Abstract
Background:
Human milk oligosaccharides (HMOs) shape the intestinal microbiota in term infants. In premature infants, alterations in the intestinal microbiota (dysbiosis) are associated with risk of necrotizing enterocolitis (NEC) and sepsis, and the influence of HMOs on the microbiota is unclear.
Methods:
Milk, urine, and stool specimens from 14 mother–premature infant dyads were investigated by mass spectrometry for HMO composition. The stools were analyzed by next-generation sequencing to complement a previous analysis.
Results:
Percentages of fucosylated and sialylated HMOs were highly variable between individuals but similar in urine, feces, and milk within dyads. Differences in urine and fecal HMO composition suggest variability in absorption. Secretor status of the mother correlated with the urine and fecal content of specific HMO structures. Trends toward higher levels of Proteobacteria and lower levels of Firmicutes were noted in premature infants of nonsecretor mothers. Specific HMO structures in the milk, urine, and feces were associated with alterations in fecal Proteobacteria and Firmicutes.
Conclusion:
HMOs may influence the intestinal microbiota in premature infants. Specific HMOs, for example those associated with secretor mothers, may have a protective effect by decreasing pathogens associated with sepsis and NEC, while other HMOs may increase dysbiosis in this population.
Similar content being viewed by others
Main
Human milk oligosaccharides (HMOs) are the third largest component of human milk. These complex carbohydrates are produced in a wide variety of structures that vary from woman to woman and within a given woman over time at great nutritional cost to the mother, and yet are not digestible by the infant (1). This observation prompts the obvious evolutionary question: what is the benefit to the mother–baby dyad of these structures? HMOs appear to have multiple functions. First, they shape the composition of the infant intestinal microbiota through selective consumption by commensal bacteria (2,3). Bifidobacteria and Bacteroidetes become dominant intestinal bacteria in healthy breastfed term infants due to their ability to digest and utilize HMOs via specific glycosidases, while most pathogenic Enterobacteriaceae lack these enzymes and are unable to utilize HMOs as a food source (4). Second, HMOs have structural homology to many cell surface glycans and thus act as decoys by binding to luminal bacteria that are then unable to bind to the surface of the enterocyte (5). Third, HMOs are absorbed into the vasculature (6) and excreted into the urine (7) potentially binding to and decreasing the capacity of invasive bacteria to cause sepsis and urinary tract infections (8). Fourth, sialic acid–containing HMOs may be important in neurodevelopment (9).
Premature infants are at increased risk for necrotizing enterocolitis (NEC), sepsis, pneumonia, and neurodevelopmental delays due to immaturity and dysfunction of essentially every component of innate immunity. Premature infants who receive their mother’s own milk have lower rates of NEC and sepsis than those who receive formula (10). This is likely due to a variety of human milk components including immunoglobulins, lactoferrin, lysozyme, and HMOs. Milk produced by women who deliver preterm differs from that of women delivering at term including the amounts and types of HMOs (11). Women delivering preterm have much wider variation (both between women and for a given woman over time) in the percentage of HMOs that contain fucose or sialic acid than women who deliver at term (12).
HMO structure is influenced in part by mutations in the maternal fucosyltransferase 2 (FUT2) gene. Individuals who are homozygous for mutations in the FUT2 gene (historically referred to as “nonsecretors”) are unable to produce α1,2 fucosylated glycans in their secretions including breast milk. Nonsecretor individuals have lower levels of commensal bifidobacteria, are at decreased risk for infections with a variety of intestinal viruses, but have an increased risk for Crohn’s disease (13), celiac disease (14), and NEC (15). The influence of maternal secretor status on the infant microbiota has not been well characterized.
In this study, our objective was to determine the HMO composition of milk, urine, and feces from 14 premature infants receiving only human milk and their mothers and to correlate these with the composition of the fecal microbiota. We hypothesized that ingested HMOs influence the intestinal microbiota.
Results
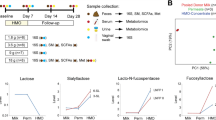
Relative abundances of the measured HMOs in milk, urine, and feces for all 14 dyads are summarized in Table 1 (schematic chemical structures for HMOs are presented in Table 3 of ref. 7, the four-digit labels correspond to the number of hexose, fucose, N-acetylhexosamine, and sialic acid residues, in that order. For example, the label 4120 (M=1220.47) means that there are 4 hexose, 1 fucose, and 2 N-acetylhexosamine residues in the structure). Representative mass spectra of milk from one of the mothers are presented in Supplementary Figures S1–S3 online with a schematic diagram of HMO structure. The high degree of interindividual variability is evident from the SDs in Table 1 . Most dyads showed similarity in HMO groups across specimen type (i.e., milk composition was similar to fecal and urine composition). Figure 1 presents relative abundance of HMO groups for each dyad and Table 2 presents the combined data for all dyads. Total fucosylated, sialylated, fucosylated–sialylated, and nonfucosylated/nonsialylated HMOs were similar across specimen types (Supplementary Figure 4 online), and correlations between milk and fecal HMO groups were strong (Supplementary Figure S5 online). Individual HMOs, however, showed considerable heterogeneity. Structures relatively more abundant in feces than in urine included F-LSTc (P < 0.01), LNFP II, 5130a, 5130c, and the group of five structures with mass 1,731 (P < 0.05) and TFLNH and 6’SL (P = 0.05–0.1). We hypothesize that these HMOs are not significantly absorbed from the gut into the bloodstream. Several structures were not present in milk but found in either urine (3’Sle), feces (TFiLNO, 5330a), or both (A-hepta, 3’SLN). The origin of these structures is unclear; it is possible that these structures were present in earlier milk samples or that they were constructed or altered in the gut lumen by gut microbial enzymes. The only structure significantly more abundant in urine than feces was 3’Sle (P < 0.01), though other structures showed a similar trend (3’SL, 3’SLN, LNFP I, LNFP V; P = 0.1–0.2). We hypothesize that these HMOs are absorbed from the gut into the bloodstream and filtered by the kidneys. The difference between 3’SL and 6’SL is striking as both structures have the same molecular weight with the difference in linkage apparently influencing absorption from the gut.
Relative abundance of groups of HMOs in the milk, urine, and feces for each of the 14 mother–infant dyads. White: fucosylated HMOs, black: sialylated HMOs, gray: fucosylated and sialylated HMOs, and hatched: HMOs with neither fucose nor sialic acid. (a) patient 4, (b) patient 6, (c) patient 9, (d) patient 10, (e) patient 13, (f) patient 14, (g) patient 16, (h) patient 18, (i) patient 20, (j) patient 21, (k) patient 22, (l) patient 26, (m) patient 27, (n) patient 30. F, feces; HMO, human milk oligosaccharide; M, milk; NA, specimen not available; U, urine.
The secretor status of the mothers was predicted based on the percentages of 2’FL and LDFT (16). Of the 14 mothers enrolled, 8 were secretors and 6 were nonsecretors. Urine and fecal HMOs suggested a similar pattern in the infant to that of the mother’s milk in all but infant number 13 ( Table 3 ). The fecal microbiota was analyzed by terminal restriction fragment length polymorphism in the original study (17). Based on the terminal restriction fragment length polymorphism data of all 14 specimens, infants of secretor mothers had lower γ-Proteobacteria (P = 0.046) and higher Lactobacillaceae (P = 0.02). For this study, a more robust approach using next-generation sequencing (NGS) was performed on the 12 fecal samples for which sufficient sample remained and is summarized in Figure 2 and Supplementary Figure S6 online). Note that the major HMO-consuming orders of bacteria (Bifidobacteriales and Bacteroidales) are present in small numbers as has been previously demonstrated in premature infants. In this small sample, the differences by NGS in overall microbial composition between infants of secretor mothers and nonsecretor mothers did not reach statistical significance; however, the following trends were noted at the phylum level: increased Firmicutes (P = 0.08) in the feces of infants of secretor mothers, and at the order level decreased Enterobacteriales (phylum Proteobacteria, P = 0.06) in the feces of infants of secretor mothers.
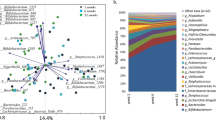
Composition of the fecal microbiota determined by next-generation sequencing in 12 of the infants (6 with secretor mothers and 6 with nonsecretor mothers). (a) Phylum level. White: Actinobacteria, black: Bacteroidetes, gray: Firmicutes, and hatched: Proteobacteria. (b) Order level. White: Bifidobacteriales, black: Bacteroidales, gray: Bacillales, hatched: Lactobacillales, dots: Clostridiales, light gray: Enterobacteriales, double hatched: Pseudomonadales, and vertical stripes: others.
Correlations between HMOs in milk and fecal bacteria by NGS are presented at the phylum level in Figure 3 (HMO groups), Supplementary Figure S7 online (HMO groups, top panel) and Table 4 (individual HMOs). For many HMOs, the associations for the dominant phyla (Firmicutes and Proteobacteria) are in opposing directions. Figure 4 (top panel) summarizes correlations at the order level with the strongest correlations between HMOs in the milk and fecal Enterobacteriales (phylum Proteobacteria, positive correlation with HMOs containing neither fucose nor sialic acid and negative correlation with fucosylated HMOs).
Linear regression associations between groups of milk HMOs and fecal microbes by phylum (n = 12). (a) R2 = 0.63; P = 0.002. (b) R2 = 0.31; P = 0.06. (c) R2 = 0.36; P = 0.04. (d) R2 = 0.48; P = 0.01. (e) R2 = 0.42; P = 0.02. HMO, human milk oligosaccharide.
Heat map demonstrating correlations between milk HMO groups and fecal bacterial orders (top) and between fecal HMO groups and fecal bacterial orders (bottom). Colors represent strength of correlations with deep blue representing Spearman’s ρ = 1 and deep red representing Spearman’s ρ = −1. HMO, human milk oligosaccharide.
HMOs in the feces must traverse the length of the gastrointestinal tract without being absorbed into the bloodstream or consumed by intestinal bacteria. In a pilot study of term breastfed infants, as HMO-consuming bacteria increased in the feces over time, fecal HMOs decreased in a structure-specific manner (i.e., structures that are able to be consumed by the dominant bacteria decrease while those HMOs that are not digestible by the dominant bacteria persist) (18). In these preterm infants, correlations are noted between groups of HMOs in the feces (Supplementary Figure S7 online, bottom panel) and individual HMO structures ( Table 5 ) and fecal bacterial composition at the phylum level. At the order level, the persistence of fucosylated HMOs in the feces shows a positive correlation with Pasteurellales, Pseudomonales, and Burkholderiales and a negative correlation with Enterobacteriales ( Figure 4 , bottom panel). A detailed heat map summarizing Spearman’s correlations for milk and fecal HMO groups and fecal bacteria at the genus level as determined by NGS in the current study is given in Supplementary Figure S8 online. While the determination of absolute abundances of HMOs in the milk and feces is beyond the scope of this article, previous studies have demonstrated that HMO content in feces is about 50% of that in milk (7).
HMOs in the urine have been absorbed from the gastrointestinal tract and filtered by the kidneys. Table 6 presents associations between urine HMOs and fecal bacteria as determined by NGS. Note that Tables 4 , 5 , and 6 include only structures correlated with changes in fecal phylum with P values <0.1, i.e., the structures not listed did not demonstrate significant correlations. While the determination of absolute abundances of HMOs in the urine is beyond the scope of this article, previous studies have demonstrated that HMO content in urine is about 20% of that in milk (7).
Discussion
Bifidobacteria and Bacteroidetes are the dominant fecal bacteria and the primary consumers of HMOs in healthy breastfed term infants but are generally rare in the feces of premature infants unless supplemented in the form of probiotics (19). In these infants, bifidobacteria and Bacteroidetes were present in very limited amounts confirming previous observations of the marked differences in the fecal microbiota between term and preterm infants (20). Bifidobacteria are associated with improved growth and vaccine responsiveness in term breastfed infants in a developing country (21). In premature infants, administration of probiotics containing bifidobacteria leads to a decreased risk of NEC, a common and devastating inflammatory disease of neonates (22,23). In animal models and in vitro, bifidobacteria decrease inflammation and gut permeability (24,25). In this small group of premature infants (none of whom received probiotic organisms), the numbers of bifidobacteria were low (<1% of all bacteria).
Enterobacteriaceae (phylum Proteobacteria) include pathogens that are associated with NEC and sepsis in premature infants such as Escherichia coli, Klebsiella, Enterobacter, Proteus, and Serratia. These organisms are rare in the healthy adult gut, but common and often dominant in the intestine of premature infants. Colonization with Klebsiella in the first week of life (26) and with Enterobacteriaceae in the second week of life (27) have both been associated with increased risk of later development of NEC (which generally presents at 2–6 wk of age). A bloom of these organisms has been demonstrated just prior to the onset of NEC in premature infants (28). Some of these bacteria are capable of inducing an inflammatory response in the host and then utilizing the products of inflammation to outcompete other intestinal bacteria (29,30). The associations noted in these premature infants suggest that specific HMOs in human milk may decrease intestinal Enterobacteriaceae (e.g., fucosylated HMOs such as LDFT and LNFP V and structures that are both fucosylated and sialylated), while other HMOs in the milk are associated with increased intestinal Enterobacteriaceae (e.g., LNnH, 5031a, and HMOs that contain neither fucose nor sialic acid). The mechanisms underlying these associations likely include both the host and interactions with other gut microbes as Enterobacteriaceae are unable to utilize HMOs as a food source (31).
The impact of host secretor status on susceptibility to various infections is well documented and somewhat balanced in the adult population such that both secretors and no-secretors have differing advantages against specific pathogens. Recent demonstration of rapid fucosylation of glycans on the luminal surface of the enterocyte in response to stimulation of Toll-like receptors 2, 4, and 9 suggest that this response (possible only in the secretor individual) may be important in protecting and nourishing gut commensals in the midst of infection (32). In the premature infant, the inability of the nonsecretor mother to provide key fucosylated HMOs appears to significantly influence the developing microbiota. The capacity of gut microbes to deconstruct specific linkages is a second promising mechanism underlying the observed associations. Recent studies have demonstrated marked differences between bacterial species in the capacity to consume specific HMOs (4) and marked differences at the species level in production of specific glycosidases (33).
Dysbiosis is often defined as alterations in the richness and diversity of the microbiota or more recently as delayed maturation of the microbiota of the infant (34). While increased diversity of the microbiota may be beneficial in the adult, limited data suggest that prior to weaning, a less complex microbiota that is dominated by bifidobacteria is associated with improved health (e.g., growth, thymus size, and vaccine response) (21). Dysbiosis as a potentially alterable risk factor for NEC is based on several observations: increased risk of NEC with increased exposure to antibiotics (35) and acid-suppressing agents (36) (both associated with dysbiosis), decreased risk of NEC with human milk (10) and probiotics (22) (both associated with less severe dysbiosis), and from direct observations (26). While human milk decreases the risk of NEC, it appears to have less of an impact on the microbiota of the premature infant than other factors such as antibiotic exposure and environmental factors (20,37).
The observed differences in absorption of specific HMO structures is consistent with previous observations (7) and with the recent reports of HMOs in the plasma (6,38). Further exploration of this area is needed to determine any effects of HMO in the systemic circulation and/or urinary tract. In addition, if these differences are consistent, it may be possible to develop ratios of urine:fecal HMOs as a marker of gut permeability for the infant (similar to the lactulose:mannitol test in current use for children and adults).
The number of mother–infant dyads included in this study is too small to make any firm conclusions but is reasonable to generate hypotheses for testing in future larger studies. Perhaps, the most compelling hypotheses generated by these data include the following: (i) that human milk rich in fucosylated HMOs is protective against dysbiosis, while human milk rich in HMOs without fucose or sialic acid increases the risk of dysbiosis, (ii) that milk from nonsecretor mothers may not be as protective against dysbiosis and NEC as milk from secretor mothers (this hypothesis is consistent with observations of increased sepsis and NEC in nonsecretor babies who likely have nonsecretor mothers (15)), and (iii) that premature infant fecal HMO composition is a biomarker, with feces high in HMOs containing sialic acid (e.g., 3’SL) and HMOs with neither fucose nor sialic acid associated with dysbiosis and feces high in fucosylated HMOs (particularly LDFT) associated with less severe dysbiosis. Given the observation that fucosylation is an immature and inconsistent process in the mammary gland of the mother delivering prematurely, these data also suggest the hypothesis that premature infants may benefit from supplementation with fucosylated HMOs (e.g., through pooled donor human milk). Future studies to confirm these observations and to correlate milk HMO composition and fecal HMO composition with short-term outcomes (e.g., NEC and sepsis) and long-term outcomes (e.g., growth and neurodevelopmental outcomes) would be valuable.
Methods
We collected specimens of stool and urine from 14 premature infants enrolled in a clinical trial of prebiotic supplementation (17) and milk from their mothers. The study was approved by the Institutional Review Board of the University of California Davis, informed consent was obtained from at least one parent prior to enrollment, and the study was registered at clinicaltrials.gov (NCT00810160). Mean gestational age was 27 wk at birth and 30 wk at the time of enrollment. All infants were receiving enteral feedings of at least 100 ml/kg/d. Specimens described in this article were obtained at baseline (prior to initiation of any supplement) from infants receiving exclusively their mother’s own milk (no donor human milk). Specimens were frozen at −40 °C upon collection and transported on dry ice for analysis. Extraction and analysis of HMOs in milk, urine, and feces was performed as previously described (7). The nonparametric Mann–Whitney rank-sum test was used to compare abundance of specific HMO structures in urine and stool, and linear regression was used to correlate HMO abundance and fecal microbial abundance. As this study is predominantly a hypothesis-generating exercise, we reported all P values <0.1.
Bacterial DNA was extracted from the stool specimens as previously described in a clinical trial (17); however, in addition to the analyses by PCR and terminal restriction fragment length polymorphism, the samples were analyzed by NGS in this study. For these analyses, DNA library construction was carried out as previously described (39) and submitted to the UC Davis Genome Center DNA Technologies Core for sequencing on an Illumina MiSeq instrument (Illumina, San Diego CA). QIIME software package (University of Colorado, Boulder, CO; version1.7.0) was used for quality filtering and demultiplexing the resulting sequencing data (40). Operational taxonomic units were assigned using UCLUST (drive5.com, Tiburon, CA) based on 97% pairwise identity (41), and taxonomic classification was based on the Ribosomal Database Project classifier (Michigan State University, East Lansing, MI) against a representative subset of the Greengenes 16S rRNA database (Second Genome, South San Francisco, CA; gg_otus_12_10 release) (42,43). Taxonomic relative abundance data from the QIIME data analysis was used to calculate correlations between HMO consumption and fecal bacteria abundance.
To visualize correlations between HMO consumption and fecal bacteria, custom R scripts were developed. HMO or bacterial populations with a maximum abundance less than 0.02 were removed from further analysis. A Spearman’s correlation matrix was computed using the remaining HMO and bacterial features. The procedure was repeated at the phylum, order, and genus levels. We chose to use the Spearman’s correlation because it is a nonparametric measure that is robust to outliers and does not require that the data be normally distributed.
Statement of Financial Support
This work was supported by the Eunice Kennedy Shriver National Institute of Child Health & Human Development of the National Institutes of Health, Bethesda, Maryland (grant numbers R01HD059127 and R01HD061923) and the National Center for Advancing Translational Sciences, National Institutes of Health, Bethesda, Maryland (grant number UL1 TR000002).
The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. D.A.M. acknowledges support as the Peter J. Shields Endowed Chair in Dairy Food Science at UC Davis.
Disclosure
Three of the authors (J.B.G., C.B.L., and D.A.M.) are the co-founders of Evolve Biosystems, a company focused on diet-based manipulation of the gut microbiota.
References
Zivkovic AM, German JB, Lebrilla CB, Mills DA. Human milk glycobiome and its impact on the infant gastrointestinal microbiota. Proc Natl Acad Sci USA 2011;108:Suppl 1:4653–8.
Sela DA, Mills DA. Nursing our microbiota: molecular linkages between bifidobacteria and milk oligosaccharides. Trends Microbiol 2010;18:298–307.
Coppa GV, Gabrielli O, Zampini L, et al. Oligosaccharides in 4 different milk groups, Bifidobacteria, and Ruminococcus obeum. J Pediatr Gastroenterol Nutr 2011;53:80–7.
Yu ZT, Chen C, Newburg DS. Utilization of major fucosylated and sialylated human milk oligosaccharides by isolated human gut microbes. Glycobiology 2013;23:1281–92.
Coppa GV, Zampini L, Galeazzi T, et al. Human milk oligosaccharides inhibit the adhesion to Caco-2 cells of diarrheal pathogens: Escherichia coli, Vibrio cholerae, and Salmonella fyris. Pediatr Res 2006;59:377–82.
Ruhaak LR, Stroble C, Underwood MA, Lebrilla CB. Detection of milk oligosaccharides in plasma of infants. Anal Bioanal Chem 2014;406:5775–84.
De Leoz ML, Wu S, Strum JS, et al. A quantitative and comprehensive method to analyze human milk oligosaccharide structures in the urine and feces of infants. Anal Bioanal Chem 2013;405:4089–105.
Lin AE, Autran CA, Espanola SD, Bode L, Nizet V. Human milk oligosaccharides protect bladder epithelial cells against uropathogenic Escherichia coli invasion and cytotoxicity. J Infect Dis 2014;209:389–98.
Wang B. Molecular mechanism underlying sialic acid as an essential nutrient for brain development and cognition. Adv Nutr 2012;3:465S–72S.
Meinzen-Derr J, Poindexter B, Wrage L, Morrow AL, Stoll B, Donovan EF. Role of human milk in extremely low birth weight infants’ risk of necrotizing enterocolitis or death. J Perinatol 2009;29:57–62.
Gabrielli O, Zampini L, Galeazzi T, et al. Preterm milk oligosaccharides during the first month of lactation. Pediatrics 2011;128:e1520–31.
De Leoz ML, Gaerlan SC, Strum JS, et al. Lacto-N-tetraose, fucosylation, and secretor status are highly variable in human milk oligosaccharides from women delivering preterm. J Proteome Res 2012;11:4662–72.
Maroni L, van de Graaf SF, Hohenester SD, Oude Elferink RP, Beuers U. Fucosyltransferase 2: a genetic risk factor for primary sclerosing cholangitis and Crohn’s disease–a comprehensive review. Clin Rev Allergy Immunol 2015;48:182–91.
Parmar AS, Alakulppi N, Paavola-Sakki P, et al. Association study of FUT2 (rs601338) with celiac disease and inflammatory bowel disease in the Finnish population. Tissue Antigens 2012;80:488–93.
Morrow AL, Meinzen-Derr J, Huang P, et al. Fucosyltransferase 2 non-secretor and low secretor status predicts severe outcomes in premature infants. J Pediatr 2011;158:745–51.
Totten SM, Zivkovic AM, Wu S, et al. Comprehensive profiles of human milk oligosaccharides yield highly sensitive and specific markers for determining secretor status in lactating mothers. J Proteome Res 2012;11:6124–33.
Underwood MA, Kalanetra KM, Bokulich NA, et al. Prebiotic oligosaccharides in premature infants. J Pediatr Gastroenterol Nutr 2014;58:352–60.
De Leoz ML, Kalanetra KM, Bokulich NA, et al. Human milk glycomics and gut microbial genomics in infant feces show a correlation between human milk oligosaccharides and gut microbiota: a proof-of-concept study. J Proteome Res 2015;14:491–502.
Underwood MA, Kalanetra KM, Bokulich NA, et al. A comparison of two probiotic strains of bifidobacteria in premature infants. J Pediatr 2013;163:1585–1591.e9.
Arboleya S, Binetti A, Salazar N, et al. Establishment and development of intestinal microbiota in preterm neonates. FEMS Microbiol Ecol 2012;79:763–72.
Huda MN, Lewis Z, Kalanetra KM, et al. Stool microbiota and vaccine responses of infants. Pediatrics 2014;134:e362–72.
Deshpande G, Rao S, Patole S, Bulsara M. Updated meta-analysis of probiotics for preventing necrotizing enterocolitis in preterm neonates. Pediatrics 2010;125:921–30.
Jacobs SE, Tobin JM, Opie GF, et al.; ProPrems Study Group. Probiotic effects on late-onset sepsis in very preterm infants: a randomized controlled trial. Pediatrics 2013;132:1055–62.
Ganguli K, Meng D, Rautava S, Lu L, Walker WA, Nanthakumar N. Probiotics prevent necrotizing enterocolitis by modulating enterocyte genes that regulate innate immune-mediated inflammation. Am J Physiol Gastrointest Liver Physiol 2013;304:G132–41.
Bergmann KR, Liu SX, Tian R, et al. Bifidobacteria stabilize claudins at tight junctions and prevent intestinal barrier dysfunction in mouse necrotizing enterocolitis. Am J Pathol 2013;182:1595–606.
Torrazza RM, Ukhanova M, Wang X, et al. Intestinal microbial ecology and environmental factors affecting necrotizing enterocolitis. PLoS One 2013;8:e83304.
Morrow AL, Lagomarcino AJ, Schibler KR, et al. Early microbial and metabolomic signatures predict later onset of necrotizing enterocolitis in preterm infants. Microbiome 2013;1:13.
Mai V, Young CM, Ukhanova M, et al. Fecal microbiota in premature infants prior to necrotizing enterocolitis. PLoS One 2011;6:e20647.
Winter SE, Thiennimitr P, Winter MG, et al. Gut inflammation provides a respiratory electron acceptor for Salmonella. Nature 2010;467:426–9.
Winter SE, Winter MG, Xavier MN, et al. Host-derived nitrate boosts growth of E. coli in the inflamed gut. Science 2013;339:708–11.
Marcobal A, Barboza M, Froehlich JW, et al. Consumption of human milk oligosaccharides by gut-related microbes. J Agric Food Chem 2010;58:5334–40.
Pickard JM, Maurice CF, Kinnebrew MA, et al. Rapid fucosylation of intestinal epithelium sustains host-commensal symbiosis in sickness. Nature 2014;514:638–41.
Sela DA, Mills DA. The marriage of nutrigenomics with the microbiome: the case of infant-associated bifidobacteria and milk. Am J Clin Nutr 2014;99:697S–703S.
Subramanian S, Huq S, Yatsunenko T, et al. Persistent gut microbiota immaturity in malnourished Bangladeshi children. Nature 2014;510:417–21.
Cotten CM, Taylor S, Stoll B, et al.; NICHD Neonatal Research Network. Prolonged duration of initial empirical antibiotic treatment is associated with increased rates of necrotizing enterocolitis and death for extremely low birth weight infants. Pediatrics 2009;123:58–66.
Chung EY, Yardley J. Are there risks associated with empiric acid suppression treatment of infants and children suspected of having gastroesophageal reflux disease? Hosp Pediatr 2013;3:16–23.
Taft DH, Ambalavanan N, Schibler KR, et al. Intestinal microbiota of preterm infants differ over time and between hospitals. Microbiome 2014;2:36.
Goehring KC, Kennedy AD, Prieto PA, Buck RH. Direct evidence for the presence of human milk oligosaccharides in the circulation of breastfed infants. PLoS One 2014;9:e101692.
Bokulich NA, Mills DA. Facility-specific “house” microbiome drives microbial landscapes of artisan cheesemaking plants. Appl Environ Microbiol 2013;79:5214–23.
Caporaso JG, Lauber CL, Walters WA, et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci USA 2011;108:Suppl 1:4516–22.
Edgar RC. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010;26:2460–1.
Wang Q, Garrity GM, Tiedje JM, Cole JR. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 2007;73:5261–7.
DeSantis TZ, Hugenholtz P, Larsen N, et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 2006;72:5069–72.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Figures
(PDF 584 kb)
Rights and permissions
About this article
Cite this article
Underwood, M., Gaerlan, S., De Leoz, M. et al. Human milk oligosaccharides in premature infants: absorption, excretion, and influence on the intestinal microbiota. Pediatr Res 78, 670–677 (2015). https://doi.org/10.1038/pr.2015.162
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/pr.2015.162
- Springer Nature America, Inc.
This article is cited by
-
The contribution of maternal oral, vaginal, and gut microbiota to the developing offspring gut
Scientific Reports (2023)
-
Creation of a milk oligosaccharide database, MilkOligoDB, reveals common structural motifs and extensive diversity across mammals
Scientific Reports (2023)
-
Gut Microbiota and Its Role in Anti-aging Phenomenon: Evidence-Based Review
Applied Biochemistry and Biotechnology (2023)
-
Safety and efficacy of a probiotic-containing infant formula supplemented with 2’-fucosyllactose: a double-blind randomized controlled trial
Nutrition Journal (2022)
-
Infant behavioral state and stool microbiome in infants receiving Lactocaseibacillus rhamnosus GG in formula: randomized controlled trial
BMC Pediatrics (2022)