Abstract
The orexin (also known as hypocretin) G protein–coupled receptors (GPCRs) regulate sleep and other behavioral functions in mammals, and are therapeutic targets for sleep and wake disorders. The human receptors hOX1R and hOX2R, which are 64% identical in sequence, have overlapping but distinct physiological functions and potential therapeutic profiles. We determined structures of hOX1R bound to the OX1R-selective antagonist SB-674042 and the dual antagonist suvorexant at 2.8-Å and 2.75-Å resolution, respectively, and used molecular modeling to illuminate mechanisms of antagonist subtype selectivity between hOX1R and hOX2R. The hOX1R structures also reveal a conserved amphipathic α-helix, in the extracellular N-terminal region, that interacts with orexin-A and is essential for high-potency neuropeptide activation at both receptors. The orexin-receptor crystal structures are valuable tools for the design and development of selective orexin-receptor antagonists and agonists.





Similar content being viewed by others
References
Marcus, J.N. et al. Differential expression of orexin receptors 1 and 2 in the rat brain. J. Comp. Neurol. 435, 6–25 (2001).
Li, J., Hu, Z. & de Lecea, L. The hypocretins/orexins: integrators of multiple physiological functions. Br. J. Pharmacol. 171, 332–350 (2014).
Sakurai, T. et al. Orexins and orexin receptors: a family of hypothalamic neuropeptides and G protein-coupled receptors that regulate feeding behavior. Cell 92, 573–585 (1998).
de Lecea, L. et al. The hypocretins: hypothalamus-specific peptides with neuroexcitatory activity. Proc. Natl. Acad. Sci. USA 95, 322–327 (1998).
Lin, L. et al. The sleep disorder canine narcolepsy is caused by a mutation in the hypocretin (orexin) receptor 2 gene. Cell 98, 365–376 (1999).
Chemelli, R.M. et al. Narcolepsy in orexin knockout mice: molecular genetics of sleep regulation. Cell 98, 437–451 (1999).
Willie, J.T. et al. Distinct narcolepsy syndromes in Orexin receptor-2 and Orexin null mice: molecular genetic dissection of non-REM and REM sleep regulatory processes. Neuron 38, 715–730 (2003).
Zheng, H., Patterson, L.M. & Berthoud, H.-R. Orexin signaling in the ventral tegmental area is required for high-fat appetite induced by opioid stimulation of the nucleus accumbens. J. Neurosci. 27, 11075–11082 (2007).
Harris, G.C., Wimmer, M. & Aston-Jones, G. A role for lateral hypothalamic orexin neurons in reward seeking. Nature 437, 556–559 (2005).
Boutrel, B. et al. Role for hypocretin in mediating stress-induced reinstatement of cocaine-seeking behavior. Proc. Natl. Acad. Sci. USA 102, 19168–19173 (2005).
Bingham, S. et al. Orexin-A, an hypothalamic peptide with analgesic properties. Pain 92, 81–90 (2001).
Johnson, P.L. et al. A key role for orexin in panic anxiety. Nat. Med. 16, 111–115 (2010).
Winrow, C.J. & Renger, J.J. Discovery and development of orexin receptor antagonists as therapeutics for insomnia. Br. J. Pharmacol. 171, 283–293 (2014).
Funato, H. et al. Enhanced orexin receptor-2 signaling prevents diet-induced obesity and improves leptin sensitivity. Cell Metab. 9, 64–76 (2009).
Xie, X. et al. Hypocretin/orexin and nociceptin/orphanin FQ coordinately regulate analgesia in a mouse model of stress-induced analgesia. J. Clin. Invest. 118, 2471–2481 (2008).
Aston-Jones, G., Smith, R.J., Moorman, D.E. & Richardson, K.A. Role of lateral hypothalamic orexin neurons in reward processing and addiction. Neuropharmacology 56 (suppl. 1), 112–121 (2009).
Cui, J.J. et al. Structure based drug design of crizotinib (PF-02341066), a potent and selective dual inhibitor of mesenchymal-epithelial transition factor (c-MET) kinase and anaplastic lymphoma kinase (ALK). J. Med. Chem. 54, 6342–6363 (2011).
Rosenbaum, D.M. et al. GPCR engineering yields high-resolution structural insights into beta2-adrenergic receptor function. Science 318, 1266–1273 (2007).
Caffrey, M. Crystallizing membrane proteins for structure determination: use of lipidic mesophases. Annu. Rev. Biophys. 38, 29–51 (2009).
White, J.F. et al. Structure of the agonist-bound neurotensin receptor. Nature 490, 508–513 (2012).
Yin, J., Mobarec, J.C., Kolb, P. & Rosenbaum, D.M. Crystal structure of the human OX2 orexin receptor bound to the insomnia drug suvorexant. Nature 519, 247–250 (2015).
Cox, C.D. et al. Discovery of the dual orexin receptor antagonist [(7R)-4-(5-chloro-1,3-benzoxazol-2-yl)-7-methyl-1,4-diazepan-1-yl][5-methyl-2-(2H-1,2,3-triazol-2-yl)phenyl]methanone (MK-4305) for the treatment of insomnia. J. Med. Chem. 53, 5320–5332 (2010).
Caffrey, M. & Cherezov, V. Crystallizing membrane proteins using lipidic mesophases. Nat. Protoc. 4, 706–731 (2009).
Palczewski, K. et al. Crystal structure of rhodopsin: A G protein-coupled receptor. Science 289, 739–745 (2000).
Hanson, M.A. et al. Crystal structure of a lipid G protein-coupled receptor. Science 335, 851–855 (2012).
Cox, C.D. et al. Conformational analysis of N,N-disubstituted-1,4-diazepane orexin receptor antagonists and implications for receptor binding. Bioorg. Med. Chem. Lett. 19, 2997–3001 (2009).
Langmead, C.J. et al. Characterisation of the binding of [3H]-SB-674042, a novel nonpeptide antagonist, to the human orexin-1 receptor. Br. J. Pharmacol. 141, 340–346 (2004).
Tran, D.-T. et al. Chimeric, mutant orexin receptors show key interactions between orexin receptors, peptides and antagonists. Eur. J. Pharmacol. 667, 120–128 (2011).
Putula, J. & Kukkonen, J.P. Mapping of the binding sites for the OX1 orexin receptor antagonist, SB-334867, using orexin/hypocretin receptor chimaeras. Neurosci. Lett. 506, 111–115 (2012).
Heifetz, A. et al. Study of human Orexin-1 and -2 G-protein-coupled receptors with novel and published antagonists by modeling, molecular dynamics simulations, and site-directed mutagenesis. Biochemistry 51, 3178–3197 (2012).
Malherbe, P. et al. Mapping the binding pocket of dual antagonist almorexant to human orexin 1 and orexin 2 receptors: comparison with the selective OX1 antagonist SB-674042 and the selective OX2 antagonist N-ethyl-2-[(6-methoxy-pyridin-3-yl)-(toluene-2-sulfonyl)-amino]-N-pyridin-3-ylmethyl-acetamide (EMPA). Mol. Pharmacol. 78, 81–93 (2010).
Müller, C.E. & Jacobson, K.A. in Methylxanthines Vol. 200 (ed. Fredholm, B.B.) 151–199 (Springer, 2011).
Huang, S.-Y. & Zou, X. Scoring and lessons learned with the CSAR benchmark using an improved iterative knowledge-based scoring function. J. Chem. Inf. Model. 51, 2097–2106 (2011).
Li, Y., Han, L., Liu, Z. & Wang, R. Comparative assessment of scoring functions on an updated benchmark: 2. Evaluation methods and general results. J. Chem. Inf. Model. 54, 1717–1736 (2014).
Friesner, R.A. et al. Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein-ligand complexes. J. Med. Chem. 49, 6177–6196 (2006).
Lyne, P.D., Lamb, M.L. & Saeh, J.C. Accurate prediction of the relative potencies of members of a series of kinase inhibitors using molecular docking and MM-GBSA scoring. J. Med. Chem. 49, 4805–4808 (2006).
Kim, H.-Y., Hong, E., Kim, J.-I. & Lee, W. Solution structure of human orexin-A: regulator of appetite and wakefulness. J. Biochem. Mol. Biol. 37, 565–573 (2004).
Lee, J.H. et al. Solution structure of a new hypothalamic neuropeptide, human hypocretin-2/orexin-B. Eur. J. Biochem. 266, 831–839 (1999).
Isberg, V. et al. GPCRDB: an information system for G protein-coupled receptors. Nucleic Acids Res. 42, D422–D425 (2014).
Rosenbaum, D.M. et al. Structure and function of an irreversible agonist-β2 adrenoceptor complex. Nature 469, 236–240 (2011).
Katritch, V., Cherezov, V. & Stevens, R.C. Structure-function of the G protein-coupled receptor superfamily. Annu. Rev. Pharmacol. Toxicol. 53, 531–556 (2013).
Ye, N., Neumeyer, J.L., Baldessarini, R.J., Zhen, X. & Zhang, A. Update 1 of: recent progress in development of dopamine receptor subtype-selective agents: potential therapeutics for neurological and psychiatric disorders. Chem. Rev. 113, PR123–PR178 (2013).
Granier, S. et al. Structure of the δ-opioid receptor bound to naltrindole. Nature 485, 400–404 (2012).
German, N.A., Decker, A.M., Gilmour, B.P., Thomas, B.F. & Zhang, Y. Truncated orexin peptides: structure-activity relationship studies. ACS Med. Chem. Lett. 4, 1224–1227 (2013).
Kufareva, I. et al. Stoichiometry and geometry of the CXC chemokine receptor 4 complex with CXC ligand 12: molecular modeling and experimental validation. Proc. Natl. Acad. Sci. USA 111, E5363–E5372 (2014).
Ahmed, S.S. et al. Antibodies to influenza nucleoprotein cross-react with human hypocretin receptor 2. Sci. Transl. Med. 7, 294ra105 (2015).
Zhao, H., Piszczek, G. & Schuck, P. SEDPHAT: a platform for global ITC analysis and global multi-method analysis of molecular interactions. Methods 76, 137–148 (2015).
Kobilka, B.K. Amino and carboxyl terminal modifications to facilitate the production and purification of a G protein-coupled receptor. Anal. Biochem. 231, 269–271 (1995).
Horcajada, C., Guinovart, J.J., Fita, I. & Ferrer, J.C. Crystal structure of an archaeal glycogen synthase: insights into oligomerization and substrate binding of eukaryotic glycogen synthases. J. Biol. Chem. 281, 2923–2931 (2006).
Karplus, P.A. & Diederichs, K. Linking crystallographic model and data quality. Science 336, 1030–1033 (2012).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data. Methods Enzymol. 276, 307–326 (1997).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Cowtan, K. Fitting molecular fragments into electron density. Acta Crystallogr. D Biol. Crystallogr. 64, 83–89 (2008).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Painter, J. & Merritt, E.A. TLSMD web server for the generation of multi-group TLS models. J. Appl. Crystallogr. 39, 109–111 (2006).
Schüttelkopf, A.W. & van Aalten, D.M.F. PRODRG: a tool for high-throughput crystallography of protein-ligand complexes. Acta Crystallogr. D Biol. Crystallogr. 60, 1355–1363 (2004).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Sastry, G.M., Adzhigirey, M., Day, T., Annabhimoju, R. & Sherman, W. Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J. Comput. Aided Mol. Des. 27, 221–234 (2013).
Jakubík, J., El-Fakahany, E.E. & Doležal, V. Towards predictive docking at aminergic G-protein coupled receptors. J. Mol. Model. 21, 284 (2015).
Bowers, K.J., Chow, E., Xu, H. & Dror, R.O. Scalable algorithms for molecular dynamics simulations on commodity clusters. Proc. 2006 ACM/IEEE Conf. Supercomputing (IEEE, 2006).
Shrake, A. & Rupley, J.A. Environment and exposure to solvent of protein atoms. Lysozyme and insulin. J. Mol. Biol. 79, 351–371 (1973).
Friesner, R.A. et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J. Med. Chem. 47, 1739–1749 (2004).
Hojayev, B., Rothermel, B.A., Gillette, T.G. & Hill, J.A. FHL2 binds calcineurin and represses pathological cardiac growth. Mol. Cell. Biol. 32, 4025–4034 (2012).
Keller, S. et al. High-precision isothermal titration calorimetry with automated peak-shape analysis. Anal. Chem. 84, 5066–5073 (2012).
Acknowledgements
We acknowledge support from the Welch Foundation (I-1770 to D.M.R.), the Searle Scholars Program (D.M.R.) and a Packard Foundation Fellowship (D.M.R.). The National Institute of General Medical Sciences and National Cancer Institute Structural Biology Facility at the Advanced Photon Source (GM/CA@APS) is funded in whole or in part with Federal funds from the US National Institutes of Health, National Cancer Institute (ACB-12002) and National Institute of General Medical Sciences (AGM-12006). This research used resources of the Advanced Photon Source, a US Department of Energy (DOE) Office of Science User Facility operated for the DOE Office of Science by Argonne National Laboratory under contract no. DE-AC02-06CH11357. We thank B. Rothermel, L. Lum and L. Zhang (University of Texas Southwestern Medical Center) for materials and assistance with the NFAT-luciferase assay, and S. Lee (University of Texas Southwestern Medical Center) for help with immunofluorescence.
Author information
Authors and Affiliations
Contributions
J.Y. expressed, purified and crystallized the hOX1R-PGS fusion protein, collected diffraction data, solved the structures, carried out ITC experiments and performed NFAT-luciferase assays. K.B. carried out computational docking and simulation experiments for orexin-receptor antagonists. C.A.B. and T.H.S. supervised, designed and analyzed ITC experiments. L.C. and Z.S. assisted with expression and purification of the hOX1R-PGS fusion protein. C.M.H. performed and analyzed radioligand binding assays to measure orexin receptor–ligand affinities. A.J.R. designed and synthesized 2-SORA-DMP. A.L.G., C.J.W., J.J.R. and P.J.C. analyzed docking and simulation experiments for orexin-receptor antagonists and devised experiments to measure receptor function. D.M.R. supervised the overall project, assisted with collection of diffraction data and wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
K.B., C.M.H., A.L.G., A.J.R., C.J.W., J.J.R. and P.J.C. are employees of Merck & Co., receive salary and research support from the company and may own stock and/or stock options in the company.
Integrated supplementary information
Supplementary Figure 1 Purification and crystallization of hOX1R-PGS.
(a) Superdex 200 size exclusion profiles of suvorexant-bound hOX1R-PGS (black) and SB-674042-bound hOX1R-PGS (purple).
(b) Coomassie stained PAGE gel of the isolated peak fractions from size exclusion chromatography. Left of markers: SB-674042-bound hOX1R-PGS; Right of markers: suvorexant-bound hOX1R-PGS.
(c) Lipidic cubic phase (LCP) crystallization setup for suvorexant-bound hOX1R-PGS
(d) Equivalent LCP setup for SB-674042-bound hOX1R-PGS.
Supplementary Figure 2 Electron density maps for hOX1R structures.
(a) Stereo view of 2Fo-Fc map, contoured at 1.5 σ, for the region of suvorexant-bound hOX1R containing the ligand, TM2, and TM5.
(b) Stereo view of 2Fo-Fc map, contoured at 1.5 σ, for the equivalent region of the SB-674042-bound hOX1R structure.
Supplementary Figure 3 Electron density and contact maps for the OXR binding pockets.
(a) Stereo view of 2Fo-Fc map, contoured at 1.5 σ, for the residues within 4 Å contact of the ligand in the hOX1R structure bound to SB-674042. Receptor is in blue sticks, ligand is in magenta. At right is a contact map for the antagonist made using LIGPLOT (Wallace, A. et al., Protein Eng. 8, 127-134, 1995).
Average B-factor for contact residues = 41.4 Å2, average B-factor for receptor = 52.8 Å2.
(b) Same as in (a), except for hOX1R (blue) bound to suvorexant (yellow).
Average B-factor for contact residues = 40.9 Å2, average B-factor for receptor = 51.8 Å2.
(c) Same as in (a), except for hOX2R (orange) bound to suvorexant (yellow), pdb 4S0V (Yin, J. et al., Nature 519, 247-250, 2015).
Average B-factor for contact residues = 30.5 Å2, average B-factor for receptor = 41.2 Å2.
Supplementary Figure 4 Molecular dynamics (MD) simulations of suvorexant in hOX1R and hOX2R.
(a) MD simulation trajectory for suvorexant-bound hOX1R. RMSD versus Time is shown for the Cα protein backbone (blue) and the ligand heavy atoms (magenta).
(b) Same as in a, except for suvorexant-bound hOX2R.
Supplementary Figure 5 Binding of 2-SORA-DMP to the orexin receptors.
Radioligand competition assays for 2-SORA-DMP binding to hOX1R (left) or hOX2R (right) membranes. For both panels, n = 3, replicates are from three separate competition assays on the same cell membrane stock. Error bars represent ± s.e.m.
Supplementary Figure 6 Sequence alignment of orexin-receptor N termini.
Alignment includes OX1R and OX2R sequences from Mus musculus (mouse), human, Bos taurus (bovine), Danio rerio (fish), Xenopus laevis (frog), and Canis lupus (dog). The magenta sequences comprise the N-terminal region that is ordered in the hOX1R structures. Sequences were downloaded from the GPCRDB (http://www.gpcr.org/7tm), and alignment was carried out with ClustalW2 (http://www.ebi.ac.uk/Tools/msa/clustalw2).
Supplementary Figure 7 Cell-surface immunofluorescence staining of orexin-receptor constructs.
Images for transfected CHO-K1 cell surface staining with Alexa488-conjugated M1-anti-FLAG antibody (Sigma) for different orexin receptor constructs (as in Rosenbaum, D.M. et al., Science 318, 1266-1273, 2007). Three separate fields from the same pool of stained cells are shown for each construct.
Supplementary Figure 8 Functional comparison of hOX1R I319 and hOX1R V319.
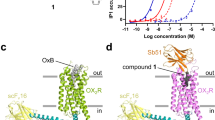
(a) Dose-response to orexin-A in CHO-K1 cell NFAT-luciferase activation assay. For a and b, n = 3, replicates are from three separate assays on the same transfected cells. Error bars represent ± s.d.
(b) Inhibition of orexin-A (20 nM) signal propagation by increasing concentrations of the antagonist SB-674042.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Table 1 (PDF 1643 kb)
Rights and permissions
About this article
Cite this article
Yin, J., Babaoglu, K., Brautigam, C. et al. Structure and ligand-binding mechanism of the human OX1 and OX2 orexin receptors. Nat Struct Mol Biol 23, 293–299 (2016). https://doi.org/10.1038/nsmb.3183
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3183
- Springer Nature America, Inc.
This article is cited by
-
Estradiol-dependent hypocretinergic/orexinergic behaviors throughout the estrous cycle
Psychopharmacology (2023)
-
Characteristic structural difference between inactive and active states of orexin 2 receptor determined using molecular dynamics simulations
Biophysical Reviews (2022)
-
Structural basis for ligand recognition of the neuropeptide Y Y2 receptor
Nature Communications (2021)
-
Structures of active-state orexin receptor 2 rationalize peptide and small-molecule agonist recognition and receptor activation
Nature Communications (2021)
-
Structure of an antagonist-bound ghrelin receptor reveals possible ghrelin recognition mode
Nature Communications (2020)