Abstract
The number of naive T cells decreases and susceptibility to new microbial infections increases with age. Here we describe a previously unknown subset of phenotypically naive human CD8+ T cells that rapidly secreted multiple cytokines in response to persistent viral antigens but differed transcriptionally from memory and effector T cells. The frequency of these CD8+ T cells, called 'memory T cells with a naive phenotype' (TMNP cells), increased with age and after severe acute infection and inversely correlated with the residual capacity of the immune system to respond to new infections with age. CD8+ TMNP cells represent a potential new target for the immunotherapy of persistent infections and should be accounted for and subtracted from the naive pool if truly naive T cells are needed to respond to antigens.
Similar content being viewed by others
Main
Protective immunity to new infections requires a sufficient number and diversity of naive T lymphocytes (TN lymphocytes), with strong expansion and effector-differentiation potential1. With age, the human TN cell pool shrinks2 and may or may not lose diversity3,4, and old TN cells exhibit proliferation and effector-differentiation defects5,6,7,8. This probably precipitates the vulnerability of older adults to new and re-emerging infections, such as infection with influenza virus, West Nile virus (WNV) and so on, and limits the efficacy of vaccination against infectious diseases9,10.
Drivers that contribute to the age-related diminishment in the homeostasis and function of TN cells include thymic involution11, impaired peripheral maintenance of T cells12, 'homeostatic' conversion to memory phenotype(s)12 and repeated antigen exposure due to persistent infections3,13. However, the extent of the qualitative and quantitative age-related diminishment in the homeostasis and function of TN cells remains incompletely understood.
T cell phenotype has long been used as means of functionally classifying T cell subsets14. For example, TN cells exhibit no immediate effector functions14, whereas effector plus effector-memory T cells (TE+EM cells), effector-memory T cells that re-express the naive-cell marker CD45RA (TEMRA cells) and, to a lesser extent, central-memory T cells (TCM cells) can rapidly express multiple different effector molecules (cytokines and cytotoxic molecules such as granzymes and perforin) after being stimulated with antigen, which enables rapid control of reinfection. TCM cells, which are less polyfunctional, reside mainly in secondary lymphoid organs and maintain high proliferative potential15,16. Memory T cells (TM cells) and TN cells are maintained by interleukin 7 (IL-7) and IL-15, respectively17.
While testing the function of human T cells with aging, we discovered a subset of phenotypically TN cells able to produce effector cytokines immediately after stimulation via the T cell antigen receptor (TCR). These memory T cells with a naive phenotype (called 'TMNP cells' here) were dominantly CD8+, exhibited a transcriptome distinct from that of other T cell subsets and increased in frequency with age. TMNP cells responded to antigens from persistent viruses. Their frequencies and numbers were expanded months and years after infection in patients who experienced symptomatic WNV infection, but not in those who experienced asymptomatic WNV infection, and they were the only T cell subset (among TN cells, TCM cells, effector-memory T cells (TEM cells) and TEMRA cells) that correlated with symptomatic WNV infection. Therefore, the presence of CD8+ TMNP cells could be useful in the immunotherapy of persistent infections or should be accounted for if truly naive T cells are needed to respond to antigens.
Results
Cytokine production by a subset of phenotypically naive T cells
One key age-related population change in the T cell pool is an absolute numerical decrease in blood CD8+ TN cells2. To investigate whether the peripheral blood CD8+ TN cells also showed qualitatively altered responses with aging, we obtained peripheral blood mononuclear cells (PBMCs) (which were the source of the cells used throughout this study, unless specified otherwise) from 92 subjects (43 males and 49 females 21–97 years of age), stimulated the cells for 3 h with phorbol-myristate acetate (PMA) and the calcium ionophore ionomycin and assessed intracellular interferon-γ (IFN-γ) and cytokines in the cells (Fig. 1). We performed multicolor flow cytometry to gate on the four main CD8+ T cell subsets (TN cells, TCM cells, TEM cells and TEMRA cells) defined by expression of CD45RA, the chemokine receptor CCR7, the death receptor CD95 and the co-receptor CD28. In this way, TN cells were classified as CD45RA+CCR7+CD95loCD28int, TCM cells were classified as CD45RA−CCR7+CD95hiCD28hi, TE+EM cells were classified as CD45RA−CCR7−CD95hiCD28lo, and TEMRA cells were classified as CD45RA+CCR7− CD95hiCD28lo. We used these definitions throughout this study (except where the full phenotype is provided) because they correlate well with the functional characteristics of various T cell subsets14 (Supplementary Fig. 1a,b). The total number of CD8+ TN cells diminished with age from over 250 cells per microliter of blood in subjects 20–30 years of age to less than 50 cells per microliter of blood in subjects 80 years of age (Fig. 1a and Supplementary Fig. 1c), which confirmed published observations2. However, following 3 h of stimulation of PBMCs with PMA plus ionomycin, 0.2–50% of CD8+ TN cells produced IFN-γ, compared with <0.1% of unstimulated CD8+ TN cells and >60% of stimulated TEM and TEMRA cells (Fig. 1a). This fraction increased with age, from 2.9% ± 1.7% (mean ± s.d.) of CD8+ TN cells in subjects 21–40 years of age to 8.7 ± 9.9% in subjects >65 years of age (Fig. 1b). The increase in IFN-γ+ CD8+ TN cells with aging was relative; the absolute number of these cells also diminished with age, albeit less rapidly than that of CD8+ TN cells (Supplementary Fig. 1c). A fraction (1–2%) of CD4+ TN cells stimulated with PMA plus ionomycin also produced IFN-γ (Supplementary Fig. 1d). After being stimulated with PMA plus ionomycin, freshly isolated PBMCs and sorted CD45RA+CCR7+CD95hiCD28lo CD8+ TN cells produced granzyme B (GzmB) (0.06–11.1%), IFN-γ (0.5–16.2%), IL-2 (0.4–3.8%) or tumor-necrosis factor (TNF) (1.8–22.7%); the proportion of cells that produced IFN-γ in the presence of brefeldin A (background control) was <0.37% (Fig. 1c and Supplementary Fig. 1d,e). This excluded the possibility that the IFN-γ+ CD8+ cells were TM cells that had acquired a naive phenotype during the freezing-thawing process or during the stimulation with PMA plus ionomycin. IFN-γ+ CD8+ TN cells did not exhibit homogenous expression of CD45RA, CCR7, CD28 or CD95 and were dispersed within the TN cell gate and showed phenotypic micro-heterogeneity (Fig. 1a), like other T cell subsets18. This suggested that the IFN-γ+ CD8+ TN cells were not a contaminating memory population that clustered adjacent to TM cells. Thus, a subset of phenotypically CD8+ TN cells was able produce multiple cytokines after polyclonal stimulation.
(a) Flow cytometry analyzing intracellular IFN-γ staining in PBMCs obtained from healthy adult donors 32 years of age (Adult) or 76 years of age (Old) and then activated PMA plus ionomycin. Numbers adjacent to outlined areas indicate percent IFN-γ+ (blue dots) CCR7+CD45RA+ TN cells (yellow outline) or IFN-γ+ (yellow dots) CD28intCD95lo TN cells (red outline) that were also CCR7+CD45RA+, overlaid over total CD8+ T cells (green background). (b) Frequency of IFN-γ-producing CD45+CCR7+CD95loCD28int CD8+ naive-phenotype T cells from donors (key) 21–40 years of age (<40; n = 27), 40–65 years of age (40–65; n = 44) or 66–97 years of age (>65; n = 21). (c) Production of IFN-γ, GzmB and TNF (key) by phenotypically naive CD45RA+CCR7+CD95loCD28int CD8+ T cells in freshly isolated blood (from n = 7 donors, 32–76 years of age). Each symbol (b,c) represents an individual donor; small horizontal lines (c) indicate the mean (± s.d.). NS, not significant (P > 0.05); *P < 0.05 and **P < 0.001 (t-test). Data are from one experiment (a,b) or one of two experiments (c).
TMNP cells are CD49d+CXCR3+
To determine whether the cytokine-producing CD8+ TN cells could be distinguished from other TN cells, we used SPADE software ('spanning tree progression of density-normalized events') for unsupervised clustering analysis of flow-cytometry data. The expression of five T cell–activation and T cell–differentiation markers (CD45RA, CCR7, CD95, CD28 and CD45RO) distinguished four canonical T cell subsets (TN cells, TCM cells, TE+EM cells and TEMRA) among the PBMCs stimulated with PMA plus ionomycin (Fig. 2a,b). The TN cell subset was further split into seven nodes, four of which were TN cells by the CD45RA+CCR7+CD95loCD28int phenotypic definition and one of which showed enrichment for IFN-γ-producing cells (Fig. 2a); this suggested that the IFN-γ+ CD8+ TN cells represented a distinct T cell subset.
(a) Grouping of IFN-γ+ cells from donors (n = 2) 27 years of age (Adult) or 68 years of age (Old) into four canonical subsets (black outlines) of naive CD8+ T cells, including CD45RA+CCR7+CD95loCD28int cells (red outline) and IFN-γ+ naive cells (bright green): node size indicates subset abundance relative to all CD8+ cells in the plot (total area of all nodes is the same across all samples); color indicates intensity of IFN-γ staining, presented as arbitrary relative median fluorescence intensity (key); outer circles around nodes emphasize the TMNP population. (b) Grouping of IFN-γ+ cells from two donors as in a (among nine donors (6 male; 3 female) 27–78 years of age) into subsets as in a, according to staining of the phenotypic markers CCR7, CD45RA, CD28, CD95 and CD45RO (left margin). Data are from one experiment.
When we assigned scores for the expression of a broader set of markers to stringently define CD8+ TN cell as CD27+CD45RO−CD127+CD122loCD31+CD11a−HLA-DR−, we found that the IFN-γ+ CD8+ TN cells did not diverge from IFN-γ− CD8+ TN cells, regardless of whether we initially gated through a less-stringent TN cell gate (CD45RA+CCR7+) or a more-stringent TN cell gate (CD45RA+CCR7+CD95loCD28int) (Fig. 3 and Supplementary Fig. 2a). However, analysis of mean fluorescence intensity revealed that IFN-γ+ CD8+ TN cells had slightly lower expression of CCR7 and higher expression of CD28 than that of IFN-γ− CD8+ TN cells; the expression of CD45RA and CD95 by these cells was similar (Supplementary Fig. 2b). The phenotypic profile of IFN-γ+ CD8+ TN cells was markedly distinct from that of TCM or TE+EM cells (Fig. 3b,c and Supplementary Fig. 2). The effector-memory function of IFN-γ+ CD8+ TN cells, along with their naive phenotype, led us to call these cells 'memory T cells with a naive phenotype' (TMNP cells).
(a) Flow cytometry of PBMCs obtained from one donor (54 years of age) and then activated with PMA plus ionomycin, showing gating strategy used for the identification of TMNP CD8+ T cells. Outlined areas indicate FSCint cells (top left plot); SSC−FSC+ cells among the cells at left (top middle plot); live CD8β+ cells among the cells at left (top right plot); canonical populations of CCR7+CD45RA− cells (TCM) cells (TE+EM) cells (bottom left), CCR7+CD45RA+ cells (TEMRA) cells (bottom right) or CCR7+CD45RA+ cells (TN) cells (top right; blue) among the live CD8β+ cells (bottom left plot); CD28intCD95lo cells among those CCR7+CD45RA+ cells (bottom middle plot); or IFN-γ+CD8β+ cells among those CCR7+CD45RA+CD28intCD95lo cells (bottom right). (b,c) Expression (as geometric mean fluorescence intensity (gMFI)) of the surface antigens CD27, CD45RO, CD31, CD11a, CD127, CD122 and HLA-DR (b) or CD49d, CXCR3 and CD103 (c) in various CD8+ T cell subsets (key) from the donor in a. P < 0.0001, TMNP versus TN (one-way analysis of variance (ANOVA)). (d) Expression of CD49d and CXCR3 on CD8+ TN and TMNP cells (from donors 27–85 years of age). Each symbol represents an individual donor; small horizontal lines indicate the mean (± s.e.m.). *P < 0.01 and **P < 0.001 (repeated-measures one-way ANOVA). Data are representative of two experiments (with n = 13 donors and n = 12 donors).
We next analyzed trafficking and adhesion receptors on TMNP cells. The α4 integrin CD49d associates with β-integrin subunits to form α4β7 or α4β1 heterodimers that regulate the trafficking of TEM cells19. In mice, CD49d marks exclusively antigen-experienced TM cells20. IFN-γ+CD45RA+CCR7+CD95loCD28int CD8+ TMNP cells consistently had the highest expression of CD49d relative to that of TN cells, TCM cells, TEM cells and TEMRA cells; their CD49d geometric mean fluorescence intensity was on average 2.5-fold higher than that of IFN-γ− CD8+ TN cells (Fig. 3c,d). However, while all IFN-γ+ CD8+ TMNP cells were CD49dhi, <10% of the CD49dhi CD8+ TN cells did not produce IFN-γ (data not shown), which suggested that while all CD8+ TMNP cells were in the CD49d+ TN pool, CD49d expression did not exclusively identify CD8+ TMNP cells. IFN-γ+CD45RA+CCR7+CD95loCD28int CD8+ TMNP cells also had the highest expression of the chemokine receptor CXCR3 relative to that of TN cells (defined as IFN-γ−CD45RA+CCR7+CD95loCD28int), TCM cells, TEM cells or TEMRA cells (Fig. 3c,d); this suggested an ability to respond rapidly to the chemokines CXCL9, CXCL10 and CXCL11, which would direct the cells to inflamed sites21. Because CD49d and CD103 (integrin αE) can both bind the integrin β7, and because the CD103-β7 (αEβ7) heterodimer directs the trafficking of immune cells to residence in tissues, including gut mucosa22,23, we assessed expression of CD103 and found it was similarly low on CD8+ TMNP cells and CD8+ TN cells (Fig. 3c); this suggested that these cells would be unlikely to home to gut mucosa. Therefore, while the CD8+ TMNP cells had most of the phenotypic attributes of CD8+ TN cells, they exhibited rapid functional responses and had the potential to traffic to inflamed sites.
Constitutive phosphorylation of the kinase Erk in TMNP cells
To determine whether the CD8+ TMNP cells responded to TCR-mediated activation, we stimulated PBMCs for 3 h via TCR crosslinking with plate-bound antibody to the TCR invariant chain CD3 (anti-CD3) and soluble antibody to the co-receptor CD28 (anti-CD28) with or without antibody to CD49d (anti-CD49d). TCR crosslinking induced the production of IFN-γ, TNF and GzmB by CD49dhiCD45RA+CCR7+CD95loCD28int CD8+ TMNP cells, although the frequency of IFN-γ+ CD8+ TMNP cells induced this way was 20–30% that of IFN-γ+ CD8+ TMNP cells induced by PMA plus ionomycin (Fig. 4a). Anti-CD49d did not affect the magnitude (Fig. 4a), breadth or kinetics (data not shown) of this cytokine production. Therefore, engagement of CD49d did not co-stimulate production of IFN-γ,TNF or GzmB in the CD8+ TMNP cells.
(a) Flow cytometry of CD8+ TMNP cells obtained from donors (n = 10, 26–68 years of age) and left unactivated (UA) or stimulated for 3 h with anti-CD3 and andi-CD28 (α-CD3 + α-CD28) alone or with anti-CD49d (+ α-CD49d), or with PMA and ionomycin (PMA + iono) (left). Numbers adjacent to outlined areas indicate percent IFN-γ+CD8β+ cells. Hs (left margin), human subject (identification number). Right, frequency of IFN-γ+ CD8+ naive T cells as at left, presented relative to results obtained with PMA and ionomycin, set as 100%. (b) Frequency of cells expressing various combinations of TNF, GzmB and IFN-γ (key) among various subsets (horizontal axis) of CD8+ T cells (n = 36 donors, 24–82 years of age). (c) Flow cytometry (left) of IFN-γ+CD45RA+CCR7+CD95loCD28int CD8+ TMNP cells (black) overlaid over that of total CD8+ T cells (gray) (n = 2 male donors, 52 or 63 years of age). Numbers in outlined areas indicate percent GzmB+CD107a+ cells (top row) or perforin (Perf)-producing cells (bottom row) that are phenotypically naive (TMNP cells). Right, cumulative frequency of CD107a+ or Perf+ CD8+ TMNP cells among IFN-γ+GzmB+ TMNP cells (n = 8 donors, 31–77 years of age). (d) Proliferation of sorted CD8+ T cell subsets after 7 d of treatment with various combinations (above plots) of IL-7 (50 ng/ml) and IL-15 (50 ng/ml), assessed as dilution of CellTrace Violet (CTV) (n = 2 donors, 52 or 64 years of age, among the eight donors in c; four of the eight samples included memory subsets as controls). (e) Flow cytometry assessing phenotype changes after 3 d of culture of CD49dhi or CD49dlo cells as in d (n = 2 male donors (of n = 3), 52 or 64 years of age). Numbers adjacent to outlined areas indicate percent CCR7+CD45RA+ cells. (f) T-bet expression (gMFI) in various subsets (horizontal axis) of CD8+ T cells (n = 9 donors (6 male; 3 female), 24–78 years of age) at before (UA) or after (Act) 2 h of activation with PMA plus ionomycin. (g) Time course of the phosphorylation of Erk (p-Erk) in TMNP cells (CD49dhi) and TN cells (CD49dlo) (key; as described below) (n = 3 donors (1 male; 2 female), of n = 7 donors, 67–83 years of age repeated with n = 4 donors, of n = 7 donors, 35–83 years of age) at baseline (0 min) and at 5, 10 or 15 min after stimulation with PMA plus ionomycin (left), or in TMNP (CCR7+CD45RA+CD95loCD28intCD49dhi) CD8+ T cells (CD49dhi) and TN (CCR7+CD45RA+CD95loCD28intCD49dlo) CD8+ T cells (CD49dlo) (key) (n = 1 female donor, 68 years of age) after 2 h of activation with PMA plus ionomycin (right). Numbers in plots (left) indicate gMFI (GM) of phosphorylated Erk. (h) Frequency of IFN-γ+ cells among CD8+ T cells (n = 14 donors (9 male; 5 female), 52–83 years of age) pre-incubated with various concentrations (horizontal axis) of Erk inhibitor and then stimulated with PMA plus ionomycin. Each symbol (c,f) represents an individual donor; small horizontal lines indicate the mean (± s.e.m.). *P < 0.05,**P < 0.01 and ***P < 0.0001 (repeated-measures one-way ANOVA). Data are representative of two experiments (a,c,g,h; mean + s.d. in a,g,h), one experiment (b,f) or three experiments (d,e).
The polyfunctionality of CD8+ TMNP cells, measured as the simultaneous production of IFN-γ, GzmB and TNF in response to stimulation, was similar to that of CD8+ TEMRA cells, as the TMNP cells produced all three cytokines (50–60%), TNF only (20–25%) or TNF plus IFN-γ (15–25%) at a frequency similar to that of CD8+ TEMRA cells, in contrast to results obtained for TCM cells (∼50% produced only TNF) and TEM cells (>40% produced TNF plus IFN-γ) (Fig. 4b). Therefore, the CD8+ TMNP cells were highly polyfunctional. Moreover, about 40% of the IFN-γ+GzmB+ CD8+ TMNP cells also expressed perforin (Fig. 4c), and all of the IFN-γ+GzmB+ CD8+ TMNP cells were CD107a+ and thus were functionally lytic (Fig. 4c). Therefore, CD8+ TMNP cells were in a differentiated, polyfunctional state that contrasted with that of the quiescent, effector-molecule-negative CD8+ TN cells.
We next used flow cytometry to sort CD49dhiCD45RA+ CCR7+CD95loCD28int CD8+ TMNP cells and compared their proliferation with that of CD49dloCD45RA+CCR7+CD95loCD28int CD8+ TN cells, TCM cells and TEM cells in response to the homeostatic cytokines IL-7 and IL-15. CD8+ TMNP cells divided more in the presence of IL-7 alone or IL-7 plus IL-15 than did CD8+ TN cells (Fig. 4d). In contrast to CD8+ TN cells and similar to the TCM and TEM subsets, CD8+ TMNP cells also proliferated in response to IL-15 alone (Fig. 4d), which is a cardinal characteristic of TM cells. When incubated with IL-15 for >3 d, CD49dhi CD8+ TMNP cells lost their expression of CCR7 and CD45RA faster than CD8+ TN cells did (Fig. 4e). Despite those results, expression of the chemokine receptor subunit CD122 (IL-15Rβ) in CD8+ TMNP cells (TN cells sorted as CD49hi) was similar to that in CD8+ TN cells (TN cells sorted as CD49lo) (both CD45RA+CCR7+CD95loCD28int) (Fig. 3b and Supplementary Fig. 2.). Finally, CD49hiCD45RA+CCR7+CD95loCD28int CD8+ TMNP cells did not secrete IFN-γ in response to overnight stimulation with IL-12 plus IL-18 (Supplementary Fig. 3a,b), which suggested that they were unable to undergo antigen-nonspecific bystander responses typical of other TM cells24.
The results presented above suggested that TMNP cells could be partially activated via TCR. We therefore assessed intracellular expression of T-bet (the master transcription factor of the TH1 subset of helper T cells25,26) and phosphorylation of the kinases Akt and Erk (which integrate multiple signaling cascades in T cells) in CD49dhiCD45RA+ CCR7+CD95loCD28int CD8+ TMNP and CD49dloCD45RA+CCR7+CD95loCD28int CD8+ TN cells (identified by flow cytometry) and compared it with that of CD8+ TCM, TEM or TEMRA cells. Expression of T-bet in CD8+ TMNP cells was significantly higher than that of CD8+ TN cells both at steady state and following brief activation with PMA plus ionomycin but was not significantly different from that of TCM cells (Fig. 4f). We found no difference between TMNP cells and TN cells in their phosphorylation of Akt (data not shown). CD8+ TMNP cells exhibited more phosphorylation of Erk at baseline than that of TN cells, and the difference increased significantly following stimulation with PMA plus ionomycin (Fig. 4g). Pre-incubation of CD8+ TMNP cells with the Erk inhibitor U0126 diminished their IFN-γ responses to stimulation with PMA plus ionomycin in a dose-dependent manner (Fig. 4h), which indicated that the activated Erk pathway in CD8+ TMNP cells mediated rapid cytokine production. No toxicity was observed at any dose of the inhibitor (data not shown). Together these results demonstrated that the CD8+ TMNP cells were polyfunctional, exhibited cytolytic potential, were able respond to IL-15, and constitutively activated the Erk pathway, which enabled their ability of rapid IFN-γ responsiveness.
Long telomeres and unique transcriptome of TMNP cells
To explore whether CD8+ TMNP cells were partially activated CD8+ TN cells or differentiated CD8+ TM cells that re-expressed markers of naive cells, we evaluated their proliferative history and transcriptional profile. Telomeres are repeating hexameric sequences of nucleotides at chromosomal ends that provide genomic stability and that shorten with each replication. Telomeres are longer in TN cells and shorter in TM cell subsets27 and can be used to estimate cell-division history. We used multicolor fluorescence in situ hybridization to compare telomere length in various CD8+ T cell subsets (defined by expression of CD45RA, CCR7, CD95, CD28 and CD49d as in Supplementary Fig. 4a). Telomere lengths in CD8+ TMNP cells were similar to those in CD8+ TN cells (Fig. 5a), which suggested that CD8+ TMNP cells, unlike the other memory subsets, including CD8+ TCM, TEM or TEMRA cells, did not undergo extensive division. Therefore, telomere length in CD8+ TMNP cells most closely resembled that in TN cells.
(a) Flow cytometry (left) showing telomere length of the TN (CCR7+CD45RA+CD95loCD28int CD49dlo), TMNP (CCR7+CD45RA+CD95loCD28int CD49dhi), TCM,TE+EM and TEMRA subsets of CD8+ T cells, presented as staining of telomere probes with indodicarbocyanine and peptide nucleic acid, via fluorescence label intensity based on the primer Tel-C. Numbers in plots indicate gMFI of staining. Right group analysis of, telomere length (as Tel-C gMFI) of the CD8+ T cell subsets at left presented relative to that of the TN cell subset, set as 1 (n = 9 (3 male; 6 female), 34–83 years of age). Each symbol represents an individual donor; small horizontal lines indicate the mean (± s.e.m.). (b) Gene expression in cells activated with PMA and ionomycin and then sorted as in a but with IFN-γ (not CD49d) to distinguish TMNP cells (IFN-γ+) from TN cells (IFN-γ+) (key), presented by projection of the expression values onto PC1-versus-PC2 plots with variables of all genes (adjusted P value, <0.01; false-discovery rate, <0.05) listed in the Consensus Coding Sequence project that were measured (n = 3–5 donors (distinguished by symbol shape in plot), 40–76 years of age). (c) Expression of genes (right margin) that contributed substantially to PC1 and PC2 and had large absolute value 'loadings' (0.045 (PC1) and 0.0375 (PC2)), in cell subsets as in a (below plot), standardized to a z-score and then binned to a color (adjusted P value, <0.01; false-discovery rate, <0.05). Brackets (top and left margin) indicate hierarchical agglomerative clustering; colors at top match those in a. Data are representative of two experiments (a) or one experiment (b,c).
We next used RNA sequencing to compared the transcriptome of CD8+ TMNP cells (defined as IFN-γ+CD45RA+CCR7+CD95loCD28int) with that of other CD8+ T cell subsets (IFN-γ−CD45RA+CCR7+CD95loCD28int TN cells; other subsets as defined above). Differential gene expression among the TN, TCM, TEM and TEMRA CD8+ T cell subsets was distinguished by variation in principal component 1 (PC1) (Fig. 5b). The variation captured by PC2 separated the TMNP cell subset, and a combination of variation in PC1 and PC2 indicated that TMNP cells were a unique T cell subset, with the highest expression (among all subsets) of Plaur, Ilirn, Anpep, Ccl3, Mmp19, Serpb2, Cxcl3, Il1b and Sppi, and with similarities to and differences from both CD8+ TN cells and CD8+ TCM cells on the one hand and the more differentiated CD8+ TEM cells and CD8+ TEMRA cells on the other hand (Fig. 5b). A heat map of expression values for genes with the greatest contributions to PC1 and PC2 indicated that TMNP cells clustered nearest to TN cells, followed by intermediate proximity to TCM cells and greater distance from the TEM and TEMRA subsets (Fig. 5c). The genes that placed the CD8+ TMNP cells outside the canonical CD8+ TN cell gene cluster included genes encoding effector molecules (for example, Ifng, Gzmb, Il1b and Cxcl3), and their expression in CD8+ TMNP cells was similar to or higher than that observed in CD8+ TCM cells (Fig. 5c). Conversely, genes such as Nkg7, Fasl and Il22 were not expressed in CD8+ TMNP cells, and this feature distinguished them from CD8+ TEM cells and TEMRA cells (Fig. 5c). Together the transcriptome data suggested that the CD8+ TMNP cells were a unique T cell subset.
CD8+ TMNP cells are highly proliferative
We next explored proliferative potential of CD8+ TMNP cells. We sorted IFN-γ+ CD45RA+CCR7+CD95loCD28int CD8+ TMNP cells and their IFN-γ− TN cell counterparts (from n = 3 subjects, 63, 65 and 68 years of age; Supplementary Fig. 5a) or stimulated cells that we then separated by flow cytometry into CD49dhi CD45RA+CCR7+CD95loCD28int CD8+ TMNP cells their CD49lo TN cell counterparts, as well as TCM,, TEM and TEMRA cells (n = 5 subjects, 30–80 years of age; Supplementary Fig. 5b). Cells were stimulated for 4 h with anti-CD3 plus anti-CD28 (Supplementary Fig. 5a) to induce IFN-γ and enable sorting. After being sorted, cells were stimulated for additional 3 d (Supplementary Fig. 5a), or cells without sorting were stimulated for 6 d (Supplementary Fig. 5b), with beads coated with anti-CD3 plus anti-CD2 plus anti-CD28, in IL-2 (10 U). Division rates, measured by dilution of the division-tracking dye CellTrace Violet, revealed similar proliferation of CD8+ TMNP cells, CD8+ TN cells and TCM cells (Supplementary Fig. 5a). We concluded that consistent with their long telomeres, TMNP cells had robust proliferative potential.
TMNP cells respond to persistent viruses
We next assessed the TCR diversity of CD8+ TMNP by flow cytometry using antibodies specific for the 24 different TCR β-chain variable (Vβ) regions, covering 70% of the total Vβ repertoire. CD49dhiCD45RA+CCR7+CD95loCD28int CD8+ TMNP cells showed a more restricted distribution of TCR Vβ segments than that of CD49dloCD45RA+CCR7+CD95loCD28int CD8+ TN cells, similar to the highly differentiated CD8+ TEMRA cells (Fig. 6a). This suggested that the CD8+ TMNP cells might have arisen in response to antigens that drove the generation of the CD8+ TEM cells and CD8+ TEMRA cells, rather than being due to broad, cytokine-driven memory conversion. Longitudinal comparison of the Vβ profile of CD8+ TMNP cells in one subject (50 years of age, among n = 2 subjects, 50 and 73 years of age) over 24 months showed stable dominant clonal expansion of β-chain variable region 14 (Vβ14) (Supplementary Fig. 6), which suggested that the CD8+ TMNP cells were stably maintained by TCR-driven signals.
(a) Distribution of TCR Vβ in each of the 24 TCR Vβ regions assessed (horizontal axis) in the TN (CD45RA+CCR7+CD95loCD28intCD49dlo), TMNP (CD45RA+CCR7+CD95loCD28int CD49dhi), TCM, TE+EM and TEMRA subsets of CD8+ T cells (of n = 3 donors (top, middle bottom) among n = 11 donors (6 male; 5 female), 27–78 years of age). (b) Flow cytometry of PBMCs at 3 h after stimulation with a pool of peptides of the EBV protein BLZF1, CMV protein pp65 or influenza A virus (IAV) matrix protein (b) or such cells left unstimulated (US) or stimulated for 3 h with a pool of peptides of HIV Gag protein (+ Gag) (c), showing gating strategy used for evaluation of the naive phenotype of the responding (IFN-γ producing) CD8+ T cells (n = 1 healthy donor (66 years of age) per condition among n = 8 donors, 24–80 years of age (b) or n = 3 donors (male; 35, 43 and 58 years of age), HIV+ undergoing anti-retroviral therapy with no detectable HIV in plasma (c)). Numbers adjacent to outlined areas indicate percent IFN-γ+CD8β+ cells (far left; red), CCR7+CD45RA+ cells among those IFN-γ+CD8β+ cells (middle; blue), or CD28intCD95lo cells among those IFN-γ+CD8β+CCR7+CD45RA+ cells (right; green), overlaid onto total CD8+ T cells (gray). Data are representative of two experiments.
CD8+ TEMRA cells are often specific for persistent viruses28. To assess whether the CD8+ TMNP cells shared that specificity, we stimulated PBMCs for 3 h with overlapping 15-amino-acid peptides covering the length of viral proteins. In one donor, TMNP cells made up 8.5% of the total response to protein BZLF1 of Epstein-Barr virus (EBV) and 1.4% of the response to protein pp65 of cytomegalovirus (CMV) but none of the response to matrix protein of influenza A virus (Fig. 6b). A similar fraction of CD8+ TMNP cells responded to a pool of peptides from Gag protein of human immunodeficiency virus type 1 (HIV-1) in two of three HIV-infected subjects undergoing highly active anti-retroviral therapy (Fig. 6c). In these analyses, control cells were treated with brefeldin A (negative control) or with PMA plus ionomycin (positive control).
We extended the observations reported above by flow-cytometry staining of unstimulated PBMCs and identification of antigen-specific cells with multimers of peptide and major histocompatibility complex (MHC) class I. The majority (>90%) of CD8+ T cells specific for HLA-A*0201 (MHC class I) bearing the immunodominant epitope of CMV pp65 (peptide sequence, NLVPMATM) or EBV BMLF1 (peptide sequence, GLCTLVAML) in CMV-seropositive subjects or EBV-seropositive subjects, respectively, exhibited the TEM, TCM or TTEMRA phenotype (Fig. 7a,b). However, phenotypically naive CD45RA+CCR7+CD95loCD28int CD8+ T cells also bound multimers of CMV or EBV peptide and MHC class I (Fig. 7a,b), and a substantial fraction of these (25% for CMV, and 12.4% for EBV) also produced IFN-γ following 3 h of stimulation with PMA plus ionomycin (Fig. 7a,b). Stimulation with PMA plus ionomycin did not induce downregulation of TCR expression and did not affect peptide-MHC binding (Supplementary Fig. 4b). In contrast, we found no IFN-γ+ CD8+ cells among the identically gated IFN-γ− CD8+ TN cells that bound multimers of peptide and MHC class I in CMV-seronegative or EBV-seronegative subjects (Fig. 7a,b). TMNP cells were present at a similar frequency (75% or 67% of CD45RA+CCR7+CD95loCD28int CD8+cells that bound peptide-MHC multimers) in two subjects that had a tenfold difference in their overall T cell responses to EBV (Fig. 7b). Finally, vaccination with seasonal vaccine against influenza virus did not result in nonspecific or bystander activation of TMNP cells (Supplementary Fig. 7). These results suggested that the CD8+ TMNP cells arose as a part of a response to persistent viral antigens.
(a,b) Flow cytometry of naive (CD45RA+CCR7+CD95loCD28int) CMV-specific CD8+ T cells from one CMV-seropositive (CMV+) donor (male, 72 years of age) and one CMV-seronegative (CMV−) donor (female, 65 years of age) (among n = 26 donors, 28–83 years of age) (a) or EBV-specific CD8+ T cells from two EBV-seropositive donors (high titer (EBV++) and low titer (EBV+); male, 67 or 75 years of age) (among n = 7 donors, 24–75 years of age) (b), assessed after 3 h of stimulation with PMA plus ionomycin, showing gating strategy and staining with a tetramer of HLA-A*0201 and CMV pp65 peptide (A2-CMV(NLV) tet) (a) or a dextramer of HLA-A*0201 and EBV BMLF1 peptide (A2-EBV(GLC) dex). Numbers adjacent to outlined areas indicate percent tetramer-positive CD8β+ cells (far left), CCR7+CD45RA+ cells among those cells (middle left), CD28intCD95lo cells among those cells (middle right), and IFN-γ+CD49+ cells (far right). (c,d) Flow cytometry of naive (CCR7+CD45RA+CD95loCD28int) vaccinia-virus-specific CD8+ T cells from one CMV-seropositive donor (female, 64 years of age) and one CMV-seronegative donor (female, age 36 years of age) (of n = 4 donors, 36–74 years of age) (c) and naive Melan-A-specific CD8+ T cells (n = 2 donors (female), 58 or 76 years of age (of n = 6 donors, 29–82 years of age)), assessed after 3 h of stimulation with PMA plus ionomycin, and stained with tetramer as in a (c) or with a tetramer of HLA-A*0201 and Melan-A peptide (A2-MelanA(ELA) tet) conjugated to Brilliant Violet 421 (BV421) or allophycocyanin (APC) (d); gating strategy and numbers adjacent to outlined areas as in b. Data are representative of one (a–c) or two (d) experiments.
To determine whether the CD8+ TMNP cells also responded to viral pathogens during acute infection, we analyzed PBMCs from a CMV-seropositive donor vaccinated with vaccinia virus (VACV). We stained the cells by flow cytometry with the CMV peptide–MHC tetramer pp65–HLA-A*0201 and the VACV tetramer CLT–HLA-A*0201. CD8+ TMNP cells, as assessed by production of IFN-γ following stimulation with PMA plus ionomycin, were present among the population positive for CMV peptide–MHC but not among the population positive for VACV peptide–MHC (Fig. 7c). We also failed to detect IFN-γ+ CD8+ TMNP cells specific for the melanoma self antigen Melan-A (Mart-1)29 in a healthy, melanoma-negative subjects in whom all cells positive for Melan-A (Mart-1) were CD49d−IFN-γ− (Fig. 7d). Therefore, the CD8+ TMNP cells were driven by persistent viral infections and did not respond to vaccination or to self antigens.
CD8+ TMNP cells are linked to past severe infections
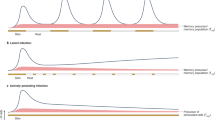
Because the size and diversity of naive T cells pool determines the ability to generate protection against new infections30,31, we investigated whether the frequency and number of CD8+ TMNP cells correlated with immunological fitness in response to acute infection. We assessed the frequency of CD8+ TMNP cells among TN cells longitudinally (from day 2 to day 90 after diagnosis) in a cohort of WNV-infected subjects who either lacked or presented with clinical signs such as fever, meningitis and/or encephalitis. Three asymptomatic subjects had 0.3–1.6% IFN-γ+CD49dhi TMNP cells and 0.3–5.9% GzB+CD49dhi TMNP cells among the CD45RA+CCR7+CD95loCD28int TN cells, whereas three symptomatic subjects had 14.2–20.8% and 12.6–19.8% of these cells, respectively (Fig. 8a,b). Furthermore, the average absolute number of CD45RA+CCR7+CD95loCD28intIFN-γ+CD49dhi CD8+ TMNP cells was <2,000 of 1 × 106 CD8+ T cells or >4,000 of 1 × 106 CD8+ T cells in asymptomatic subjects or symptomatic subjects, respectively (Fig. 8c), with no difference in the number of CD8+ TN, TCM, TEM or TEMRA cells (Supplementary Fig. 8). WNV infection resulted in the population expansion of CD8+ TMNP cells in symptomatic subjects but not in asymptomatic subjects, as shown by an increase in the abundance of CD8+ TMNP cells per 1 × 106 CD8+ T cells between days 2–7 and days 60–90 after infection in symptomatic subjects (Fig. 8d). This showed that the frequency and number of TMNP cells positively correlated with the severity of acute viral infection.
(a,b) Flow cytometry analyzing intracellular IFN-γ and GzmB in PBMCs obtained from asymptomatic WNV+ donors (n = 3, of n = 11 donors, 39–92 years of age) (a) or symptomatic WNV+ donors (n = 3, of n = 16 donors, 43–84 years of age) (b) and activated with PMA plus ionomycin, showing gating strategy for TMNP (IFN-γ+CD45RA+CCR7+CD95loCD28int) CD8+ T cells. Numbers adjacent to outlined areas indicate percent CCR7+CD45RA+ cells (far left), CD28intCD95lo cells among those cells (middle left), and IFN-γ+CD49+ cells (middle right) or GzmB+CD49+ cells (far right) among those CCR7+CD45RA+CD28intCD95lo cells. (c) Quantification of TMNP cells in donors as in a,b (n = 26). (d) Quantification of TMNP cells (as in c) in asymptomatic donors (n = 11, 39–92 years of age) or symptomatic donors (n = 16, 43–84 years of age) with WNV infection, assessed early (days 2–7) or late (days 60–90) after 'index donation'42. Each symbol (c,d) represents an individual donor; small horizontal lines indicate the mean (± s.e.m.). *P < 0.05 and **P < 0.01 (Mann-Whitney test (c) or Levene's test of equal variance assumption followed by repeated measures ANOVA, with heterogeneous compound symmetry covariance structure used to not only account for within-subject correlation but also adjust for variance heterogeneity (d)). Data are representative of QQ experiments (a) or QQ experiments (c) or are from one experiment (b,d).
Discussion
Here we characterized a previously unknown subset of T cells that seemed to be functionally differentiated and activated but that underwent only minimal phenotypic changes compared with those of CD8+ TN cells and continued to express molecules associated with a naive T cell state. The frequency of these CD8+ TMNP cells increased with age. They were polyfunctional and exhibited greater baseline phosphorylation of Erk, which suggested that they were receiving TCR-mediated signals. Their skewed utilization of TCR Vβ and their TCR specificity for persistent viral infection or vaccination but not acute viral infection or vaccination confirmed that these CD8+ TMNP cells represented a T cell–activation stage that allowed persistence of antigen-primed, effector-ready T cells in a nearly naive state.
We speculate that TMNP cells are functionally 'imprinted' at an early stage of differentiation and propose that they represent a third line of defense against persistent viral infections, behind TEM cells and TCM cells. Their in vitro differentiation with IL-7 plus IL-15 suggested that the CD8+ TMNP cells maintained their plasticity and were able to give rise to cells that resembled CD8+ TCM, TEM and TEMRA cells. This would be consistent with the observation that CD8+ TMNP cells did not respond to the inflammatory cytokines IL-12 and IL-18 by producing IFN-γ, as would be expected of highly differentiated TEM cells and TEMRA cells. Their telomere lengths were similar to those of CD8+ TN cells, which suggested they had not undergone the extensive proliferation associated with full effector and memory differentiation. Consistent with all of the results reported above, unsupervised and unbiased analysis of the CD8+ TMNP cell transcriptome classified these as a unique subset, with the highest expression (among all subsets) of Plaur, Ilirn, Anpep, Ccl3, Mmp19, Serpb2, Cxcl3, Il1b and Sppi, but otherwise with the greatest similarity to TN cells. This clustered TMNP cells between CD8+ TN cells and CD8+ TCM cells; TMNP cells were transcriptionally further apart from terminally differentiated CD8+ TEM cells and CD8+ TEMRA cells.
TMNP cells were the only CD8+ T cell subset whose number and frequency increased in the course of, and correlated with, symptomatic, severe WNV disease. This could have been because acute viral infection led to persistent viral reactivation or for other reasons that remain to be investigated. Despite this, we do not believe these cells are pathogenic or that they contribute to immunological dysfunction in a direct, antigen-specific manner. This is because CD8+ TMNP cells expressed CXCR3 and CD49d, which allowed them to traffic rapidly to inflamed tissues and engage in viral control using their polyfunctional secretion of TH1 cytokines and cytotoxic granules. However, because their numbers also correlated with deterioration of the naive T cell pool, they could serve as a biomarker for immunological vulnerability. Specifically, their presence in the CD8+ TN pool could be marker of low TN cell fitness due the fact that TMNP cells mask an even more pronounced decrease in the pool size and diversity of CD8+ TN cells, which often manifests as an impaired ability to generate optimal immunity to new infections or immunization32.
Increasing the knowledge about qualitative and quantitative changes in the aging human immune system could help improve immunological interventions for older adults by stratifying at-risk populations and customizing intervention strategies. Indeed, understanding of the complexity of T cell subset phenotype and function has greatly increased since the original classification of these cells into naive, effector and memory subsets15,33. Technical advances have allowed the identification of previously unknown T cell subpopulations28,34,35. While numerous (>200) transient populations were observed in a study that combined phenotypic, functional and antigen-specific tetramer-based markers18, some of them form functionally defined, stable subsets, such as resident memory T cells36, stem T cell memory37, recent thymic immigrants38, early memory CD4+ T cells39 and self-renewing memory CD8+ T cells resistant to chemotherapy40. CD8+ TMNP cells add to the complexity of the primary T cell response during aging and to the understanding of the generation of an immune response to persistent viruses. Additional work is needed to address whether the developmental fate of TMNP cells intersects with that of any of the subsets above. In this context, it will be of interest to directly compare CD8+ TMNP cells with CD8+ stem-memory T cells37, even if indirect comparisons (as shown here and in ref. 37) suggest important differences in the expression of CD95, the integrin CD11a (LFA-1), CD122 and the adhesion molecule CD31 (PECAM-1), proliferation history and specificity, whereby stem-memory T cells, unlike TMNP cells, respond to acute infectios37.
Overall, CD8+ TMNP cells should be considered in the development of immunological interventions that rely on the naive T cell pool41. These cells could be useful for immunotherapy aimed at targeting persistent infections and should be accounted for if truly naive T cells are needed to respond to antigens.
Methods
Study subjects and blood samples.
This study was approved by the Institutional Review Boards at the University of Arizona (Tucson, AZ; # 080000673), the Oregon Health and Science University (Portland, OR; # IRB00003007), the University of Texas HSC at Houston (Houston, TX), the Vaccine and Gene Therapy Institute-Florida (Port St. Lucie, FL) and the Blood Systems Research Institute (San Francisco, CA). Human samples were obtained from healthy donors, 21–101 years of age, as indicated in each experiment, recruited at the OHSU, the University of Arizona or the Martin Health system (Florida) over the period of 6 years. Exclusion criteria included known immunosuppressive pathology, stroke, cancer, or use of steroids within the last 5 years. WNV-infected donors were enrolled by Blood Systems Research Institute (BSRI) between 2009 and 2011 or by the University of Texas at Houston, between 2006 and 2009. Samples, demographics and symptoms data were collected and analyzed as previously described42 after the subjects provided informed consent approved by the UCSF Committee on Human Research (protocol # 10-01255) or the Committee for the Protection of Human Subjects at the University of Texas Health Science Center at Houston (HSC-SPH-03-039), respectively. Exclusion criteria included known immunosuppressive status, stroke, cancer, or the use of steroids within the last 5 years. Blood was drawn into heparinized Vacutainer CPT tubes (BD Bioscience, Franklin Lakes, NJ) and processed fresh at respective sites per manufacturer's recommendations to isolate peripheral blood mononuclear cells (PBMCs) and plasma; K2-EDTA tubes were used to determine complete blood counts. Peripheral blood mononuclear cells were also isolated from leukapheresis using Ficoll-Paque (GE Healthcare, NJ, USA) density gradient media. PBMCs were frozen in 90% FBS (FBS) and 10% DMSO. Initial observational experiment (Fig. 1a,b) was performed on subjects selected randomly to meet age criteria and ensured adequate power, as described2. Unless stated otherwise, subsequent experiments were performed using subgroups of subjects defined by age and >5% naive T cell responsiveness (measured in Fig. 1a,b) and selected by A.M.W. randomly based on these criteria, without input of V.P., who did the experiments. In these studies, we initially did not observe, and subsequently did not follow, sex differences in TMNP cell abundance. Experiments, data collection and Flow Jo analysis were performed blindly up to the point of final analysis as samples associated specific subject ID did not contain information about their age.
Antibodies, flow cytometry and cell sorting.
All antibodies, dilutions and validation are provided in Supplementary Table 1. TCR-Vβ specificity was evaluated by IOTest Beta Mark Kit (Beckman Coulter). HLA-A2 CMV-NLV, HLA-A2 VV-CLT, and HLA-A2 Melan-A-ELA tetramers were obtained from the NIH tetramer core facility at Emory University, HLA-A2 INF-GIL tetramers were provided by K. Kedzierska's laboratory at the University of Melbourne, Melbourne, Australia and HLA-A2 EBV-GLC dextramers were obtained from IMMUDEX (Copenhagen, Denmark).
Frozen PBMCs were thawed in RPMI medium supplemented with 10% FBS, penicillin and streptomycin in the presence of DNAse (Sigma, Saint Louis, MO), rested overnight in X-Vivo medium (Lonza/Basel, Switzerland) supplemented with 5% human male AB serum and used for surface and intracellular staining at 1 × 106 to 3 × 106 per sample. Cells were first stained with LIVE/DEAD Fixable Dead Cell Stain (Life technologies, Eugene, OR), and next incubated with various combinations of antibodies against T cell markers (both for 30 min at 4 °C). Tetramers were added 30 min before antibodies against other surface markers. For intracellular staining, cells were permeabilized with FACS Permeabilization solution (BD Biosciences or eBiosciences) and incubated with antibodies against various intracellular molecules for 30 min at 4 °C. Flow cytometry acquisition was performed on a custom-designed BD Biosciences Fortessa and analyzed using FlowJo software (Tree Star, Ashland, OR). T cell subsets were sorted using a modified FACSAria (BD Biosciences).
Stimulation assays.
PBMCs were stimulated with either PMA plus ionomycin (Cell stimulation cocktail, eBioscience), plate-bound CD3/CD49d/soluble CD28 (at 10 μg/ml, 5 μg/ml, and 5 μg/ml respectively) or with peptides or their mixtures [Influenza Matrix peptide57-66 (AnaSpec, San Jose, CA), EBV BZLF-1 peptide pool, CMV pp65 peptide pool (both Miltenyi Biotec, San Diego, CA) or HIV-1 PTE Gag peptide pool43 (obtained from the NIH AIDS reagent program)] for 3 h in the presence of brefeldin A (eBioscience). All peptides were used according to manufacturer's instructions at approximately 1 μg/ml of each peptide; peptide mixtures were constructed as sets of 15-mers overlapped by 9 amino acids. Phenotype and cytokine production were evaluated by flow cytometry as described above. In the phospho Erk blocking experiment, MAPK inhibitor U0126 (Tocris Bioscience, Bristol, UK) was added to PBMCs for 3 h prior stimulation with PMA plus ionomycin.
Phos-Flow.
PBMCs were stimulated with PMA plus ionomycin for 5, 10 or 15 min and simultaneously stained with antibodies to surface markers and phospho-Erk (Thr202/Tyr204) using BD Biosciences Phos-flow fixation and permeabilization solutions according to manufacturer's protocol.
Cytokine proliferation assays.
PBMCs were thawed, labeled with CellTrace Violet (CTV) proliferation dye (Molecular Probes, Eugene, OR) according to manufacturer's instructions (2 μM at 5 × 106 cells/ml for 30 min at 37 °C) and rested overnight. The next day, various T cell subsets were isolated by cell sorting and incubated in the presence of cytokines (IL-7 (50 ng/ml), IL-15 (50 ng/ml) (both R&D Systems, Minneapolis, MN), IL-12 (5 ng/ml; Peprotech, Rocky Hill, NJ) or IL-18 (5 ng/ml; Life Technologies, Eugene, OR)). On day 3 or 7, phenotype, proliferation (dilution of CTV proliferation dye) and/or cytokine production were evaluated by flow cytometry.
Sorting of IFN-γ+ cells.
PBMCs were thawed, labeled with CTV and rested overnight. The next day, phenotypically naive IFN-γ-producing cells (TVN) and IFN-γ-nonproducing (TN) CD8+ T cells were isolated using an IFN-γ secretion assay cell enrichment and detection kit (Miltenyi Biotec) and cell sorting. Cells were transferred onto plates coated with anti-CD3 (10 μg/ml; in the presence of soluble CD28 at 5 μg/ml), stimulated for 3 h at 37 °C, labeled with IFN-γ specific capture antibody reagent and incubated for 45 min at 37 °C (while slowly rotating). Next, the cells were concurrently incubated with IFN-γ detection antibody and antibodies against the remaining phenotypic T cell markers, which allowed for identification of naive T cell subset. IFN-γ-producing and IFN-γ-nonproducing naive CD8+ T cell subsets were isolated by cell sorting and incubated in the presence of CD3/CD2/CD28 coated beads (T cell activation/expansion kit, Miltenyi Biotec) at ratio 1:1 in the presence of IL-2 (100 U/ml) for 3 d, when proliferation and phenotype were evaluated by flow cytometry.
SPADE analysis.
Spanning-tree progression analysis of density-normalized events (SPADE) clustering algorithm44 on the Cytobank.org platform was used to analyze doublet-exclused, alive CD8b+ lymphocytes from nine subjects. This produced an interconnected cluster of nodes, or clusters of connected nodes, that correspond with phenotypically defined CD8 T cell populations.
Multicolor fluorescence in situ hybridization for determination of telomere length.
Experiment was performed according to modified manufacturer's protocol as described45. PBMCs were first stained with LIVE/DEAD Fixable Dead Cell Stain and then incubated with biotinylated anti-CD28 or directly conjugated antibodies to CD45RA (BV711), CCR7 (FITC), CD3 (BV570), CD95 (BV421), CD49d (BV510) and CD8α (BV650), followed by 15 min incubation with streptavidin-conjugated Cy3. Cells were fixed and permeabilized using FACS Fixation and permeabilization kit (BD Biosciences). Samples were than washed in PBS, fixed in 1 mM BS3 (30 min on ice, ThermoFisher Scientific) and quenched with 50 mM Tris-HCl in PBS (pH 7.2, 20 min, room temperature). The cells were washed twice; first in PBS, and then in hybridization buffer (70% deionized formamide, 28.5 mM Tris-HCl pH 7.2, 1.4% BSA and 0.2 M NaCl). The samples were subsequently resuspended in hybridization buffer and incubated with of the PNA TelC –Cy5 probe (200 nM, PNA Bio Inc, Thousand Oaks, CA) and heated for 10 min at 82 °C, rapidly cooled on ice and left to hybridize for 1 h at room temperature in the dark. Lastly, the samples were washed in post-hybridization buffer (70% deionized formamide, 14.25 mM Tris-HCl pH 7.2, 0.14% BSA, 0.2 M NaCl and 0.14% Tween-20) and in PBS 2% BSA before acquisition on BD Biosciences Fortessa and analyzed using FlowJo software. Unless stated otherwise, chemicals were purchased from Sigma-Aldrich (St. Louis, MO).
RNA sequencing.
T cell samples from sorted populations were obtained for genome-wide expression profiling by RNA-Seq. There was a broad range of cell numbers available from the different populations (some samples had down to a few thousand cells), so a strategy for low-input RNA-Seq across all of the samples was employed. Trizol extraction was followed by isopropanol precipitation to obtain total RNA. Each total RNA sample was re-suspended in 20 μl of Elution Solution (nuclease-free water supplemented with 0.5 U/uL RNase inhibitor (SUPERase IN, Life Technologies)). To isolate mRNA, LNA-oligo(dT) was used rather than DNA-oligo(dT). LNA is an analog of RNA with a modified sugar backbone that imparts a much higher melting point46 and therefore facilitates high sensitivity mRNA capture. Biotinylated LNA-Oligo(dT) (Exiqon) was attached to streptavidin-coated superparamagnetic beads (C1 Dynabeads, Life Technologies) in Hybridization Buffer (20 mM Tris pH 8, 1 M NaCl, 0.1% tween-20). After washing the beads three times in Wash Buffer (20 mM Tris pH 8, 50 mM NaCl, 0.1% tween-20) and re-suspending in Hybridization Buffer, 20 μl of beads were added to each total RNA sample and the mixture was incubated for 45 min at room temperature on a rotisserie. After washing the beads three times with Wash Buffer and re-suspending in 15 μl of Elution Solution, the bead mixture was heated to 75C for two minutes followed by immediate removal of the supernatant containing purified mRNA.
Strand-specific RNA-Seq libraries were constructed using the template-switching strategy implemented in the SMARTer Stranded RNA-Seq kit (Clontech). mRNA fragmentation reactions were run for four minutes and cDNA was purified twice with size selection beads (Ampure XP, Beckman) before PCR enrichment. Libraries were sequenced on an Illumina NextSeq 500 sequencer with 75-base single-end reads. The reads were mapped to the Ensembl GRCh37 human genome and transcriptome annotation (obtained from Illumina iGenomes) using Tophat2 (ref. 47) and the uniquely mapped reads associated with each gene using HTseq were counted46. Differential expression analysis between different T cell populations was constructed using DESeq2 (ref. 48). Normalization was performed with samples within cell types as biological replicates. The counts were transformed using the regularized log transformation provided in the DESeq2 package as described48. Principal component analysis was done based on all genes annotated from the Consensus Coding Sequence (CCDS) Project49. The figures were produced using ggplot2 (ref. 50) and gplots51 packages from R programming language.
Statistical analyses.
Statistical analyses were performed with Prism (GraphPad Software) and SAS 9.4 (SAS Inc. NC). The Shapiro-Wilk normality test was used to evaluate normal distribution of the data. Comparisons between two groups were performed with the paired t-test (when the groups were matched-paired and normal distribution assumption of the data was met) or the Mann-Whitney U-test (when the groups were independent and normal distribution assumption of the data was violated). ANOVA was performed to compare more than two independent group comparisons. When normal distribution assumption was violated, the Kruskal-Wallis test was used. Repeated-measures ANOVA was used to evaluate longitudinal experiment with multiple groups. Prior to the application of repeated-measures ANOVA, Levene's test was performed to test equal variance assumption for ANOVA. Heterogeneous Compound Symmetry covariance structure was applied to repeated-measures ANOVA to not only account for the within-subject correlation but also adjust for the variance heterogeneity between samples that did not meet the equal-variance assumption.
Accession codes.
GEO: RNA-Seq data set (Fig. 5), GSE80306; NIAID ImmPort database: raw data (all other results), SDY736.
Accession codes
References
Jenkins, M.K., Chu, H.H., McLachlan, J.B. & Moon, J.J. On the composition of the preimmune repertoire of T cells specific for peptide-major histocompatibility complex ligands. Annu. Rev. Immunol. 28, 275–294 (2010).
Wertheimer, A.M. et al. Aging and cytomegalovirus infection differentially and jointly affect distinct circulating T cell subsets in humans. J. Immunol. 192, 2143–2155 (2014).
Nikolich-Zugich, J. Ageing and life-long maintenance of T-cell subsets in the face of latent persistent infections. Nat. Rev. Immunol. 8, 512–522 (2008).
Qi, Q. et al. Diversity and clonal selection in the human T-cell repertoire. Proc. Natl. Acad. Sci. USA 111, 13139–13144 (2014).
Linton, P.J. & Dorshkind, K. Age-related changes in lymphocyte development and function. Nat. Immunol. 5, 133–139 (2004).
Cambier, J. Immunosenescence: a problem of lymphopoiesis, homeostasis, microenvironment, and signaling. Immunol. Rev. 205, 5–6 (2005).
Goronzy, J.J. & Weyand, C.M. T cell development and receptor diversity during aging. Curr. Opin. Immunol. 17, 468–475 (2005).
Miller, R.A. Effect of aging on T lymphocyte activation. Vaccine 18, 1654–1660 (2000).
Thompson, W.W. et al. Mortality associated with influenza and respiratory syncytial virus in the United States. J. Am. Med. Assoc. 289, 179–186 (2003).
Petersen, L.R., Roehrig, J.T. & Hughes, J.M. West Nile virus encephalitis. N. Engl. J. Med. 347, 1225–1226 (2002).
Chinn, I.K., Blackburn, C.C., Manley, N.R. & Sempowski, G.D. Changes in primary lymphoid organs with aging. Semin. Immunol. 24, 309–320 (2012).
Nikolich-Žugich, J. Aging of the T cell compartment in mice and humans: from no naive expectations to foggy memories. J. Immunol. 193, 2622–2629 (2014).
Albright, J.F. & Albright, J.W. Aging, Immunity and Infection. Humana Press: Totowa, 2003.
Maecker, H.T., McCoy, J.P. & Nussenblatt, R. Standardizing immunophenotyping for the Human Immunology Project. Nat. Rev. Immunol. 12, 191–200 (2012).
Sallusto, F., Geginat, J. & Lanzavecchia, A. Central memory and effector memory T cell subsets: function, generation, and maintenance. Annu. Rev. Immunol. 22, 745–763 (2004).
Kaech, S.M., Wherry, E.J. & Ahmed, R. Effector and memory T-cell differentiation: implications for vaccine development. Nat. Rev. Immunol. 2, 251–262 (2002).
Tan, J.T. et al. Interleukin (IL)-15 and IL-7 jointly regulate homeostatic proliferation of memory phenotype CD8+ cells but are not required for memory phenotype CD4+ cells. J. Exp. Med. 195, 1523–1532 (2002).
Newell, E.W., Sigal, N., Bendall, S.C., Nolan, G.P. & Davis, M.M. Cytometry by time-of-flight shows combinatorial cytokine expression and virus-specific cell niches within a continuum of CD8+ T cell phenotypes. Immunity 36, 142–152 (2012).
Hamann, A., Andrew, D.P., Jablonski-Westrich, D., Holzmann, B. & Butcher, E.C. Role of alpha 4-integrins in lymphocyte homing to mucosal tissues in vivo. J. Immunol. 152, 3282–3293 (1994).
Chiu, B.C., Martin, B.E., Stolberg, V.R. & Chensue, S.W. Cutting edge: Central memory CD8 T cells in aged mice are virtual memory cells. J. Immunol. 191, 5793–5796 (2013).
Xia, M.Q., Qin, S.X., Wu, L.J., Mackay, C.R. & Hyman, B.T. Immunohistochemical study of the β-chemokine receptors CCR3 and CCR5 and their ligands in normal and Alzheimer's disease brains. Am. J. Pathol. 153, 31–37 (1998).
Jaensson, E. et al. Small intestinal CD103+ dendritic cells display unique functional properties that are conserved between mice and humans. J. Exp. Med. 205, 2139–2149 (2008).
Gebhardt, T. et al. Memory T cells in nonlymphoid tissue that provide enhanced local immunity during infection with herpes simplex virus. Nat. Immunol. 10, 524–530 (2009).
Tough, D.F., Zhang, X. & Sprent, J. An IFN-γ-dependent pathway controls stimulation of memory phenotype CD8+ T cell turnover in vivo by IL-12, IL-18, and IFN-γ. J. Immunol. 166, 6007–6011 (2001).
Szabo, S.J. et al. A novel transcription factor, T-bet, directs Th1 lineage commitment. Cell 100, 655–669 (2000).
Knox, J.J., Cosma, G.L., Betts, M.R. & McLane, L.M. Characterization of T-bet and eomes in peripheral human immune cells. Front. Immunol. 5, 217 (2014).
Hodes, R.J., Hathcock, K.S. & Weng, N.P. Telomeres in T and B cells. Nat. Rev. Immunol. 2, 699–706 (2002).
Appay, V. et al. Memory CD8+ T cells vary in differentiation phenotype in different persistent virus infections. Nat. Med. 8, 379–385 (2002).
Coulie, P.G. et al. A new gene coding for a differentiation antigen recognized by autologous cytolytic T lymphocytes on HLA-A2 melanomas. J. Exp. Med. 180, 35–42 (1994).
Messaoudi, I., Guevara Patiño, J.A., Dyall, R., LeMaoult, J. & Nikolich-Zugich, J. Direct link between MHC polymorphism, T cell avidity, and diversity in immune defense. Science 298, 1797–1800 (2002).
Moon, J.J. et al. Naive CD4+ T cell frequency varies for different epitopes and predicts repertoire diversity and response magnitude. Immunity 27, 203–213 (2007).
Cicin-Sain, L. et al. Dramatic increase in naive T cell turnover is linked to loss of naive T cells from old primates. Proc. Natl. Acad. Sci. USA 104, 19960–19965 (2007).
Hamann, D. et al. Phenotypic and functional separation of memory and effector human CD8+ T cells. J. Exp. Med. 186, 1407–1418 (1997).
Bendall, S.C., Nolan, G.P., Roederer, M. & Chattopadhyay, P.K. A deep profiler's guide to cytometry. Trends Immunol. 33, 323–332 (2012).
Ornatsky, O., Baranov, V.I., Bandura, D.R., Tanner, S.D. & Dick, J. Multiple cellular antigen detection by ICP-MS. J. Immunol. Methods 308, 68–76 (2006).
Schenkel, J.M. & Masopust, D. Tissue-resident memory T cells. Immunity 41, 886–897 (2014).
Gattinoni, L. et al. A human memory T cell subset with stem cell-like properties. Nat. Med. 17, 1290–1297 (2011).
Haines, C.J. et al. Human CD4+ T cell recent thymic emigrants are identified by protein tyrosine kinase 7 and have reduced immune function. J. Exp. Med. 206, 275–285 (2009).
Song, K. et al. Characterization of subsets of CD4+ memory T cells reveals early branched pathways of T cell differentiation in humans. Proc. Natl. Acad. Sci. USA 102, 7916–7921 (2005).
Turtle, C.J., Swanson, H.M., Fujii, N., Estey, E.H. & Riddell, S.R. A distinct subset of self-renewing human memory CD8+ T cells survives cytotoxic chemotherapy. Immunity 31, 834–844 (2009).
Hinrichs, C.S. et al. Adoptively transferred effector cells derived from naive rather than central memory CD8v T cells mediate superior antitumor immunity. Proc. Natl. Acad. Sci. USA 106, 17469–17474 (2009).
Lanteri, M.C. et al. West Nile virus nucleic acid persistence in whole blood months after clearance in plasma: implication for transfusion and transplantation safety. Transfusion 54, 3232–3241 (2014).
Betts, M.R. et al. Analysis of total human immunodeficiency virus (HIV)-specific CD4+ and CD8+ T-cell responses: relationship to viral load in untreated HIV infection. J. Virol. 75, 11983–11991 (2001).
Qiu, P. et al. Extracting a cellular hierarchy from high-dimensional cytometry data with SPADE. Nat. Biotechnol. 29, 886–891 (2011).
Riddell, N.E. et al. Multifunctional cytomegalovirus (CMV)-specific CD8+ T cells are not restricted by telomere-related senescence in young or old adults. Immunology 144, 549–560 (2015).
Koshkin, A.A. et al. LNA (locked nucleic acid): an RNA mimic forming exceedingly stable LNA:LNA duplexes. J. Am. Chem. Soc. 120, 13252–13253 (1998).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Love, M.I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Pruitt, K.D. et al. The consensus coding sequence (CCDS) project: Identifying a common protein-coding gene set for the human and mouse genomes. Genome Res. 19, 1316–1323 (2009).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer Science & Business Media, 2009).
Warnes, G.R. et al. gplots: Various R Programming Tools for Plotting Data. R package version 2.17.0 (2015).
Acknowledgements
We thank P. Campbell of the Arizona Cancer Center, Arizona Research Laboratory Flow Cytometry Core Facility for cell sorting; the Arizona Cancer Center Support Grant (NCI grant CA 023074) for facility support; NIH Tetramer Facility for reagent preparation; K. Kedzierska (University of Melbourne) for tetramer reagents (HLA-A2 INF-GIL tetramers); and the members of the Nikolich, Kuhns, Frelinger, Wu and Schenten laboratories for insight and discussions. Supported by the US National Institutes of Health (NIAID (N01-AI00017), NHLBI (RC2HL101) and NIA (R01 AG-048021 and P30 AG008017) and by the Bowman Endowed Professorship in Medical Research (J.N.-Ž.).
Author information
Authors and Affiliations
Contributions
V.P. and J.N.-Ž. designed and analyzed experiments and wrote the manuscript; V.P. performed stimulation analysis, cell sorting and RNA isolation; J.S.D. performed flow cytometry and SPADE analysis; C.M., K.O.M. and A.M.W. obtained subject consent, drew blood draws, organized samples, searched human subject databases and managed samples for subjects from Arizona, Oregon and Texas; M.C.L. and M.P.B. provided WNV-exposed samples from the San Francisco Blood Bank; M.S.D. facilitated sample transfer; K.K. provided samples from HIV+ subjects; E.S.B. and P.A.S. performed the RNA-Seq experiments and, together with V.P., J.S.D., S.S. and D.B., performed RNA-Seq data analysis; E.K.H. provided subject samples and critical design input; K.O.M. provided samples from WNV-exposed subjects from the Houston area; J.S.D., M.C.L., M.P.B., M.S.D., K.K., P.A.S., D.B. and A.M.W. edited the manuscript; and J.N.-Ž. conceived of the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Identification of IFN-γ-producing CD8+ and CD4+ T cells with naive phenotype by alternative gating and sample-processing strategies.
a. Contour 5% probability plots show definition of TN, TCM, TE+EM and TEMRA subsets based on the expression of pairs of CD45RA and CCR7 or CD95 and CD28 molecules by total CD8+ T cells from a representative donor (age 71). b. Gating strategies for identification of TN (CD45RA+CCR7+CD95loCD28int)(top row), TCM (middle row), TE+EM and TEMRA (both bottom row) CD8+ T cell subsets, using gating in reverse compared to Fig. 1, by first applying the CD95/28 gate and then the CD45RA/CCR7 gate. c. Age dependent decrease in phenotypically naive CD8+ T cells (shown in black) and IFN-γ producing phenotypically naive (CD45RA+CCR7+CD95loCD28int CD8+ T cells (in green) shown as cell number (Log10)/µL of blood (n=92). d. Graph shows flow cytometric analysis of percentages of IFN-γ+CD4+ T cells with naive phenotype (gated through CD45RA+CCR7+CD28intCD95lo) with age. Student’s t-test was applied to analyze significance in the age-related increase between the group of donors <40 years old (n=27), 40 – 65y old (n=44) or >65 years old (n=21; *p<0.05). Data are representative of one experiment. e. Intracellular cytokine staining of human PMA+iono- activated fresh (not cryopreserved) PBMCs from healthy donors. Dot plots show IFN-γ, TNF-α, and GzmB production by phenotypically naïve (CD45RA+CCR7+CD95loCD28int CD8+ T cells from freshly isolated blood of 7 donors (age 32-76 as indicated). Unstimulated (US) PBMCs were used as control. f. Intracellular cytokine staining of sorted naïve (CD45RA+CCR7+CD95loCD28int CD8+ T cells after 3 hr activation with PMA-iono. Bar graphs show percentage of cytokine producing naive CD8+ T cells in healthy subjects age <40 and > 65, with n=6/group (*p<0.05, experiment repeated once with different subjects, n=4/group).
Supplementary Figure 2 Expression of phenotypic markers on TMNP cells relative that on other CD8+ T cell subsets.
Graphs show flow cytometry-based analysis of diversity (mean gMFI ± SD) in expression of a given molecule between different CD8+ T cell subsets from 12 donors. Repeated measures ANOVA was used to quantify differences in CD45RA, CCR7, CD95 and CD28 expression between TN and TMNP, TCM, TEM or TEMRA CD8 T cells, defined as in Supplemental Fig. 1d (*p<0.05,**p<0.0001, n=12, ages 27-85, 5 males, 7 females).
Supplementary Figure 3 TMNP cells do not respond to bystander stimulation by IL-12 and IL-18.
a. Flow cytometry-based evaluation IFNγ production in sorted naïve CD45RA+CCR7+CD95loCD28int CD49dhigh and CD49dlow CD8+ T cells after overnight IL-12 (5ng/mL) + IL-18 (5ng/mL) culture. Data are representatives of 2 subjects, males ages 52, 64 (of n=3). b. IFN-γ production by different PBMC CD8+ T cell subsets (TN and TMNP, TCM, TEM or TEMRA, defined as in Supplemental Fig. 1d) after overnight IL-12+IL-18 culture and 3hr presence of protein transport inhibitors shown as % of IFN-g+ cells (repeated measures ANOVA,*P<0.05, n=9, ages 35-77).
Supplementary Figure 4 Flow FISH analysis and stability of phenotype after stimulation with PMA plus ionomycin.
a. Flow cytometry analysis of human PBMCs from a representative donor (male, age 68). Contour 5% probability plots and histogram show gating strategy used to identify different CD8+ T cell populations (TN, TCM, TE+EM and TEMRA. As shown in the final graph, 4n cells were excluded from the analysis. b. Contour 5% probability plots and histograms show expression of naive phenotypic markers and intensity of CD8 and A2 CMV-NLV tetramer staining after 3hr PMA+iono stimulation (in right column or in red) or in the presence of BFA alone in one representative subject (left column or in blue) of n=8 (age 35-83).
Supplementary Figure 5 TMNP cells undergo robust proliferation in response to TCR stimulation.
a,b. Overlaid histograms show proliferation (dilution of CTV proliferation dye) of a. sorted TMNP (FN-γ+ CD45RA+CCR7+CD95loCD28int) cells (red line), isolated using IFN-γ capture assay (as in methods) and TN (naive IFN-γ- CD45RA+CCR7+CD95loCD28int) CD8+ T cells(blue line), 3 days after activation with anti-CD3+anti-CD2+anti-CD28 Ab-coated beads (two donors of n=3, 2 females, 1 male, ages 65-73, are shown, t-test, NS) and b. sorted TMNP (CD45RA+CCR7+CD95loCD28intCD49dhi), TN (CD45RA+ CCR7+CD95loCD28intCD49dlo), TCM (CD45RA- CCR7+CD95loCD28hi) and TEM+TEMRA (CD45RA-or+ CCR7+CD95hiCD28int or lo) cells at 6 days after activation with anti-CD3+ anti-CD2+anti-CD28 Ab-coated beads, two representative donors, age 32 and 35 are shown. Proliferation and division indices are shown by scatter graphs (shown as mean ±SEM; repeated measures one-way ANOVA,*P<0.05, n=5, 4 females, 1 male, age 32-69).
Supplementary Figure 6 TCR Vβ distribution in various subsets of CD8+ T cells from a healthy donor (female, 65 years of age), isolated 24 months apart.
3D line graphs showing TCR Vβ distribution in different CD8+ T cells subsets (indicated in the legend). Percentages of each of the 24 tested TCR Vβ (marked on the x axis) are shown. Total analyzed were n=2 (1 female, 1 male, age 65,81). Data are representative of one experiment.
Supplementary Figure 7 Flow cytometry analyzing IFN-γ production by naive IAV-specific CD8+ T cells before and 14 d after vaccination against IAV shows no bystander or nonspecific activation of TMNP cells.
PBMCs were activated briefly with PMA+Iono. Contour 5% probability plots show gating strategy to evaluate the naive phenotype (CD45RA+CCR7+CD95loCD28int) and IFN-γ production by IAV matrix GIL peptide:HLA-A*0201 or CMV pp65 NLV: HLA-A*0201 tetramer+ CD8+ T cells. Numbers represent percentages of the population within each gate after subsequent gating. Data from two representative donors (one CMV seropositive and one seronegative (of n=7, ages 65-80).
Supplementary Figure 8 Major CD8+ T cell subsets do not change in number regardless of the severity of symptoms in WNV-exposed subjects.
Quantification of CD8+ T cell subsets in WNV asymptomatic and symptomatic subjects is represented as absolute number of TN, TE+EM, TEMRA and TCM subset/106 CD8+ T cells within each subject. Repeated measures one-way ANOVA was used to quantify the subsets’ differences between asymptomatic (n=11,,39-92) and symptomatic (n=16, ages 43-84).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 (PDF 2988 kb)
Supplementary Table 1
Antibody validation Table (PDF 118 kb)
Rights and permissions
About this article
Cite this article
Pulko, V., Davies, J., Martinez, C. et al. Human memory T cells with a naive phenotype accumulate with aging and respond to persistent viruses. Nat Immunol 17, 966–975 (2016). https://doi.org/10.1038/ni.3483
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3483
- Springer Nature America, Inc.
This article is cited by
-
CD40 ligand stimulation affects the number and memory phenotypes of human peripheral CD8+ T cells
BMC Immunology (2023)
-
Aging-associated HELIOS deficiency in naive CD4+ T cells alters chromatin remodeling and promotes effector cell responses
Nature Immunology (2023)
-
Real clinical outcomes of nivolumab plus ipilimumab for renal cell carcinoma in patients over 75 years old
International Journal of Clinical Oncology (2023)
-
The Effects of Aging on Solid Organ Transplantation—Characteristics and Consequences of Immunosenescence
Current Transplantation Reports (2023)
-
The interplay between oxidative stress and autophagy: focus on the development of neurological diseases
Behavioral and Brain Functions (2022)