Abstract
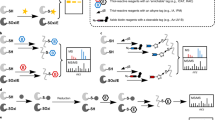
Protein S-sulfenylation is the reversible oxidative modification of cysteine thiol groups to form cysteine S-sulfenic acids. Mapping the specific sites of protein S-sulfenylation onto complex proteomes is crucial to understanding the molecular mechanisms controlling redox signaling and regulation. This protocol describes global, in situ, site-specific analysis of protein S-sulfenylation using sulfenic acid–specific chemical probes and mass spectrometry (MS)-based proteomics. The major steps in this protocol are as follows: (i) optimization of conditions for selective labeling of cysteine S-sulfenic acids in intact cells with the commercially available dimedone-based probe, DYn-2; (ii) tagging the modified cysteines with a functionalized biotin reagent containing a cleavable linker via Cu(I)-catalyzed azide-alkyne cycloaddition reaction; (iii) enrichment of the biotin-tagged tryptic peptides with streptavidin; (iv) liquid chromatography-tandem MS (LC-MS/MS)-based shotgun proteomics; and (v) computational data analysis. We also outline strategies for quantitative analysis of this modification in cells responding to redox perturbations and discuss special issues pertaining to experimental design of thiol redox studies. Our chemoproteomic platform should be broadly applicable to the investigation of other bio-orthogonal chemically engineered post-translational modifications. The entire analysis protocol takes ∼1 week to complete.





Similar content being viewed by others
References
Roos, G. & Messens, J. Protein sulfenic acid formation: from cellular damage to redox regulation. Free Radic. Biol. Med. 51, 314–326 (2011).
Poole, L.B. & Nelson, K.J. Discovering mechanisms of signaling-mediated cysteine oxidation. Curr. Opin. Chem. Biol. 12, 18–24 (2008).
Kettenhofen, N.J. & Wood, M.J. Formation, reactivity, and detection of protein sulfenic acids. Chem. Res. Toxicol. 23, 1633–1646 (2010).
Gupta, V. & Carroll, K.S. Sulfenic acid chemistry, detection and cellular lifetime. Biochim. Biophys. Acta 1840, 847–875 (2014).
Lee, S.R., Kwon, K.S., Kim, S.R. & Rhee, S.G. Reversible inactivation of protein-tyrosine phosphatase 1B in A431 cells stimulated with epidermal growth factor. J. Biol. Chem. 273, 15366–15372 (1998).
van Montfort, R.L., Congreve, M., Tisi, D., Carr, R. & Jhoti, H. Oxidation state of the active-site cysteine in protein tyrosine phosphatase 1B. Nature 423, 773–777 (2003).
Anastasiou, D. et al. Inhibition of pyruvate kinase M2 by reactive oxygen species contributes to cellular antioxidant responses. Science 334, 1278–1283 (2011).
Sobotta, M.C. et al. Peroxiredoxin-2 and STAT3 form a redox relay for H2O2 signaling. Nat. Chem. Biol. 11, 64–70 (2015).
Guo, Z., Kozlov, S., Lavin, M.F., Person, M.D. & Paull, T.T. ATM activation by oxidative stress. Science 330, 517–521 (2010).
Poole, L.B. The basics of thiols and cysteines in redox biology and chemistry. Free Radic. Biol. Med. 80, 148–157 (2015).
Claiborne, A., Miller, H., Parsonage, D. & Ross, R.P. Protein-sulfenic acid stabilization and function in enzyme catalysis and gene regulation. FASEB J. 7, 1483–1490 (1993).
Willett, W.S. & Copley, S.D. Identification and localization of a stable sulfenic acid in peroxide-treated tetrachlorohydroquinone dehalogenase using electrospray mass spectrometry. Chem. Biol. 3, 851–857 (1996).
Depuydt, M. et al. A periplasmic reducing system protects single cysteine residues from oxidation. Science 326, 1109–1111 (2009).
Wood, Z.A., Poole, L.B. & Karplus, P.A. Peroxiredoxin evolution and the regulation of hydrogen peroxide signaling. Science 300, 650–653 (2003).
Paulsen, C.E. & Carroll, K.S. Cysteine-mediated redox signaling: chemistry, biology, and tools for discovery. Chem. Rev. 113, 4633–4679 (2013).
Brandes, N. et al. Time line of redox events in aging postmitotic cells. Elife 2, e00306 (2013).
Rosenwasser, S. et al. Mapping the diatom redox-sensitive proteome provides insight into response to nitrogen stress in the marine environment. Proc. Natl. Acad. Sci. USA 111, 2740–2745 (2014).
Su, D. et al. Proteomic identification and quantification of S-glutathionylation in mouse macrophages using resin-assisted enrichment and isobaric labeling. Free Radic. Biol. Med. 67, 460–470 (2014).
Garcia-Santamarina, S. et al. Monitoring in vivo reversible cysteine oxidation in proteins using ICAT and mass spectrometry. Nat. Protoc. 9, 1131–1145 (2014).
Derakhshan, B., Wille, P.C. & Gross, S.S. Unbiased identification of cysteine S-nitrosylation sites on proteins. Nat. Protoc. 2, 1685–1691 (2007).
Hao, G., Derakhshan, B., Shi, L., Campagne, F. & Gross, S.S. SNOSID, a proteomic method for identification of cysteine S-nitrosylation sites in complex protein mixtures. Proc. Natl. Acad. Sci. USA 103, 1012–1017 (2006).
Leichert, L.I. et al. Quantifying changes in the thiol redox proteome upon oxidative stress in vivo. Proc. Natl. Acad. Sci. USA 105, 8197–8202 (2008).
Guo, J. et al. Resin-assisted enrichment of thiols as a general strategy for proteomic profiling of cysteine-based reversible modifications. Nat. Protoc. 9, 64–75 (2014).
Su, D. et al. Quantitative site-specific reactivity profiling of S-nitrosylation in mouse skeletal muscle using cysteinyl peptide enrichment coupled with mass spectrometry. Free Radic. Biol. Med. 57, 68–78 (2013).
Brandes, N., Reichmann, D., Tienson, H., Leichert, L.I. & Jakob, U. Using quantitative redox proteomics to dissect the yeast redoxome. J. Biol. Chem. 286, 41893–41903 (2011).
Devarie Baez, N.O., Reisz, J.A. & Furdui, C.M. Mass spectrometry in studies of protein thiol chemistry and signaling: opportunities and caveats. Free Radic. Biol. Med. 80, 191–211 (2014).
Reisz, J.A., Bechtold, E., King, S.B., Poole, L.B. & Furdui, C.M. Thiol-blocking electrophiles interfere with labeling and detection of protein sulfenic acids. FEBS J. 280, 6150–6161 (2013).
Benitez, L.V. & Allison, W.S. The inactivation of the acyl phosphatase activity catalyzed by the sulfenic acid form of glyceraldehyde 3-phosphate dehydrogenase by dimedone and olefins. J. Biol. Chem. 249, 6234–6243 (1974).
Poole, L.B. et al. Fluorescent and affinity-based tools to detect cysteine sulfenic acid formation in proteins. Bioconjug. Chem. 18, 2004–2017 (2007).
Wani, R. et al. Isoform-specific regulation of Akt by PDGF-induced reactive oxygen species. Proc. Natl. Acad. Sci. USA 108, 10550–10555 (2011).
Paulsen, C.E. et al. Peroxide-dependent sulfenylation of the EGFR catalytic site enhances kinase activity. Nat. Chem. Biol. 8, 57–64 (2012).
Truong, T.H. & Carroll, K.S. Bioorthogonal chemical reporters for analyzing protein sulfenylation in cells. Curr. Protoc. Chem. Biol. 4, 101–122 (2012).
Kulathu, Y. et al. Regulation of A20 and other OTU deubiquitinases by reversible oxidation. Nat. Commun. 4, 1569 (2013).
Yang, J., Gupta, V., Carroll, K.S. & Liebler, D.C. Site-specific mapping and quantification of protein S-sulfenylation in cells. Nat. Commun. 5, 4776 (2014).
Kim, H.Y., Tallman, K.A., Liebler, D.C. & Porter, N.A. An azido-biotin reagent for use in the isolation of protein adducts of lipid-derived electrophiles by streptavidin catch and photorelease. Mol. Cell. Proteomics 8, 2080–2089 (2009).
Wang, Z. et al. Enrichment and site mapping of O-linked N-acetylglucosamine by a combination of chemical/enzymatic tagging, photochemical cleavage, and electron transfer dissociation mass spectrometry. Mol. Cell. Proteomics 9, 153–160 (2010).
Speers, A.E. & Cravatt, B.F. A tandem orthogonal proteolysis strategy for high-content chemical proteomics. J. Am. Chem. Soc. 127, 10018–10019 (2005).
Weerapana, E., Speers, A.E. & Cravatt, B.F. Tandem orthogonal proteolysis-activity-based protein profiling (TOP-ABPP)—a general method for mapping sites of probe modification in proteomes. Nat. Protoc. 2, 1414–1425 (2007).
Zheng, T., Jiang, H. & Wu, P. Single-stranded DNA as a cleavable linker for bioorthogonal click chemistry-based proteomics. Bioconjug. Chem. 24, 859–864 (2013).
Szychowski, J. et al. Cleavable biotin probes for labeling of biomolecules via azide-alkyne cycloaddition. J. Am. Chem. Soc. 132, 18351–18360 (2010).
Qian, Y. et al. An isotopically tagged azobenzene-based cleavable linker for quantitative proteomics. Chembiochem 14, 1410–1414 (2013).
Hulce, J.J., Cognetta, A.B., Niphakis, M.J., Tully, S.E. & Cravatt, B.F. Proteome-wide mapping of cholesterol-interacting proteins in mammalian cells. Nat. Methods 10, 259–264 (2013).
Codreanu, S.G., Zhang, B., Sobecki, S.M., Billheimer, D.D. & Liebler, D.C. Global analysis of protein damage by the lipid electrophile 4-hydroxy-2-nonenal. Mol. Cell. Proteomics 8, 670–680 (2009).
Codreanu, S.G. et al. Alkylation damage by lipid electrophiles targets functional protein systems. Mol. Cell. Proteomics 13, 849–859 (2014).
Lanning, B.R. et al. A road map to evaluate the proteome-wide selectivity of covalent kinase inhibitors. Nat. Chem. Biol. 10, 760–767 (2014).
Martin, B.R. & Cravatt, B.F. Large-scale profiling of protein palmitoylation in mammalian cells. Nat. Methods 6, 135–138 (2009).
Martin, B.R., Wang, C., Adibekian, A., Tully, S.E. & Cravatt, B.F. Global profiling of dynamic protein palmitoylation. Nat. Methods 9, 84–89 (2012).
Weerapana, E. et al. Quantitative reactivity profiling predicts functional cysteines in proteomes. Nature 468, 790–795 (2010).
Wang, C., Weerapana, E., Blewett, M.M. & Cravatt, B.F. A chemoproteomic platform to quantitatively map targets of lipid-derived electrophiles. Nat. Methods 11, 79–85 (2014).
Deng, X. et al. Proteome-wide quantification and characterization of oxidation-sensitive cysteines in pathogenic bacteria. Cell Host Microbe 13, 358–370 (2013).
Presolski, S.I., Hong, V., Cho, S.H. & Finn, M.G. Tailored ligand acceleration of the Cu-catalyzed azide-alkyne cycloaddition reaction: practical and mechanistic implications. J. Am. Chem. Soc. 132, 14570–14576 (2010).
Lin, D., Li, J., Slebos, R.J. & Liebler, D.C. Cysteinyl peptide capture for shotgun proteomics: global assessment of chemoselective fractionation. J. Proteome Res. 9, 5461–5472 (2010).
Martinez-Acedo, P., Gupta, V. & Carroll, K.S. Proteomic analysis of peptides tagged with dimedone and related probes. J. Mass Spectrom. 49, 257–265 (2014).
Mertins, P. et al. Integrated proteomic analysis of post-translational modifications by serial enrichment. Nat. Methods 10, 634–637 (2013).
Alfaro, J.F. et al. Tandem mass spectrometry identifies many mouse brain O-GlcNAcylated proteins including EGF domain-specific O-GlcNAc transferase targets. Proc. Natl. Acad. Sci. USA 109, 7280–7285 (2012).
Doulias, P.T. et al. Structural profiling of endogenous S-nitrosocysteine residues reveals unique features that accommodate diverse mechanisms for protein S-nitrosylation. Proc. Natl. Acad. Sci. USA 107, 16958–16963 (2010).
Glatter, T. et al. Large-scale quantitative assessment of different in-solution protein digestion protocols reveals superior cleavage efficiency of tandem Lys-C/trypsin proteolysis over trypsin digestion. J. Proteome Res. 11, 5145–5156 (2012).
Guo, X., Trudgian, D.C., Lemoff, A., Yadavalli, S. & Mirzaei, H. Confetti: a multiprotease map of the HeLa proteome for comprehensive proteomics. Mol. Cell. Proteomics 13, 1573–1584 (2014).
Huesgen, P.F. et al. LysargiNase mirrors trypsin for protein C-terminal and methylation-site identification. Nat. Methods 12, 55–58 (2015).
Besanceney-Webler, C. et al. Increasing the efficacy of bioorthogonal click reactions for bioconjugation: a comparative study. Angew. Chem. Int. Ed. Engl. 50, 8051–8056 (2011).
Wang, W. et al. Sulfated ligands for the copper(I)-catalyzed azide-alkyne cycloaddition. Chem. Asian J. 6, 2796–2802 (2011).
Olsen, J.V. et al. Higher-energy C-trap dissociation for peptide modification analysis. Nat. Methods 4, 709–712 (2007).
Ma, Z.Q. et al. Supporting tool suite for production proteomics. Bioinformatics 27, 3214–3215 (2011).
Zhang, B. et al. Proteogenomic characterization of human colon and rectal cancer. Nature 513, 382–387 (2014).
Holman, J.D., Ma, Z.Q. & Tabb, D.L. Identifying proteomic LC-MS/MS data sets with Bumbershoot and IDPicker. Curr. Protoc. Bioinform. 37, 13.17.1–13.17.15 (2012).
Holman, J.D., Dasari, S. & Tabb, D.L. Informatics of protein and posttranslational modification detection via shotgun proteomics. Methods Mol. Biol. 1002, 167–179 (2013).
Tabb, D.L., Ma, Z.Q., Martin, D.B., Ham, A.J. & Chambers, M.C. DirecTag: accurate sequence tags from peptide MS/MS through statistical scoring. J. Proteome Res. 7, 3838–3846 (2008).
Dasari, S. et al. TagRecon: high-throughput mutation identification through sequence tagging. J. Proteome Res. 9, 1716–1726 (2010).
Ma, Z.Q. et al. IDPicker 2.0: improved protein assembly with high discrimination peptide identification filtering. J. Proteome Res. 8, 3872–3881 (2009).
Tabb, D.L., Friedman, D.B. & Ham, A.J. Verification of automated peptide identifications from proteomic tandem mass spectra. Nat. Protoc. 1, 2213–2222 (2006).
Chen, Y., Kwon, S.W., Kim, S.C. & Zhao, Y. Integrated approach for manual evaluation of peptides identified by searching protein sequence databases with tandem mass spectra. J. Proteome Res. 4, 998–1005 (2005).
Schilling, B. et al. Platform-independent and label-free quantitation of proteomic data using MS1 extracted ion chromatograms in skyline: application to protein acetylation and phosphorylation. Mol. Cell. Proteomics 11, 202–214 (2012).
MacLean, B. et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26, 966–968 (2010).
Ong, S.E. et al. Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol. Cell. Proteomics 1, 376–386 (2002).
Gygi, S.P. et al. Quantitative analysis of complex protein mixtures using isotope-coded affinity tags. Nat. Biotechnol. 17, 994–999 (1999).
Ross, P.L. et al. Multiplexed protein quantitation in Saccharomyces cerevisiae using amine-reactive isobaric tagging reagents. Mol. Cell. Proteomics 3, 1154–1169 (2004).
Grammel, M. & Hang, H.C. Chemical reporters for biological discovery. Nat. Chem. Biol. 9, 475–484 (2013).
Simon, G.M., Niphakis, M.J. & Cravatt, B.F. Determining target engagement in living systems. Nat. Chem. Biol. 9, 200–205 (2013).
Acknowledgements
This work was supported by US National Institutes of Health (NIH) grants U24CA159988 and R01GM102187.
Author information
Authors and Affiliations
Contributions
J.Y. designed the protocol, optimized experimental conditions for applications and wrote the manuscript; V.G. synthesized the DYn-2-d6; K.A.T. synthesized Az-UV-biotin; and N.A.P., K.S.C. and D.C.L. supervised the project and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
D.C.L. is a stockholder in Protypia, Incorporated.
Integrated supplementary information
Supplementary Figure 1 Spin columns and sample processing devices.
(a) MacroSpin SCX column. The Eppendorf centrifuge device (b) and VIAFLO electronic multi-channel pipettes are used for multiple-sample processing.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Discussion (PDF 1404 kb)
Rights and permissions
About this article
Cite this article
Yang, J., Gupta, V., Tallman, K. et al. Global, in situ, site-specific analysis of protein S-sulfenylation. Nat Protoc 10, 1022–1037 (2015). https://doi.org/10.1038/nprot.2015.062
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2015.062
- Springer Nature Limited
This article is cited by
-
A modification-centric assessment tool for the performance of chemoproteomic probes
Nature Chemical Biology (2022)
-
Wittig reagents for chemoselective sulfenic acid ligation enables global site stoichiometry analysis and redox-controlled mitochondrial targeting
Nature Chemistry (2021)
-
The redox language in neurodegenerative diseases: oxidative post-translational modifications by hydrogen peroxide
Cell Death & Disease (2021)
-
A quantitative thiol reactivity profiling platform to analyze redox and electrophile reactive cysteine proteomes
Nature Protocols (2020)
-
Prediction of S-Sulfenylation Sites Using Statistical Moments Based Features via CHOU’S 5-Step Rule
International Journal of Peptide Research and Therapeutics (2020)