Abstract
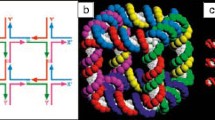
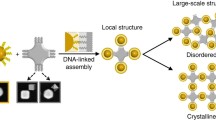
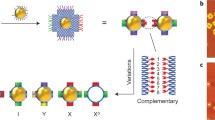
Advances in self-assembly over the past decade have demonstrated that nano- and microscale particles can be organized into a large diversity of ordered three-dimensional (3D) lattices. However, the ability to generate different desired lattice types from the same set of particles remains challenging. Here, we show that nanoparticles can be assembled into crystalline and open 3D frameworks by connecting them through designed DNA-based polyhedral frames. The geometrical shapes of the frames, combined with the DNA-assisted binding properties of their vertices, facilitate the well-defined topological connections between particles in accordance with frame geometry. With this strategy, different crystallographic lattices using the same particles can be assembled by introduction of the corresponding DNA polyhedral frames. This approach should facilitate the rational assembly of nanoscale lattices through the design of the unit cell.





Similar content being viewed by others
References
Talapin, D. V., Lee, J.-S., Kovalenko, M. V. & Shevchenko, E. V. Prospects of colloidal nanocrystals for electronic and optoelectronic applications. Chem. Rev. 110, 389–458 (2010).
Hu, T. et al. Self-organization of plasmonic and excitonic nanoparticles into resonant chiral supraparticle assemblies. Nano Lett. 14, 6799–6810 (2014).
Cargnello, M. et al. Substitutional doping in nanocrystal superlattices. Nature 524, 450–453 (2015).
Kuzyk, A. et al. DNA-based self-assembly of chiral plasmonic nanostructures with tailored optical response. Nature 483, 311–314 (2012).
Xiong, H., Sfeir, M. Y. & Gang, O. Assembly, structure and optical response of three-dimensional dynamically tunable multicomponent superlattices. Nano Lett. 10, 4456–4462 (2010).
Alder, B. J. & Wainwright, T. E. Phase transition for a hard sphere system. J. Chem. Phys. 27, 1208–1209 (1957).
Zhu, J. X. et al. Crystallization of hard-sphere colloids in microgravity. Nature 387, 883–885 (1997).
Casey, M. T. et al. Driving diffusionless transformations in colloidal crystals using DNA handshaking. Nature Commun. 3, 1209 (2012).
Travesset, A. Binary nanoparticle superlattices of soft-particle systems. Proc. Natl Acad. Sci. USA 112, 9563–9567 (2015).
Kranendonk, W. G. T. & Frenkel, D. Simulation of the adhesive-hard-sphere model. Mol. Phys. 64, 403–424 (1988).
Damasceno, P. F., Engel, M. & Glotzer, S. C. Predictive self-assembly of polyhedra into complex structures. Science 337, 453–457 (2012).
Leunissen, M. E. et al. Ionic colloidal crystals of oppositely charged particles. Nature 437, 235–240 (2005).
Shevchenko, E. V., Talapin, D. V., Kotov, N. A., O’Brien, S. & Murray, C. B. Structural diversity in binary nanoparticle superlattices. Nature 439, 55–59 (2006).
Nykypanchuk, D., Maye, M. M., van der Lelie, D. & Gang, O. DNA-guided crystallization of colloidal nanoparticles. Nature 451, 549–552 (2008).
Park, S. Y. et al. DNA-programmable nanoparticle crystallization. Nature 451, 553–556 (2008).
Auyeung, E. et al. Synthetically programmable nanoparticle superlattices using a hollow three-dimensional spacer approach. Nature Nanotech. 7, 24–28 (2012).
Talapin, D. V. et al. Quasicrystalline order in self-assembled binary nanoparticle superlattices. Nature 461, 964–967 (2009).
Rechtsman, M. C., Stillinger, F. H. & Torquato, S. Synthetic diamond and wurtzite structures self-assemble with isotropic pair interactions. Phys. Rev. E 75, 031403 (2007).
Jain, A., Errington, J. R. & Truskett, T. M. Dimensionality and design of isotropic interactions that stabilize honeycomb, square, simple cubic, and diamond lattices. Phys. Rev. X 4, 031049 (2014).
Tkachenko, A. V. Morphological diversity of DNA-colloidal self-assembly. Phys. Rev. Lett. 89, 148303 (2002).
Angioletti-Uberti, S., Varilly, P., Mognetti, B. M. & Frenkel, D. Mobile linkers on DNA-coated colloids: valency without patches. Phys. Rev. Lett. 113, 128308 (2014).
Glotzer, S. C. & Solomon, M. J. Anisotropy of building blocks and their assembly into complex structures. Nature Mater. 6, 557–562 (2007).
Jones, M. R. et al. DNA-nanoparticle superlattices formed from anisotropic building blocks. Nature Mater. 9, 913–917 (2010).
Zhang, Y., Lu, F., Lelie, D. v. d. & Gang, O. Continuous phase transformation in nanocube assemblies. Phys. Rev. Lett. 107, 135701 (2011).
Xiong, H. M., van der Lelie, D. & Gang, O. Phase behavior of nanoparticles assembled by DNA linkers. Phys. Rev. Lett. 102, 015504 (2009).
van Anders, G. et al. Entropically patchy particles: engineering valence through shape entropy. ACS Nano 8, 931–940 (2014).
Macfarlane, R. J. et al. Nanoparticle superlattice engineering with DNA. Science 334, 204–208 (2011).
Paik, T. & Murray, C. B. Shape-directed binary assembly of anisotropic nanoplates: a nanocrystal puzzle with shape-complementary building blocks. Nano Lett. 13, 2952–2956 (2013).
Romano, F., Sanz, E. & Sciortino, F. Phase diagram of a tetrahedral patchy particle model for different interaction ranges. J. Chem. Phys. 132, 184501 (2010).
Russo, J., Tartaglia, P. & Sciortino, F. Association of limited valence patchy particles in two dimensions. Soft Matter 6, 4229–4236 (2010).
Chen, Q., Bae, S. C. & Granick, S. Directed self-assembly of a colloidal kagome lattice. Nature 469, 381–384 (2012).
Wang, Y. et al. Colloids with valence and specific directional bonding. Nature 491, 51–56 (2012).
Lu, F., Yager, K. G., Zhang, Y., Xin, H. & Gang, O. Superlattices assembled through shape-induced directional binding. Nature Commun. 6, 6912 (2015).
O’Brien, M. N., Jones, M. R., Lee, B. & Mirkin, C. A. Anisotropic nanoparticle complementarity in DNA-mediated co-crystallization. Nature Mater. 14, 833–839 (2015).
Edwardson, T. G. W., Carneiro, K. M. M., McLaughlin, C. K., Serpell, C. J. & Sleiman, H. F. Site-specific positioning of dendritic alkyl chains on DNA cages enables their geometry-dependent self-assembly. Nature Chem. 5, 868–875 (2013).
Suzuki, K., Hosokawa, K. & Maeda, M. Controlling the number and positions of oligonucleotides on gold nanoparticle surfaces. J. Am. Chem. Soc. 131, 7518–7519 (2009).
Ye, X. et al. Shape alloys of nanorods and nanospheres from self-assembly. Nano Lett. 13, 4980–4988 (2013).
Zheng, J. P. et al. From molecular to macroscopic via the rational design of a self-assembled 3D DNA crystal. Nature 461, 74–77 (2009).
Rogers, W. B. & Manoharan, V. N. Programming colloidal phase transitions with DNA strand displacement. Science 347, 639–642 (2015).
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009).
Zhao, Z., Jacovetty, E. L., Liu, Y. & Yan, H. Encapsulation of gold nanoparticles in a DNA origami cage. Angew. Chem. Int. Ed. 50, 2041–2044 (2011).
Schreiber, R. et al. Hierarchical assembly of metal nanoparticles, quantum dots and organic dyes using DNA origami scaffolds. Nature Nanotech. 9, 74–78 (2014).
Seeman, N. C. Nucleic-acid junctions and lattices. J. Theor. Biol. 99, 237–247 (1982).
Douglas, S. M. et al. Rapid prototyping of 3D DNA-origami shapes with caDNAno. Nucleic Acids Res. 37, 5001–5006 (2009).
Tian, Y. et al. Prescribed nanoparticle cluster architectures and low-dimensional arrays built using octahedral DNA origami frames. Nature Nanotech. 10, 637–644 (2015).
Yager, K. G., Zhang, Y. G., Lu, F. & Gang, O. Periodic lattices of arbitrary nano-objects: modeling and applications for self-assembled systems. J. Appl. Crystallogr. 47, 118–129 (2014).
Zhang, Y. G. et al. Selective transformations between nanoparticle superlattices via the reprogramming of DNA-mediated interactions. Nature Mater. 14, 840–847 (2015).
Mao, X., Souslov, A., Mendoza, C. I. & Lubensky, T. C. Mechanical instability at finite temperature. Nature Commun. 6, 5968 (2015).
Haji-Akbari, A., Chen, E. R., Engel, M. & Glotzer, S. C. Packing and self-assembly of truncated triangular bipyramids. Phys. Rev. B 88, 012127 (2013).
Acknowledgements
We thank W. Shih and Y. Ke for help with DNA octahedra design and useful discussions. We thank L. Bai for help with the cryo-STEM sample preparations. We thank D. Chen for assistance with schematic drawing. Research carried out at the Center for Functional Nanomaterials, Brookhaven National Laboratory was supported by the US Department of Energy, Office of Basic Energy Sciences, under Contract No. DE-SC0012704. H.L. and T.W. were supported by a National Institute of Health R01 grant (AG029979).
Author information
Authors and Affiliations
Contributions
Y.T. and O.G. conceived and designed the experiments. Y.T. performed the experiments. Y.Z. contributed to the model fitting of the SAXS data. Y.T., T.W. and H.L. contributed to the cryo-EM imaging and reconstruction. Y.T. and H.L.X. contributed to the STEM imaging of the superlattice. Y.T., Y.Z. and O.G. analysed the data. Y.T. and O.G. wrote the paper. O.G. supervised the project. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 5159 kb)
Rights and permissions
About this article
Cite this article
Tian, Y., Zhang, Y., Wang, T. et al. Lattice engineering through nanoparticle–DNA frameworks. Nature Mater 15, 654–661 (2016). https://doi.org/10.1038/nmat4571
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmat4571
- Springer Nature Limited
This article is cited by
-
Supercrystal engineering of atomically precise gold nanoparticles promoted by surface dynamics
Nature Chemistry (2023)
-
Three-dimensional electron ptychography of organic–inorganic hybrid nanostructures
Nature Communications (2022)
-
Open-channel metal particle superlattices
Nature (2022)
-
DNA origami
Nature Reviews Methods Primers (2021)
-
Designed and biologically active protein lattices
Nature Communications (2021)