Abstract
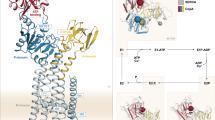
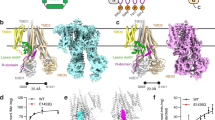
The P1B-ATPases, which couple cation transport across membranes to ATP hydrolysis, are central to metal homeostasis in all organisms. An important feature of P1B-ATPases is the presence of soluble metal binding domains (MBDs) that regulate transport activity. Only one type of MBD has been characterized extensively, but bioinformatics analyses indicate that a diversity of MBDs may exist in nature. Here we report the biochemical, structural and functional characterization of a new MBD from the Cupriavidus metallidurans P1B-4-ATPase CzcP (CzcP MBD). The CzcP MBD binds two Cd2+, Co2+ or Zn2+ ions in distinct and unique sites and adopts an unexpected fold consisting of two fused ferredoxin-like domains. Both in vitro and in vivo activity assays using full-length CzcP, truncated CzcP and several variants indicate a regulatory role for the MBD and distinct functions for the two metal binding sites. Taken together, these findings elucidate a previously unknown MBD and suggest new regulatory mechanisms for metal transport by P1B-ATPases.




Similar content being viewed by others
Accession codes
References
Palmgren, M.G. & Nissen, P. P-type ATPases. Annu. Rev. Biophys. 40, 243–266 (2011).
Argüello, J.M. Identification of ion-selectivity determinants in heavy-metal transport P1B-type ATPases. J. Membr. Biol. 195, 93–108 (2003).
Argüello, J.M., Eren, E. & González-Guerrero, M. The structure and function of heavy metal transport P1B-type ATPases. Biometals 20, 233–248 (2007).
Smith, A.T., Smith, K.P. & Rosenzweig, A.C. Diversity of the metal-transporting P1B-type ATPases. J. Biol. Inorg. Chem. 19, 947–960 (2014).
Argüello, J.M., González-Guerrero, M. & Raimunda, D. Bacterial transition metal P1B-ATPases: transport mechanism and roles in virulence. Biochemistry 50, 9940–9949 (2011).
Rosenzweig, A.C. & Arguello, J.M. Toward a molecular understanding of metal transport by P1B-type ATPases. in Metal Transporters Vol. 69 (eds. Lutsenko, S. & Argüello, J.M.) 113–136 (Elsevier, San Diego, 2012).
Williams, L.E. & Mills, R.F. P1B-ATPases—an ancient family of transition metal pumps with diverse functions in plants. Trends Plant Sci. 10, 491–502 (2005).
Gupta, A. & Lutsenko, S. Evolution of copper transporting ATPases in eukaryotic organisms. Curr. Genomics 13, 124–133 (2012).
Lutsenko, S., Gupta, A., Burkhead, J.L. & Zuzel, V. Cellular multitasking: the dual role of human Cu-ATPases in cofactor delivery and intracellular copper balance. Arch. Biochem. Biophys. 476, 22–32 (2008).
Andersson, M. et al. Copper-transporting P-type ATPases use a unique ion-release pathway. Nat. Struct. Mol. Biol. 21, 43–48 (2014).
Gourdon, P. et al. Crystal structure of a copper-transporting PIB-type ATPase. Nature 475, 59–64 (2011).
Wang, K. et al. Structure and mechanism of Zn2+-transporting ATPase. Nature 514, 518–522 (2014).
Boal, A.K. & Rosenzweig, A.C. Structural biology of copper trafficking. Chem. Rev. 109, 4760–4779 (2009).
Singleton, C. & LeBrun, N.E. Atx1-like chaperones and their cognate P-type ATPases: copper-binding and transfer. Biometals 20, 275–289 (2007).
Banci, L. et al. A new zinc-protein coordination site in intracellular metal trafficking: solution structure of the apo and Zn(II) forms of ZntA(46–118). J. Mol. Biol. 323, 883–897 (2002).
Fu, Y. et al. A new structural paradigm in copper resistance in Streptococcus pneumoniae. Nat. Chem. Biol. 9, 177–183 (2013).
Mana-Capelli, S., Mandal, A.K. & Argüello, J.M. Archaeoglobus fulgidus CopB is a thermophilic Cu2+-ATPase. J. Biol. Chem. 278, 40534–40541 (2003).
Traverso, M.E. et al. Identification of a hemerythrin-like domain in a P1B-type transport ATPase. Biochemistry 49, 7060–7068 (2010).
Zielazinski, E.L., Cutsail, G.E. III, Hoffman, B.M., Stemmler, T.L. & Rosenzweig, A.C. Characterization of a cobalt-specific P1B-ATPase. Biochemistry 51, 7891–7900 (2012).
Raimunda, D., Long, J.E., Sassetti, C.M. & Arguello, J.M. Role in metal homeostasis of CtpD, a Co2+ transporting P1B4-ATPase of Mycobacterium smegmatis. Mol. Microbiol. 84, 1139–1149 (2012).
Raimunda, D., Long, J.E., Padilla-Benavides, T., Sassetti, C.M. & Arguello, J.M. Differential roles for the Co2+/Ni2+ transporting ATPases, CtpD and CtpJ, in Mycobacterium tuberculosis virulence. Mol. Microbiol. 91, 185–197 (2014).
Scherer, J. & Nies, D.H. CzcP is a novel efflux system contributing to transition metal resistance in Cupriavidus metallidurans CH34. Mol. Microbiol. 73, 601–621 (2009).
Whitmore, L. & Wallace, B.A. DICHROWEB, an online server for protein secondary structure analyses from circular dichroism spectroscopic data. Nucleic Acids Res. 32, W668–W673 (2004).
Whitmore, L. & Wallace, B.A. Protein secondary structure analyses from circular dichroism spectroscopy: methods and reference databases. Biopolymers 89, 392–400 (2008).
Zelazowski, A.J., Szymanska, J.A., Law, A.Y.C. & Stillman, M.J. Spectroscopic properties of the α fragment of metallothionein. J. Biol. Chem. 259, 12960–12963 (1984).
Frankel, A.D., Berg, J.M. & Pabo, C.O. Metal-dependent folding of a single zinc finger from transcription factor IIIA. Proc. Natl. Acad. Sci. USA 84, 4841–4845 (1987).
Giedroc, D.P., Keating, K.M., Martin, C.T., Williams, K.R. & Coleman, J.E. Zinc metalloproteins involved in replication and transcription. J. Inorg. Biochem. 28, 155–169 (1986).
Maret, W. & Vallee, B.L. Cobalt as probe and label of proteins. Methods Enzymol. 226, 52–71 (1993).
DeSilva, T.M., Veglia, G. & Opella, S.J. Solution structures of the reduced and Cu(I) bound forms of the first metal binding sequence of ATP7A associated with Menkes disease. Proteins 61, 1038–1049 (2005).
Banci, L., Bertini, I., Ciofi-Baffoni, S., Huffman, D.L. & O'Halloran, T.V. Solution structure of the yeast copper transporter domain Ccc2a in the apo and Cu(I) loaded states. J. Biol. Chem. 276, 8415–8426 (2001).
Banci, L. et al. Structural basis for metal binding specificity: the N-terminal cadmium binding domain of the P1-type ATPase CadA. J. Mol. Biol. 356, 638–650 (2006).
Achila, D. et al. Structure of human Wilson protein domains 5 and 6 and their interplay with domain 4 and the copper chaperone HAH1 in copper uptake. Proc. Natl. Acad. Sci. USA 103, 5729–5734 (2006).
Mandal, A.K., Cheung, W.D. & Argüello, J.M. Characterization of a thermophilic P-type Ag+/Cu+-ATPase from the extremophile Archaeglobus fulgidus. J. Biol. Chem. 277, 7201–7208 (2002).
Sharma, R., Rensing, C., Rosen, B.P. & Mitra, B. The ATP hydrolytic activity of purified ZntA, a Pb(Ii)/Cd(Ii)/Zn(Ii)-translocating ATPase from Escherichia coli. J. Biol. Chem. 275, 3873–3878 (2000).
Mandal, A.K. & Argüello, J.M. Functional roles of metal binding domains of the Archaeoglobus fulgidus Cu+-ATPase CopA. Biochemistry 42, 11040–11047 (2003).
Mattle, D. et al. On allosteric modulation of P-type Cu+-ATPases. J. Mol. Biol. 425, 2299–2308 (2013).
Eren, E., Kennedy, D.C., Maroney, M.J. & Argüello, J.M. A novel regulatory metal binding domain is present in the C terminus of Arabidopsis Zn2+-ATPase HMA2. J. Biol. Chem. 281, 33881–33891 (2006).
Bækgaard, L. et al. A combined zinc/cadmium sensor and zinc/cadmium export regulator in a heavy metal pump. J. Biol. Chem. 285, 31243–31252 (2010).
Monchy, S. et al. Plasmids pMOL28 and pMOL30 of Cupriavidus metallidurans are specialized in the maximal viable response to heavy metals. J. Bacteriol. 189, 7417–7425 (2007).
Zoropogui, A., Gambarelli, S. & Covés, J. CzcE from Cupriavidus metallidurans CH34 is a copper-binding protein. Biochem. Biophys. Res. Commun. 365, 735–739 (2008).
Petit-Haertlein, I. et al. Evidence for conformational changes upon copper binding to Cupriavidus metallidurans CzcE. Biochemistry 49, 1913–1922 (2010).
Lanzetta, P.A., Alvarez, L.J., Reinach, P.S. & Candia, O.A. Improved assay for nanomole amounts of inorganic-phosphate. Anal. Biochem. 100, 95–97 (1979).
Bencze, K.Z., Kondapalli, K.C. & Stemmler, T.L. X-ray absorption spectroscopy. in Applications of Physical Methods to Inorganic and Bioinorganic Chemistry: Handbook, Encyclopedia of Inorganic Chemistry (eds. Scott, R.A. & Lukehart, C.M.) 513–528 (John Wiley & Sons, Ltd., Chicester, UK, 2007).
Ankudinov, A.L. & Rehr, J.J. Relativistic calculations of spin-dependent X-ray absorption spectra. Phys. Rev. B 56, R1712–R1715 (1997).
Lee, P.A., Citrin, P.H., Eisenberger, P. & Kincaid, B.M. Extended X-ray absorption fine structure— its strengths and limitations as a structural tool. Rev. Mod. Phys. 53, 769–806 (1981).
Wang, B. et al. Structure and ubiquitin interactions of the conserved zinc finger domain of Np14. J. Biol. Chem. 278, 20225–20234 (2003).
Winter, G. xia2: an expert system for macromolecular crystallography data reduction. J. Appl. Crystallogr. 43, 186–190 (2010).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Winn, M.D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
Murshudov, G.N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D Biol. Crystallogr. 67, 355–367 (2011).
Lebedev, A.A. et al. JLigand: a graphical tool for the CCP4 template-restraint library. Acta Crystallogr. D Biol. Crystallogr. 68, 431–440 (2012).
Acknowledgements
This work was supported by US National Institutes of Health grants GM58518 (A.C.R.), DK068139 (T.L.S.), F32GM105339 (A.T.S.) and T32HL120822 (D.B.). Sequence searches used both database and analysis functions of the Universal Protein Resource (UniProt) Knowledgebase and Reference Clusters (http://www.uniprot.org/) and the US National Center for Biotechnology Information (http://www.ncbi.nlm.nih.gov/). Portions of this research were carried out at the National Synchrotron Light Source (NSLS). NSLS, located at Brookhaven National Laboratory, is supported by the US Department of Energy, Division of Materials Sciences and Division of Chemical Sciences under contract no. DE-AC02-98CH10886.
Author information
Authors and Affiliations
Contributions
A.T.S. and D.B. conducted all of the experiments, and A.C.R. and T.L.S. directed the research. A.T.S., D.B., T.L.S. and A.C.R. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–10 and Supplementary Tables 1–4. (PDF 33719 kb)
Rights and permissions
About this article
Cite this article
Smith, A., Barupala, D., Stemmler, T. et al. A new metal binding domain involved in cadmium, cobalt and zinc transport. Nat Chem Biol 11, 678–684 (2015). https://doi.org/10.1038/nchembio.1863
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1863
- Springer Nature America, Inc.
This article is cited by
-
Stabilization of supramolecular membrane protein–lipid bilayer assemblies through immobilization in a crystalline exoskeleton
Nature Communications (2021)
-
Using Chemical Modifiers and Increasing the Pyrolysis Temperature for High-Sensitivity Spectrometric Determination of Cadmium in Dairy Products
Journal of Applied Spectroscopy (2020)
-
Characterization of a promiscuous cadmium and arsenic resistance mechanism in Thermus thermophilus HB27 and potential application of a novel bioreporter system
Microbial Cell Factories (2018)
-
Enhanced expression of SaHMA3 plays critical roles in Cd hyperaccumulation and hypertolerance in Cd hyperaccumulator Sedum alfredii Hance
Planta (2016)
-
Novel and efficient screening of PQQ high-yielding strains and subsequent cultivation optimization
Applied Microbiology and Biotechnology (2016)