Abstract
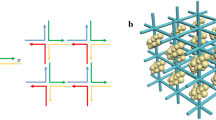
The science of self-assembly has undergone a radical shift from asking questions about why individual components self-organize into ordered structures, to manipulating the resultant order. However, the quest for far-reaching nanomanufacturing requires addressing an even more challenging question: how to form nanoparticle (NP) structures with designed architectures without explicitly prescribing particle positions. Here we report an assembly concept in which building instructions are embedded into NPs via DNA frames. The integration of NPs and DNA origami frames enables the fabrication of NPs with designed anisotropic and selective interactions. Using a pre-defined set of different DNA-framed NPs, we show it is possible to design diverse planar architectures, which include periodic structures and shaped meso-objects that spontaneously emerge on mixing of the different topological types of NP. Even objects of non-trivial shapes, such as a nanoscale model of Leonardo da Vinci's Vitruvian Man, can be self-assembled successfully.




Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.References
Damasceno, P. F., Engel, M. & Glotzer, S. C. Predictive self-assembly of polyhedral into complex structures. Science 337, 453–457 (2012).
Agarwal, U. & Escobedo, F. A. Mesophase behaviour of polyhedral particles. Nature Mater. 10, 230–235 (2011).
Whitelam, S., Tamblyn, I., Garrahan, J. P. & Beton, P. H. Emergent rhombus tilings from molecular interactions with M-fold rotational symmetry. Phys. Rev. Lett. 114, 115702 (2015).
Jain, A., Errington, J. R. & Truskett, T. M. Inverse design of simple pairwise interactions with low-coordinated 3D lattice ground states. Soft Matter 9, 3866–3870 (2013).
Nykypanchuk, D., Maye, M. M., van der Lelie, D. & Gang, O. DNA-guided crystallization of colloidal nanoparticles. Nature 451, 549–552 (2008).
Park, S. Y. et al. DNA-programmable nanoparticle crystallization. Nature 451, 553–556 (2008).
Podsiadlo, P., Krylova, G. V., Demortiere, A. & Shevchenko, E. V. Multicomponent periodic nanoparticle superlattices. J. Nanopart. Res. 13, 15–32 (2011).
Xu, L. et al. Nanoparticle assemblies: dimensional transformation of nanomaterials and scalability. Chem. Soc. Rev. 42, 3114–3126 (2013).
Talapin, D. V. et al. Quasicrystalline order in self-assembled binary nanoparticle superlattices. Nature 461, 964–967 (2009).
Whitelam, S., Schulman, R. & Hedges, L. Self-assembly of multicomponent structures in and out of equilibrium. Phys. Rev. Lett. 109, 265506 (2012).
Barish, R. D., Schulman, R., Rothemund, P. W. K. & Winfree, E. An information-bearing seed for nucleating algorithmic self-assembly. Proc. Natl Acad. Sci. USA 106, 6054–6059 (2009).
Romano, F. & Sciortino, F. Two dimensional assembly of triblock Janus particles into crystal phases in the two bond per patch limit. Soft Matter 7, 5799–5804 (2011).
Smallenburg, F. & Sciortino, F. Liquids more stable than crystals in particles with limited valence and flexible bonds. Nature Phys. 9, 554–558 (2013).
Yi, G. R., Pine, D. J. & Sacanna, S. Recent progress on patchy colloids and their self-assembly. J. Phys. Condensed Matter 25, 193101 (2013).
van Ravensteijn, B. G. P., Kamp, M., van Blaaderen, A. & Kegel, W. K. General route toward chemically anisotropic colloids. Chem. Mater. 25, 4348–4353 (2013).
Wang, Y. F. et al. Colloids with valence and specific directional bonding. Nature 491, 51–61 (2012).
Ye, X. et al. Competition of shape and interaction patchiness for self-assembling nanoplates. Nature Chem. 5, 466–473 (2013).
Klinkova, A., Therien-Aubin, H., Choueiri, R. M., Rubinstein, M. & Kumacheva, E. Colloidal analogs of molecular chain stoppers. Proc. Natl Acad. Sci. USA 110, 18775–18779 (2013).
Lau, K. L., Hamblin, G. D. & Sleiman, H. F. Gold nanoparticle 3D-DNA building blocks: high purity preparation and use for modular access to nanoparticle assemblies. Small 10, 660–666 (2014).
Sun, D. Z. et al. Heterogeneous nanoclusters assembled by PNA-templated double-stranded DNA. Nanoscale 4, 6722–6725 (2012).
Jones, M. R. et al. DNA-nanoparticle superlattices formed from anisotropic building blocks. Nature Mater. 9, 913–917 (2010).
Chen, Q., Bae, S. C. & Granick, S. Directed self-assembly of a colloidal Kagome lattice. Nature 469, 381–384 (2011).
Knorowski, C. & Travesset, A. Self-assembly and crystallization of hairy (f-star) and DNA-grafted nanocubes. J. Am. Chem. Soc. 136, 653–659 (2014).
Liu, W. et al. Diamond family of nanoparticle superlattices. Science 351, 582–586 (2016).
Tian, Y., Wang, T., Zhang, Y., Li, H. & Gang, O. Lattice engineering via nanoparticle-DNA frameworks. Nature Mater. 15, 654–661 (2016).
Halverson, J. D. & Tkachenko, A. V. DNA-programmed mesoscopic architecture. Phys Rev. E 87, 062310 (2013).
Tkachenko, A. V. Theory of programmable hierarchic self-assembly. Phys. Rev. Lett. 106, 255501 (2011).
Murugan, A., Zou, J. & Brenner, M. P. Undesired usage and the robust self-assembly of heterogeneous structures. Nature Commun. 6, 6203 (2015).
Jacobs, W. M., Reinhardt, A. & Frenkel, D. Rational design of self-assembly pathways for complex multicomponent structures. Proc. Natl Acad. Sci. USA 112, 6313–6318 (2015).
Murugan, A., Zeravcic, Z., Brenner, M. P. & Leibler, S. Multifarious assembly mixtures: systems allowing retrieval of diverse stored structures. Proc. Natl Acad. Sci. USA 112, 54–59, (2015).
Ruzicka, B. et al. Observation of empty liquids and equilibrium gels in a colloidal clay. Nature Mater. 10, 56–60 (2011).
Feng, L., Dreyfus, R., Sha, R. J., Seeman, N. C. & Chaikin, P. M. DNA patchy particles. Adv. Mater. 25, 2779–2783 (2013).
Kraft, D. J. et al. Patchy polymer colloids with tunable anisotropy dimensions. J. Phys. Chem. B 115, 7175–7181 (2011).
Romano, F., Sanz, E. & Sciortino, F. Crystallization of tetrahedral patchy particles in silico. J. Chem. Phys. 134, 174502 (2011).
Hamblin, G. D., Rahbani, J. F. & Sleiman, H. F. Sequential growth of long DNA strands with user-defined patterns for nanostructures and scaffolds. Nature Commun. 6, 7065 (2015).
Alivisatos, A. P. et al. Organization of ‘nanocrystal molecules’ using DNA. Nature 382, 609–611 (1996).
Maye, M. M., Nykypanchuk, D., Cuisinier, M., van der Lelie, D. & Gang, O. Stepwise surface encoding for high-throughput assembly of nanoclusters. Nature Mater. 8, 388–391 (2009).
Kim, J. W., Kim, J. H. & Deaton, R. DNA-linked nanoparticle building blocks for programmable matter. Angew. Chem. Int. Ed. 50, 9185–9190 (2011).
Winfree, E., Liu, F., Wenzler, L. A. & Seeman, N. C. Design and self-assembly of two-dimensional DNA crystals. Nature 394, 539–544 (1998).
Yan, H., Park, S. H., Finkelstein, G., Reif, J. H. & LaBean, T. H. DNA-templated self-assembly of protein arrays and highly conductive nanowires. Science 301, 1882–1884 (2003).
He, Y., Chen, Y., Liu, H., Ribbe, A. E. & Mao, C. Self-assembly of hexagonal DNA two-dimensional (2D) arrays. J. Am. Chem. Soc. 127, 12202–12203 (2005).
Liu, W., Zhong, H., Wang, R. & Seeman, N. C. Crystalline two-dimensional DNA-origami arrays. Angew. Chem. Int. Ed. 50, 264–267 (2011).
Liu, Y., Ke, Y. & Yan, H. Self-assembly of symmetric finite-size DNA nanoarrays. J. Am. Chem. Soc. 127, 17140–17141 (2005).
Zheng, J. et al. From molecular to macroscopic via the rational design of a self-assembled 3D DNA crystal. Nature 461, 74–77 (2009).
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Woo, S. & Rothemund, P. W. K. Self-assembly of two-dimensional DNA origami lattices using cation-controlled surface diffusion. Nature Commun. 5, 4889 (2014).
Ke, Y., Ong, L. L., Shih, W. M. & Yin, P. Three-dimensional structures self-assembled from DNA bricks. Science 338, 1177–1183 (2012).
Flory, P. J. Principles of Polymer Chemistry (Cornell Univ. Press, 1953).
Zhang, Y. G., Lu, F., Yager, K. G., van der Lelie, D. & Gang, O. A general strategy for the DNA-mediated self-assembly of functional nanoparticles into heterogeneous systems. Nature Nanotech. 8, 865–872 (2013).
Zhang, C. et al. A general approach to DNA-programmable atom equivalents. Nature Mater. 12, 741–746 (2013).
Seeman, N. C. De novo design of sequences for nucleic acid structural engineering. J. Biomol. Struct. Dyn. 8, 573–581 (1990).
Acknowledgements
Research carried out at the Center for Functional Nanomaterials, Brookhaven National Laboratory, was supported by the US Department of Energy, Office of Basic Energy Sciences, under Contract No. DE-AC02-98CH10886.
Author information
Authors and Affiliations
Contributions
W.L. and O.G. conceived and designed the experiments. W.L. performed the experiments. W.L., Y.T. and O.G. analysed the data. J.H. and A.V.T. contributed to the theoretical/numerical analysis. W.L. and O.G. wrote the paper. O.G. supervised the projects. All the authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Liu, W., Halverson, J., Tian, Y. et al. Self-organized architectures from assorted DNA-framed nanoparticles. Nature Chem 8, 867–873 (2016). https://doi.org/10.1038/nchem.2540
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.2540
- Springer Nature Limited
This article is cited by
-
Chirally assembled plasmonic metamolecules from intrinsically chiral nanoparticles
Nano Research (2022)
-
Designed and biologically active protein lattices
Nature Communications (2021)
-
Self-regulated co-assembly of soft and hard nanoparticles
Nature Communications (2021)
-
DNA origami
Nature Reviews Methods Primers (2021)
-
How to design an icosahedral quasicrystal through directional bonding
Nature (2021)