Abstract
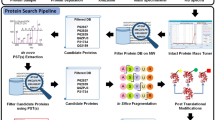
Frequently, proteomic LC-MS/MS data may contain sets of modifications that evade identification during standard database search. For many laboratories, the standard technique to seek posttranslational modifications (PTMs) adds a short list of specified mass shifts to database search configuration. This technique provides information for only the specified PTMs, takes substantial time to run, and drives false discoveries upward through an exponential expansion of search space. This protocol describes a more structured approach to blind PTM discovery through reducing protein lists, targeting attention to a data-driven list of mass shifts, and seeking the resulting short list of modifications through targeted search.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Eng JK, McCormack AL, Yates JR (1994) An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J Am Soc Mass Spectrom 5:976–989
Yates JR, Eng JK, McCormack AL, Schieltz D (1995) Method to correlate tandem mass spectra of modified peptides to amino acid sequences in the protein database. Anal Chem 67:1426–1436
Craig R, Beavis RC (2003) A method for reducing the time required to match protein sequences with tandem mass spectra. Rapid Commun Mass Spectrom 17:2310–2316
Mann M, Wilm M (1994) Error-tolerant identification of peptides in sequence databases by peptide sequence tags. Anal Chem 66:4390–4399
Tabb DL, Saraf A, Yates JR (2003) GutenTag: high-throughput sequence tagging via an empirically derived fragmentation model. Anal Chem 75:6415–6421
Tanner S, Shu H, Frank A et al (2005) InsPecT: identification of posttranslationally modified peptides from tandem mass spectra. Anal Chem 77:4626–4639
Tsur D, Tanner S, Zandi E, Bafna V, Pevzner PA (2005) Identification of post-translational modifications by blind search of mass spectra. Nat Biotechnol 23:1562–1567
Dasari S, Chambers MC, Codreanu SG et al (2011) Sequence tagging reveals unexpected modifications in toxicoproteomics. Chem Res Toxicol 24:204–216
Tabb DL, Fernando CG, Chambers MC (2007) MyriMatch: highly accurate tandem mass spectral peptide identification by multivariate hypergeometric analysis. J Proteome Res 6:654–661
Tabb DL, Ma Z-Q, Martin DB, Ham A-JL, Chambers MC (2008) DirecTag: accurate sequence tags from peptide MS/MS through statistical scoring. J Proteome Res 7:3838–3846
Dasari S, Chambers MC, Slebos RJ, Zimmerman LJ, Ham A-JL, Tabb DL (2010) TagRecon: high-throughput mutation identification through sequence tagging. J Proteome Res 9:1716–1726
Zhang B, Chambers MC, Tabb DL (2007) Proteomic parsimony through bipartite graph analysis improves accuracy and transparency. J Proteome Res 6:3549–3557
Ma Z-Q, Dasari S, Chambers MC et al (2009) IDPicker 2.0: improved protein assembly with high discrimination peptide identification filtering. J Proteome Res 8:3872–3881
Holman JD, Ma Z-Q, Tabb DL (2012) Identifying proteomic LC-MS/MS data sets with Bumbershoot and IDPicker, Current protocols in bioinformatics/editorial board, Andreas D. Baxevanis. Chapter 13, Unit 13.17
Sprung RW Jr, Brock JWC, Tanksley JP et al (2009) Equivalence of protein inventories obtained from formalin-fixed paraffin-embedded and frozen tissue in multidimensional liquid chromatography-tandem mass spectrometry shotgun proteomic analysis. Mol Cell Proteomics 8:1988–1998
Acknowledgments
The algorithms described in this protocol were developed through support to all three authors by R01 CA126218. In addition, J.D.H. was supported by U01 CA08402.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Holman, J.D., Dasari, S., Tabb, D.L. (2013). Informatics of Protein and Posttranslational Modification Detection via Shotgun Proteomics. In: Zhou, M., Veenstra, T. (eds) Proteomics for Biomarker Discovery. Methods in Molecular Biology, vol 1002. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-360-2_14
Download citation
DOI: https://doi.org/10.1007/978-1-62703-360-2_14
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-359-6
Online ISBN: 978-1-62703-360-2
eBook Packages: Springer Protocols