Abstract
Pyruvate kinase muscle isozymes (PKMs) have crucial roles in regulating metabolic changes during carcinogenesis. A switch from PKM1 to PKM2 isoform was thought to lead to aerobic glycolysis promoting carcinogenesis, and was considered as one of the cancer signatures. However, recent evidence has argued against the existence of PKM isoform switch and related metabolic effects during cancer progression. We compared the effects of PKM1 and PKM2 in cell invasiveness and metastasis of pancreatic ductal adenocarcinoma (PDAC). Both PKM1 and PKM2 expression affected cell migration and invasion abilities of PDAC cells, but only knockdown of PKM2 suppressed metastasis in a xenograft model. By comparing the established PKM2 mutants in the regulation of cell invasion, we found that PKM2 may control cell mobility through its protein kinase and phopho-peptide binding abilities. Further survey for PKM2-associated proteins identified PAK2 as a possible phosphorylation target in PDAC. In vitro binding and kinase assays revealed that PKM2 directly phosphorylated PAK2 at Ser20, Ser141, and Ser192/197. Knockdown of PKM2 decreased PAK2 protein half-life by increasing ubiquitin-dependent proteasomal degradation. Moreover, we identified PAK2 as an HSP90 client protein and the mutation at Ser192/197 of PAK2 reduced PAK2–HSP90 association. Knockdown of PAK2 diminished in vitro cell mobility and in vivo metastatic ability of PKM2 overexpressed PDAC cells. PKM2 and PAK2 protein expression also positively correlated with each other in PDAC tissues. Our findings indicate that PKM2–PAK2 regulation is critical for developing metastasis in PDAC, and suggest that targeting the PKM2/HSP90/PAK2 complex has a potential therapeutic value in this deadly disease.
Similar content being viewed by others
Introduction
The rapid proliferating property of cancer cells makes it highly demand the glycolysis intermediates to support cell growth and generate biomass [1]. Pyruvate kinases (PKs) are rate-limiting glycolytic enzymes catalyzing the phosphotransfer reaction between phosphoenolpyruvate (PEP) and adenosine diphosphate (ADP), producing pyruvate and adenosine triphosphate (ATP). Theoretically, decrease of PK activity could provide most abundant glycolysis intermediates to supply biosynthetic pathways. Indeed, PKs activity was under tightly control at different levels. Four isoforms of PKs are identified and expressed in tissues with different metabolic demands. PKL and PKR derived from the PKLR gene are regulated by tissue-specific promoters [2], while PKM1 and PKM2 derived from the PKM gene are regulated by alternative splicing [3]. PKL has the lowest affinity to PEP and is expressed in tissues with high rates of gluconeogenesis, such as liver and kidney. PKR also has low PEP affinity and is exclusively expressed in red blood cells [2]. PKM1 is the only PK without allosteric regulations, and expressed in tissues with high energy consumption, such as muscle and brain. PKM2 is expressed in early embryonic cells, normal proliferating cells, and cancer cells [4]. Because only PKM2 can change between a highly active tetrameric and an inactive dimeric forms during glycolysis, PKM2 has the most complicated ways of regulation. Unique post-translational modification and allosteric regulation of PKM2 in cancer cells are also well-documented [5,6,7].
In recent years, more and more reports have documented the non-glycolytic functions of PKM2. PKM2 can act as a transcriptional coactivator of HIF-1α and β-catenin in the nucleus to induce the expression of downstream metabolic and proliferation-related genes [8, 9]. PKM2 can be a protein kinase in the nucleus and directly phosphorylates STAT3 and histone H3 to promote tumor transformation [10]. PKM2 also can phosphorylate the spindle checkpoint protein Bub3, and regulate correct chromosome segregation of tumor cells [11]. Besides, mitochondrial PKM2 is capable to phosphorylate Bcl-2, prevent Bcl-2 degradation, and thus inhibit apoptosis in glioblastoma [12]. Recently, we have also identified a novel function of PKM2, which works together with HSP90 to stabilize mutated EGFR and contribute to EGFR-dependent tumorigenesis of non-small cell lung cancer (NSCLC) cells [13]. These findings not only reveal the non-glycolytic functions of PKM2 but also support the roles of PKM2 in regulating rapid cell proliferation and promoting tumorigenesis.
Paradoxically, isoform-specific deletion of PKM2 in a mouse model of breast cancer does not reduce or delay but accelerates tumor formation, suggesting that PKM2 is dispensable for tumor growth. Tumors from those PKM2 knockout mice express variable amounts of PKM1, which inversely correlate with cell proliferation. Furthermore, despite selection for PKM2 during cell line establishment, there was no selective pressure to retain PKM2 expression in allograft tumors [14]. Besides, both the isoform analysis with The Cancer Genome Atlas (TCGA) data sets and absolute quantification by mass spectrometry do not support that PKM2 is specific for proliferating tissue. These data argue against the isoform switch hypothesis from PKM1 to PKM2 during tumorigenesis [15,16,17].
The aforementioned contradictory results indicate that the roles of PKM2 in cancer cells remain elusive. In this study, we attempted to evaluate the role of PKM1 and PKM2 in regulating invasiveness of pancreatic ductal adenocarcinoma (PDAC). We have found that both PKM1 and PKM2 regulate cell migration and invasion in vitro; however, only PKM2 overexpression promotes metastasis of PDAC cells in vivo. We further evaluated the mechanistic regulation of PKM2 and identified its protein kinase substrate PAK2 to be responsible for related PDAC cancer invasion and metastasis. Besides, PKM2 could maintain PAK2 stability via phosphorylation-dependent HSP90 recruitment. PKM2 and PAK2 expression also correlate with each other in human PDAC specimens. Our results indicate a pivotal role of PKM2 in the regulation of PDAC metastasis, suggesting that PKM2 overexpression may be a more critical event during the tumor progression rather than tumor initiation.
Results
Both PKM1 and PKM2 regulate migration and invasion of PDAC cells
To assess the roles of PKM1 and PKM2 in PDAC progression, we first knocked down PKM1 and PKM2 with subtype-specific shRNAs in two highly invasive PDAC cells (Miapaca-2 and BxPC-3) (Fig. 1a, left panel). Both PKM1 and PKM2 depletion resulted in the decrease of cell migration and invasion abilities, but the suppressive effects were more prominent in PKM2-depleted cells (Fig. 1b and c). We next overexpressed PKM1 and PKM2 in an immortalized pancreatic epithelial cell (hTERT-HPNE) and a lowly invasive PDAC cell (AsPC-1) (Fig. 1a, right panel). In hTERT-HPNE cells, forced expression of PKM1 resulted in an increase of cell migration ability, while invasion ability remained unchanged. In AsPC-1 cells, forced expression of PKM1 induced a significantly increase of invasion ability but to a lesser extent on cell migration. On the contrary, forced expression of PKM2 showed a more consistent effect on increasing both migration and invasion abilities of both hTERT-HPNE and AsPC-1 cells (Fig. 1d and e). These results suggested a more important role of PKM2 in the regulation of cellular invasiveness during PDAC progression. Consistently, results from wound healing assays also supported a more critical role of PKM2 in regulating cell mobility (Supplementary Figure S1).
PKM2 expression promotes cell migration and invasion abilities of PDAC cells. a Left: Western blot analysis of PKM1 and PKM2 in PKM1/PKM2 knockdown Miapaca-2 or BxPC-3 cells. Right: Western blot analysis of PKM1 and PKM2 in HA-PKM1/PKM2 overexpressed hTERT-HPNE or AsPC-1 cells. b Transwell migration assays of PKM1 or PKM2 knockdown PDAC cells. Left: Representative micrographs. Right: Quantitative plot shown as % of control. c Transwell invasion assays of PKM1 or PKM2 knockdown PDAC cells. Left: Representative micrographs. Right: Quantitative plot shown as % of control. d Transwell migration assays of HA-PKM1 or HA-PKM2 overexpressed PDAC cells. Left: Representative micrographs. Right: Quantitative plot shown as % of control. e Transwell invasion assays of HA-PKM1 or HA-PKM2 overexpressed PDAC cells. Left: Representative micrographs. Right: Quantitative plot shown as % of control. Scale bar, 100 μm. Statistical significance was determined by paired, two-tailed Student's t-test. Error bars, SD. *p < 0.05; **p < 0.01
PKM2 promotes distant metastasis in a spontaneous metastasis model
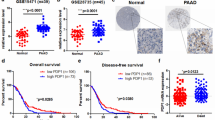
PKM2 expression was associated with survival outcome of PDAC patients [18, 19]. However, since PDAC patients diagnosed with metastatic disease usually did not receive surgery, the role of PKM2 during metastasis was hard to define by surgical specimens. We therefore examined the possible involvement of PKM1 and PKM2 in this vital process by using an orthotopic xenograft model. 106 Miapaca-2 cells stably expressed luciferase and scrambled PKM1- or PKM2-specific shRNA were injected into the pancreas of NOD/SCID mice. After seven weeks of inoculation, the primary tumor was slightly, but not significantly, smaller in the group with Miapaca-2/shPKM2 cells in comparison with the control group by measuring luciferase activity and tumor weight (Fig. 2a). In contrast to the primary tumor, metastases in the lung, liver, and mesentery were all significantly reduced in the group with Miapaca-2/shPKM2 cells compared to the group with control cells. Metastatic signals were slightly higher in mice implanted with Miapaca-2/shPKM1 cells (Fig. 2b–d). Furthermore, four out of six and four out of five mice injected with control and PKM1-depleted Miapaca-2 cells respectively form detectable amount of ascites when sacrificed, while no ascites formation was noted in the group with Miapaca-2/shPKM2 cells (Fig. 2h). These results suggested PKM2 as an important role in promoting metastases of PDAC in mice.
PKM2 promotes metastasis of PDAC cells in an orthotopic xenograft model. a Miapaca-2 stable clones with shCtrl, shPKM1, or shPKM2 expression were inoculated into the pancreas of 8-week SCID mice for 7 weeks. Left: Representative graphs show luciferase activity taken at weeks 1 and 7. Right: Primary tumor weight was measured immediately after sacrificed. b Top: Representative graphs show luciferase activity in the lung between groups taken immediately after sacrificed. Bottom: Quantification of lung metastases shown as photon radiance (p/s/cm2/sr). c Top: Representative graphs show luciferase activity in the liver between groups taken immediately after sacrificed. Bottom: Quantification of liver metastases shown as photon radiance (p/s/cm2/sr). d Top: Representative graphs show luciferase activity in the mesentery between groups taken immediately after sacrificed. Bottom: Quantification of mesentery metastases shown as photon radiance (p/s/cm2/sr). e Quantification of ascites volume (μl) taken after sacrificed between groups. Statistical significance was determined by paired, two-tailed Student's t-test. * p < 0.05
PKM2 associates and phosphorylates PAK2
PKM2 is a multifunctional protein with both metabolic and non-metabolic activities [20]. To evaluate the molecular mechanism of PKM2-regulated cellular invasiveness and tumor metastasis, we compared the effects of PKM2 overexpression among wild-type and mutants by cell invasion assay. Forced expression of PKM2R399E, a dimeric mutant, showed similar invasion-promoting effect as wild-type PKM2. Since dimeric PKM2 decreased its PK activity and could act as a protein kinase [10], the result suggested a non-glycolytic mechanism may exist. Moreover, a kinase-dead mutant PKM2K367M and a phospho-peptide binding mutant PKM2K433E suppressed invasion abilities of both hTERT-HPNE and AsPC-1 cells, further supporting the involvement of PKM2 protein kinase activity in regulating cell invasion (Fig. 3a). For identifying novel PKM2 substrates as mediators of cell invasion, we started with a list of 152 potential PKM2 substrates derived from an in vitro phosphoproteomic assessment with use of a human protein microarray including 9479 proteins [21]. Next, we cross-referenced the listed proteins with the Ingenuity Pathway Analysis database and yielded 46 proteins functionally annotated as “invasion of cells”. The remaining proteins were rescreened with the overall survival analysis of PDAC patients in the TCGA database. We identified six proteins whose expression levels correlated with decreased overall survival in PDAC patients (Supplementary Figure S2a). On testing whether PKM2 involved in the phosphorylation process, only PAK2, one member of p21-activating kinases (PAKs) mediating multiple oncogenic effects including cell invasion and metastasis [22], increased on overexpressing PKM2 and decreased on knocking down PKM2 among the hit proteins and PAKs (Fig. 3b, Supplementary Figure S2b). Phosphorylated PAK2 expression levels at serine 20, 141, 192/197, and threonine 402 changed accordingly under PKM2 manipulations in some pancreatic cell lines examined (Fig. 3c). PKM1 expression also affected PAK2 expression and phosphorylation, although to a lesser extent in comparison with PKM2 (Fig. 3b and c). We next confirmed PAK2 as a potential PKM2 substrate by examining their interaction. Endogenous interaction between PKM2 and PAK2 was illustrated by co-immunoprecipitation (Fig. 3d). Both PKM1 and PKM2 could bind to PAK2, but the PKM1–PAK2 interaction was much less prominent in comparison with PKM2–PAK2 interaction (Supplementary Figure S3). Ni-pull down with recombinant His-tagged PKM2 and GST-tagged PAK2 also showed a direct interaction between them (Fig. 3e). On examining the phosphorylation sites of PAK2 catalyzed by PKM2, we found that recombinant PKM2 phosphorylated protein phosphatase 1 (PP1) pre-treated PAK2 at the serine 20, 141, and 192/197, but not threonine 402 sites (Fig. 3f). When co-transfecting PAK2 or its phosphorylation mutants with the increasing amount of PKM2 plasmids, the protein expression levels of wild-type PAK2, PAK2S20A, and PAK2S141A increased in accordance with increasing PKM2 levels. Only the PAK2S192/197A expression level was almost constant, suggesting the serine 192/197 as the most important phosphorylation site for maintaining PAK2 protein levels on interacting with PKM2 (Fig. 3g).
PKM2 phosphorylate PAK2. a hTERT-HPNE (left) or AsPC-1 (right) cells were transfected with an empty vector (Ctrl vector), wild-type PKM2 (wt), or mutant PKM2s (K433E, R399E, or K367M). Top: Western blot analysis of HA-PKM2 expression. Bottom: Transwell invasion assays of hTERT-HPNE (left) or AsPC-1 (right) cells. b Western blot analysis of PAKs in PKM1/2 knockdown or overexpressed pancreatic cells. c Western blot analysis of phosphorylated PAK2 including pSer20, pSer141, pSer192/197 and pThr402 in PKM1/2 knockdown or overexpressed pancreatic cells. d Lysates from BxPC-3 cells were subjected to co-immunoprecipitation using PKM2 or normal IgG antibodies followed by western blot analysis of PKM2 and PAK2. e Recombinant GST-PAK2 and His-PKM2 were incubated for 16 h followed by Ni-pull down and western blot analysis of PKM2 and PAK2. f In vitro kinase assay was performed using His-PKM2 and protein phosphatase 1 (PP1) pre-treated recombinant GST-PAK2. PAK2 phosphorylation was detected by phospho-specific antibodies. g Western blot analysis of Flag-PAK2 in AsPC-1 cells co-transfected with 1 μg Flag-tagged wild-type PAK2 (wt), or mutant PAK2s (S20A, S141A, or S192/197A) and an increasing amount (0, 1, 3 μg) of HA-PKM2 plasmid
PKM2 maintains PAK2 protein stability via phosphorylation-dependent HSP90 recruitment
Though PAK2 protein expression level would follow the change of the PKM2 protein in the same direction, PAK2 mRNA levels remained unchanged in the absence of PKM2 expression (Supplementary Figure S4). The estimated PAK2 protein half-life was 4.5 h after PKM2 depletion, while the half-life was longer than 24 h in control cells (Fig. 4a). These results suggested PKM2 may modulate PAK2 expression by post-transcriptional regulation. We attempted to verify whether PKM2 enhanced PAK2 protein stability by preventing protein degradation. Decreased PAK2 protein expression after knocking down PKM2 was rescued by the proteasome inhibitor, MG132 (Fig. 4b). Consistent to the above findings, the poly-ubiquitination of PAK2 also increased in PKM2-depleted Miapaca-2 cells in comparison with the control cells (Fig. 4c). Altogether, these data suggested that PKM2-mediated PAK2 phosphorylation at serine 192/197 stabilized PAK2 protein by preventing proteasomal degradation. We have previously identified PKM2 as a co-chaperone of HSP90 to increase mutant EGFR protein stability [13]. We found that HSP90 also involved in PAK2 protein stability since the PAK2 protein expression level decreased similar to EGFR and AKT, two known HSP90 clients, after the treatment of an HSP90 inhibitor, 17-allyl-aminogeldanamycin (17-AAG) (Fig. 4d). Recombinant GST-tagged PAK2 could pull down His-tagged HSP90 in an in vitro condition (Fig. 4e). Although PKM2 still bound to PAK2S192/197A, the associated HSP90 in the PKM2–PAK2 immunocomplex diminished significantly in comparison with wild-type PAK2 (Fig. 4f). These results indicated that PKM2–PAK2 phosphorylated at serine 192/197 was responsible for HSP90 recruitment and thus preventing proteasomal degradation of PAK2. Besides, phosphorylating serine 20 and 141 sites of PAK2 by PKM2 might further strength the association between PKM2 and PAK2 since the amount of PKM2 decreased on co-immunoprecipitating with either PAK2S20A or PAK2S141A in comparison with wild-type PAK2 (Supplementary Figure S5).
PKM2 protect PAK2 protein from proteasomal degradation via HSP90 recruitment. a Protein stability of PAK2 proteins in Miapaca-2 cells expressing shCtrl or shPKM2. Cells were treated with 20 μg/ml cycloheximide (CHX) at indicated intervals and analyzed in western blot analysis. Protein expression levels were normalized with an internal control and quantified as a ratio compared with hour 0. b Western blot analysis of PAK2 and PKM2 in Miapaca-2 cells expressing shCtrl or shPKM2 with or without 10 μM MG132 treatment for 6 h before protein harvested. c Ubiquitination assay of PAK2 in response to PKM2 knockdown. Miapaca-2 cells were transiently transfected with HA-ubiquitin. Ubiquitinated PAK2 was immunoprecipitated (IP) and subjected to western blot analysis with the anti-HA antibody. d Western blot analysis of EGFR, PAK2, or AKT in AsPC-1 cells with or without 100 nM 17-AAG treatment for 24 or 48 h. e Recombinant His-HSP90 was incubated with or without recombinant GST-PAK2 and pull down with Glutathione-Sepharose beads followed by western blot analysis. f Miapaca-2 cells were transfected with HA-PKM2, Flag-PAK2, or Flag-PAK2-S192/197A as indicated followed by immunoprecipitation with the anti-HA antibody. PKM2-associated HSP90 and Flag-PAK2 were analyzed by western blotting. Ten percent input was used as a loading control
PAK2 is responsible for PKM2-regulated cell invasiveness and tumor metastasis
As we confirmed PAK2 to be a PKM2 substrate, the possible role of PAK2 in mediating PKM2-regulated cellular invasiveness of PDAC cells should be clarified. We knocked down PAK2 in AsPC-1 cells overexpressed with PKM2, and the increased migration and invasion abilities induced by PKM2 overexpression were suppressed (Fig. 5a). We then generated luciferase-expressing AsPC-1 cells with the same grouping strategies in the orthotopic xenograft model. The primary tumor weight did not reach significant difference between groups, although PKM2 overexpression and PAK2 knockdown groups slightly increased and decreased tumor weight respectively (Fig. 5b). Increased number of metastatic nodules in the lung, liver, and mesentery were observed in the PKM2 overexpressed group in comparison with the control group, whereas few nodules were detected in the PAK2 knockdown group. Most importantly, PAK2 depletion significantly reduced metastasis induced by PKM2 overexpression in the lung, liver, and mesentery (Fig. 5c–e). We also observed one and two mice forming a detectable amount of ascites in the control and PKM2 overexpression group, respectively, but none in the other groups (Fig. 5f). Collectively, these findings suggested that induction of PAK2 expression is one important pathway by which PKM2 exerted its pro-metastatic activity.
PAK2 mediates PKM2-enhanced metastatic abilities of PDAC cells. a Top: Western blot analysis of PAK2 and PKM2 in AsPC-1 cells stably expressed HA-PKM2, shPAK2, and corresponding controls. Bottom: Transwell migration and invasion assays shown as % of control. b AsPC-1 stable clones as indicated were inoculated into the pancreas of 8-week SCID mice for 7 weeks. Primary tumor weight was measured immediately after sacrificed. c Top: Representative graphs show luciferase activity in the lung between groups as indicated taken immediately after sacrificed. Bottom: Quantification of lung metastases shown as photon radiance (p/s/cm2/sr). d Top: Representative graphs show luciferase activity in the liver between groups as indicated taken immediately after sacrificed. Bottom: Quantification of liver metastases shown as photon radiance (p/s/cm2/sr). e Top: Representative graphs show luciferase activity in the mesentery between groups as indicated taken immediately after sacrificed. Bottom: Quantification of mesentery metastases shown as photon radiance (p/s/cm2/sr). f Quantification of ascites volume (μl) taken after sacrificed between groups as indicated. Statistical significance was determined by paired, two-tailed Student's t-test. *p < 0.05; **p < 0.01
Co-expression of PKM2 and PAK2 in PDAC
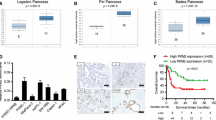
As PAK2 mediated the pro-metastatic effect of PKM2 in PDAC cells, we next investigated this relationship in clinical samples. We examined tumor specimens from 54 PDAC patients for expression of PKM2 and PAK2, and observed a significantly positive correlation between them (p < 0.0005) (Fig. 6a and b). Moreover, although PKM2 and PAK2 alone could be used as prognostic markers for PDAC patients (p = 0.007 and 0.003, respectively), co-existence of PKM2 and PAK2 showed a more significant prognostic value (p < 0.0001) in predicting overall survival of PDAC patients (Supplementary Figure S6). These data suggested that there is a significantly clinical association between PKM2 and PAK2, further supporting the experimental findings of biochemical and functional linkage between PKM2 and PAK2 (Fig. 6c).
Co-expression of PKM2 and PAK2 in human PDAC. a Representative pictures of IHC staining with anti-PKM2 or anti-PAK2 antibody on PDAC and adjacent normal tissues. Scale bars, 100 μm. b Analysis of the correlation between PKM2 and PAK2 levels on the basis of immunohistochemistry results from 54 PDAC tissue samples. Statistical significance was determined by chi-squared test. c A schematic diagram illustrating the regulation of PAK2 protein stability via PKM2-mediated S192/197 phosphorylation in PDAC cells
Discussion
PKM2 is recognized initially as the key isoform of PK contributing to aerobic glycolysis, the Warburg effect, which is critical for supporting tumor growth after switching from PKM1 [23]. Recently, more and more data suggest that the abundance of PKM2 in cancer cells plays much more roles in cancer initiation and progression beyond regulating glucose metabolism [11, 20]. In this study, we have found PKM2 expression played an important role in tumor invasion of PDAC. In comparison to PKM1, PKM2 had a greater and more consistent impact to migration/invasion abilities of PDAC cells. The metastasis promoting effect of PDAC cells in the mice model was evident in PKM2 but not in PKM1. These results supported that PKM2 was not only a regulator of glucose metabolism but also a crucial mediator of tumor invasiveness in PDAC.
Plasma PKM2 has been identified as a diagnostic biomarker with comparable sensitivity and specificity to serum CA19-9 in PDAC [24,25,26]. Consistent with our findings, Goonetilleke et al. [26] observed that elevated plasma PKM2 strongly correlated with the metastatic disease and strongly predicted survival in PDAC patients. However, there are contradictory results about clinical impacts of tissue PKM2 levels on overall survival in PDAC patients [18, 19, 27,28,29,30,31]. While most clinical evaluations support that PDAC patients with high PKM2 expression in tumors have worse overall survival in comparison to those with low expression [28,29,30,31], Lockney et al. [19] reported that positive PKM2 expression predicted better overall survival in PDAC patients. Our analysis of TCGA database also supported PKM2 as a poor prognostic marker in PDAC. PKM2 may play an important role in promoting PDAC cell invasiveness and consequently result in poorer overall survival.
The metastatic signals were slightly higher in mice with PKM1-depleted cell implantation in comparison with the control group and the PKM2-depleted cell group although it did not reach statistical significance (Fig. 2b–d). Since the effects after differential expression of PKM1 among in vitro migration/invasion assays were less effective than those of PKM2, it was possible that the migratory/invasive effects of PKM1 were not powerful enough to increase tumor metastasis in in vivo experiments. Besides, previous reports have shown forced expression of PKM1 inhibits the growth of xenograft tumors either with or without suppression of PKM2 expression [23, 32]. Constitutively forced expression of PKM1 in mammary tumors-derived cells with isoform-specific deletion of PKM2 can suppress tumor formation in vivo [14]. These data suggest that increased PKM1 levels are detrimental for cancer cell proliferation. The slightly increased metastatic signals in the PKM1 depletion group may be due to the proliferative benefit after reduced PKM1 expression in cancer cells. More sophisticated studies are required to define the oncosuppressive role of PKM1 in the future.
HSP90 is an important chaperone protein responsible for regulating proteostasis under both physiological and stress conditions [33]. Though heat-shock proteins are differentially expressed in human gastrointestinal cancers, HSP90 is constitutively expressed in these gastrointestinal cancers including PDAC [34]. Blocking HSP90 with 17-dimethylaminoethylamino-17-demethoxygeldanamycin (17-DMAG) can inhibit epidermal growth factor receptor (EGFR)-dependent signaling in gastric cancer and reduce the growth and vascularization of gastric cancer in a murine xenograft model [35]. Our recent study in NSCLC cells showed PKM2 can interact directly with EGFR and HSP90, and stabilize EGFR protein expression by maintaining its binding with HSP90 and co-chaperones. The process of stabilization relied on dimeric form of PKM2 without phosphorylation. When PKM2 was forced to express in its tetrameric form, the half-life of mutant EGFR decreased [13, 27]. In this study, we have also identified HSP90 as a stabilizer for PAK2 protein expression. In contrast to phosphorylation-independent regulation during EGFR stabilization, PAK2 needs phosphorylation by PKM2 to facilitate the association with HSP90 and subsequent protein stabilization.
Recently, Li and colleagues also have demonstrated that PKM2 promoted cell invasion in PDAC. However, they concluded that PKM2 regulated PDAC cell invasion along with survival by enhancing aerobic glycolysis, the Warburg effect. The statement was based on the results after 2-deoxy-d-glucose (2-DG) treatment, which diminished the difference of invasion between cells with and without downregulating PKM2 [31]. In this study, we used a series of PKM2 mutants to demonstrate that PKM2 may regulate cell invasion of PDAC via its protein kinase activity (Fig. 3a). We further demonstrated that PKM2 may promote PDAC cell invasion through a non-metabolic mechanism with phosphorylation and stabilization of PAK2. The inconsistent results may be related to the different strategies used between experiments. 2-DG inhibits hexokinase and completely blocks glycolysis from the first reaction. But 2-DG has wide-ranging pharmacological properties in addition to metabolic blockade of aerobic glycolysis. For example, 2-DG can interrupt the N-linked glycosylation of proteins, which may affect the activities of invasion-regulated molecules [36, 37]. Collectively, our results based on PKM2 mutants and functional rescue experiments demonstrated a novel non-metabolic mechanism involving PAK2 by which PKM2 regulated PDAC cell invasion.
Protein kinase activity of PKM2 is well documented in recent years. PKM2 could transfer the phosphate group of PEP to serine, threonine, or tyrosine residues of protein substrates. Although recent evidence provided by Hosios et al. [38] argue against the protein kinase function for PKM2, more and more phosphorylated protein substrates of PKM2 have been identified by different laboratories. PKM2 can phosphorylate histone H3, Bcl-2, and SNAP-23 at their serine or threonine residues [12, 39, 40], and phosphorylate STAT3, Bub3, MLC2 at their tyrosine residues [10, 11, 41]. Like the tightly controlled PK activity of PKM2, the protein kinase activity of PKM2 may also be tightly regulated. Yang et al. [39] showed that only the PKM2 purified from cancer cells, but not purified from E. coli, phosphorylated histone H3 in vitro. Phosphorylation of Bcl2 by PKM2 was found to be stimulated by oxidative stress [12]. Keller et al. [21] reported that the PKM2-SAICAR interaction was necessary and sufficient to induce robust protein kinase activity from PKM2 in vitro and in cancer cells. They also identified more than 100 potential substrates, including PAK2, 6, and 7, by proteomic screening with SAICAR-associated PKM2. Thus, protein kinase activity of PKM2 may be dependent on alternative modifications or ligand binding. Although the detailed activation process of PKM2 protein kinase activity in PDAC remains to be elucidated, our study confirmed that PAK2 was phosphorylated by PKM2 in PDAC cells, and the phosphorylation sites included Ser20, Ser141, and Ser192/197.
PAK2 is a serine–threonine kinase with a major role in regulating cell mobility. PAK2 is activated by binding with the small G protein Cdc42 or Rac, leading to autophosphorylation at multiple Ser/Thr sites, including Ser20, Ser139, Ser141, Ser144, Ser192/197, Thr402, Thr421, and Thr423 [42, 43]. PAK2 can also be activated by an alternative pathway to generate the constitutively active PAK2p34 fragment through proteolytic cleavage by caspase-3 or similar proteases. The levels of PAK2p34 are tightly regulated through ubiquitination and proteasomal degradation [44]. In this study, we have identified Ser20, Ser141, and Ser192/197 of PAK2 to be phosphorylated by PKM2 in vitro, suggesting that PKM2 is a novel regulator of PAK2. Furthermore, we have found that PKM2-mediated phosphorylation of Ser192/197 to be critical for maintaining PAK2 level in PDAC cells. Ser192/197 phosphorylation facilitated HSP90 association to PAK2 and thus prevented ubiquitination and proteasomal degradation of PAK2. To our knowledge, the proteasomal regulation of the full-length PAK2 has never been reported. Besides, our data also suggest that the phosphorylation of Ser141 and/or Ser20 may further strengthen the PKM2–PAK2 interaction. These results have established a novel regulatory axis that PKM2 can regulate PAK2 activation through control of protein stability.
Our data have shown PKM2 to be a critical regulator of cell invasion and metastasis of PDAC. Through phosphorylation of PAK2 and stabilization of PAK2 by facilitating PAK2–HSP90 association, PKM2 can increase the cell invasion ability and promote tumor metastasis of PDAC. All these results suggest that the regulatory interplay between PKM isoforms contributes substantially to cancer progression of PDAC, and targeting the PKM2/HSP90/PAK2 complex may be beneficial for disease control of PDAC.
Methods
Patients, specimens, and immunohistochemical staining
This study included 54 patients with newly diagnosed PDAC undergoing surgical resection. Patients receiving preoperative anticancer therapies were excluded. Formalin-fixed and paraffin-embedded tumor tissues were sectioned and processed for standard immunohistochemical staining procedures. The primary antibodies for PAK2 (1:150) and PKM2 (1:400) were used to detect the expression of PAK2 and PKM2 in the tumor tissues. The results of immunohistochemical staining were scored by multiplying the percentage score by the intensity score. The percentage score 1–4 was classified using extent of tumor cells stained (1, <25%; 2, 25–50%; 3, 50–75%; 4, >75%). The intensity score 1–3 was classified using intensity of tumor cells stained (1, weak; 2, moderate; 3, strong). Institutional Review Board approval was obtained to procure and analyze the tissues used in this study.
Cell culture
Human cell lines AsPC1, Miapaca-2, BxPC-3, and hTERT-HPNE were purchased from the American Type Culture Collection (authenticated using STR profile analysis, Manassas, VA, USA). These cell lines were cultured in RPMI-1640 medium containing 10% fetal bovine serum in a humidified atmosphere containing 5% CO2 at 37 °C. All cell lines were confirmed to be Mycoplasma negative.
Antibodies and reagents
Antibodies and reagents used were as follows: PKM1, PKM2, PAK1/2/3/4, Ser20-/Ser141-/Thr423-/Ser192/197-phosphorylated PAK2, and HSP90 (Cell Signaling Technology Inc., Danvers, MA), PAK6/7 (Origene Technology, Inc., Rockville, MD).
Western blot analysis
Protein lysates were prepared as previously described [13]. Western blot analysis was performed with primary antibodies for PKM1, PKM2, PAK1/2/3/4/6/7, Ser20-/Ser141-/Thr423-/Ser192/197-phosphorylated PAK2, HSP90, and GAPDH.
Construction of plasmids and transient transfection
Plasmids bearing HA-PKM2, K433E-PKM2, and K367M-PKM2 were modified from the plasmids we constructed previously [13]. pECE-PAK2 was purchased from Addgene, and mutant constructs were obtained by using QuikChange® Site-Directed Mutagenesis Kit. The shRNA constructs against PAK2 and PKM2 were purchased from National RNAi Core Facility. The target sequences were: shPAK2: 5′-CTC TAG GAA CCA AAG TGA TTT-3′, shPKM2: 5′-CCA TAA TCG TCC TCA CCA A-′, shPKM1: 5′-CGC AAG CTG TTT GAA GAA CTT-3′. The expression vectors were transfected transiently into test cells with Lipofectamine 2000 transfection reagents.
Two-chamber migration/invasion assay
Cell migration and invasion abilities were assessed by the modified two-chamber migration assay. 106 cells were seeded into the upper chamber, and migrated cells in the lower chamber were counted after 24 h of incubation. We used the upper chambers coated with 40 μg matrigel and counted cells 48 h after incubation for invasion assays. Migrated cells at the bottom of the membrane were fixed with 20% methanol and stained with 0.2% crystal violet. Quantification analysis was done by counting those stained cells. Each data point is derived from three independent experiments and is shown as mean ± SD.
In vitro phosphatase and PKM2 kinase assay
For in vitro phosphatase assays, 1 μg GST-PAK2 protein (Invitrogen) was incubated in 50 µl containing 1× protein metallophosphatases supplemented, 1 mM MnCl2 and PP1 (NEB) for 1 h at 30 °C. Phosphatase reactions were stopped with heated at 65 °C for 30 min and then GST-PAK2 were further immunoprecipitated by glutathione-sepharose beads. After three times wash, the protein was add with PK enzyme mix (Sigma, MAK072) for 15 min at 25 °C for PKM2 kinase assay. All samples were analyzed by western blotting.
In vitro binding assay
GST-PAK2 fusion proteins were incubated with His-PKM2 or His-HSP90 proteins, in ice-cold protein binding buffer (50 mM NaH2PO4, 500 mM NaCl, 1% NP-40) at 4 °C with gentle rocking overnight, then add glutathione-sepharose beads for an hour incubation. After the beads were washed five times with 1 ml binding buffer, bound proteins were resolved by immunoblotting with indicated antibodies.
Animal study of orthotopic implantation
Seven-week-old male SCID mice (NOD.CB17-Prkdcscid mice) were randomly grouped and acclimatized over 1 week while they were caged in groups of five. After indicated treatment, 2 × 105 cells were inoculated orthotopically into the pancreas of test mice. The mice were sacrificed 7 weeks later. Primary tumors, ascites, and metastases were analyzed. All mouse studies were performed using protocols approved by the Institutional Animal Care and Use Committee of the College of Medicine, National Taiwan University.
Statistical analysis
The Student’s t-test was used for comparison between the treatment group and the control group. ANOVA was used to evaluate the statistical significance of the mean values. Data were analyzed with the SPSS program for Windows, version 11.0 (SPSS Inc., Chicago, IL, USA). All statistical tests included two-way analysis of variance. Statistical significance was assumed at p-values of less than 0.05.
References
Vander Heiden MG, Cantley LC, Thompson CB. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science. 2009;324:1029–33.
Imamura K, Tanaka T. Multimolecular forms of pyruvate kinase from rat and other mammalian tissues. I. Electrophoretic studies. J Biochem. 1972;71:1043–51.
Noguchi T, Inoue H, Tanaka T. The M1- and M2-type isozymes of rat pyruvate kinase are produced from the same gene by alternative RNA splicing. J Biol Chem. 1986;261:13807–12.
Mazurek S. Pyruvate kinase type M2: a key regulator of the metabolic budget system in tumor cells. Int J Biochem Cell Biol. 2011;43:969–80.
Hitosugi T, Kang S, Vander Heiden MG, Chung TW, Elf S, Lythgoe K, et al. Tyrosine phosphorylation inhibits PKM2 to promote the Warburg effect and tumor growth. Sci Signal. 2009;2:ra73.
Mellati AA, Yucel M, Altinors N, Gunduz U. Regulation of M2-type pyruvate kinase from human meningioma by allosteric effectors fructose 1,6 diphosphate and L-alanine. Cancer Biochem Biophys. 1992;13:33–41.
Nakatsu D, Horiuchi Y, Kano F, Noguchi Y, Sugawara T, Takamoto I, et al. L-cysteine reversibly inhibits glucose-induced biphasic insulin secretion and ATP production by inactivating PKM2. Proc Natl Acad Sci USA. 2015;112:E1067–76.
Yang W, Xia Y, Ji H, Zheng Y, Liang J, Huang W, et al. Nuclear PKM2 regulates beta-catenin transactivation upon EGFR activation. Nature. 2011;480:118–22.
Luo W, Hu H, Chang R, Zhong J, Knabel M, O’Meally R, et al. Pyruvate kinase M2 is a PHD3-stimulated coactivator for hypoxia-inducible factor 1. Cell. 2011;145:732–44.
Gao X, Wang H, Yang JJ, Liu X, Liu ZR. Pyruvate kinase M2 regulates gene transcription by acting as a protein kinase. Mol Cell. 2012;45:598–609.
Jiang Y, Li X, Yang W, Hawke DH, Zheng Y, Xia Y, et al. PKM2 regulates chromosome segregation and mitosis progression of tumor cells. Mol Cell. 2014;53:75–87.
Liang J, Cao R, Wang X, Zhang Y, Wang P, Gao H, et al. Mitochondrial PKM2 regulates oxidative stress-induced apoptosis by stabilizing Bcl2. Cell Res. 2017;27:329–51.
Yang YC, Cheng TY, Huang SM, Su CY, Yang PW, Lee JM, et al. Cytosolic PKM2 stabilizes mutant EGFR protein expression through regulating HSP90-EGFR association. Oncogene. 2016;35:3387–98.
Israelsen WJ, Dayton TL, Davidson SM, Fiske BP, Hosios AM, Bellinger G, et al. PKM2 isoform-specific deletion reveals a differential requirement for pyruvate kinase in tumor cells. Cell. 2013;155:397–409.
Bluemlein K, Gruning NM, Feichtinger RG, Lehrach H, Kofler B, Ralser M. No evidence for a shift in pyruvate kinase PKM1 to PKM2 expression during tumorigenesis. Oncotarget. 2011;2:393–400.
Desai S, Ding M, Wang B, Lu Z, Zhao Q, Shaw K, et al. Tissue-specific isoform switch and DNA hypomethylation of the pyruvate kinase PKM gene in human cancers. Oncotarget. 2014;5:8202–10.
Zhan C, Yan L, Wang L, Ma J, Jiang W, Zhang Y, et al. Isoform switch of pyruvate kinase M1 indeed occurs but not to pyruvate kinase M2 in human tumorigenesis. PLoS ONE. 2015;10:e0118663.
Wu J, Hu L, Chen M, Cao W, Chen H, He T. Pyruvate kinase M2 overexpression and poor prognosis in solid tumors of digestive system: evidence from 16 cohort studies. Onco Targets Ther. 2016;9:4277–88.
Lockney NA, Zhang M, Lu Y, Sopha SC, Washington MK, Merchant N, et al. Pyruvate kinase muscle isoenzyme 2 (PKM2) expression is associated with overall survival in pancreatic ductal adenocarcinoma. J Gastrointest Cancer. 2015;46:390–8.
Li Z, Yang P, Li Z. The multifaceted regulation and functions of PKM2 in tumor progression. Biochim Biophys Acta. 2014;1846:285–96.
Keller KE, Doctor ZM, Dwyer ZW, Lee YS. SAICAR induces protein kinase activity of PKM2 that is necessary for sustained proliferative signaling of cancer cells. Mol Cell. 2014;53:700–9.
Radu M, Semenova G, Kosoff R, Chernoff J. PAK signalling during the development and progression of cancer. Nat Rev Cancer. 2014;14:13–25.
Christofk HR, Vander Heiden MG, Harris MH, Ramanathan A, Gerszten RE, Wei R, et al. The M2 splice isoform of pyruvate kinase is important for cancer metabolism and tumour growth. Nature. 2008;452:230–3.
Kumar Y, Gurusamy K, Pamecha V, Davidson BR. Tumor M2-pyruvate kinase as tumor marker in exocrine pancreatic cancer a meta-analysis. Pancreas. 2007;35:114–9.
Joergensen MT, Heegaard NH, Schaffalitzky de Muckadell OB. Comparison of plasma Tu-M2-PK and CA19-9 in pancreatic cancer. Pancreas. 2010;39:243–7.
Goonetilleke KS, Mason JM, Siriwardana P, King NK, France MW, Siriwardena AK. Diagnostic and prognostic value of plasma tumor M2 pyruvate kinase in periampullary cancer: evidence for a novel biological marker of adverse prognosis. Pancreas. 2007;34:318–24.
Zhu H, Luo H, Zhu X, Hu X, Zheng L, Zhu X. Pyruvate kinase M2 (PKM2) expression correlates with prognosis in solid cancers: a meta-analysis. Oncotarget. 2017;8:1628–40.
Mohammad GH, Olde Damink SW, Malago M, Dhar DK, Pereira SP. Pyruvate kinase M2 and lactate dehydrogenase A are overexpressed in pancreatic cancer and correlate with poor outcome. PLoS ONE. 2016;11:e0151635
Ogawa H, Nagano H, Konno M, Eguchi H, Koseki J, Kawamoto K, et al. The combination of the expression of hexokinase 2 and pyruvate kinase M2 is a prognostic marker in patients with pancreatic cancer. Mol Clin Oncol. 2015;3:563–71.
Calabretta S, Bielli P, Passacantilli I, Pilozzi E, Fendrich V, Capurso G, et al. Modulation of PKM alternative splicing by PTBP1 promotes gemcitabine resistance in pancreatic cancer cells. Oncogene. 2016;35:2031–9.
Li C, Zhao Z, Zhou Z, Liu R. PKM2 promotes cell survival and invasion under metabolic stress by enhancing Warburg effect in pancreatic ductal adenocarcinoma. Dig Dis Sci. 2016;61:767–73.
Anastasiou D, Yu Y, Israelsen WJ, Jiang JK, Boxer MB, Hong BS, et al. Pyruvate kinase M2 activators promote tetramer formation and suppress tumorigenesis. Nat Chem Biol. 2012;8:839–47.
Schopf FH, Biebl MM, Buchner J, The HSP90 chaperone machinery. Nat Rev Mol Cell Biol. 2017;18:345–60.
Ehrenfried JA, Herron BE, Townsend CM Jr, Evers BM. Heat shock proteins are differentially expressed in human gastrointestinal cancers. Surg Oncol. 1995;4:197–203.
Lang SA, Klein D, Moser C, Gaumann A, Glockzin G, Dahlke MH, et al. Inhibition of heat shock protein 90 impairs epidermal growth factor-mediated signaling in gastric cancer cells and reduces tumor growth and vascularization in vivo. Mol Cancer Ther. 2007;6:1123–32.
Jiang K, Li W, Zhang Q, Yan G, Guo K, Zhang S, et al. GP73 N-glycosylation at Asn144 reduces hepatocellular carcinoma cell motility and invasiveness. Oncotarget. 2016;7:23530–41.
Vargas DA, Sun M, Sadykov K, Kukuruzinska MA, Zaman MH. The integrated role of Wnt/beta-Catenin, N-glycosylation, and E-Cadherin-mediated adhesion in network dynamics. PLoS Comput Biol. 2016;12:e1005007
Hosios AM, Fiske BP, Gui DY, Vander Heiden MG. Lack of evidence for PKM2 protein kinase activity. Mol Cell. 2015;59:850–7.
Yang W, Xia Y, Hawke D, Li X, Liang J, Xing D, et al. PKM2 phosphorylates histone H3 and promotes gene transcription and tumorigenesis. Cell. 2012;150:685–96.
Wei Y, Wang D, Jin F, Bian Z, Li L, Liang H, et al. Pyruvate kinase type M2 promotes tumour cell exosome release via phosphorylating synaptosome-associated protein 23. Nat Commun. 2017;8:14041.
Jiang Y, Wang Y, Wang T, Hawke DH, Zheng Y, Li X, et al. PKM2 phosphorylates MLC2 and regulates cytokinesis of tumour cells. Nat Commun. 2014;5:5566.
Coniglio SJ, Zavarella S, Symons MH. Pak1 and Pak2 mediate tumor cell invasion through distinct signaling mechanisms. Mol Cell Biol. 2008;28:4162–72.
Kumar R, Sanawar R, Li X, Li F. Structure, biochemistry, and biology of PAK kinases. Gene. 2017;605:20–31.
Jakobi R, McCarthy CC, Koeppel MA, Stringer DK. Caspase-activated PAK-2 is regulated by subcellular targeting and proteasomal degradation. J Biol Chem. 2003;278:38675–85.
Acknowledgements
We thank Dr. Zhi-Min Lu for the Flag-tagged K367M, K433E, and wild-type PKM2; Dr. Zhi-Ren Liu for HA-tagged R399E and wild-type PKM2. This work was supported by grants from the Ministry of Science and Technology of Taiwan, 104-0210-01-09-02, 105-0210-01-13-01, and 106-0210-01-15-02 (to M.H.); 105-2320-B-002-035-MY3 (to K.-T.H.); and 104-2314-B-002-027 (to T.-Y.C.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Cheng, TY., Yang, YC., Wang, HP. et al. Pyruvate kinase M2 promotes pancreatic ductal adenocarcinoma invasion and metastasis through phosphorylation and stabilization of PAK2 protein. Oncogene 37, 1730–1742 (2018). https://doi.org/10.1038/s41388-017-0086-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-017-0086-y
- Springer Nature Limited
This article is cited by
-
Hypoxia-Challenged Pancreatic Adenocarcinoma Cell-Derived Exosomal circR3HCC1L Drives Tumor Growth Via Upregulating PKM2 Through Sequestering miR-873-5p
Molecular Biotechnology (2024)
-
Arginine methylation of MTHFD1 by PRMT5 enhances anoikis resistance and cancer metastasis
Oncogene (2022)
-
APC loss induces Warburg effect via increased PKM2 transcription in colorectal cancer
British Journal of Cancer (2021)
-
The Chk2-PKM2 axis promotes metabolic control of vasculogenic mimicry formation in p53-mutated triple-negative breast cancer
Oncogene (2021)
-
Protein kinase function of pyruvate kinase M2 and cancer
Cancer Cell International (2020)