experiments is shown.
The Golgi apparatus is a key player in the cell migration process and is oriented towards the direction of migration [14]. To further confirm the role of ZNF750 in cell migration, we assessed the localization of the Golgi apparatus in a wound-scratch assay [15]. As shown in Fig. 2f, g, inhibition of ZNF750 expression (Supplementary Fig. 3d) significantly increased the number of cells with the Golgi apparatus facing the scratch. Focal adhesions (FAs) are required for interaction with the extracellular matrix and generate the forces to pull the cell forward; formation and turnover of FAs are indeed required for proper cell migration [16]. To verify whether ZNF750 is involved in FAs, we stained MDA-MB-468 cells with phalloidin and vinculin. As shown in Fig. 2h, inhibition of ZNF750 expression resulted in an increase in focal adhesion formation, as displayed by the localization of vinculin at the edge of stress fibres.
We next tested whether the effect of ZNF750 on migration correlates with the regulation of EMT [17]. However, ZNF750 silencing did not significantly affect the expression of the EMT markers E-Cadherin, Vimentin, Snail and Zeb1 at the mRNA level, as shown in Supplementary Fig. 3e.
Overall, these data suggest that ZNF750 inhibits the migration and invasion of breast cancer cell lines.
ZNF750 regulates the WNT pathway and focal adhesion
To delineate the molecular mechanism by which ZNF750 regulates cell migration and invasion, we performed a gene expression microarray analysis in MDA-MB-468 cells upon ZNF750 depletion (Supplementary Fig. 4a). This analysis revealed that ZNF750 controls either directly or indirectly the expression of 616 genes (285 upregulated and 331 downregulated genes) (Fig. 3a). In keeping with the tumour suppressor role of ZNF750, gene ontology analysis (Fig. 3b), using DAVID software (https://david.ncifcrf.gov/), revealed enrichment in pathways involved both in cell proliferation and cell migration, such as the WNT pathway (p = 5.40E−03) and pathways related to focal adhesion (p = 3.50E−03). Real-time PCR validation confirmed that ZNF750 regulated the expression of genes belonging to the WNT pathway, such as GPC4, JUN, PPARD, TCF7, WNT10a and FGFR2, and the focal adhesion pathway, including LAMB3, MYLK2, RAC1 and MAPK9 (Supplementary Fig. 4b). Next, to identify genes that are directly regulated by ZNF750 at the transcriptional level, we compared the genes identified in our microarray with those from a previously generated ChIP-seq analysis [6]. Our analysis showed that 90 potential genes could be directly regulated by ZNF750 (Fig. 3c). Among those genes, we identified and validated two putative target genes, LAMB3 and CTNNAL1, that played a role in the regulation of migration in cancer cells [18,19,20,21]. Indeed, depletion of ZNF750 with two different siRNAs significantly upregulated the expression of LAMB3 and CTNNAL1 at both the mRNA and protein levels in several breast cancer cell lines (Fig. 3d, e and Supplementary Fig. 4c, d).
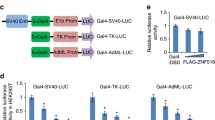
a Heat map showing the gene expression changes after ZNF750 depletion in MDA-MB-468 cells (n = 4, biological replicates). b Gene ontology analysis of the most representative pathways regulated by ZNF750. The analysis was performed with DAVID (https://david.ncifcrf.gov/). c Venn diagram showing the overlap between genes regulated by ZNF750 (our microarray analysis) and genes from the ChIP-seq analysis of ZNF750 [6]. d, e Depletion of endogenous ZNF750 by specific siRNA resulted in an increase in LAMB3 and CTNNAL1 expression in the indicated breast cancer cell lines at both the mRNA and protein levels. mRNA levels of the indicated genes were assessed by real-time PCR. Data were normalized to the housekeeping gene β-actin and relative to the scrambled sample (Scr). Data represent the mean ± SD of three independent experiments. Representative western blot image of three independent experiments. f LAMB3 and CTNNAL1 are direct target genes of ZNF750. A schematic representation of the LAMB3 and CTNNAL1 promoters with putative ZNF750 binding sites. The binding sites are indicated as “L1” and “L2” for LAMB3 and “C1” and “C2” for CTNNAL1. g A chromatin immunoprecipitation (ChIP) assay was performed in MDA-MB-468 cells. Protein–chromatin complexes were precipitated using a ZNF750 antibody, and the DNA sequences were amplified by real-time PCR. Statistical significance was calculated with a two-sided Student’s t test.
These results suggest that ZNF750 may negatively regulate cell migration and invasion by inhibiting the expression of LAMB3 and CTNNAL1.
CTNNAL1 and LAMB3 are direct targets of ZNF750
Next, we verified whether ZNF750 directly regulates the transcription of LAMB3 and CTNNAL1. Using MatInspector software, we searched for potential ZNF750 binding sites in the promoter sequence of the CTNNAL1 and LAMB3 genes (Supplementary Fig. 4e). We found two putative ZNF750 binding sites in the LAMB3 promoter located at −513 and −6 bp from the transcriptional start site (TSS) (Fig. 3f, indicated as L1 and L2, respectively) and two putative ZNF750 binding sites in the CTNNAL1 promoter located at −217 and −8 bp from the TSS (Fig. 3f, indicated as C1 and C2, respectively). We then performed chromatin immunoprecipitation (ChIP) assays using an antibody against endogenous ZNF750. As shown in Fig. 3g, ZNF750 binds the LAMB3 and CTNNAL1 promoters at the L1, L2, and C2 sites. Overall, this finding indicates that ZNF750 binds to the LAMB3 and CTNNAL1 promoters and represses their expression.
ZNF750 opposes cell migration by repressing the expression of the LAMB3 and CTNNAL1 genes
Our findings lead to the hypothesis that ZNF750 inhibits cell migration by regulating the expression of LAMB3 and CTNNAL1. To verify this, we transfected MDA-MB-468 cell lines with siRNAs targeting ZNF750 alone or in combination with either LAMB3 or CTNNAL1 siRNA to revert the migratory phenotype (Supplementary Fig. 4f). As predicted, depletion of CTNNAL1 or LAMB3 abrogated the increase in cell migration triggered by ZNF750 silencing (Fig. 4a, b). A consistent effect was also observed on the Golgi orientation (Fig. 4c). Interestingly, LAMB3 or CTNNAL1 knockdown in the absence of ZNF750 silencing resulted in a reduction of MDA-MB-468 cell migratory capacity.
a, b A transwell migration assay was performed in MDA-MB-468 cells transfected with the indicated targeting oligos alone or in combination. Bars represent the mean ± SD of three independent experiments. A representative micrograph of three independent experiments is shown. c ZNF750 inhibits Golgi orientation by repressing LAMB3 and CTNNAL1 expression. d, e ZNF750 expression inversely correlated with LAMB3 and CTNNAL1 expression in cancer patient datasets. Coexpression analysis of ZNF750 with the target genes was performed with the indicated cancer datasets specifically for breast cancers classified as either “basal-like” or luminal B according to the PAM50 classification. The analysis was carried out using the cBioPortal for Cancer Genomics (http://www.cbioportal.org/). f Overexpression of ZNF750 in the indicated breast cancer cell lines inhibited the expression of LAMB3 and CTNNAL1. Data represent the mean of three technical replicates (n = 3, PCR runs) ± SD and are representative of one experiment. Statistical significance was calculated with a two-sided Student’s t test.
To support these data, we correlated the expression of ZNF750 with its direct targets using publicly available gene expression datasets from cancer patients (TCGA; see “Materials and methods”) [22, 23]. As predicted, we found that expression of both CTNNAL1 and LAMB3 (Fig. 4e) was negatively correlated with ZNF750 expression in basal-like breast cancer, luminal B breast cancer and TNBC (Fig. 4d, e and Supplementary Fig. 5) (p values and q value <0.05). No correlation between ZNF750 and the target genes was observed in breast cancers classified as luminal A or Her2-enriched according to the PAM50 classification [24], suggesting a specific role of the ZNF750/CTNNAL1-LAMB3 axis in selective breast cancer subtypes.
To further support the role of ZNF750 in breast cancer cell migration, we also overexpressed ZNF750 in three different breast cancer cell lines characterized by low levels of ZNF750 expression. As shown in Fig. 4f, ZNF750 inhibited the expression of LAMB3 and CTNNAL1 in all three cell lines. Taken together, these results clearly demonstrate that ZNF750 suppresses breast cancer cell migration by inhibiting the expression of the LAMB3 and CTNNAL1 genes.
ZNF750 recruits KDM1A and HDAC1 to the LAMB3 and CTNNAL1 promoters
ZNF750 has been shown to act in a complex with KDM1A, inhibiting the transcription of genes involved in the maintenance of an undifferentiated state [6]. Therefore, we hypothesized that ZNF750 might also interact with KDM1A in breast cancer cells and repress the expression of LAMB3 and CTNNAL1. Indeed, co-immunoprecipitation showed an interaction between endogenous ZNF750 and KDM1A (Fig. 5a). In addition, a ChIP assay showed that KDM1A bound to the promoters of CTNNAL1 and LAMB3 (Fig. 5b). KDM1A is an integral component of the CoREST complex [25], which exerts histone deacetylation activity due to the presence of its components HDAC1 and HDAC2. Hence, we asked whether ZNF750 might also recruit HDACs. As shown in Fig. 5a, endogenous ZNF750 interacted with HDAC1, and importantly, HDAC1 bound the promoters of CTNNAL1 and LAMB3 (Fig. 5c). To verify whether HDAC1 is involved in the repression of LAMB3 and CTNNAL1 expression, MDA-MB-468 cells were treated with givinostat, a histone deacetylase inhibitor [26]. As shown in Fig. 5d, e, LAMB3 and CTNNAL1 mRNA and protein were increased upon histone deacetylase inhibition. To determine whether the interaction of ZNF750 with KDM1A and HDAC1 is important for the repression of LAMB3 and CTNNAL1, we performed a ChIP assay after depletion of ZNF750. Depletion of ZNF750 resulted in reduced binding of HDAC1 and KDM1A to both promoters (Fig. 5f). Consequentially, and in line with the reduced HDAC1 and KDM1A binding, we also observed a significant increase in H3 acetylation (AcH3) and methylation of H3K4 (Me2H3K4) on both promoters (Fig. 5g). In conclusion, these findings indicate that ZNF750 represses the expression of LAMB3 and CTNNAL1 by recruiting histone modifiers to their promoters.
a Co-immunoprecipitation analysis of endogenous ZNF750 in the MDA-MB-468 cell line and western blot for ZNF750, KDM1A and HDAC1. A representative image of three independent experiments is shown. b, c A ChIP assay was performed in MDA-MB-468 cells. Protein–chromatin complexes were precipitated using either HDAC1 or KDM1A antibodies. DNA sequences were amplified by real-time PCR. A representative experiment of two independent experiments is shown. IgG antibody was used as a negative control. d, e Expression of LAMB3 and CTNNAL1 increased following treatment with the HDAC inhibitor givinostat (GN) at both the mRNA and protein levels. Bars represent the mean ± SD of three independent experiments. Statistical significance was calculated with a two-sided Student’s t test. f, g ZNF750 was depleted in the MDA-MB-468 cell line, and ChIP and real-time PCR were performed after immunoprecipitation with antibodies against the following targets: HDAC1, AcH3 KDM1A and Me2H3K4. h Schematic representation of the proposed molecular mechanism by which ZNF750 represses the expression of LAMB3 and CTNNAL1. Statistical significance was calculated with a two-sided Student’s t test.
Discussion
Our study shed light on the role of the transcription factor ZNF750 in the regulation of migration and invasion of breast cancer cells, suggesting its function as a metastasis suppressor. ZNF750 is located on chromosome 17, which is rearranged in at least 30% of breast cancers [27,28,29]. Remarkably, several important oncogenes/tumour suppressors, such as p53, BRCA1 and ERBB2, are located on this chromosome [30,31,32,33]. Our analyses indicate that the ZNF750 locus is frequently amplified in breast cancer, although its expression is generally downregulated. Although gene amplification is often associated with overexpression, some evidence indicates that it can also be associated with decreased expression [34, 35]. Epigenetic mechanisms, post-transcriptional RNA modifications and non-coding RNAs might control ZNF750 expression. For example, recent evidence has suggested that the downregulated status of ZNF750 mRNA could be also due to m6A modification [36]; hence, several layers of control might be responsible for the expression level of ZNF750. Importantly, deregulation of ZNF750 has a role in predicting patient survival. Patients with low expression of ZNF750 show worse disease-free survival than patients with high expression of ZNF750. Therefore, it appears to be likely that amplification is a passenger event associated with the complexity of chromosome 17 during breast cancer progression.
We identified downstream regulators of the ZNF750 antimigratory effect. The ZNF750/LAMB3 axis is involved in breast cancer cell migration and invasion. The LAMB3 gene encodes the beta chain of laminin 332 (also known as laminin 5), which also consists of alpha and gamma chains coded by the LAMA3 and LAMC2 genes, respectively [37]. Laminin 332 plays a key role in cell adhesion and has been shown to be extensively involved in cancer invasion and tumour development. Although deregulation of LAMB3 has been reported in several cancers, the molecular mechanism has not been fully investigated [38, 39]. Our data indicate that ZNF750 controls epigenetic modifications of LAMB3 promoter. Indeed, depletion of ZNF750 deregulates the recruitment of HDAC1 and KDM1A, thus increasing the acetylation and methylation of the LAMB3 promoter. LAMC2 is also transcriptionally regulated by ZNF750 [7], and therefore, ZNF750 appears to be a potential key transcriptional regulator of laminin 332.
We also demonstrated that ZNF750 activity in breast cancer is related to its ability to repress the expression of CTNNAL1. The CTNNAL1 gene encodes α-catulin, a protein that regulates Rho signalling, which is a key pathway for cytoskeletal reorganization [18]. Moreover, CTNNAL1 expression is upregulated in human cancers and correlates with tumour size and cancer progression [40]. Our data suggest that ZNF750 alteration is a plausible molecular mechanism for CTNNAL1 deregulation in cancer. Similar to the ZNF750/LAMB3 axis, ZNF750 represses the expression of CTNNAL1 by recruiting KDM1A and HDAC1 to its promoter, reducing the migratory capacity of breast cancer cells. Our in vitro data were also confirmed in two breast cancer patient datasets. The expression of CTNNAL1 and LAMB3 inversely correlated with ZNF750 expression in breast cancer. In particular, this negative correlation was only statistically significant in breast cancers classified as basal-like or luminal B and not in breast cancers classified as luminal A or Her2+ [41].
In conclusion, we show that ZNF750 acts as a tumour suppressor gene in breast cancer by inhibiting cell migration and invasion. ZNF750 expression could potentially serve as a novel biomarker for staged breast cancers. Overall, these findings improve our understanding of the molecular mechanisms underlying the acquisition of a metastatic phenotype in breast cancer.
Materials and methods
Cell culture and transfection
All cell lines used were obtained from American Type Culture Collection and maintained at 37 °C in 5% CO2 in culture medium. MCF-10A (fibrocystic disease), MCF7 (adenocarcinoma), BT-549 (ductal carcinoma, mesenchymal), MDA-MB-468 (ductal carcinoma, basal like-1), MDA-MB-453 (carcinoma, LAR) HCC1143 (ductal carcinoma, basal like-1), HCC1954 (ductal carcinoma, basal-like), HCC1937 (ductal carcinoma, basal like-1), MDA-MB-231 (adenocarcinoma, mesenchymal-stem-like), FADU, Detroit and A253 cells were grown in RPMI medium supplemented with 250 μM l-glutamine (Gibco), penicillin/streptomycin 1 U/ml (Gibco), 10% FBS (Invitrogen) or Dulbecco’s modified eagle’s medium with 4,5 g/l glucose and l-glutamine (Lonza), penicillin/streptomycin 1 U/ml (Gibco), and 10% FBS (Invitrogen). Cells were transfected with: (i) 50 nM siRNA of Qiagen by lipofectamine RNAiMAX, according to the manufacturer’s instructions or (ii) plasmids by lipofectamine 2000, according to the manufacturer’s instructions.
Bioinformatics analysis
Bioinformatics analysis was carried out by using the following website: http://www.cbioportal.org. [8, 9] UALCAN, http://ualcan.path.uab.edu, and R2, https://hgserver1.amc.nl/cgi-bin/r2/main.cgi, were also used. The following datasets were examined: TCGA Nature 2012 Breast Invasive Carcinoma and TCGA Cell 2015 Invasive Lobular Carcinoma (GSE69031 and GSE12093, respectively).
Tissue microarray
Tumour samples were fixed in formalin and then embedded in paraffin. Sections were deparaffinized, and after antigen retrieval (in HIER solution at pH 6; 98 °C for 30 min), sections were blocked. Then, sections were incubated for 1 h with anti-ZNF750 (Sigma HPA023012 and HPA021573) diluted 1:25. The Novocastra Novolink™ (Leica) system was used for detection. Ethical approval of this study was obtained from Tor Vergata Hospital, approval number 129/18.
Cloning expression vectors and lentiviral transduction
The ZNF750 coding sequence (CDS) was subcloned into the pLenti-III-HA vector (ABM) derived from the pCMV6-entry-ZNF750 vector (Origene) using KpnI and EcoRV restriction enzymes (NEB). For lentiviral transduction, 2 × 106 HEK293T cells, seeded in a 10-cm dish, were transfected with 2.5 μg of either pLenti-III-HA empty vector (as control) or pLenti-III-HA-ZNF750 vector, 0.5 μg of VSV-G (ABM) and 2.5 μg of pCMV-ΔR8.2 (packaging vector). Forty-eight hours after transfection, the lentiviral supernatant was collected and used to transduce the MCF7, MDA-MB-231 and BT-549 cell lines overnight and in the presence of 8 μg/ml polybrene. Forty-eight hours after transduction, cells were selected with puromycin (1 μg/ml) for 7 days. ZNF750 overexpression was tested by qRT-PCR and western blot analysis.
Site-directed mutagenesis
For the mutagenesis of the zinc-finger motif, specific primers (Supplementary Table 1) were used to perform PCR on the pCDNA3.1-HA vector carrying the ZNF750 coding sequence. The PCR product was then digested using DpnI (NEB). The presence of the mutated nucleotides was confirmed through sequencing.
RNA isolation and real-time PCR
Total RNA from cells was isolated using an RNeasy mini kit (Qiagen) according to the manufacturer’s instructions. RNA samples were treated with RNase-free DNase I (Qiagen), and RNA was quantified using a NanoDrop spectrophotometer (Thermo Scientific). Total RNA was reverse transcribed using the GoScriptTM Reverse Transcription System (Promega) according to the manufacturer’s protocol. qRT-PCR was performed using GoTaq qPCR Mastermix (Promega) with SYBR Green ready mix. The specific primers used are listed in Supplementary table 1.
The expression of each gene was defined according to the threshold cycle (Ct), and relative expression levels were calculated by using the 2−ΔΔCt method after normalization to the expression of the housekeeping genes TBP and/or β-actin.
Western blot
Proteins were extracted with RIPA buffer containing cocktail inhibitors (Roche) and separated on SDS-polyacrylamide gels and then transferred onto PVDF membranes by the wet-transfer system. Membranes were blocked with TBS-0.1% Tween and 5% milk and incubated overnight with primary antibody, washed and then incubated with the appropriate horseradish peroxidase-conjugated secondary antibody. Proteins were visualized with the Super Signal chemiluminescence kit (Thermo Scientific). The following antibodies were used: anti-ZNF750 (1:1,000; Sigma HPA023012), anti-LAMB3 (1:200; Santa Cruz sc-133178), anti-CTNNAL1, (1:250; Abcam ab96184), anti-p63 (1:300; Abcam ab735), anti-KDM1A (1:1,000; Cell Signalling #2139), anti-HDAC1 (1:1,000; Abcam ab7028), and anti-β-actin (1:50,000; Sigma A-5441).
Co-immunoprecipitation
MDA-MB 468 cells were lysed in Triton buffer (50 mM Tris-HCl pH 7.5, 250 mM NaCl, 50 mM NaF, 1 mM EDTA 1 pH 8, 0.1% Triton) supplemented with proteases and phosphatases inhibitors. After preclearing for 1 h at 4 °C, immunoprecipitation was performed by incubating 1,5 mg of whole-cell protein extracts with 1.5 μg of anti- ZNF750 rabbit (Sigma HPA023012) and with anti-IgG antibody at the same concentration, with rocking at 4 °C overnight. The immune complexes were collected by incubation with protein A-sepharose 4 fast flow (G&E Healthcare) for 1 h and washed with Triton buffer. The beads were then resuspended in 25 μl SDS Laemmli sample buffer, subjected to SDS-PAGE (10% polyacrylamide) analysis, and electrotransferred onto PVDF membranes. Membranes were probed with primary antibodies as described above.
Microarray analysis
Seventy-two hours after ZNF750 depletion (with four different biological replicates for each condition), total RNA was extracted from MDA-MB-468 cells using TRIzol (Invitrogen). RNA was reverse transcribed, converted to cRNA, amplified, and labelled with a cyanine-3 dye using a Low-Input Quick Amp labelling kit from Agilent. Then, labelled cRNAs were hybridized to human gene expression microarrays (Agilent, catalogue number G4851B). The slides were washed using Agilent wash buffer reagents and scanned with a G2505C Agilent Microarray Scanner (scan control version A.8.4.1). The data were extracted using the Agilent Feature Extraction software (version 10.7.3.1) and analysed using Agilent GeneSpring GX software (version 12.1). To analyse significant differential expression, an unpaired Student’s t test with a Benjamini–Hochberg multiple testing correction was applied. mRNAs with a P value of less than or equal to 0.05 and a fold change of greater than 1.5 were considered to be both statistically and biologically significant.
Confocal microscopy
Immunofluorescence was performed as previously described [10]. Briefly, MDA-MB-468 cells were transfected and seeded into 12-well plates onto glass coverslips and scratched after 24 h. Six hours after the scratch, cells were fixed in 4% paraformaldehyde and then permeabilized with 0.5% Triton X-100. The following antibody was used: anti-GM130 (1:200; BD Transduction Laboratories). Actin filaments were stained with GFP-phalloidin for 30 min. Afterwards, nuclei were stained with DAPI. The coverslips were then mounted with aquapolymount antifading solution (Polysciences) onto glass slides and observed under a confocal microscope (Nikon A1).
Chromatin Immunoprecipitation (ChIP)
MDA-MB-468 or A253 cells (2 × 106) were crosslinked for 10 min in a solution containing 1% formaldehyde, and ChIP assays were performed using a MAGnify ChIP system (Invitrogen) according to the manufacturer’s instructions. Briefly, cells were lysed and then sonicated to obtain chromatin fragments of ~400 bp.
The lysates were immunoprecipitated using specific antibodies, anti-ZNF750 (Sigma HPA023012), anti-HDAC1 (Abcam ab7028), anti-KDM1a (Cell Signalling #2139), and anti-histone H3 (acetyl K9, K14, K18, K23, and K27; Abcam ab47915; or dimethyl K4; Abcam ab7766), and nonspecific IgG as a negative control. Collected DNA fragments were tested by real-time PCR using the primers listed in Supplementary table 1.
Migration and invasion assays
Breast cancer cell lines were transfected, and after 48 h, cell migration and invasion were assessed by using the CytoSelect™ 24-Well Cell Migration and Invasion Assay Combo Kit (Cell Biolabs, INC). Briefly, after transfection, cells were starved and then seeded on polycarbonate membrane inserts, and medium containing 10% FBS was added to the lower well of the migration plate. Cells were incubated for 12 h, non-migratory cells were removed with cotton-tipped swabs, and then cells on the insert were stained with cell stain solution. Pictures of at least three fields per insert were taken.
Statistical analysis
All results are expressed as the means ± s.ds. p < 0.05 was considered significant.
References
Yeo SK, Guan JL. Breast cancer: multiple subtypes within a tumor? Trends Cancer. 2017;3:753–60.
Polyak K. Breast cancer: origins and evolution. J Clin Investig. 2007;117:3155–63.
Sen GL, Boxer LD, Webster DE, Bussat RT, Qu K, Zarnegar BJ, et al. ZNF750 is a p63 target gene that induces KLF4 to drive terminal epidermal differentiation. Dev Cell. 2012;22:669–77.
Cassandri M, Smirnov A, Novelli F, Pitolli C, Agostini M, Malewicz M, et al. Zinc-finger proteins in health and disease. Cell Death Discov. 2017;3:17071.
Lin DC, Hao JJ, Nagata Y, Xu L, Shang L, Meng X, et al. Genomic and molecular characterization of esophageal squamous cell carcinoma. Nat Genet. 2014;46:467–73.
Boxer LD, Barajas B, Tao S, Zhang J, Khavari PA. ZNF750 interacts with KLF4 and RCOR1, KDM1A, and CTBP1/2 chromatin regulators to repress epidermal progenitor genes and induce differentiation genes. Genes Dev. 2014;28:2013–26.
Hazawa M, Lin DC, Handral H, Xu L, Chen Y, Jiang YY, et al. ZNF750 is a lineage-specific tumour suppressor in squamous cell carcinoma. Oncogene. 2017;36:2243–54.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6:pl1.
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012;2:401–4.
Tucci P, Agostini M, Grespi F, Markert EK, Terrinoni A, Vousden KH, et al. Loss of p63 and its microRNA-205 target results in enhanced cell migration and metastasis in prostate cancer. Proc Natl Acad Sci USA. 2012;109:15312–7.
Memmi EM, Sanarico AG, Giacobbe A, Peschiaroli A, Frezza V, Cicalese A, et al. p63 Sustains self-renewal of mammary cancer stem cells through regulation of Sonic Hedgehog signaling. Proc Natl Acad Sci USA. 2015;112:3499–504.
Su X, Napoli M, Abbas HA, Venkatanarayan A, Bui NHB, Coarfa C, et al. TAp63 suppresses mammary tumorigenesis through regulation of the Hippo pathway. Oncogene. 2017;36:2377–93.
Chakrabarti R, Wei Y, Hwang J, Hang X, Andres Blanco M, Choudhury A, et al. DeltaNp63 promotes stem cell activity in mammary gland development and basal-like breast cancer by enhancing Fzd7 expression and Wnt signalling. Nat Cell Biol. 2014;16:1004–15. 1001–13.
Jiang P, Enomoto A, Takahashi M. Cell biology of the movement of breast cancer cells: intracellular signalling and the actin cytoskeleton. Cancer Lett. 2009;284:122–30.
Millarte V, Farhan H. The Golgi in cell migration: regulation by signal transduction and its implications for cancer cell metastasis. ScientificWorldJournal. 2012;2012:498278.
Maziveyi M, Alahari SK. Cell matrix adhesions in cancer: The proteins that form the glue. Oncotarget. 2017;8:48471–87.
Ye X, Weinberg RA. Epithelial-mesenchymal plasticity: a central regulator of cancer progression. Trends Cell Biol. 2015;25:675–86.
Kreiseder B, Orel L, Bujnow C, Buschek S, Pflueger M, Schuett W, et al. alpha-Catulin downregulates E-cadherin and promotes melanoma progression and invasion. Int J Cancer. 2013;132:521–30.
Zhang Z, Yang XF, Huang KQ, Ren L, Zhao S, Gou WF, et al. The upregulated alpha-catulin expression was involved in head-neck squamous cell carcinogenesis by promoting proliferation, migration, invasion, and epithelial to mesenchymal transition. Tumour Biol. 2016;37:1671–81.
Zhang H, Pan YZ, Cheung M, Cao M, Yu C, Chen L, et al. LAMB3 mediates apoptotic, proliferative, invasive, and metastatic behaviors in pancreatic cancer by regulating the PI3K/Akt signaling pathway. Cell Death Dis. 2019;10:230.
Kinoshita T, Hanazawa T, Nohata N, Kikkawa N, Enokida H, Yoshino H, et al. Tumor suppressive microRNA-218 inhibits cancer cell migration and invasion through targeting laminin-332 in head and neck squamous cell carcinoma. Oncotarget. 2012;3:1386–1400.
Ciriello G, Gatza ML, Beck AH, Wilkerson MD, Rhie SK, Pastore A, et al. Comprehensive molecular portraits of invasive lobular breast. Cancer Cell. 2015;163:506–19.
Cancer Genome Atlas N. Comprehensive molecular portraits of human breast tumours. Nature. 2012;490:61–70.
Hu Z, Fan C, Oh DS, Marron JS, He X, Qaqish BF, et al. The molecular portraits of breast tumors are conserved across microarray platforms. BMC Genomics. 2006;7:96.
Wang Y, Zhang H, Chen Y, Sun Y, Yang F, Yu W, et al. LSD1 is a subunit of the NuRD complex and targets the metastasis programs in breast cancer. Cell. 2009;138:660–72.
Jain N, Rossi A, Garcia-Manero G. Epigenetic therapy of leukemia: an update. Int J Biochem Cell Biol. 2009;41:72–80.
Zhang W, Yu Y. The important molecular markers on chromosome 17 and their clinical impact in breast cancer. Int J Mol Sci. 2011;12:5672–83.
Courjal F, Theillet C. Comparative genomic hybridization analysis of breast tumors with predetermined profiles of DNA amplification. Cancer Res. 1997;57:4368–77.
Orsetti B, Nugoli M, Cervera N, Lasorsa L, Chuchana P, Ursule L, et al. Genomic and expression profiling of chromosome 17 in breast cancer reveals complex patterns of alterations and novel candidate genes. Cancer Res. 2004;64:6453–60.
Kaiser AM, Attardi LD. Deconstructing networks of p53-mediated tumor suppression in vivo. Cell Death Differ. 2018;25:93–103.
Engeland K. Cell cycle arrest through indirect transcriptional repression by p53: I have a DREAM. Cell Death Differ. 2018;25:114–32.
Sullivan KD, Galbraith MD, Andrysik Z, Espinosa JM. Mechanisms of transcriptional regulation by p53. Cell Death Differ. 2018;25:133–43.
Aubrey BJ, Kelly GL, Janic A, Herold MJ, Strasser A. How does p53 induce apoptosis and how does this relate to p53-mediated tumour suppression? Cell Death Differ. 2018;25:104–13.
Jia Y, Chen L, Jia Q, Dou X, Xu N, Liao DJ. The well-accepted notion that gene amplification contributes to increased expression still remains, after all these years, a reasonable but unproven assumption. J Carcinog. 2016;15:3.
Sugimoto Y, Roninson IB, Tsuruo T. Decreased expression of the amplified mdr1 gene in revertants of multidrug-resistant human myelogenous leukemia K562 occurs without loss of amplified DNA. Mol Cell Biol. 1987;7:4549–52.
Zhang P, He Q, Lei Y, Li Y, Wen X, Hong M, et al. m(6)A-mediated ZNF750 repression facilitates nasopharyngeal carcinoma progression. Cell Death Dis. 2018;9:1169.
Marinkovich MP. Tumour microenvironment: laminin 332 in squamous-cell carcinoma. Nat Rev Cancer. 2007;7:370–80.
Kwon OH, Park JL, Kim M, Kim JH, Lee HC, Kim HJ, et al. Aberrant up-regulation of LAMB3 and LAMC2 by promoter demethylation in gastric cancer. Biochem Biophys Res Commun. 2011;406:539–45.
Wang XM, Li J, Yan MX, Liu L, Jia DS, Geng Q, et al. Integrative analyses identify osteopontin, LAMB3 and ITGB1 as critical pro-metastatic genes for lung cancer. PLoS ONE. 2013;8:e55714.
Fan LC, Chiang WF, Liang CH, Tsai YT, Wong TY, Chen KC, et al. alpha-catulin knockdown induces senescence in cancer cells. Oncogene. 2011;30:2610–21.
Sweeney C, Bernard PS, Factor RE, Kwan ML, Habel LA, Quesenberry CP Jr., et al. Intrinsic subtypes from PAM50 gene expression assay in a population-based breast cancer cohort: differences by age, race, and tumor characteristics. Cancer Epidemiol Biomark Prev. 2014;23:714–24.
Acknowledgements
We would like to thank Dr Annalisa Conti for helpful comments and suggestions and K. Dudek and C. Jones for assistance in gene microarray analysis. This work has been supported by the Medical Research Council (to GM) UK, the Associazione Italiana per la Ricerca contro il Cancro (AIRC) (to GM; IG#20473 (2018–2022)), the AIRC (to EC; IG#22206 (2019–2023)), the AIRC (Start-Up to IA; ID 23219), the Ministero Salute, IDI-IRCCS (RC to EC), and the Italian Ministry of Foreign Affairs and International Cooperation (to EC). The work has also been supported by Regione Lazio through the LazioInnova Progetto Gruppo di Ricerca n 85-2017-14986 (to GM).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Cassandri, M., Butera, A., Amelio, I. et al. ZNF750 represses breast cancer invasion via epigenetic control of prometastatic genes. Oncogene 39, 4331–4343 (2020). https://doi.org/10.1038/s41388-020-1277-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-020-1277-5
- Springer Nature Limited
This article is cited by
-
A BRCA2 germline mutation and high expression of immune checkpoints in a TNBC patient
Cell Death Discovery (2023)
-
A primary luminal/HER2 negative breast cancer patient with mismatch repair deficiency
Cell Death Discovery (2023)
-
Single-cell Analysis Reveals Inter- and Intratumour Heterogeneity in Metastatic Breast Cancer
Journal of Mammary Gland Biology and Neoplasia (2023)
-
Long non-coding RNAs affecting cell metabolism in cancer
Biology Direct (2022)
-
Catulin reporter marks a heterogeneous population of invasive breast cancer cells with some demonstrating plasticity and participating in vascular mimicry
Scientific Reports (2022)